Abstract
The tissue-specific gene expression at the brain microvasculature, which forms the blood–brain barrier (BBB) in vivo, can be elucidated with a brain vascular genomics program, which starts with the isolation of gene products derived from purified brain microvessels. Genes commonly expressed in peripheral organs are subtracted with the suppression subtractive hybridization method using driver cDNA produced from a pool of rat liver/kidney–derived poly A+RNA. From a screen of 480 clones in the subtracted tester cDNA library, 156 clones were sequenced. The clones fell into 3 groups: known genes (51%), rat expressed sequence tags (31%), and novel rat genes not found in databases (18%). The known genes could be categorized into families of common function including vascular remodeling, signal transduction, transcription factors, biologic transport, vascular amyloid, hemostasis, myelin, lipids, secretion, cytoskeleton, and junctional complexes. Brain vascular genomics, or BBB genomics, allows for an accelerated discovery of the gene families that are differentially expressed at the microvasculature in brain.
The capillary endothelial cells of the vertebrate brain are unique, as compared with endothelial cells in other tissues, since brain capillary endothelial cells express epithelial-like high-resistance tight junctions that eliminate interendothelial pores in the capillary wall (Brightman, 1977). These tight junctions, and other brain endothelial properties, form the anatomic basis of the blood–brain barrier (BBB) in the vertebrate brain. The microvasculature of the brain provides the interface between blood and brain and serves multiple functions in both health and disease. One approach to investigating the multiple functions of the brain microvasculature is to examine the profiles of genes that are differentially expressed in the microvasculature of brain (Li et al., 2001). A brain vascular genomics program should be distinct from a whole brain genomics analysis, because many vascular-specific transcripts will not be detected in whole brain analyses. The microvasculature occupies a very small volume within the brain, which is less than 10−3 parts of whole brain. Since the sensitivity of gene microarray methodologies is on the order of 10−4 (Schena et al., 1995), most brain microvascular-specific transcripts will be missed in the whole-brain gene microarray. A brain vascular genomics program starts with the isolation of brain capillary–derived poly A+ RNA (Li et al., 2001). The specificity of gene profiling of the brain microvasculature is increased with subtractive hybridization methodologies such as suppression subtractive hybridization (SSH) (Diatchenko et al., 1996). In this approach, brain capillary–derived poly A+RNA is used to prepare tester cDNA and a pool of liver and kidney–derived poly A+RNA is used to prepare driver cDNA. The present investigation extends prior work (Li et al., 2001), and profiles genes that are differentially expressed at the microvasculature of control rat brain. Approximately half of the genes profiled are genes of known function, and about half of the genes are either rat expressed sequence tags (ESTs) or novel genes. The known genes that are differentially expressed at the rat brain microvasculature can be grouped on the basis of common functions ranging from biologic transport to vascular remodeling.
MATERIALS AND METHODS
Rat brain capillary cDNA library
Brain microvessels were isolated from control male Sprague-Dawley rats of 250- to 300-g body weight, and brain poly A+RNA was purified from these brain capillaries as described previously (Li et al., 2001). The rat brain capillary poly A+RNA was used to produce 2.5 μg of tester cDNA, and the subtraction procedure was completed using both rat liver and kidney mRNA to generate driver cDNA. In a prior screening of this library, a total of 50 clones were isolated and sequenced (Li et al., 2001). In the present study, this same library is screened, and an additional 157 clones were sequenced.
Suppression subtractive hybridization and subtracted cDNA screening
Suppression subtractive hybridization was performed using the polymerase chain reaction (PCR)-Select cDNA subtraction system according to the manufacturer's instructions (Clontech, Palo Alto, CA, U.S.A.) previously described (Li et al., 2001; Shusta et al., 2002a). Positive clones were identified by differential hybridization. Membranes were hybridized with [32P]-labeled subtracted or unsubtracted rat brain capillary tester cDNA in the presence of PCR-select Differential Screening Blocking Solution (Clontech). Film autoradiography was performed with X-Omat Blue XB-1 film (Kodak, Rochester, NY, U.S.A.) for 4 days at −20°C. cDNA was purified with Qiaquick PCR purification kit (Qiagen, Santa Clarita, CA, U.S.A.) and labeled as previously described (Boado et al., 1999). The 96-well x-ray films were scanned with a flatbed scanner and transparency adapter, and the hybridization signal was quantified with National Institutes of Health (NIH) Image software. Positive clones were divided into two groups according to the integrated density of the signal of the [32P]-subtracted tester cDNA hybridization with the subtracted tester cDNA library. Those clones with an integrated density more than 160,000 arbitrary units were considered high intensity, and this group was designated the LKH group for liver (L)/kidney (K) subtracted with high (H) intensity. Out of 480 clones screened, a total of 66 clones in the LKH group were selected for DNA sequence analysis. Those clones with an integrated density between 60,000 to 160,000 arbitrary units were considered medium intensity, and this group was designated the LKM group for liver (L)/kidney (K) subtracted with medium (M) intensity. Out of 500 clones screened, a total of 103 clones in the LKM group were selected for DNA sequence analysis. The separation of the clones into the H and M hybridization intensity is arbitrary, and done to simplify sequencing of a large number of clones. In the H and M groups combined, the ratio of the integrated density of the hybridization signal for the subtracted tester cDNA library relative to the unsubtracted tester cDNA library ranged from 6 to 200 with a mean of 64. Clones with a ratio less than 6 were not selected for DNA sequencing.
