Abstract
Transforming growth factor-β1 (TGF-β1) plays a central role in the response of the brain to different types of injury. Increased TGF-β1 has been found in the central nervous system of patients with acute or chronic disorders such as stroke or Alzheimer disease. To further define the molecular targets of TGF-β1 in cerebral tissues, a selection of high-density cDNA arrays was used to characterize the mRNA expression profile of 7,000 genes in transgenic mice overexpressing TGF-β1 from astrocytes as compared with the wild-type line. Selected findings were further evaluated by reverse transcription-polymerase chain reactions from independent transgenic and wild-type mice. Furthermore, the expression pattern of seven selected genes such as Delta-1, CRADD, PRSC-1, PAI-1, Apo-1/Fas, CTS-B, and TβR-II were confirmed in either cultured cortical neurons or astrocytes following TGF-β1 treatment. The authors' observations enlarge the repertoire of known TGF-β1–modulated genes and their possible involvement in neurodegenerative processes.
Many diseases of the central nervous system are initiated by a large cascade of events that lead to neuronal cell death, which occurs through necrotic and apoptotic mechanisms. Although considerable interest exists in developing effective methods to modulate these two fatal pathways, most of the drugs tested so far have shown either no significant protective activity or sufficient harmful side effects so as to be inappropriate for use in patients (del Zoppo et al., 1997). In contrast, the use of neurotrophic factors could represent an interesting alternative therapeutic strategy against both acute and chronic brain neurodegenerative processes. Based on these observations, there is considerable interest to further understand the molecular mechanisms through which cytokines may modulate many neuropathologic processes.
Among these cytokines, transforming growth factor-β1 (TGF-β1) is a prototype of various multifunctional cytokines and has been previously characterized as an injury-related molecule in both rodents and man (Flanders et al., 1998; Krieglstein and Krieglstein, 1997). Transforming growth factor-β has been characterized for its pleiotropic activity and its ability to control cellular functions that underlie embryonic development and tissue homeostasis. The basic signaling machinery consists of two serine/threonine kinase receptors and a family of transcription factors: the Smad family (Piek et al., 1999; Massagué, 2000). Such complexes recognizing the TGF-β–inducible element CAGAC contribute to the transcriptional activity of this cytokine. Smad proteins are often found to be associated with other transcription factors such as CBP/p300 (Massagué and Wotton, 2000), Sp1 (Moustakas and Kardassis, 1998; Brodin et al., 2000; Pardali et al., 2000; Feng et al., 2000; Datta et al., 2000), and AP-1 (i.e., c-fos/c-jun complex) (Zhang et al., 1998; Wong et al., 1999; Liberati et al., 1999). However, the diversity of effects displayed by this simple system remains largely unknown.
In vivo, several studies have reported a marked increase in TGF-β expression after an ischemic insult to the brain (Wiessne et al., 1993; Lehrmann et al., 1995; Krupinski et al., 1996). In vitro, we have demonstrated that TGF-β1 exerts a neuroprotective activity against NMDA-induced neuronal death via an upregulation of the type I plasminogen activator inhibitor (PAI-1) in astrocytes (Docagne et al., 1999). PAI-1 is markedly induced by TGF-β1 in many cell models (Matrisian and Hogan, 1990; Westerhausen et al., 1991; McCaffrey, 2000), and multiple TGF-β–responsive elements have been described in its promoter (Dennler et al., 1998). It has also been shown that an overproduction of TGF-β1 is associated with the pathology of Alzheimer disease and that 18-month-old transgenic mice overexpressing TGF-β1 in astrocytes elicit enhanced amyloid-β (Aβ) deposition in cerebral blood vessels and meninges (Wyss-Coray et al., 1997). In addition to this proamyloidogenic role of TGF-β1, the same group demonstrated, in a recent report, that 12- to 15-month-old double transgenic mice expressing TGF-β1 and human amyloid-β precursor protein (hAPP) display reduced numbers of parenchymal plaques (Wyss-Coray et al., 2001). Despite unknown Aβ-promoting mechanisms in these transgenic models, these data suggest that TGF-β1 is an important regulator of Aβ deposition in vivo.
Many genes have been reported to be modulated after TGF-β exposure, such as PAI-1 (Westerhausen et al., 1991; Keeton et al., 1991; Riccio et al., 1992), TGF-β1 (Van Obberghen-Schilling et al., 1988; Kim et al., 1989), the cyclin-dependent kinases inhibitors p21 and p15 (Elbendary et al., 1994; Li et al., 1995; Datto et al., 1997), fibronectin (Ignotz et al., 1987) and collagen a2(I) (Inagaki et al., 1994). Based on the pleiotropic activities of this cytokine, these genes represent probably a minor fraction of all genes induced or repressed following a TGF-β signal. Identification of these genes and the ultimate characterization of their functions are crucial to understand the pathology of brain diseases in which TGF-β is believed to play a critical role.
To further investigate the molecular events through which TGF-β1 influences the evolution of threatened brain tissue, we used cDNA arrays to analyze the expression pattern of 7,104 genes from brains of both wild-type (WT) and transgenic (Tg) mice overexpressing TGF-β1 in astrocytes. These mice express a bioactive form of TGF-β1 by astrocytes and were previously characterized and used as a transgenic model of amyloid deposition (Wyss-Coray et al., 1995, 1997, 2001). Some selected TGF-β1–regulated genes potentially related in neurodegenerative diseases were then further analyzed in TGF-β1–treated cultures of cortical neurons and astrocytes.
