Abstract
High density oligonucleotide arrays offer tremendous potential to study gene changes occurring in disease states. The authors described the first case of using a custom designed high density oligonucleotide probe array containing 750 genes to monitor the changes in mRNA transcript levels occurring after focal ischemia for a period of 3 hours. Permanent middle cerebral artery occlusion in the rat resulted in neuronal degeneration in the dorsolateral cortex and striatum over a time course of 24 hours. Comparing the changes in hybridization levels in the frontal and parietal cortices and the striatum, between the ipsilateral and contralateral sides of the brain using the probe arrays resulted in the up-regulation of 24 genes, which showed greater than a twofold change. Very few genes were found to be downregulated after the ischemic insult. Many of the immediate early genes (IEGs) such as c-fos, NGFI-A, NGFI-C, and Krox-20 were found to be robustly up-regulated in the three different regions studied. Other genes that were up-regulated in perifocal regions included Arc, Inhibin-β-A, and the phosphatases MKP-1 and MKP-3. The hybridization signal intensity from the probe arrays enabled quantification of many genes relative to one another, and robust changes in expression were obtained with very little interanimal variability. Furthermore, the authors were able to validate the increased expression of NGFI-C and Arc using in situ hybridization. This represented the first example of using high density oligonucleotide probe arrays in studying the expression of many genes in parallel and in discrete brain regions after focal ischemia.
Over the past few years, studies using animal models of cerebral ischemia have demonstrated that after the onset of ischemia a whole cascade of changes in gene expression occurs (Kogure and Kato 1993; Koistinaho and Hokfelt, 1997). It is now accepted that neurons and glial cells respond to ischemia, stress, or traumatic brain injury with a complex response involving differential gene expression. This response can be measured using in situ hybridization, real time quantitative reverse transcription polymerase chain reaction, and differential display to identify unknown genes (Liang and Pardee, 1992; Wang et al., 1996; Livesey and Hunt, 1996; Van Lookeren Campagne and Gill, 1998a, b ). The techniques used thus far to investigate changes in gene expression have been quite slow and laborious, enabling only a few genes to be measured at a time. Although the differential display technique can potentially pick up many novel genes, major drawbacks of this technique include artifacts and limited reproducibility (Liang and Pardee, 1992; Wang et al., 1996). Other techniques that have been used include the differential screening of cDNA libraries and serial analysis of gene expression (SAGE; Velculescu et al., 1995), but both of these techniques are difficult and time consuming.
Over the last few years the detection of differentially expressed genes has improved immensely with the introduction of DNA microarrays, either with immobilized cDNA's or with thousands of photochemically synthesized oligonucleotides (Lockhart et al., 1996; Lipshutz et al., 1999). This technology allows parallel expression monitoring of thousands of genes arrayed in a small area of about 1 cm2. The computer-assisted analysis of the data gives precise information about the expression level of a given gene and whether it is differentially expressed. Although this technology has been successfully used with yeast, cell lines, bacteria, and an inflammatory disease (Heller et al., 1997; DeRisi et al, 1997; de Saizieu et al., 1998; Wodicka et al., 1997), its impact on neuroscience research is still in its infancy. Therefore, the authors sought to assess the feasibility of using the high density oligonucleotide probe arrays to monitor the levels of hundreds of transcripts simultaneously in a well-defined model of cerebral ischemia in rats. Among the challenges that the authors faced were the quantity of tissue required, the uncertainty of success in comparing discrete brain regions from different animals, the speed of processing the tissue to minimize RNA degradation, and the interanimal variability. The production process for oligonucleotides minimized array to array variation and differences arose mainly from heterogeneity of the biological sample.
The model of permanent middle cerebral artery (MCA) occlusion used in the current studies has been very well characterized in terms of pathologic changes, development of infarct, and neuroprotective strategies (Tamura et al., 1981; Gill et al., 1991, 1992a, 1995). In many respects, this model of focal ischemia replicates the neuropathologic changes seen after a stroke in humans. In animals, this pathophysiology has been described to be associated with changes in gene expression (Kogure and Kato 1993; Koistinaho and Hokfelt, 1997; Van Lookeren Campagne and Gill, 1998a). The current studies have examined changes in gene expression 3 hours after permanent MCA occlusion, a time when the brain infarct is dynamically evolving (Gill et al., 1995). The pattern of gene expression between discrete brain regions on the ipsilateral side was compared with changes in the same regions on the contralateral side of the brain, which was not effected by the ischemia. These first experiments were run using an oligonucleotide array (Roche Bioscience; Palo Alto, CA, U.S.A.) originally designed to analyze rat genes associated with bone and cartilage.
MATERIALS AND METHODS
Surgical procedures for MCA occlusion
All experiments were performed in accordance with the Swiss Federal act on animal protection. Male Sprague—Dawley (320 to 350 g) rats were maintained on a 12-hour light—dark cycle and allowed food and water ad libitum.
The animals were anesthetized with a mixture of 3% isoflurane, 70% nitrous oxide, and 30% oxygen. They were then intubated and ventilated. Anesthesia was maintained using 1% to 2% isoflurane, 70% nitrous oxide, and 30% oxygen. The femoral artery was cannulated to enable monitoring of physiologic variables such as mean arterial blood pressure and blood gases. The temperature of the animals was maintained strictly between 37°C ± 0.1°C. The animals were subjected to permanent occlusion of the left middle cerebral artery as described previously (Tamura et al., 1981; Gill et al., 1992a). The left MCA was occluded permanently using bipolar coagulation, afterwards the skin incision was sutured and the femoral catheter removed. Then the anesthetic was turned off, the animals were allowed to breathe air, and once consciousness was regained they were extubated and allowed to breathe spontaneously. The animals were decapitated 3 hours after permanent MCA occlusion for gene chip analysis and at 20 minutes to 24 hours for the remaining procedures.
Gene and oligonucleotide probe selection
The custom designed rat chip ROEZ002 (Roche) originally designed for bone and cartilage research was used for these feasibility studies. This rat chip contained 750 probe sets for bone and cartilage genes, and more general genes such as cytokines or oncogenes (a complete listing of the genes is available from the authors) were used for each transcript 40 oligonucleotide probe. Approximately 70% of the genes are known to be expressed in the central nervous system and were thus relevant to the current study.
RNA preparation
The rats (n = 5) were decapitated 3 hours after permanent MCA occlusion, the brains were rapidly removed, and the different brain areas, frontal, parietal cortices, and striatum were dissected from the contralateral and ipsilateral sides. The tissue samples were rapidly frozen in liquid nitrogen and kept at −80°C until required for RNA isolation.