DNA sequence analysis
DNA sequencing of isolated clones was performed with M13 reverse primer using a CEQ2000XL DNA Sequence Analysis System available in this laboratory, according to the manufacturer's instructions (Beckman Coulter, Fullerton, CA, U.S.A.). Sequencing reactions were carried out with M13 reverse primer and 100 fmol plasmid DNA in the pCRII vector using Dye Terminator Cycle Sequencing (DTCS) Quick Start Kit in Thermowell 96-well sample plates (Corning Inc., Corning, NY, U.S.A.). The reaction mixture was amplified over 30 cycles in an iCycler Thermal Cycler (Bio-Rad, Hercules, CA, U.S.A.). The amplification conditions were: 96°C for 20 seconds, 50°C for 20 seconds, and 60°C for 4 minutes. Sequencing products were precipitated with 60 μL of 95% ethanol and centrifuged at 5,700 g for 2 minutes at 4°C using an Allegra 25R plate centrifuge with a S5700 rotor (Beckman Coulter). Samples were run in the CEQ2000XL using the manufacturer's LFR-1 method at the following conditions: capillary temperature 50°C; denaturation, 120 seconds at 90°C; injection, 15 seconds at 2 kV; separation, 85 minutes at 4.2 kV). The DNA sequence between the EcoRI insert sites of the selected clones was compared with DNA sequence in GenBank using the BLAST program (NCBI, NIH).
RESULTS
A 96-well Southern blot shows the hybridization of the radiolabeled subtracted tester cDNA with individual clones randomly picked from the subtracted tester cDNA library (Fig. 1). The clones that exhibited very high hybridization intensity were selected for the LKH group (see Materials and Methods), and representative clones are identified in Fig. 1. Clones with intermediate hybridization intensity were selected for the LKM group (see Materials and Methods) and representative clones are shown in Fig. 1. Out of 480 clones screened (see Materials and Methods), a total of 169 clones fit the LKH or LKM criteria, and DNA sequence analysis was obtained on 156 of these clones. Based on the DNA sequence, the clones were segregated into one of three groups: known genes, rat ESTs, and novel genes not found in the databases, as listed in Tables 1, 2, and 3, respectively. There were 36 known genes identified (Table 1), 22 rat ESTs isolated (Table 2), and 12 novel