MATERIALS AND METHODS
The reverse transcriptase system kit and the polymerase chain reaction kit were obtained from Promega (Paris, France). Dulbecco's Modified Eagle's medium (DMEM), cytosine arabinoside, horse serum and fetal bovine serum, poly-
Cell cultures
Cortical cultures of astrocytes
Cortical cultures of astrocytes were prepared from 1- to 3-day-old postnatal mice (Rose et al., 1993). Cerebral cortices were microdissected and incubated in DMEM containing 2 mmol/L glutamine supplemented with 10% fetal bovine (FCS) and 10% horse serum (HS). Cultures were kept at 37°C in a humidified 5% CO2-containing atmosphere. After 3 days in vitro (DIV), astrocytes were vigorously washed 3 times with phosphate-buffered saline, and this process was repeated every 3 days to prevent contamination of cell cultures by microglia. Purity of culture has been estimated under our conditions, where 99% of cells were glial fibrillary acidic protein (GFAP)-positive. Experiments were performed on confluent cultured cortical astrocytes after 10 DIV.
Cultures of cortical neurons
Cultures of cortical neurons were prepared from 14- to 15-day-old mice embryos as previously described (Rose et al., 1993), plated in 2-cm2 dishes previously coated with poly-
Transgenic mice
To determine TGF-β1-related cerebral gene expression, we used brains from Tg mice overexpressing TGF-β1 in astrocytes from the GFAP promoter (line T65) generated by Lennart Mucke's laboratory similarly to the previously described low-expressing GFAP-TGF-β1 transgenic lines, T64 and T115 (Wyss-Coray et al., 1995). These mice express large amounts of the transgene associated with enhanced expression of endogenous TGF-β1 as determined either by RT-PCR or by immunoblotting (data not shown).
Gene expression profiling
Human brain cDNA arrays
Human brain cDNA arrays were prepared with a set of 7,104 cDNA clones selected from a normalized infant brain library (library 1NIB, [Soares et al., 1994]). The 3' and/or 5' ends of these 7,104 cDNA clones have been previously sequenced (Houlgatte et al., 1995). The sequences, registered in Genbank (http://www.ncbi.nlm.nih.gov/Genbank), have been compared with those in public databases, permitting a tentative identification of the corresponding gene transcripts. Each insert was amplified by PCR with vector-specific primers and spotted at medium density (25 PCR products/cm2) onto nylon membranes (Hybond-N+, Amersham Pharmacia Biotech U.K., Ltd., Little Chalfont, Buckinghamshire, U.K.) by a robot (Flexis, Perkin Elmer), as previously described (Piétu et al., 1996; Decraene et al., 1999). A set of 137 known genes and 84 negative and positive controls were added to the filters. The entire collection was spotted on a set of 4 filters.
Preparation and labeling of complex cDNA targets
Preparation and labeling of complex cDNA targets were based on [α-33P]-labeled complex cDNA targets which were synthesized by reverse transcription of 500-ng poly(A)+ brain mRNA extracted from Tg and WT lines (3 animals for each group). The reaction was performed with the SuperScript Preamplification System (Invitrogen, SARL, Cergy Pontoise, France) in the presence of 50-μCi [α-33P]dATP, 3,000 Ci/mmol (Amersham Pharmacia Biotech) as previously described (Decraene et al., 1999).
Hybridization and signal quantitation
Hybridization and signal quantitation were performed under nonstringent conditions to allow optimal hybridization between the human cDNA probes and the mouse cDNA targets. Membranes were prehybridized at 68°C for 30 minutes in an ExpressHyb solution (Clontech, Palo Alto, CA, U.S.A.), hybridized for 2 hours in the same solution to which the radiolabeled complex cDNA target was added, then washed twice for 30 minutes at 25°C in standard saline citrate (SSC)/0.1% sodium dodecyl sulfate (SDS), and twice for 30 minutes at 25°C in SSC/0.1% SDS. The washed filters were exposed to phosphorus screens (Amersham Biosciences, Sunnyvale, CA, U.S.A.) for 16 hours.
Under such conditions, among the 7,104 cDNA clones, 4,651 and 4,525 clones were significantly detected in WT or Tg brain tissues, respectively. Differential gene expression was determined by calculating the ratio of the normalized hybridization signal intensity for each clone obtained with the WT brain cDNA target to that obtained with the Tg brain cDNA target. Genes were considered to be differentially expressed in Tg brain compared with WT brain if the ratio between the corresponding hybridization intensity values was greater than 1.67 (upregulated clones) or less than 0.60 (downregulated clones), the limits of confidence for the method. Applying these selection criteria, the vast majority of genes (95%) showed relatively little change in mRNA expression levels.
Image acquisition was performed on a PhosphorImager (Amersham Biosciences). Identification and quantitation of the spots were performed by the XdotsReader software (Cose, Le Bourget, France). After subtraction of the background, normalized intensity values for each clone in each experimental condition were determined as the average of the intensity of the hybridization signal for one clone divided by the intensity obtained for all clones deposited on the filters. Duplicate filters were hybridized with an initial preparation of each target; the entire set of hybridization was then repeated with novel target preparations, so that for each clone/target combination, four hybridization data points were collected. The four values were compared and validated if at least three of the four values were similar (SD < 25%). The final value assigned to each clone was the average of the validated values.