Isolation of total RNA
The brain tissue was homogenized in FastPrep tubes for 20 seconds in a Savant homogenizer (Bio101, Buena Vista, CA, U.S.A.) containing teflon beads and 1 mL RNAzol™ B reagent (Biotecx, Friendswood, TX, U.S.A.). Total RNA was isolated according to the manufacturer's instruction and quality was assessed using gel electrophoresis.
cDNA synthesis and in vitro transcription
Double strand cDNA was synthesized from 20 μg of total RNA, using a polyT primer with a T7 promoter sequence (5′ GGCCAGTGAATTGTAATACGACTCAC TATAGGGAGGCGG(dT)24VN- 3′) as described by Wodicka et al. (1997). The cDNA was phenol-chloroform extracted, purified using phase lock gel (5 Prime to 3 Prime, Boulder, CO, U.S.A.), and ethanol precipitated.
The MEGA Script T7 kit (Ambion, 5′-3′ Austin, TX, U.S.A.) was used to transcribe the cDNA into cRNA in the presence of Biotin-11-CTP and Biotin-16-UTP (Enzo, Farmingdale, NY, U.S.A.) using the instructions supplied with the kit. After purification with the RNAeasy kit (Qiagen, Hilden, Germany), integrity of the cRNA was checked using gel electrophoresis before fragmentation. Afterwards, 10 μg fragmented cRNA were used for hybridization to the array. All subsequent steps were done according to Affymetrix (GeneChip version 3.0, Affymetrix, Santa Clara, CA, U.S.A.) (Lockhart et al., 1996; de Saizieu et al., 1998).
Data processing
The expression data from the gene chips were quantified using commercial GeneChip software from Affymetrix (EDMT/LIMS Affymetrix; Lockhart et al., 1996). The values for average differences for each brain region between the contralateral and ipsilateral side were compared using an analysis of variance (ANOVA) test with repeated measures followed by a post hoc contrast test that corrects for multiple comparisons (BMDP Statistical Software, Cork, Ireland).
In situ hybridization for NGFI-C and Arc mRNAs
Animals were decapitated at 20 minutes, 1, 3, 6, and 24 hours after permanent MCA occlusion (n = 3 for each time point) and after sham operation (total n = 5). Control adult rats were processed to determine the basal expression of the genes of interest (n = 2). The brains were rapidly removed, frozen on dry ice, sectioned into 12-μm-thick sections on a cryostat, and then fixed with 4% paraformaldehyde. The 60-mer antisense synthetic oligonucleotide probes were obtained commercially (Genosys Biotechnology's, Woodlands, TX, U.S.A.). Three specific sequences were designed from the GeneBank and EMBL for each gene. The complementary sense sequence of each oligonucleotide was used as a negative control. The hybridization signal between the different antisense probes for each gene was similar. All of the results presented in this study were performed with the oligo complementary to nucleotides 601 to 660 for NGFI-C mRNA and the oligo complementary to nucleotides 1313 to 1372 for Arc mRNA. No signal was observed with sense probes. The oligonucleotides were labeled at the 3' end using terminal deoxynucleotidyl-transferase (Life Technology's, Basel, Switzerland) and [35S]dATP (New England Nuclear, Boston, MA, U.S.A.). The protocol for labeling and in situ hybridization has been described previously (Van Lookeren Campagne et al., 1998a). Sections were exposed for up to 4 weeks at 4°C to β-Max Hyperfilm (Amersham, Zurich, Switzerland). Quantification of hybridization signal was done using storage phosphor screen imaging. Image analysis was performed using M2-Image Analysis MCID Software (Imaging Research, St. Catherine's, Ontario, Canada). Photostimulated luminescence per mm2 values were obtained by the image analysis software and indicated the concentration of radioactivity in the original region of interest. Five areas were selected for quantification in the ipsilateral and contralateral side—infarct core, parietal, frontal and cingulate cortices, and nucleus accumbens. The data were expressed as the ratio of the ipsilateral versus the contralateral side. For each time point, values from the ipsilateral side were compared statistically with those of the contralateral side using a paired t-test.
Western blot analysis for Arc protein
Rats were decapitated at 1, 3, 6, and 24 hours after MCA occlusion (n = 3, per each time point) or after sham operation (total n = 4). The brains were removed, parietal and frontal cortices and striatum were dissected for each hemisphere and then frozen in liquid nitrogen. Protein homogenates were prepared as described previously by Van Lookeren Campagne and Gill (1998a). Thirty μg of each sample was diluted with 4X NuPAGE buffer (Novex, San Diego, CA, U.S.A.) containing 8% β-mercaptoethanol and then boiled for 10 minutes and allowed to cool down. Samples were loaded in 4% to 12% Bis-Tris Gel (Novex) and run with MOPS buffer (Novex) and then blotted on to a nitrocellulose membrane. Membranes were incubated overnight with a monoclonal mouse IgG against Arc (1:1000; Transduction Laboratories, Lexington, KY, U.S.A.) and then incubated for 1 hour with goat anti-mouse IgG HRP-conjugated (BioRad, Glattbrugg, Switzerland). The peroxidase reaction was visualized by enhanced chemiluminiscence (ECL, Amersham). The image of each film was captured with a high resolution digital camera (Model 1400, Xillix Technologies, Vancouver, Canada). Arc immunoreactivity was quantified by measuring optical densities of the 53 kDa band on the film using M2-Image Analysis MCID Software. Values from the dissected region in the ischemic hemisphere were compared with those in the contralateral hemisphere using a t-test.
RESULTS
Focal ischemia induced by permanent MCA occlusion resulted in ischemic damage in the dorsolateral cortex and striatum. At 3 hours after MCA occlusion the damage was restricted to the striatum and parietal cortex, which represent the densely ischemic areas (Gill et al., 1995). At this time, the frontal cortex did not show any signs of ischemic damage, however, by 24 hours some of the tissue in the frontal cortex became incorporated into the infarct. To identify genes differentially expressed as a consequence of the ischemia, the authors analyzed the three different regions (Fig. 3B)—frontal and parietal cortices and striatum—from the ipsilateral side and compared these with the same regions from the contralateral side. The different regions responded to the ischemic insult with a different time course of changes.
The physiologic variables were monitored regularly during the ischemia, the blood pressure was maintained at approximately 100mm Hg, and the blood gases were maintained at approximately 100mm Hg for PaO2 and at approximately 35mm Hg for Pa
Using different rat tissues the authors were able to show that as little as 30 mg of rat brain tissue was sufficient to generate meaningful data on oligonucleotide arrays. Therefore, discrete brain regions were dissected out and processed for probe array analysis. From pilot experiments (n = 3 animals; data not shown) using the frontal cortex, dissected 3 hours after permanent MCA occlusion, the authors were able to demonstrate good reproducibility in terms of the numbers of genes detected and differentially expressed between the ipsilateral and contralateral sides of the brain. Therefore, a second series of animals were prepared to study the changes in gene expression in the different brain regions (n = 5).