genes cloned (Table 3). Several genes were isolated from multiple clones and the number of isolates for these genes are shown in Tables 1 to 3. The rat ESTs and the novel genes represent 31% and 18%, respectively, of the total genes cloned. The known genes represent 51% of the total, and these genes could be classified on the basis of common functions, as shown in Fig. 2.
Known genes selectively enriched at the rat brain microvasculature
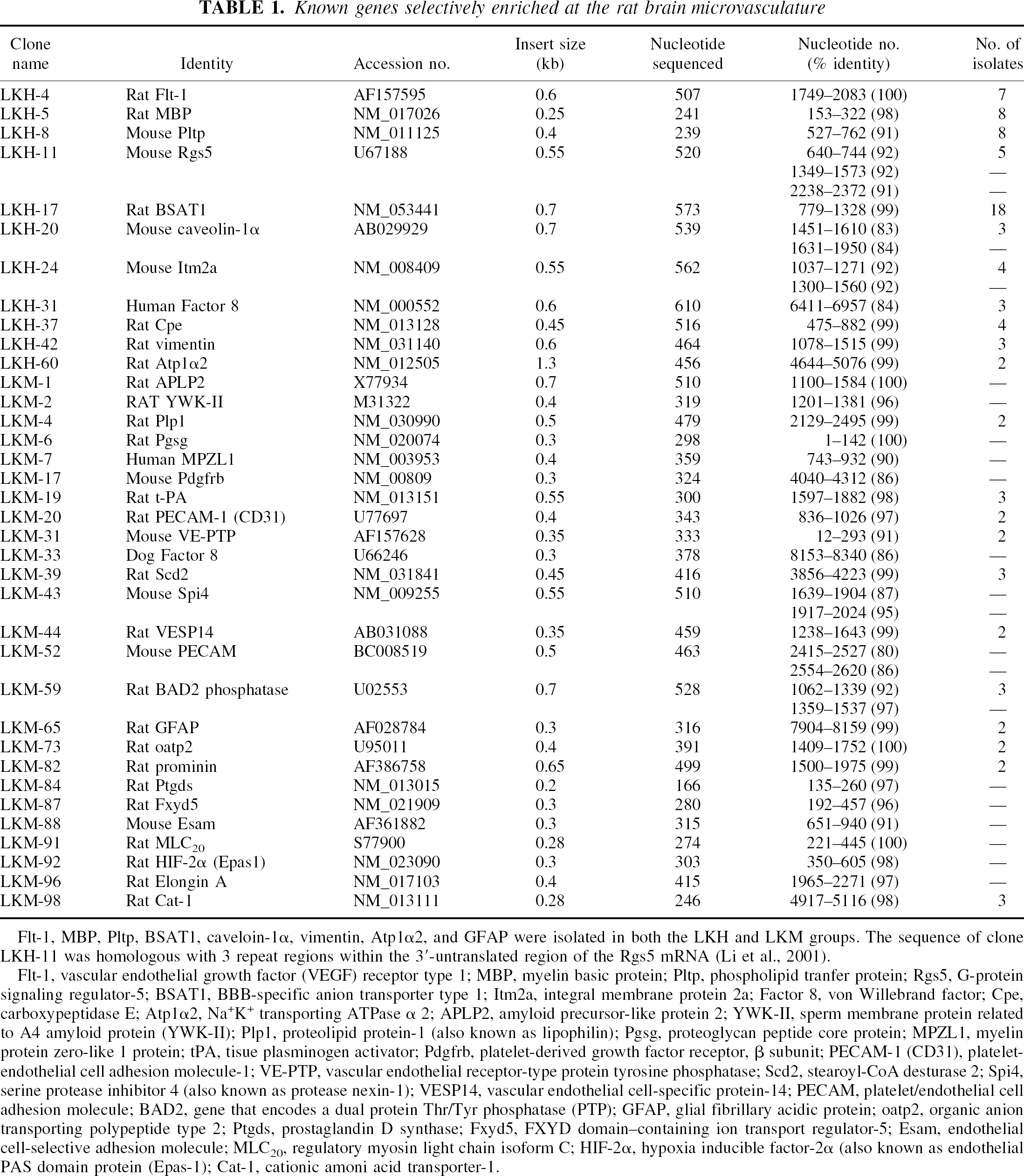
Flt-1, MBP, Pltp, BSAT1, caveloin-1α, vimentin, Atp1α2, and GFAP were isolated in both the LKH and LKM groups. The sequence of clone LKH-11 was homologous with 3 repeat regions within the 3′-untranslated region of the Rgs5 mRNA (Li et al., 2001).
Flt-1, vascular endothelial growth factor (VEGF) receptor type 1; MBP, myelin basic protein; Pltp, phospholipid tranfer protein; Rgs5, G-protein signaling regulator-5; BSAT1, BBB-specific anion transporter type 1; Itm2a, integral membrane protein 2a; Factor 8, von Willebrand factor; Cpe, carboxypeptidase E; Atp1α2, Na+K+ transporting ATPase α 2; APLP2, amyloid precursor-like protein 2; YWK-II, sperm membrane protein related to A4 amyloid protein (YWK-II); Plp1, proteolipid protein-1 (also known as lipophilin); Pgsg, proteoglycan peptide core protein; MPZL1, myelin protein zero-like 1 protein; t-PA, tisue plasminogen activator; Pdgfrb, platelet-derived growth factor receptor, β subunit; PECAM-1 (CD31), plateletendothelial cell adhesion molecule-1; VE-PTP, vascular endothelial receptor-type protein tyrosine phosphatase; Scd2, stearoyl-CoA desturase 2; Spi4, serine protease inhibitor 4 (also known as protease nexin-1); VESP14, vascular endothelial cell-specific protein-14; PECAM, platelet/endothelial cell adhesion molecule; BAD2, gene that encodes a dual protein Thr/Tyr phosphatase (PTP); GFAP, glial fibrillary acidic protein; oatp2, organic anion transporting polypeptide type 2; Ptgds, prostaglandin D synthase; Fxyd5, FXYD domain–containing ion transport regulator-5; Esam, endothelial cell-selective adhesion molecule; MLC20, regulatory myosin light chain isoform C; HIF-2α, hypoxia inducible factor-2α (also known as endothelial PAS domain protein (Epas-1); Cat-1, cationic amoni acid transporter-1.