Semiquantitative reverse transcriptase-polymerase chain reaction
Total RNAs (Figs. 2 to 4) or polyadenylated mRNAs (Fig. 1), as indicated in the Figures legends, were prepared from cerebral cortex, cultured neurons, and astrocytes by either the phenol/chloroform extraction method using the RNAXEL extraction kit (Eurobio, Paris, France) or the RNAeasy system (Qiagen S.A., Courtaboeuf, France). One microgram of total RNA was transcribed into cDNA via poly-dT oligonucleotides. An aliquot of the cDNAs so produced was amplified using sense and antisense primers for each selected gene and β-actin respectively. For semiquantitative experiments, an aliquot of cDNA libraries was amplified by PCR with specific oligonucleotides (Table 1) or for β-actin (25 cycles of PCR were chosen, corresponding to the 50%-of-the-saturation curve of the PCR products). For each chosen gene, PCR cycles indicated in Table 1 correspond to the 50%-of-the-saturation curve of the PCR product. Amplified products were then separated by agarose gel electrophoresis and visualized by ethidium bromide staining. PCR experiments were performed only if the expression pattern obtained for the housekeeping gene, β-actin, failed to show any significant modification between different samples. We also verified that quantitation of ethidium bromide-stained PCR bands was accurate. To this end, we amplified β-actin mRNA by using increasing amounts of cDNA and found a direct correlation between data obtained from densitometry analysis and amounts of cDNA used.
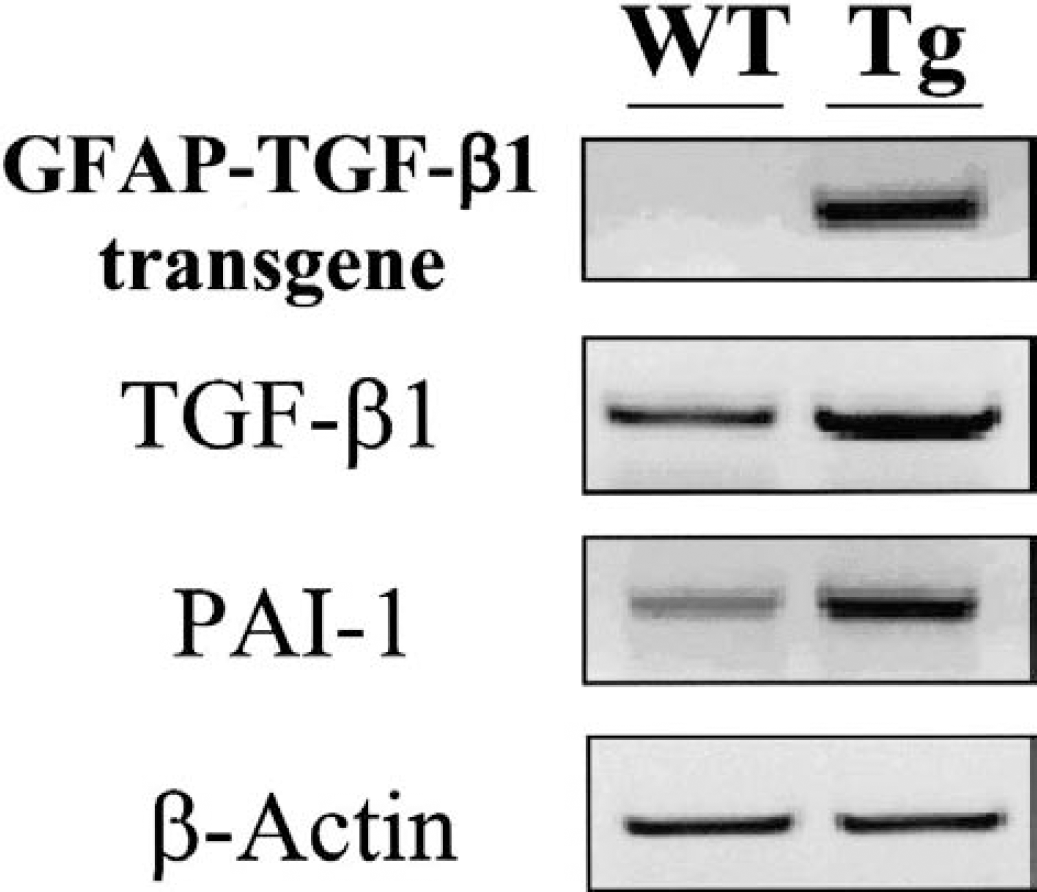
Characterization of GFAP-TGF-β1 transgenic mice. Polyadenylated mRNAs encoding for either the transgene, transcripts for TGF-β1 and type I plasminogen activator inhibitor (PAI-1) in addition to the housekeeping gene β-actin, harvested from cortices of both transgenic (Tg) and wild-type (WT) mice, were reverse transcribed as described in the methods section and amplified by using corresponding oligonucleotides prior to agarose gel electrophoresis and ethidium bromide staining. GFAP, glial fibrillary acidic protein; TGF-β1, transforming growth factor-β1.