Hybridization experiments to oligonucleotide microarrays
After probe array hybridization and scanning, the fluorescence intensity of each gene was calculated by subtracting the mismatch from the match signal. The experiment took approximately 5 days from MCA occlusion to obtaining expression data. The greater the average differences, the higher the probability that a given mRNA was present in the hybridization target. This, and the number of probe pairs showing positive hybridization, was part of the decision matrix used to decide whether a gene was absent or present. In each sample the analysis software (GeneChip, Affymetrix) called, on average, only 15% of the genes represented on the array as expressed. This was not surprising as the array used was biased for muscle and cartilage genes. A typical image of a scanned chip from the frontal cortex of an animal subjected to 3 hours of ischemia is shown in Fig. 1A. In Fig. 1B, the average intensities of all genes from the ipsilateral and contralateral RNA sample are plotted. This display provides a simple representation of complex data and allows electronic selection of genes for further analysis.
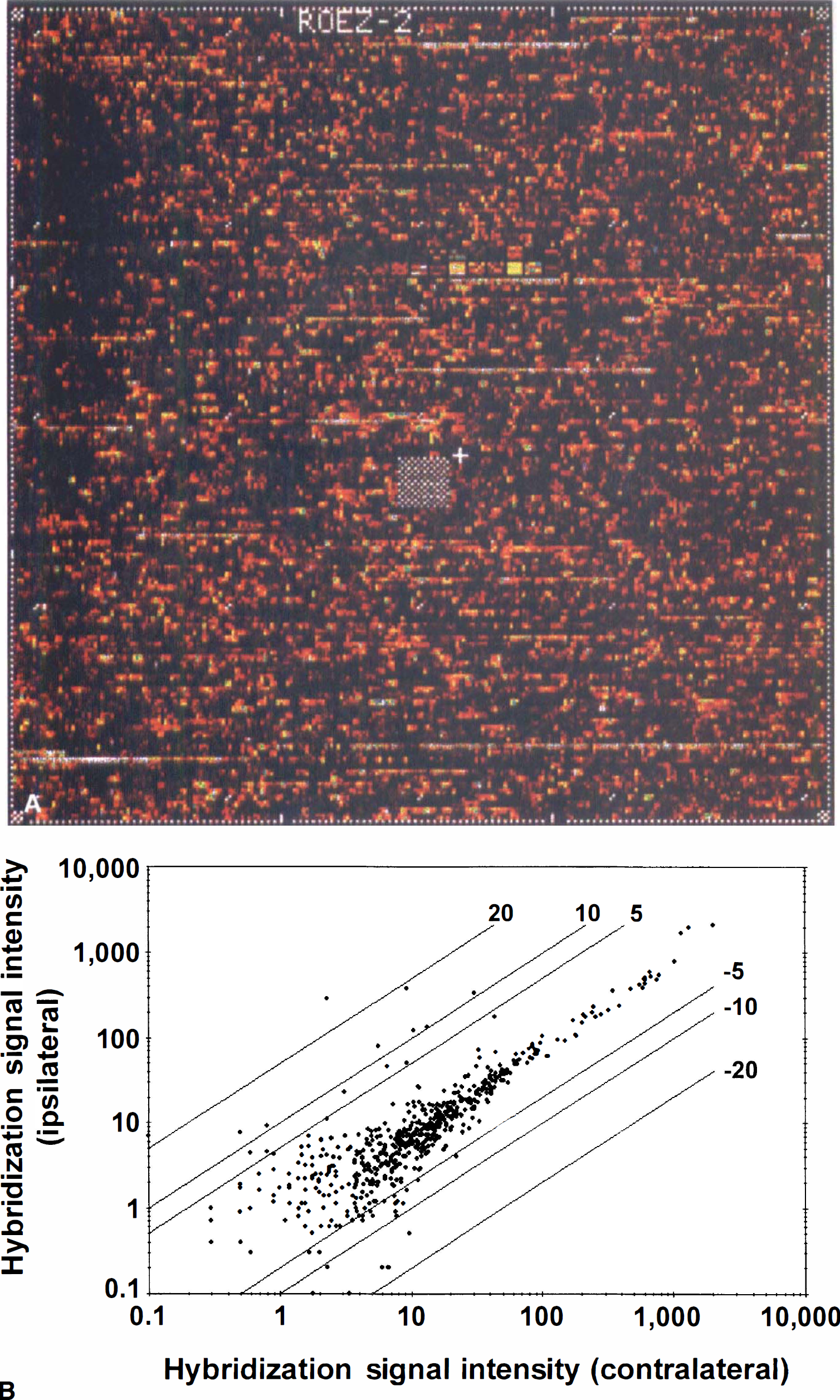
(A) Image of an oligonucleotide array containing probe sets for approximately 750 rat genes after hybridization with biotin-labeled cRNA obtained from the ipsilateral frontal cortex after MCA occlusion.
The comparative analysis of the same brain region from a single animal demonstrated the increases and decreases in the expression of specific genes compared with the levels of other genes that do not change after the ischemic insult (Fig. 1B). The alteration in the transcription of one gene was considered significant if the increase or decrease was at least twofold compared with the contralateral hemisphere. The fold change was based on the analysis of the gene expression pattern from the five animals used for each region. The gene expression profile showing the mean fold change and the hybridization signal intensity in the frontal cortex from five animals is shown in Fig. 2. Twenty different transcripts were up-regulated and another five were downregulated at 3 hours after permanent MCA occlusion in the frontal cortex. The same analysis for the parietal cortex and striatum demonstrated that 16 and 15 genes, respectively, were up-regulated, and 4 genes were downregulated in the striatum. In the same analysis, two housekeeping genes, GAPDH and β-actin, were examined and neither showed a significant change in the ipsilateral side compared with the contralateral side in any of the three regions studied (data not shown).