Rat expressed sequence tags selectively enriched at the rat brain microvasculature
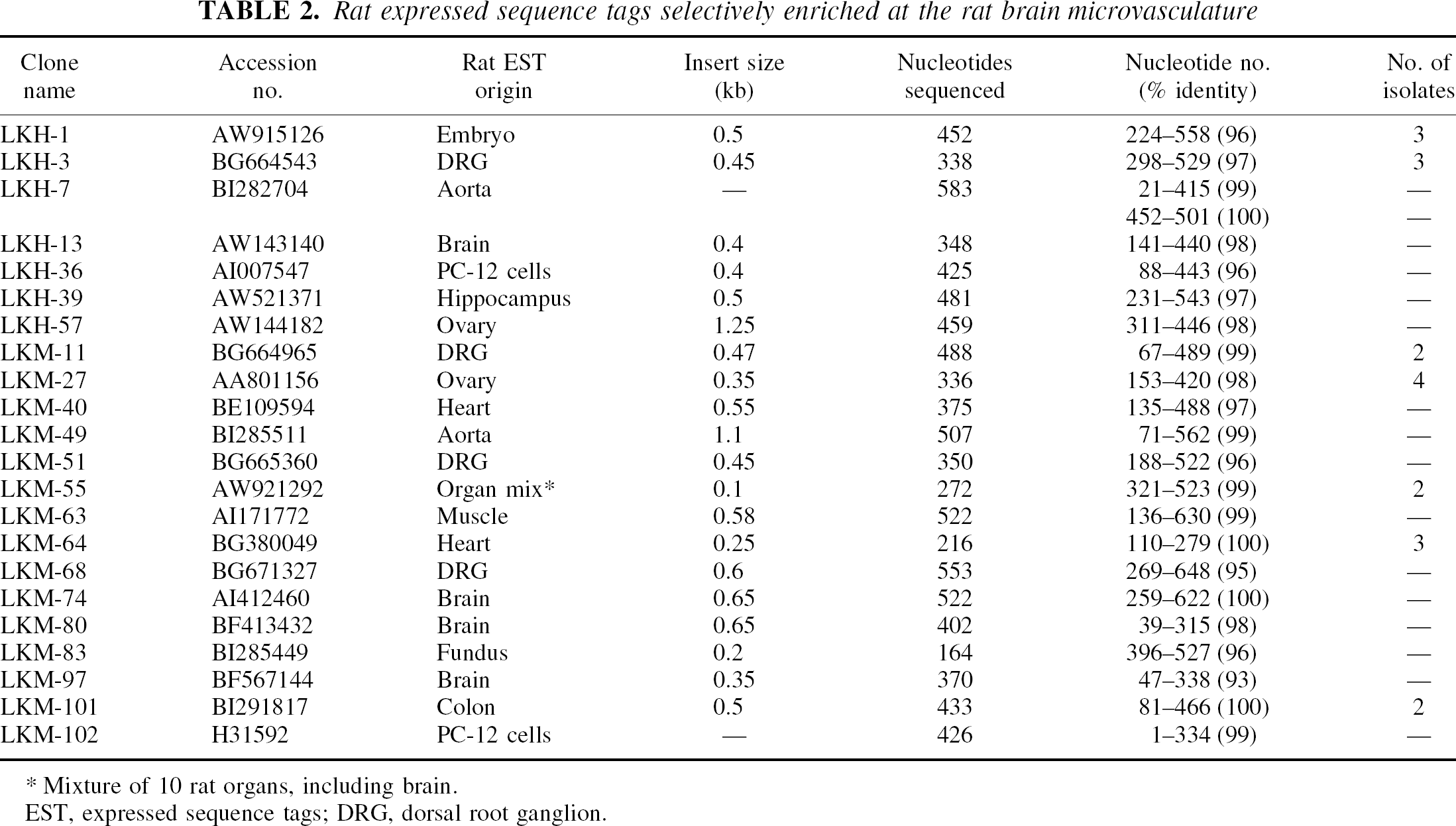
Mixture of 10 rat organs, including brain.
EST, expressed sequence tags; DRG, dorsal root ganglion.
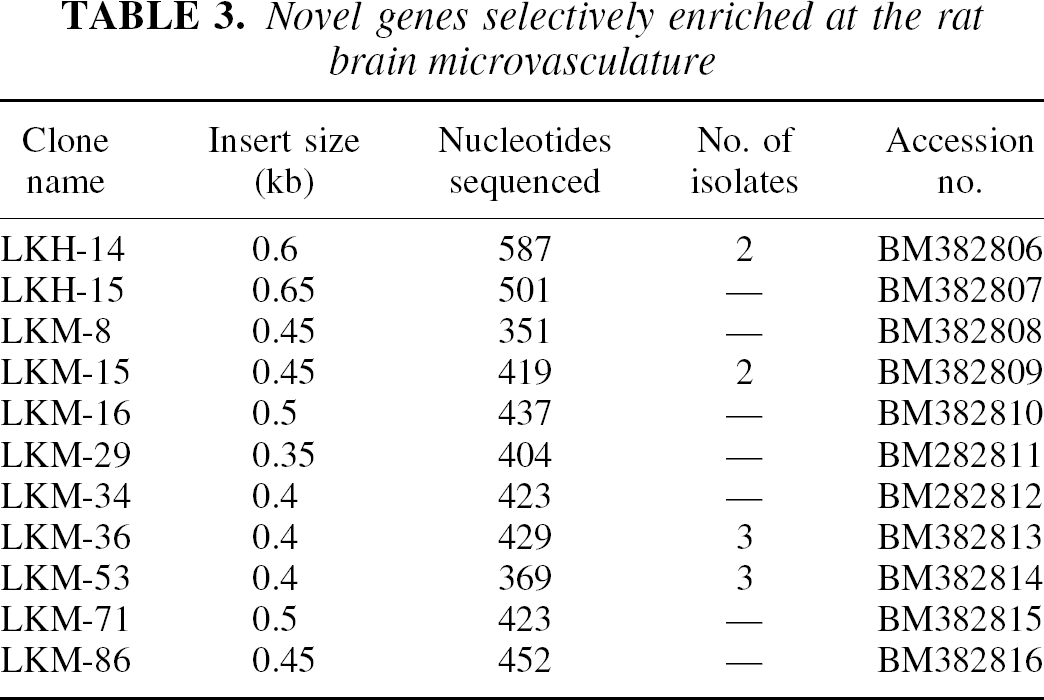
Novel genes selectively enriched at the rat brain microvasculature
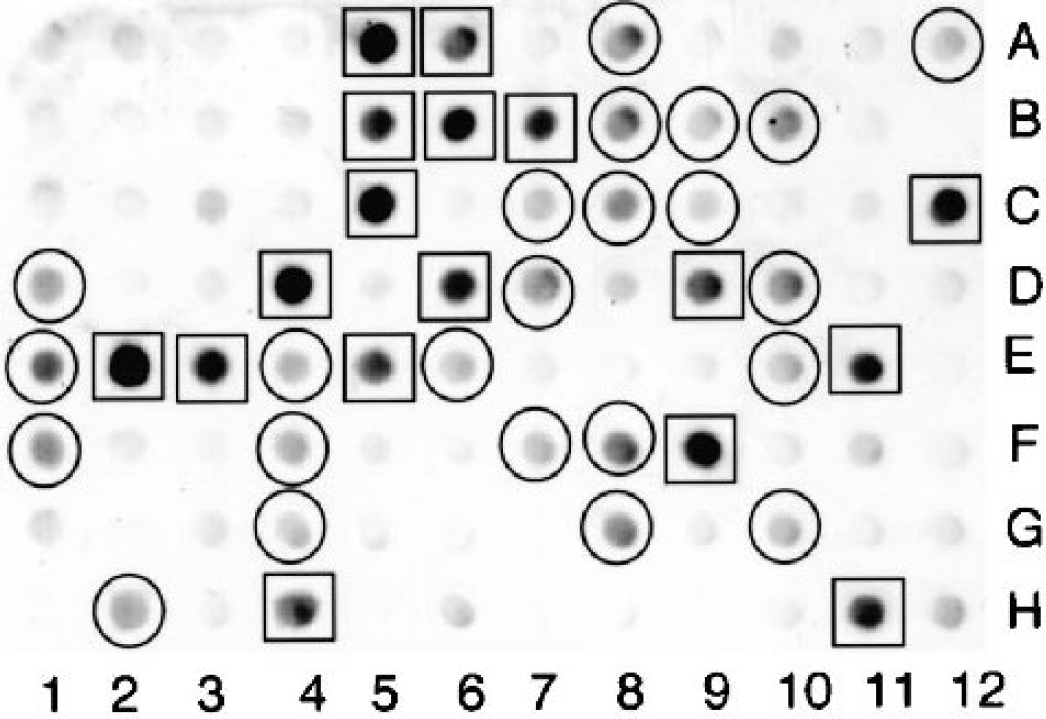

Ninety-six-well microarray analysis of subtracted tester (rat brain microvascular) cDNA library. Filters containing identical arrays of individual subtracted clones were probed with either [32P]-labeled subtracted tester cDNA (shown) or unsubtracted tester cDNA (not shown). Brain microvascular-specific genes enriched in the suppression subtractive hybridization process are identified as those clones that hybridize at high levels with the subtracted tester cDNA compared to the unsubtracted tester cDNA. The x-ray films were scanned and the hybridization density was quantified. This scanning densitometry was performed on the films obtained with both the subtracted tester cDNA (shown here) and the unsubtracted tester cDNA (not shown), and only those clones that had a ratio greater than 6 for the hybridization signal of the subtracted tester cDNA relative to the unsubstracted tester cDNA were selected for sequencing. Therefore, those clones shown in the figure with a relatively high hybridization signal, for example, 3C, were not selected for sequencing because this clone also gave a high signal with the unsubtracted tester cDNA, indicating a low enrichment at the blood–brain barrier. Those clones that had a hybridization signal greater than 160,000 are denoted by the square and were placed in the LKH group; those clones with an intermediate hybridization signal and integrated density between 60,000 and 160,000 arbitrary units are denoted by the circle and were placed in the LKM group (see Materials and Methods). LKH, LKM, liver (L)/kidney (K), subtracted with high (H) or medium (M) intensity, respectively.