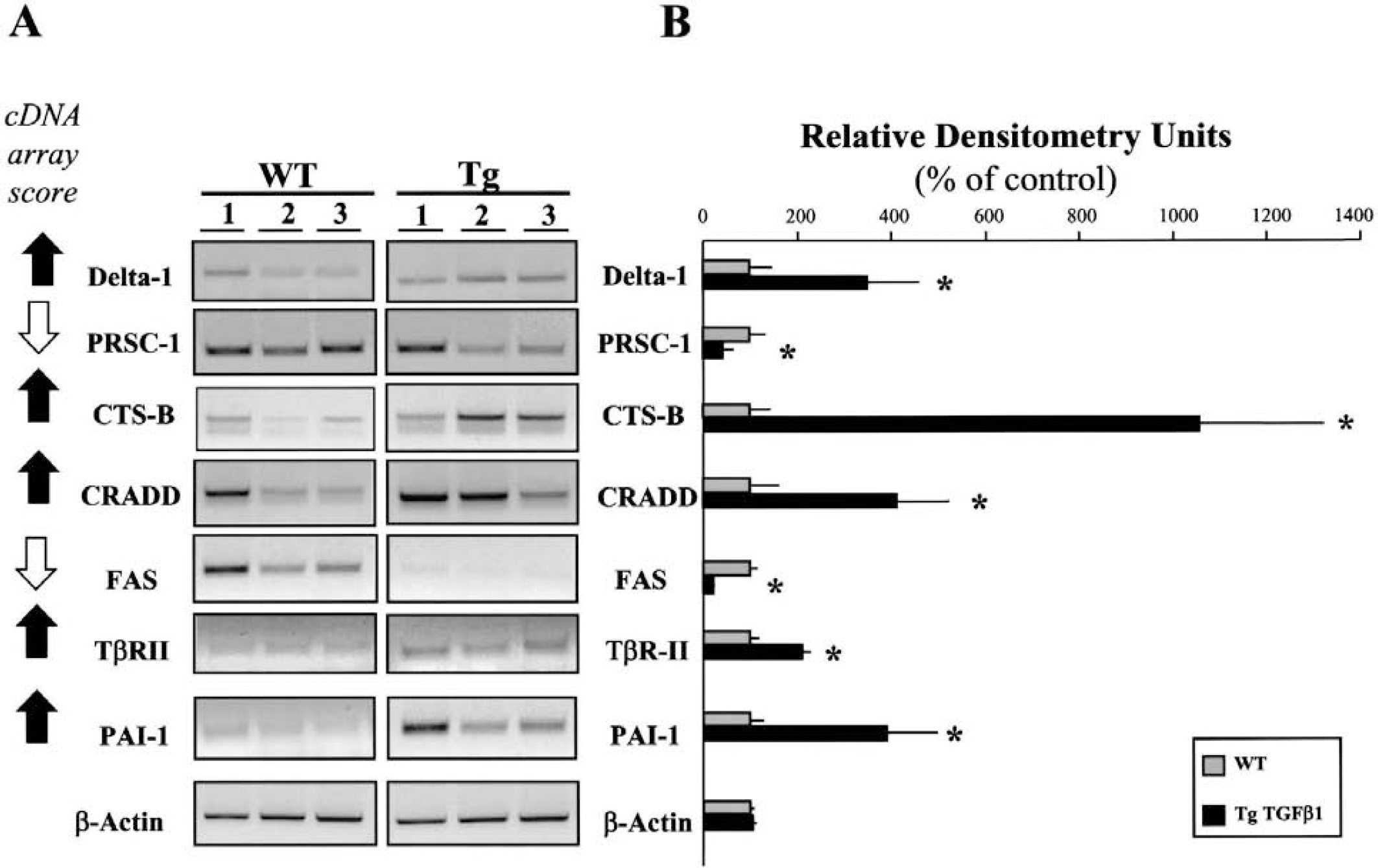
Reverse transcription-polymerase chain reaction (RT-PCR) analysis of cDNA array-selected genes from the brains of both wild-type (WT) and GFAP-TGF-β transgenic (Tg) mice.
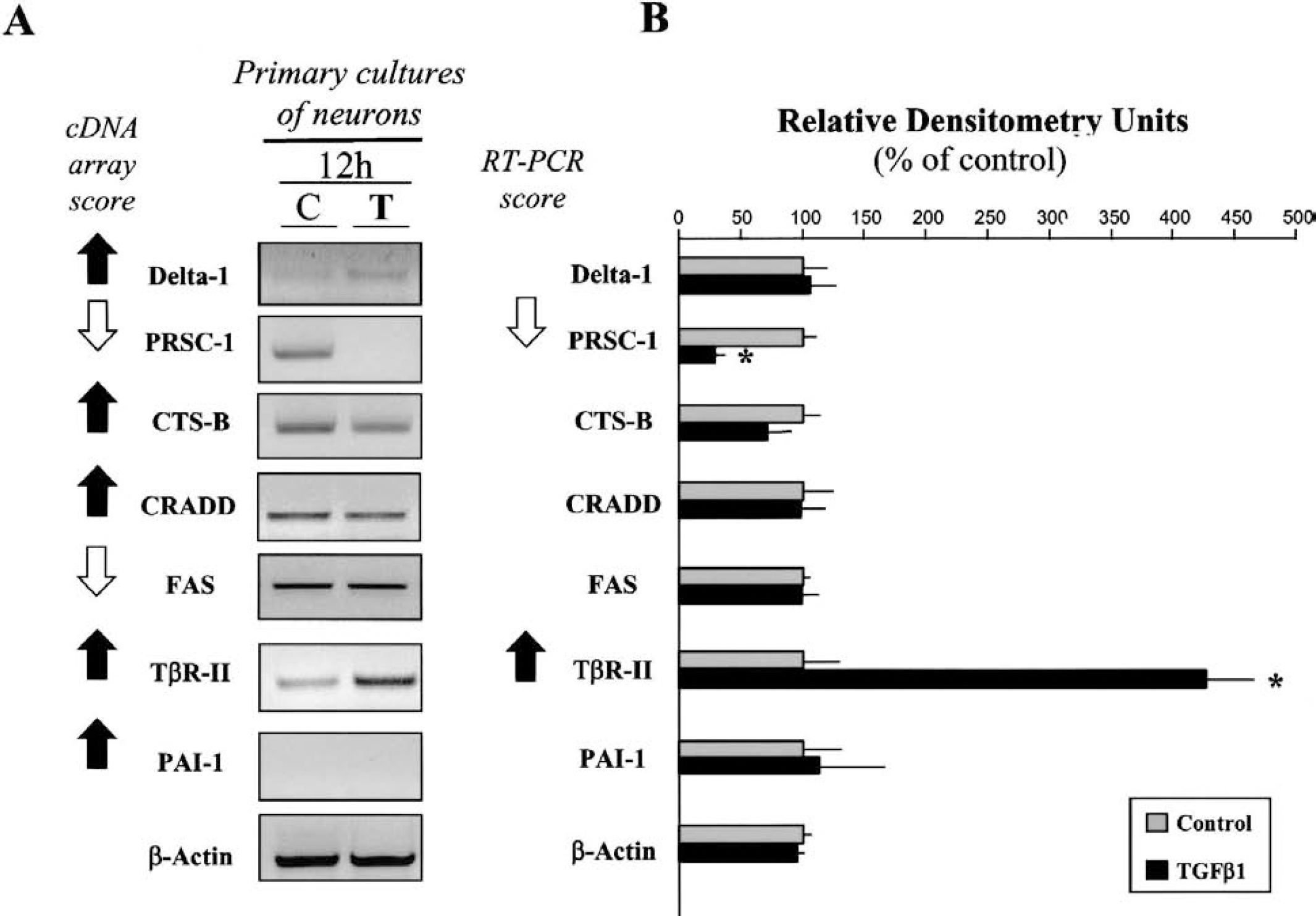
Reverse transcription-polymerase chain reaction (RT-PCR) analysis of cDNA array-selected genes from primary cultures of cortical neurons following exposure to transforming growth factor-β1 (TGF-β1).