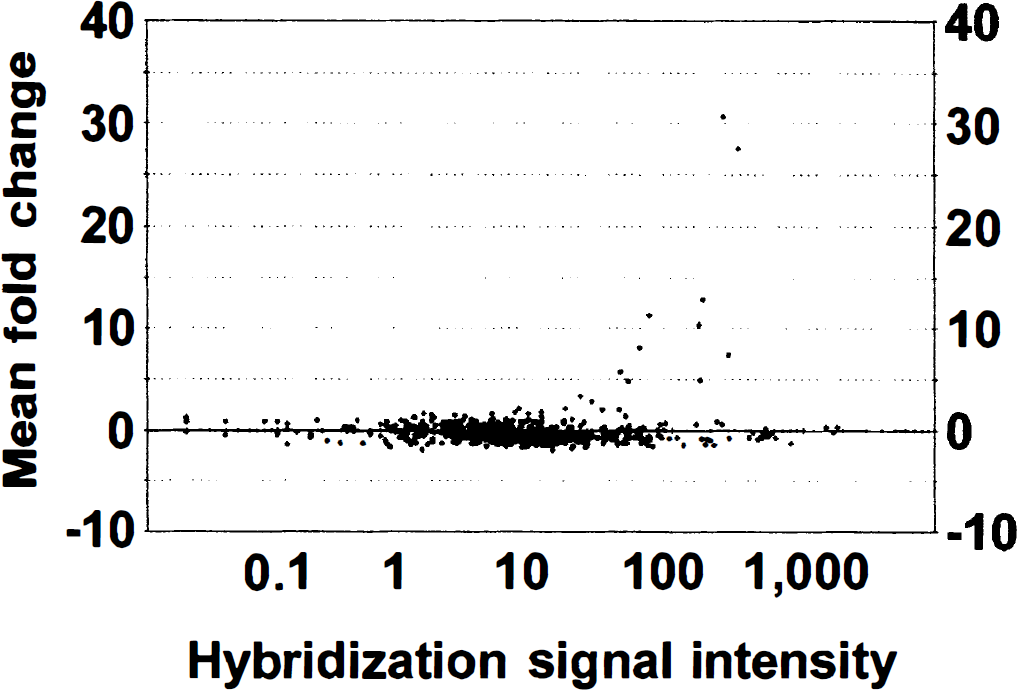
Scatter plot indicates the relative mRNA levels in frontal cortex of the ipsilateral versus the contralateral side. Each dot corresponds to the mean fold change factor and the mean hybridization signal intensity (average difference) on the ipsilateral side of the occlusion. Selected genes with mean fold change values of > 2 or < −2 were chosen for further analysis.
Up-regulation of gene expression in different brain regions
The robust changes in gene expression seen in the three different brain regions are illustrated in Fig. 3. Immediate early genes (IEGs) such as c-fos, NGFI-A, NGFI-C, Krox20, NGFI-B, Nor-1, Cyclooxygenase-2 (Cox-2) and activity-regulated cytoskeleton associated protein (Arc) were found to be up-regulated in all three brain regions (Fig. 3A). Statistical analysis revealed that for the frontal cortex there was an overall significant effect for (ANOVA F = 158, P < 0.0001) up-regulation of different genes when the ipsilateral and contralateral regions were compared. The genes demonstrating a significant (P < 0.05) change, which was greater than twofold, between the ipsilateral and contralateral frontal cortex are shown in Fig. 3A. There was a strong hybridization signal intensity for the transcript of c-fos and it showed a 100-fold increase in expression in the frontal cortex.
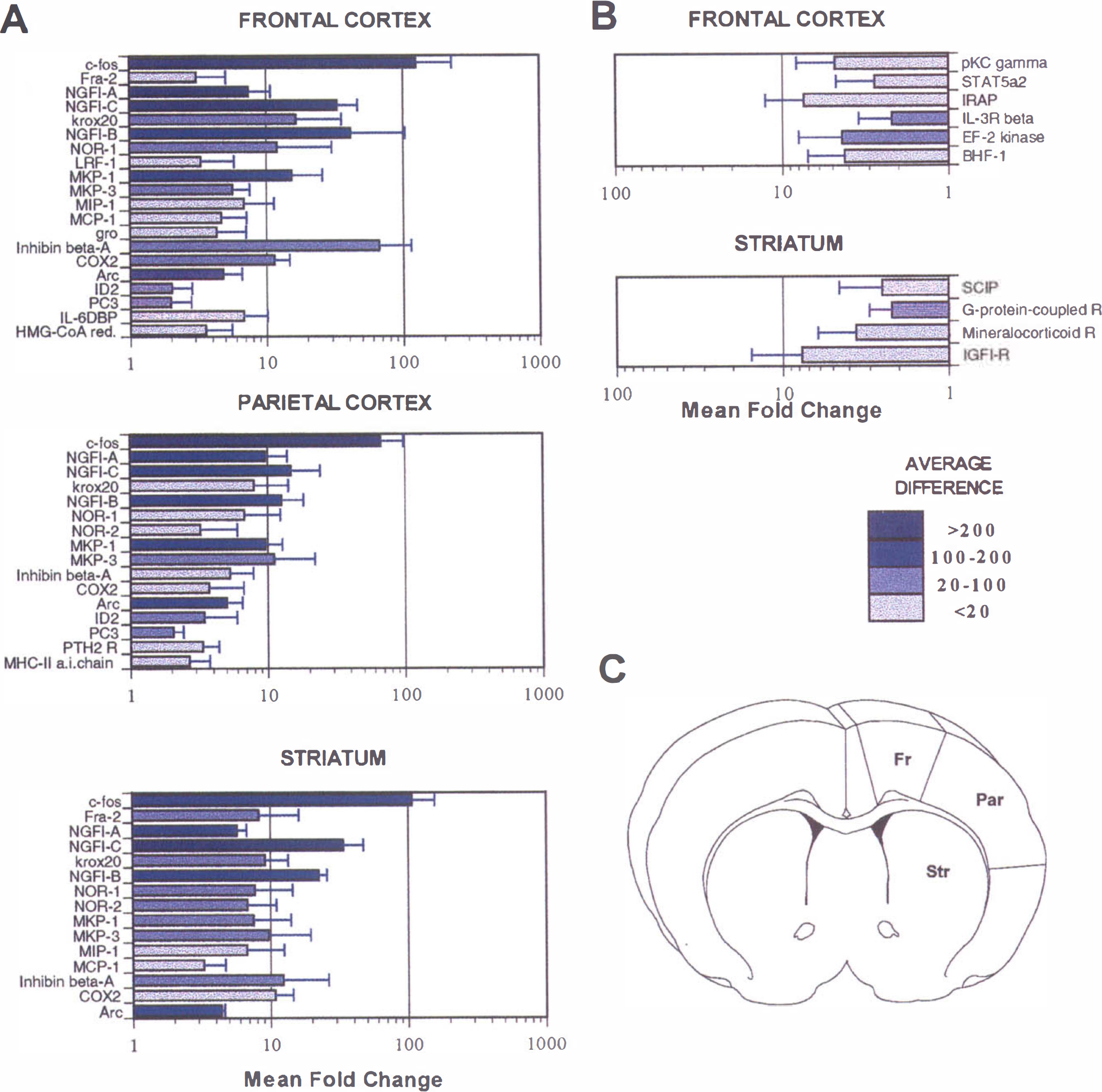
Changes in gene expression measured at 3 hours after permanent MCA occlusion.