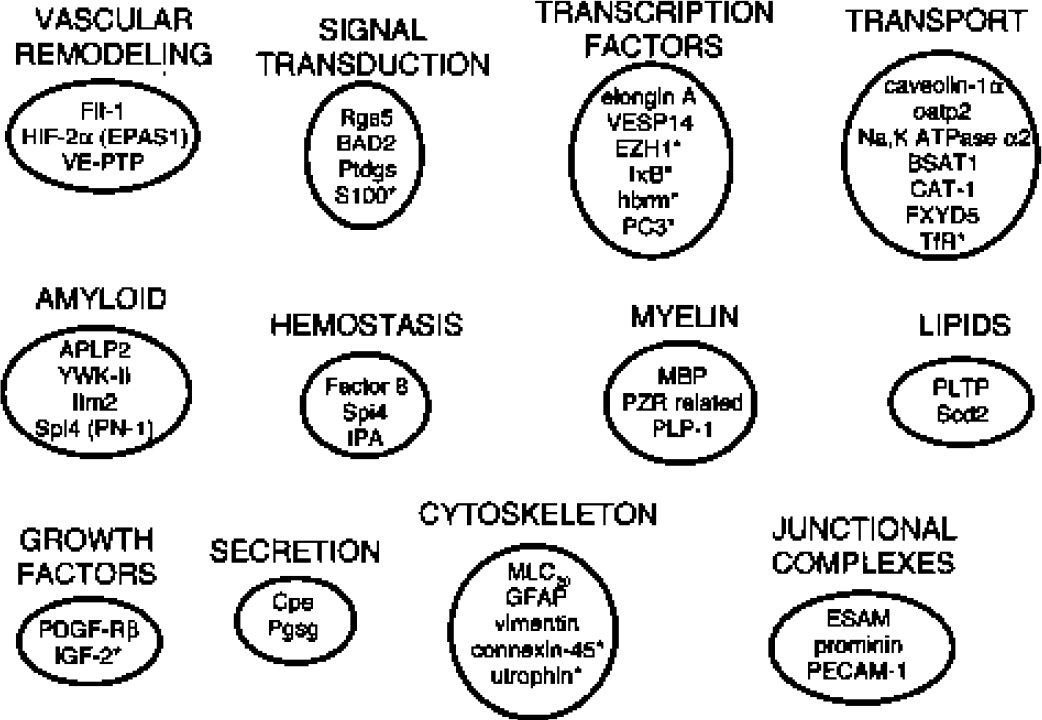
Clustering of rat brain microvascular genes by common function. The rat brain microvascular gene products detected in a prior study (Li et al., 2001) are noted by asterisks.
DISCUSSION
The present study uses the SSH subtraction cloning procedure to identify genes differentially expressed at the rat brain microvasculature. When the driver cDNA is derived from a pool of rat liver/kidney transcripts, nearly all of the genes cloned from the subtracted tester cDNA library are differentially expressed at the brain microvasculature (Li et al., 2001; Shusta et al., 2002a). The generation of driver cDNA from a mix of rat liver and kidney subtracts genes commonly expressed in all cells, including common endothelial genes, since kidney and liver are highly vascularized. In the prior study of this series (Li et al., 2001), Northern blotting was exhaustively performed on all cloned gene products, and this demonstrated that more than 90% of the gene products are differentially expressed in brain at the microvasculature. Some of the genes are also expressed in peripheral tissues of high vascularity such as heart or lung, suggesting that some of the genes are common to microvascular cells, including endothelium or pericytes. In situ hybridization was performed previously on selected genes to demonstrate gene expression in the endothelium (Li et al., 2001).
The genes identified in this study include known genes and novel genes. About half of the genes identified are genes of known function, and about half are either rat ESTs or novel genes not present in the rat EST database (Tables 1 to 3). The known genes that are differentially expressed at the rat BBB (Table 1) can be categorized on the basis of common function as shown in Fig. 2. Genes associated with brain vascular modeling or angiogenesis include Flt-1 (Table 1), which is the type 1 receptor for vascular endothelial growth factor (Breir et al., 1995). Clone LKM-31 is the rat homologue of mouse vascular endothelial receptor type protein tyrosine phosphatase (Table 1), which is differentially expressed at the brain microvasculature; this enzyme dephosphorylates the Tie-2 angiopoietin receptor (Fachinger et al., 1998), which plays a role in vascular remodeling (Evans et al., 2000). The hypoxia inducible factor-2α (HIF-2α), also known as the endothelial Per-Arnt-Sim domain protein-1, is enriched at the brain microvasculature (Table 1), and is a transcription factor that also participates in vascular remodeling (Peng et al., 2000). HIF-2α forms a complex with the von Hippel-Lindau tumor suppressor gene product, pVHL, which also forms a complex with elongin (Clifford et al., 2001), a transcription factor differentially expressed at the brain microvasculature (Table 1). Mutations in the pVHL gene are associated with the development of hemangioblastoma tumors of the brain vasculature (Clifford et al., 2001). HIF-2α also regulates the expression of the Glut1 glucose transporter (Maemura et al., 1999), which is upregulated in the vasculature of hemangioblastoma (Cornford et al., 1995), and is preferentially expressed in endothelial cells of the brain (Pardridge et al., 1990). LKM-17 is the rat homologue of the mouse platelet-derived growth factor receptor β subunit (Table 1), which plays a role in the recruitment of pericytes to endothelial cells to form capillaries (Hellstrom et al., 1999). In addition to elongin A and HIF-2α (Table 1), another transcription factor differentially expressed at the rat brain microvasculature is vascular endothelial cell specific protein-14, which was recently isolated from a rat liver cDNA library (AB031088). BLASTP analysis shows that this protein is 94% related to human ERG, a transcriptional factor related to the ets oncogene (P11308).