Oligonucleotides and amplification conditions used for reverse transcription-polymerase chain reaction analysis
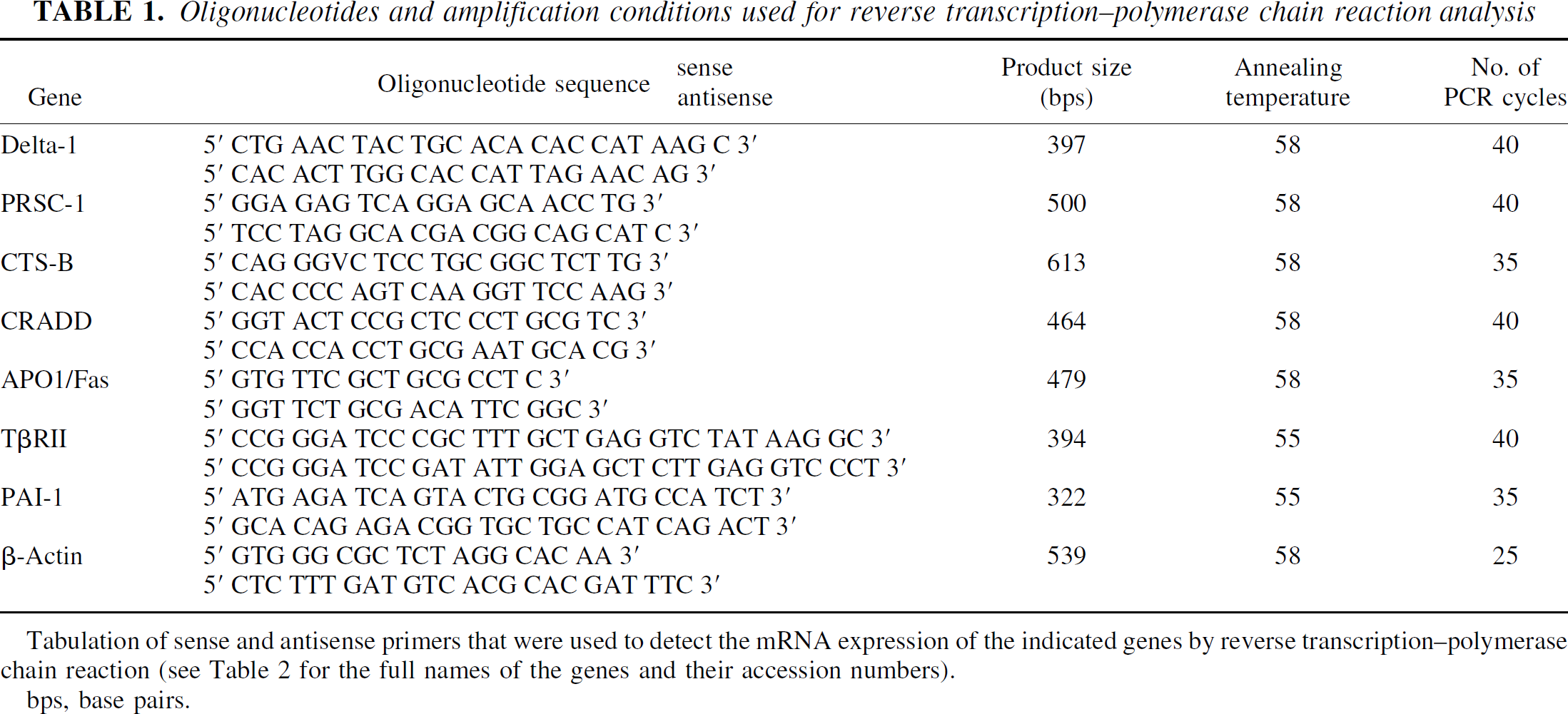
Tabulation of sense and antisense primers that were used to detect the mRNA expression of the indicated genes by reverse transcription-polymerase chain reaction (see Table 2 for the full names of the genes and their accession numbers).
bps, base pairs.