The magnitude of changes in gene expression in the parietal cortex was less than that seen in the frontal cortex or striatum, which was densely ischemic at 3 hours. There was no overall significant difference between the ipsilateral and contralateral side in terms of up-regulation of gene expression (ANOVA F = 0.90, P > 0.3) for the parietal cortex. However, a post hoc contrast test revealed that some genes were significantly (P < 0.05) up-regulated as illustrated in Fig. 3A. One of the most marked changes was in the IEG, c-fos, which increased 60-fold in the parietal cortex.
There were overall significant (ANOVA F = 16.9, P < 0.01) changes in gene expression in the striatum when the ipsilateral and contralateral regions were compared for the different genes. The genes that were significantly (P < 0.05) up-regulated and showed a greater than or equal to twofold change are shown in Fig. 3A. Once again the highest change was seen in c-fos, which increased almost 90-fold in the ipsilateral striatum compared with the contralateral side. Some of the genes, for example Fra-2 and c-jun, showed quite low hybridization intensities, but significant up-regulation could still be detected. Three genes of the early growth response family of transcription regulatory factors, NGFI-A, NGFI-C and Krox20, were significantly induced in all three regions. NGFI-B, Nor-1, and Nor-2 have structural features of ligand-activated transcriptional regulators and constitute the NGFI-B subfamily within the nuclear receptor superfamily; they are also known as orphan nuclear receptors because specific ligands for these molecules are not known. These genes were up-regulated in all of the three regions studied, although Nor-2 was not up-regulated in the frontal cortex. NGFI-B showed the most robust increases (>40-fold increase in the frontal cortex) with a medium to high hybridization signal intensity compared with the other members of this family of genes.
Other genes that are considered to be IEGs because of their rapid and transient activation are Cox-2 and Arc. Cox-2 was up-regulated over 10-fold in the frontal cortex and striatum, and changed over threefold in the parietal cortex; although in the latter two regions low hybridization intensity, which was at the limits of detection, was shown. There was a fivefold increase in the expression of Arc in all of the three brain regions studied. MKP-1 and MKP-3 are stress-inducible members of the mitogen-activated protein kinase phosphatase family of dual specificity protein-tyrosine phosphatases. They were both up-regulated in all three brain regions. Inhibin β-A, a member of the TGF-β gene superfamily which is not constitutively expressed in the brain, was dramatically up-regulated in frontal cortex (over 60-fold change). In the parietal cortex (> 5-fold) and striatum (> 10-fold), this transcript showed a high induction with a moderate hybridization intensity. Chemokines, monocyte chemoattractant protein-1 (MCP-1), macrophage inflammatory protein-1 (MIP-1), and gro were also found to be up-regulated, although the mRNA of these chemokines was increased in the frontal cortex and striatum, with a low hybridization signal intensity; no changes were observed in the parietal cortex. Increases of two additional transcripts, IL-6 dependent binding protein and in 3-hydroxy-3-methylglutaryl-coenzyme A (HMG-CoA) reductase, were noted in the frontal cortex.
Downregulation of gene expression in different brain regions
There was very little downregulation of genes seen after MCA occlusion in the regions studied. There was an overall significant change (ANOVA, F = 15.04, P < 0.01) for downregulation of genes in the frontal cortex, however, the magnitude of these changes was small as compared with changes seen in up-regulation of genes in the same area. The genes that were significantly (P < 0.05) downregulated by a factor of 2 or more are shown in Fig. 3B. They were mostly related to signal transduction, for example IL-3 receptor β, STAT5 α2, and the γ subunit of protein Kinase C. Other downregulated genes were IRAP, (insulin-regulated membrane aminopeptidase); the elongation factor-2 kinase, involved in protein synthesis; and BHF-1, a brain basic helix-loop helix factor. All of these transcripts had a low hybridization signal intensity apart from IL-3 Rβ. There was no overall significant downregulation of gene expression in the parietal cortex (ANOVA F = 1.6, P > 0.2). There were no transcripts that demonstrated downregulation by a factor of 2 or more in the parietal cortex. There was a significant downregulation in the expression of some genes in the ipsilateral striatum compared with the contralateral striatum (ANOVA F = 35.8, P < 0.001). The genes that were significantly (P < 0.05) downregulated by a factor of 2 or more in the striatum are shown in Fig. 3B. Interestingly, the largest decrease of sevenfold was seen in the insulinlike growth factor-1 receptor.
In situ hybridization studies with NGFI-C and Arc mRNAs
To further validate that the described up-regulation of genes the authors detected on the oligonucleotide probe arrays could be reproduced using in situ hybridization, the authors performed a time course study of NGFI-C and Arc mRNA expression after focal ischemia. Sham-operated and control rats showed no induction of NGFI-C mRNA (data not shown). At 3 hours after MCA occlusion there was an increase in NGFI-C mRNA expression, which appeared to be restricted to the “penumbral” areas such as the frontal and cingulate cortices and the nucleus accumbens (Fig. 4A). Quantification of the hybridization signal of the specific radiolabeled probe for NGFI-C showed that the peak of expression of this gene was at 3 hours, coinciding with the time when the analysis of gene expression by oligonucleotide probe arrays was performed (Fig. 4B). In the parietal cortex the increase of NGFI-C mRNA was lower and its peak was delayed up to 6 hours. The level of this transcript returned to the basal level at 24 hours after MCA occlusion in all of the regions evaluated.