Several genes differentially expressed at the rat brain microvasculature are known factors involved in signal transduction (Fig. 2). Clone LKH-11 is the rat homologue of mouse G-protein signaling regulator-5 (Rgs5), which is differentially expressed at the rat brain microvasculature (Li et al., 2001), and is downregulated in stroke-prone spontaneously hypertensive rats (Kirsch et al., 2001). The BAD2 gene encodes a dual threonine/tyrosine phosphatase that is also expressed in brain cells and is upregulated in seizures and long-term potentiation (Qian et al., 1994). Prostaglandin D synthase synthesizes prostaglandin D2, which is a precursor to ligands for the peroxisome proliferator-activated receptor-γ, and is upregulated in endothelial cells subjected to shear stress (Taba et al., 2000).
Genes differentially expressed at the rat brain microvasculature that are involved in lipid metabolism include LKH-8 (Table 1), which is the rat homologue of mouse phospholipid transfer protein. This protein mediates the exchange of phospholipids and other amphipathic compounds between lipoproteins in the plasma compartment and the endothelium (Desrumaux et al., 1999). The Scd2 gene encodes stearoyl-CoA desaturase-2, an enzyme involved in fatty acid desaturation (Tabor et al., 1999). Myelin-related proteins are also expressed at the brain microvasculature, including myelin basic protein (Table 1). Prior work using in situ hybridization with cytocentrifuged isolated rat brain capillaries showed that the myelin basic protein transcript is produced within the endothelial cell (Li et al., 2001). Another myelin-related gene differentially expressed at the rat brain microvasculature is clone LKM-7, which is the rat homologue of human myelin protein zero-related (PZR) protein-1 (Table 1). PZR is a transmembrane protein that recruits tyrosine phosphatase SHP-2 to the plasma membrane (Zhao and Zhao, 1998). PZR is not only a structural protein of the myelin sheath but may also play a role in signal transduction at the brain microvasculature. Clone LKM-4 corresponds to rat proteolipid protein (Plp), which is also a major myelin constituent (Laman et al., 2001). The administration of either myelin basic protein or Plp-derived peptides to mice results in experimental autoimmune encephalomyelitis, which is characterized by demyelination that follows an initial inflammatory reaction at the microvasculature of brain (Laman et al., 2001). Plp is also called lipophilin and is expressed at the choroid plexus of brain (Gow, 1997), in addition to the microvasculature (Table 1). Immune reactions against myelinrelated proteins, such as myelin basic protein or Plp, may originate at the brain microvasculature, where these myelin-related proteins are also expressed.
Amyloid-related proteins are also differentially expressed at the rat brain microvasculature (Table 1), including amyloid precursor-like protein-2 (APLP2). APLP2, like the amyloid peptide precursor (APP) of Alzheimer disease (AD), is a transmembrane protein of unknown function, and APLP2 and APP are members of the same gene family (Sester et al., 2000). The abnormal processing of APP results in the formation of the amyloid plaque of AD, which deposits around blood vessels in brain (Miyakawa, 1997). Clone LKM-2 corresponds to rat YWK-II protein (Table 1), which is another transmembrane protein originally identified in human sperm (Yan et al., 1990), and is also a member of the APP/APLP2 gene family (Huang et al., 2000). The YWK-II protein is a G0-coupled receptor that is involved in signal transduction (Huang et al., 2000). Clone LKM-43 is the rat homologue of the mouse Spi4 gene, which expresses the serine protease inhibitor-4 (Vassalli et al., 1993). Spi4 is homologous to the rat glial-derived nexin, a neurite-promoting factor and a member of the serine protease inhibitor (SERPIN) family. Spi4, also known as protease nexin-1 (Docagne et al., 1999), is an inhibitor of the factor XIa coagulation protein (Knauer et al., 2000), and is a ligand of tissue plasminogen activator (Docagne et al., 1999), which is also expressed at the brain microvasculature (Table 1). Since factor XIa forms a complex with APP (Bornebroek et al., 2000), the Spi4/PN-1 may play a role in the development of amyloid formation of AD. A disease analogous to AD is familial Danish dementia, which is associated with progressive neurodegeneration, dementia and amyloid angiopathy. The amyloid plaque of familial Danish dementia is formed by a 4-KD peptide designated Adan. The Adan peptide is produced by a point mutation in the BRI gene, which encodes a transmembrane type II protein that is also called the Itm2b protein (Vidal et al., 2000), which has a 39% amino acid identity with the human Itm2a membrane protein (Pittois et al., 1999). Clone LKH-24 is the rat homologue of mouse Itm2a, which is differentially expressed at the brain microvasculature in both the rat (Table 1) and in humans (Shusta et al., 2002a). The differential expression of multiple amyloid-related genes at the brain microvasculature (Table 1) is consistent with electron microscopy showing that most amyloid plaques in Alzheimer disease arise from the microvasculature (Miyakawa, 1997). The finding that genes involved in either multiple sclerosis or AD are selectively expressed at the brain microvasculature is of interest, since both of these disorders are considered to involve pathogenetic mechanisms originating in brain parenchyma. It is possible that early pathogenetic events in either multiple sclerosis or AD take place at the microvasculature in brain.