Transforming growth factor-β1–regulated genes identified by cDNA array and selected for their link with neurodegenerative disorders

Genes selected from a Medline analysis that shows a high score for both keywords (“TGF-β” and “brain”) are presented with their cDNA probe access number, their full names and standard abbreviation, their GeneBank access number, their function, and the normalized ratio, obtained by cDNA array.
Genes selected for further reverse transcription-polymerase chain reaction analyses.
TGF-β, transforming growth factor-β.
RESULTS
Identification of brain-derived genes differentially expressed between wild-type and glial fibrillary acidic protein-transforming growth factor-β1 transgenic mice
Mice used for this study were previously characterized as expressing the GFAP-TGF-β1 transgene (Wyss-Coray et al., 1995) and their corresponding WT line. In the Tg line, transgene-derived bioactive TGF-β1 is produced by astrocytes from birth based on a transcriptional control by the modified astrocytic GFAP promoter. To assess the transgenic properties of our murine line, we performed semiquantitative RT-PCR for TGF-β1 transgene, endogenous TGF-β1 mRNA, and a gene previously characterized as inducible by TGF-β1, PAI-1 (Westerhausen et al., 1991; Dennler et al., 1998) from both Tg and WT lines (Fig. 1). All our PCR reactions showed the same level of expression of chosen housekeeping gene (β-actin). Each set of oligonucleotides gave products of the expected size for the GFAP-TGF-β1 transgene and for PAI-1. Specificity of PCR products has been previously demonstrated by sequencing (Wyss-Coray et al., 1997; Docagne et al., 1999). Assessment of cerebral TGF-β1 and PAI-1 confirmed the robust association between TGF-β1 expression and cerebral production of PAI-1 in Tg animals when compared with their non-Tg controls. These data indicate that GFAP-TGF-β1 Tg mice are an appropriate model to determine the pattern of genes that are under the control of TGF-β1 in cerebral tissues.
We used the cDNA arrays corresponding to 7,104 cDNA clones to identify genes differentially expressed from the brain tissues of WT mice when compared with Tg mice expressing TGF-β1. Spots with variable intensities were observed. The mean differences in hybridization signal intensity reflect a difference in the steady-state level of transcription. To avoid a misinterpretation due to lowly expressed genes, only those clones that gave a hybridization signal two-fold greater than background were taken into consideration. Under such conditions, there were 239 genes that displayed a modulated expression; 174 were identified as upregulated in Tg versus WT mice, whereas 65 were downregulated. The results of expression profiling are detailed in Table A and Table B at the web site on the following address: http://www.unicaen.fr/unicaen/ufr/medecine/neurosciences/cdnatgfbeta1.html.
Partial 5' and 3' sequences of the 239 clones were compared with all the sequences present in the UniGene database (http://www.ncbi.nlm.nih.gov/UniGene/). From the 174 genes found to be upregulated in Tg (Table A on the web site), a sequence homology search revealed that 59 of them correspond to sequences that are related to human genes that encode proteins with a known function. Thirty-seven clones were found to be similar to genes encoding proteins without known function, and 34 clones corresponded to poorly characterized genes that demonstrated either no or a weak similarity with sequences registered in the databases.
From the 65 genes found to be downregulated in Tg (Table B on the web site), 23 corresponded to human genes encoding proteins with a known function. In addition, 12 clones were found to be identical with sequences of unknown function. Finally, 12 clones corresponded to poorly characterized genes with no similarity with the sequences found in the databases.
Accordingly, 28 clones corresponding to 37 differentially expressed genes encode proteins that have been reported to play a role in the physiopathology of various CNS disease states (Table 2). Among these genes, some of them are directly related with the TGF-β1 signaling pathway (TGF-β type II receptor, the FK506-binding protein, and PAI-1), others are associated with cell death in both necrotic and apoptotic cascades (CRADD, Apo-1/Fas, requiem HREQ), while yet others are characteristic of neurologic disorders (Delta-1, Notch 3, Huntingtin, cathepsin B, putative cysteine protease: PRSC-1). Although the majority of these genes are upregulated in Tg mice when compared with the WT line, Apo-1/Fas and the cysteine protease PRSC-1 are downregulated with increased TGF-β1 expression.