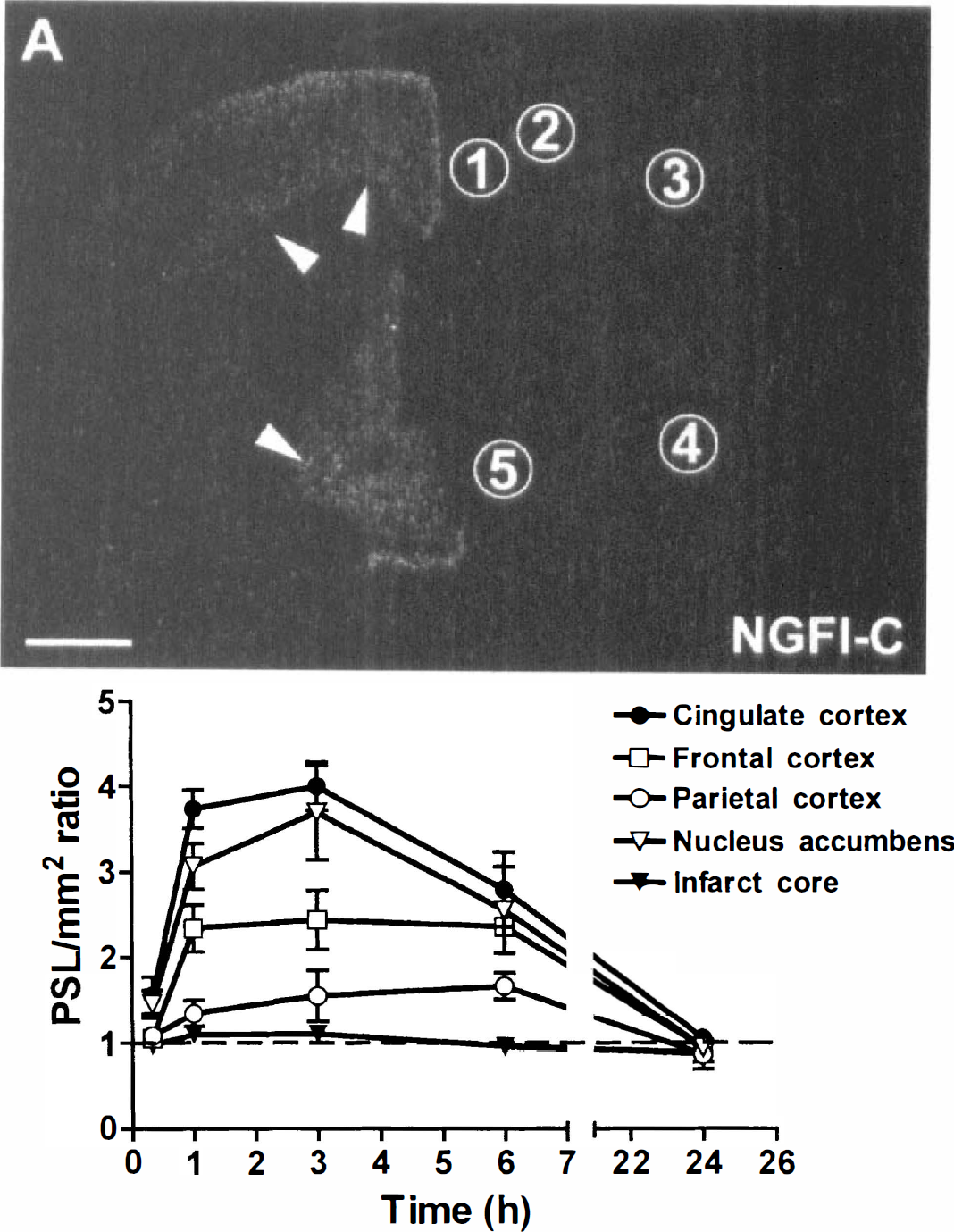
In situ hybridization of NGFI-C mRNA on coronal brain section 3 hours after permanent MCA occlusion.
There was a constitutive expression of Arc mRNA observed by in situ hybridization in the cortex and striatum (Fig. 5), which is in accordance with previous studies. Focal ischemia induced Arc mRNA in several perifocal regions of the infarct. Hybridization signals were observed at 20 minutes after MCA occlusion in medial striatum, cingulate cortex, and nucleus accumbens; stronger induction was detected from 1 hour in the same regions and in frontal and parietal cortices. Transcript levels of Arc decreased to basal expression levels by 24 hours after focal ischemia. Quantification of hybridization signal by phosphor screen imaging demonstrated that Arc mRNA expression peaked at 1 hour after MCA occlusion in penumbral areas such as cingulate and frontal cortices and nucleus accumbens (Fig. 5B). In the parietal cortex, Arc mRNA levels, like NGFI-C mRNA expression, increased slowly reaching peak levels at 3 hours.
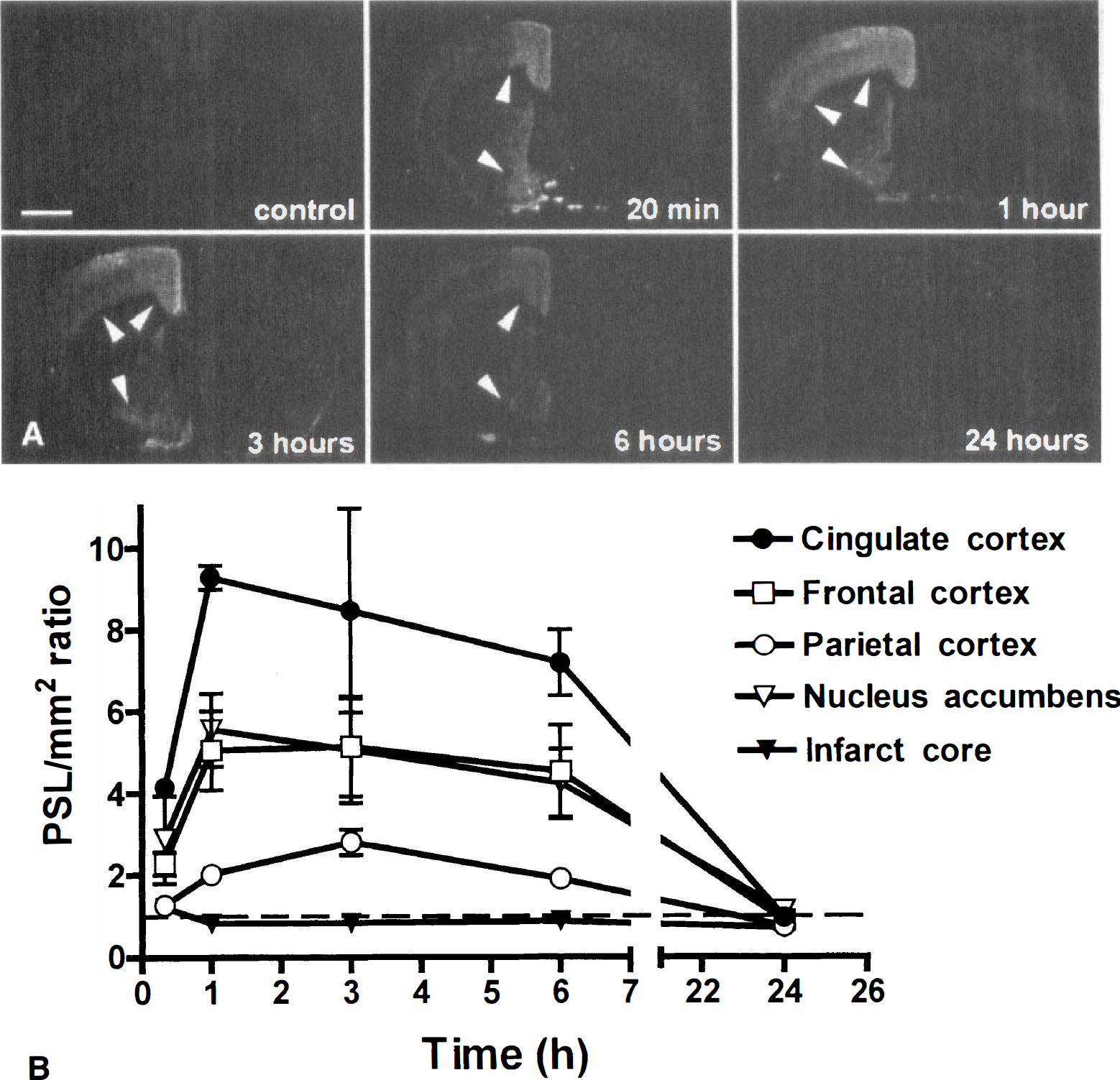
Arc mRNA expression after in situ hybridization in coronal sections of the control rat brain and rats subjected to 20 minutes, 1, 3, 6, and 24 hours of MCA occlusion.