Transport-related proteins that are differentially expressed at the rat brain microvasculature include clone LKH-20, which is the rat homologue of mouse caveolin-1α (Table 1). Caveolin-1 is a principal protein component of the caveolae organelle (Couet et al., 2001), and is enriched at the brain microvasculature (Ikezu et al., 1998). The CAT1 gene encodes the cationic amino acid transporter (Aulak et al., 1996), which mediates the BBB transport of basic amino acids such as arginine, lysine, and ornithine and is differentially expressed at the BBB in vivo (Smith and Stoll, 1998). Organic anions are transported by the organic anion transporting polypeptide type 2 (oatp2), which is clone LKM-73 (Table 1), and oatp2 has an amino acid homology with BBB specific anion transporter type 1 or BSAT1 (Li et al., 2001), which corresponds to clone LKH-17 (Table 1). BSAT1 is an orphan transporter member of the oatp2 family. The sodium potassium ATPase α2 subunit is differentially expressed at the rat brain microvasculature (Table 1), which confirms earlier Western blot studies (Zlokovic et al., 1993). The FXYD5 gene encodes the FXYD domain containing ion transport regulator-5, which is identical to the related ion channel, a protein of unknown function that is abundant in high transporting tissues (Sweadner and Rael, 2000).
The transport properties of the BBB are regulated by the endothelial tight junctions, and a number of genes involved in junctional complex or membrane structure are differentially expressed at the rat brain microvasculature including clone LKM-88 (Table 1). This clone is the rat homologue of mouse endothelial cell-selective adhesion molecule, a protein that colocalizes with cadherins (Hirata et al., 2001). The platelet/endothelial cell adhesion molecule (PECAM-1) is differentially expressed at the rat brain microvasculature (Table 1) and is also associated with cadherins (Shaw et al., 2001). PECAM-1, also known as CD31, is a marker of acute brain endothelial cell injury (Minagar et al., 2001), and exposure to inflammatory-related cytokines such tumor necrosis factor-α causes a redistribution of PECAM-1 and cadherins away from the lateral junctions of endothelial cells (Shaw et al., 2001). Prominin is another glycoprotein differentially expressed at the rat brain microvasculature (Table 1) as well as the brain ependymal epithelium, where it is associated with the microvilli on the apical border of the epithelial cell (Weigmann et al., 1997).
Cytoskeleton-related proteins are differentially expressed at the rat brain microvasculature including MLC20, which is the regulatory myosin light chain isoform C (Birukov et al., 2001). MLC20 is phosphorylated at a serine residue by the myosin light chain kinase, and this phosphorylation is essential for cell contractility in nonmuscle cells including endothelium (Birukov et al., 2001). Cytoskeletal microfilament proteins in astrocyte foot processes include both vimentin and glial fibrillary acidic protein (Table 1), and these proteins are specific to the astrocyte foot processes (Bertossi et al., 1999). Fragments of astrocyte foot processes remain adhered to the basement membrane surface of brain microvessels after the isolation procedure (White et al., 1981). Therefore, the finding of astrocyte foot process neurofilament transcripts in a brain microvessel preparation indicates that the astrocyte mRNA molecules are transported to the foot processes that envelop the capillary endothelial cell. Alternatively, the astrocyte-derived transcripts could originate from astrocyte impurities in the rat brain microvessel preparation. Microscopic examination of the brain capillaries, however, shows no astrocyte cell bodies in the preparation (Li et al., 2001).
Clones LKH-31 and LKM-33 are the rat homologues of the human or dog Factor 8 coagulation protein, respectively (Table 1). Little is known about the brain endothelial Factor 8 secretory processes, but the present studies shows that the Psgs secretory granule proteoglycan core protein precursor, which is a chondroitin sulfate proteoglycan (Oynebraten et al., 2000), is differentially expressed at the rat brain microvasculature (Table 1). In addition, carboxypeptidase E (CpE), which may be involved in the secretory processing of the cell (Coo et al., 1997), is also expressed at the rat brain microvasculature (Table 1). There presently is little information on the secretory role played in the brain by the microvasculature.
Rat ESTs, as well as novel genes not found in EST databases, are also differentially expressed at the rat brain microvasculature (Tables 2 and 3) in addition to the genes of known function (Table 1). The availability of these partial EST or novel clones will allow for screening of full-length cDNAs from rat brain capillary cDNA libraries as the first step toward elucidating the functional role that these unknown genes play in the regulation of the rat brain microvasculature. The rat BBB genomics described here and in Li et al. (2001) can be correlated with either a human BBB genomics program (Shusta et al., 2002a), as well as with a BBB proteomics analysis (Shusta et al., 2002b). The pattern of BBB gene expression in control states, such as that described here, can be correlated with an analysis of brain microvascular gene expression in pathologic conditions. These molecular biologic approaches to brain microvascular function can accelerate the discovery of genes that are responsible for the maintenance of the microvasculature in the brain in health and disease.