To confirm the data obtained with the cDNA arrays, we assessed the expression of a set of seven genes by RT-PCR from WT and Tg mice. These seven genes were chosen for their involvement in neurodegenerative diseases including acute (cerebral ischemia, trauma) and progressive (CADASIL [cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy], Alzheimer disease) disorders (Peter and Krammer, 1998; Pinkoski and Green, 1999; Schneider and Tschopp, 2000; Vivien and Buisson, 2000; Mattson, 2000; Hughes and Olson, 2001). Oligonucleotides primers were designed for several genes, such as Delta-1, CRADD, PRSC-1, PAI-1, Apo-1/Fas, CTS-B, and TβR-II (Table 1). RT-PCR experiments were performed on total mRNA harvested from three WT and three Tg mice. For the seven genes, mRNA expression was estimated by densitometric analysis of corresponding PCR products following ethidium bromide staining of three independent experiments performed from three animals (N = 3; n = 3). All expression patterns matched with the data obtained from the cDNA array analysis (Figs. 2A and 2B). Based on these results, we are able to confirm the predictive power of the cDNA array screening experiments in our system.
Expression pattern of differentially expressed genes in cultured cortical neurons and astrocytes after exposure to transforming growth factor-β1
To further investigate the expression pattern of these seven selected genes, we have performed semiquantitative RT-PCR experiments from primary cultures of cortical astrocytes and neurons exposed to treatment with TGF-β1 (1 ng/mL) for 12 hours. In these models, we have studied the expression of the mRNA encoding for Delta-1, CRADD, PRSC-1, PAI-1, Apo-1/Fas, CTS-B, and TβR-II (see Table 1). Transcriptional expression was assessed by the densitometric analysis of corresponding PCR products following ethidium bromide staining in three independent experiments (n = 3). We showed that the mRNAs of all selected genes were expressed in cultured astrocytes. As shown in Fig. 3, the addition of TGF-β1 promoted the transcription of the mRNAs encoding for Delta-1, CRADD, PAI-1, and CTS-B in cultured astrocytes as illustrated by the cDNA array analysis. Similarly, TGF-β1 reduced the expression of the mRNA for PRSC-1 and Apo-1/Fas in cultured astrocytes, whereas the expression of TβR-II mRNA was not affected.
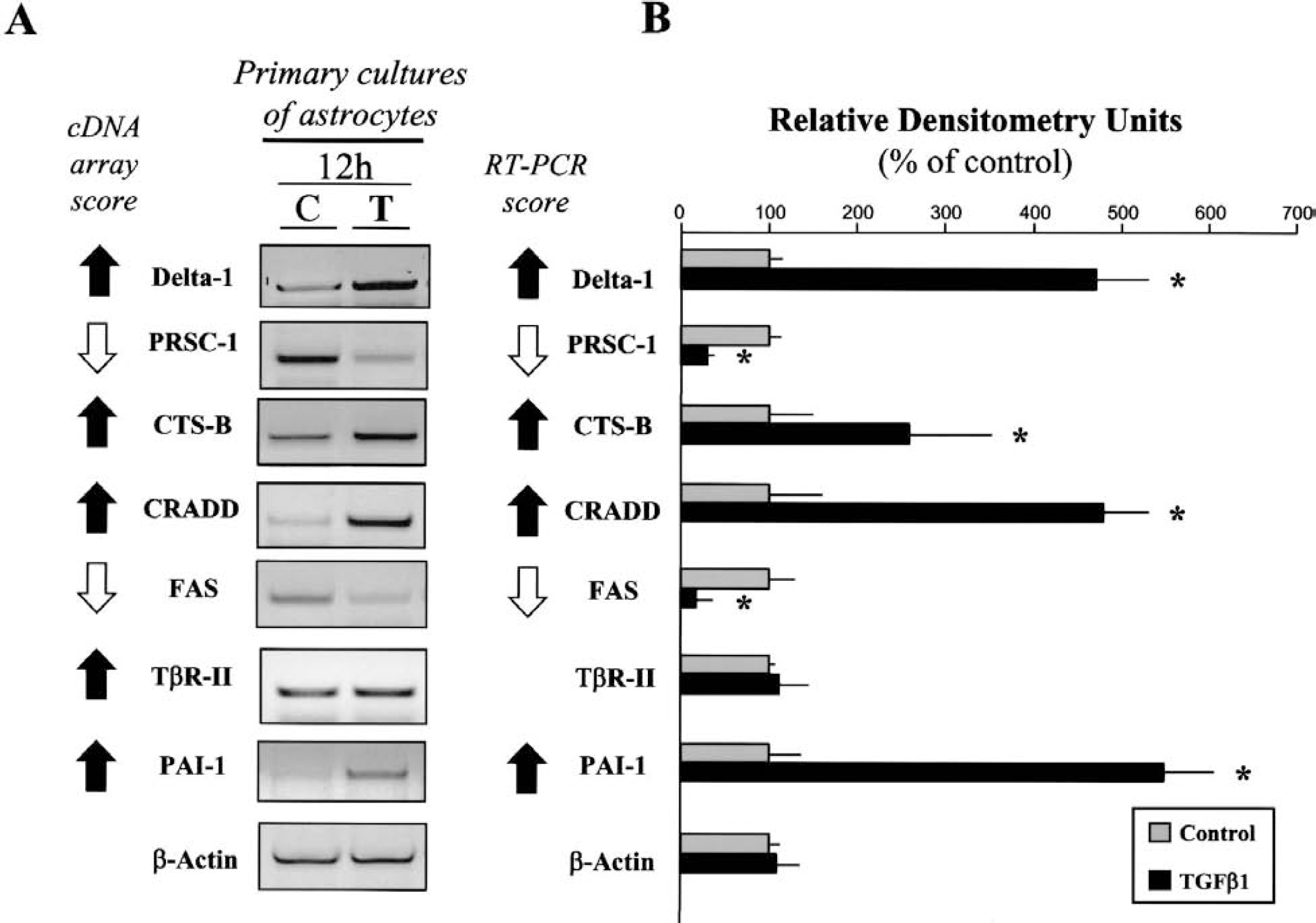
Reverse transcription-polymerase chain reaction (RT-PCR) analysis of cDNA array-selected genes from primary cultures of cortical astrocytes following exposure to transforming growth factor-β1 (TGF-β1).