It is well known that cerebral ischemia inhibits global protein synthesis, although some specific transcripts can be translated. Therefore, the authors further characterized the induction of Arc using a monoclonal antibody to assess the protein levels of Arc (Fig. 6A). There was a slight increase in Arc immunoreactivity in the striatum and parietal cortex after MCA occlusion, however, there was a marked change observed from 6 hours in the three regions studied. Semiquantification of the Western blots corroborated the time course of the induction of Arc immunoreactivity (Fig. 6B). The induction of Arc protein evolved with some delay compared with the time course of Arc mRNA expression.
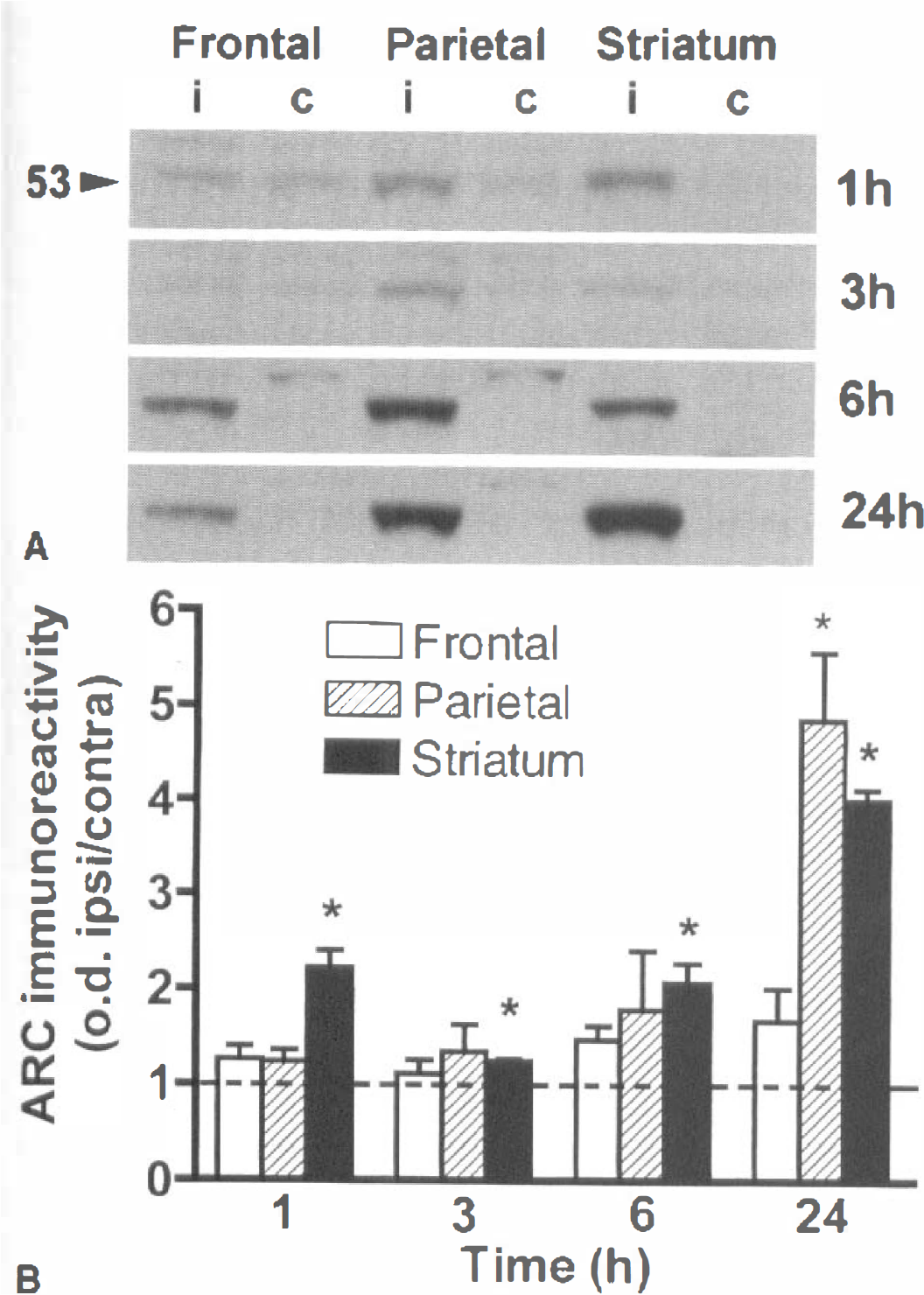
Arc immunoreactivity in the rat brain after MCA occlusion.
DISCUSSION
The present studies have illustrated that sophisticated oligonucleotide probe arrays can be used as a powerful and reliable technique for the analysis of ischemia specific gene expression in the brain. This was the first study demonstrating the utility of oligonucleotide arrays in fingerprinting changes in gene expression in the brain after disease modification. The initial problems encountered included the quantity of tissue required; however, improved hybridization techniques reduced the quantity of brain tissue required for each probe array to 30 mg, thus enabling discrete brain regions to be analyzed for parallel changes in gene expression. In total, we detected approximately 24 up-regulated genes, and of these approximately 20 showed greater than or equal to a twofold change. Interestingly, we did not see downregulation of many genes which suggests that the brain tends not to switch off gene expression in response to acute stress, but rather up-regulates many IEGs. The limits of hybridization for this technique were approximately a twofold change in regulation of the genes. Some of the up-regulated genes had an average hybridization signal intensity of < 20 as shown in Fig. 3 and these were at the limits of detection in this assay.