In contrast, the application of TGF-β1 to cultured neurons failed to influence the expression pattern of most of the genes tested, such as Delta-1, CRADD, PAI-1, Apo-1/Fas and CTS-B (Figs. 4A and 4B). However, the expression of the mRNA for PRSC-1 was reduced. As previously shown (Docagne et al., 1999), cultured cortical neurons do not exhibit mRNA that encodes for PAI-1. It is interesting to note that, although TGF-β1 did not influence the expression pattern of the mRNA for TβR-II in cultured astrocytes, it clearly induced an overexpression of the mRNA for TβR-II in cultured neurons.
DISCUSSION
Based on the results obtained in this study, we suggest that brain-derived TGF-β1 within the CNS would be a key element in the physiopathology of a number of disease processes. Our data evidence that TGF-β1 modulates the transcriptional activity of genes directly involved in the apoptotic cascade, such as Apo-1/Fas, CRADD, and Requiem. Apo-1/Fas, also called the CD95 receptor or tumor necrosis factor-related receptor (TNF-α R), is critically involved in the induction of apoptosis (Peter and Krammer, 1998; Pinkoski and Green, 1999; Schneider and Tschopp, 2000). An ischemic insult to the brain evokes a cellular stress, which involves the expression and the activation of the Apo-1/Fas receptor pathway (Matsuyama et al., 1994, 1995; Martin-Villalba et al., 1999; Rosenbaum et al., 2000). Fas-induced apoptosis involves the recruitment of procaspase-8 (cysteine aspartate-specific proteinase) through death domain-containing adaptor/signaling proteins such as CRADD or FADD to form the death-inducing signaling complex (DISC) (Boldin et al., 1996; Muzio et al., 1998). Accordingly, CRADD could play a role in the regulation of apoptosis in mammalian cells. The DISC formation leads to the proteolytic activation of procaspase-8 into caspase-8, which is released finally from the DISC into the cytoplasm (Muzio et al., 1996). Active caspase-8 cleaves various proteins including procaspase-3, and results in its activation and the completion of the cell death program (Stennicke et al., 1998). Interestingly, although the function of TGF-β1 is not so far characterized, we have shown that the overexpression of TGF-β1 in brain tissue leads to the activation of the transcription of a putative cysteine protease, PRSC-1.
The overexpression of PAI-1 observed in the GFAP-TGF-β1 transgenic mice is instructive. We have recently demonstrated that an upregulation of the expression of this serine protease inhibitor (PAI-1) in astrocytes mediates the neuroprotective activity of TGF-β1 against a variety of excitotoxic challenges in cultures cortical neurons (Wiessner et al., 1993; Buisson et al., 1998). These data are reinforced by the demonstration that blocking TGF-β signaling in cerebral tissue, by using a chimerical soluble TGF-β type II receptor, induced a marked increase in the ischemic lesion that follows the transient occlusion of the middle cerebral artery in rats (Ruocco et al., 1999). Modulation of the expression of genes such as Notch-3 or Delta-1 is also of considerable interest in neuropathologic processes. The fate of neurons is thought to be determined by a series of local cell-cell interactions. Many reports would suggest that Notch-mediated intercellular signaling is involved in the events that control the spatial and temporal aspects of cell-fate determination (Weinmaster, 1997; Artavanis-Tsakonas et al., 1999; Perron and Harris, 2000; Selkoe, 2000). The modulation or dysfunction of such mechanisms may lead to severe brain disorders such as CADASIL (Joutel et al., 1996). Although these mechanisms need to be further clarified, our study suggests that cytokines, such as TGF-β1, may influence such processes. Similar comments can be made concerning the overexpression of cathepsin B. Although its mechanisms of action remain largely unknown, the expression of cathepsin B, which is largely confined to endosomes or lysosomes of neuronal cells (Cataldo et al., 1997), has been associated with amyloid-β accumulation (Frautschy et al., 1998) and neuronal cell shrinkage following transient cerebral ischemia (Hill et al., 1997). Furthermore, cathepsin B released by activated microglial cells have been shown to trigger apoptosis in primary neuronal cultures (Kingham and Pocok, 2001). Since cathepsin B expression is upregulated in brains of patients with Alzheimer disease, this may have significance for tissue damage and neuronal loss in the neuropathology of Alzheimer disease.
Of the genes that we have analyzed by RT-PCR, only two are modulated by TGF-β1 in cultured neurons. These observations suggest that astrocytes could be the major target for the responses to TGF-β1 in brain tissue. Altogether, these data may be related to studies that have described elevated concentrations of TGF-β1 in brain from patients with Parkinson disease (Lippa et al., 1995; Nagatsu et al., 2000), Alzheimer disease (van der Wal et al., 1993; Flanders et al., 1995), and stroke (Krupinski et al., 1996). Although it is necessary to be circumspect with such data, the use of a cDNA array appears to be a potent tool to identify novel targets for the understanding and eventual treatment of neurodegenerative conditions.