The changes in gene expression detected on the probe arrays after 3 hours of focal ischemia were similar to changes reported using traditional methods such as in situ hybridization and reverse transcription polymerase chain reaction techniques (Koistinaho and Hokfelt, 1997). A consistent up-regulation of the IEG c-fos has been widely described after focal ischemia, and in the current study we saw a robust increase in c-fos expression in the frontal cortex, an area that was not damaged by the ischemic insult. We also saw marked increases in c-fos in the parietal cortex and striatum in which the ischemia was quite dense at 3 hours after MCA occlusion. Interestingly, Fra-2, another gene related to Fos, was up-regulated in the frontal cortex and striatum that border the infarct, but not in parietal cortex. We did not see a robust increase in expression of c-jun at 3 hours in the frontal or parietal cortex, and although we did see an increase in the striatum, the hybridization signal was low and at the limits of detection. This observation was similar to findings using in situ hybridization studies in which only weak changes in c-jun were reported in the striatum after permanent MCA occlusion (Collaco-Moraes et al., 1994). Therefore, the probe arrays were sensitive to changes, which were at the borderline of detection using in situ hybridization techniques.
Expression of these IEGs was regulated at the level of transcription and was independent of new protein synthesis. This was quite important as global protein synthesis was rapidly perturbed after focal ischemia, especially in the densely ischemic areas such as the striatum and parietal cortex (Jacewicz et al., 1986; Mies et al., 1991; Hossmann, 1993). The current results that showed an induction of mRNA of several genes in the parietal cortex and striatum further support earlier observations that inhibition of protein synthesis after focal ischemia is not caused by inhibition of transcriptional processes (Abe et al., 1988) but is caused by inhibition at the translational level (Hossmann, 1993). With respect to these observations it is interesting that elongation factor-2 kinase, which is involved in protein synthesis, was found to be downregulated in the frontal cortex.
Other genes that were robustly up-regulated were the IEGs encoding the zinc finger transcription factors, NGFI-A, C, and krox 20 (egr-2). Changes in expression of these genes have also been documented previously (Koistinaho and Hokfelt, 1997; Honkaniemi et al., 1997). However, using the probe array technology we were able to compare the relative changes of these genes in parallel. We were able to confirm the robust expression of NGFI-C using in situ hybridization, and a time course study revealed that peak changes were seen at 3 hours in the cingulate cortex, frontal cortex, and nucleus accumbens areas which were not damaged by the ischemic insult. Thus, the up-regulation of NGFI-C might be a defensive mechanism of these areas in response to the ischemic insult. It has also been speculated that these genes were up-regulated in these areas because of spreading depressionlike activity that had been reported in the cortex and penumbral areas after focal ischemia (Gill et al., 1992b). Alternatively, these genes had Ca2+ response elements and the high levels of Ca2+ in the brain postischemically might have caused this up-regulation.
The phosphatases MKP-1 and MKP-3 (Muda et al., 1996a; Boschert et al., 1998) were also found to be up-regulated after the ischemic insult. It has been suggested that MKP-1 is up-regulated in surviving neurons after an ischemic insult, kainate toxicity, or axotomy (Wiessner et al., 1995; Gass et al, 1996; Winter et al., 1998). However, in the current study we obtained an increase in mRNA of MKP-1 in all three areas of the brain with the highest increase seen in the frontal cortex. This was the first report of an increase in expression of the mRNA for MKP-3 three hours after focal ischemia. Transcriptional activation of MKP-1 and MKP-3, in response to diverse environmental stresses, antagonizes activation of the mitogen-activated protein kinases ERK and JNK by dephosphorylation of the threonine and tyrosine residues, which are critical for the activation of these kinases (Sun et al., 1993; Muda et al., 1996b). In the current study we did not see any up-regulation of ERK or JNK (these genes were present on the probe array), so it is possible that under ischemic conditions MKP-1 and MKP-3 inhibit ERK or JNK activation.
We also detected changes in the chemokines MCP-1, MIP-1, and gro, although these changes were at the limits of detection because low hybridization intensity was seen. Previous studies have also reported weak induction of MCP-1 and MIP-1 in ischemic areas using in situ hybridization techniques (Kim et al., 1995). Other inflammatory genes such as IL-1, IL-6, ICAM, and LECAM were not up-regulated at this time point after ischemia.
Some of the more novel genes that we found to be up-regulated after ischemia in the frontal cortex, an area which survived the ischemic insult, were inhibin-β-A, Arc, and HMG-CoA-reductase. Inhibin-β-A is a growth factor belonging to the family of inhibin-activin growth factors that are related to the TGF-β family. There was quite a marked up-regulation of inhibin-β-A at 3 hours after ischemia. It has been suggested that this growth factor may be acting as a neurotrophic agent for neurons that are trying to survive the ischemic insult although little is known about the role of Inhibin-β-A in the brain (Lai et al., 1996).
We further investigated the changes in expression of Arc using in situ hybridization and Western blotting. There was a very robust increase of Arc in perifocal regions suggesting it might play a role in adaptive synaptic plasticity after focal cerebral ischemia. Arc was discovered using differential screening of cDNA libraries prepared from the hippocampus of rats subjected to electroconvulsive seizures (Lyford et al., 1995). Immunohistochemical studies demonstrated that Arc protein is expressed in neurons located in the cytoplasm of the cell body, and is enriched in neuronal dendrites; it exhibits some homology with α-spectrin, a cytoskeleton-associated protein, and in vitro studies have demonstrated that Arc is associated with the actin cytoskeleton (Lyford et al., 1995; Fosnaugh et al., 1995). Therefore, the increased expression of Arc after focal ischemia suggested that this gene might mediate cytoskeletal changes, which might underlie the spreading depressionlike activity and neuronal plasticity in these perifocal regions. Arc is also an IEG but the localization to dendrites suggested it may serve as an effector molecule rather than a transcriptional factor (Fosnaugh et al., 1995). Another “effector” IEG induced after focal ischemia, Cox-2 (Yamagata et al., 1993), was responsible for metabolizing arachidonic acid to prostaglandins. There were robust changes in the frontal cortex of Cox-2, however, changes in Cox-2 expression in the parietal cortex and striatum were at the limits of detection. The induction of Cox-2 has been reported to occur after both transient and permanent occlusion of the MCA (Planas et al., 1995; Collaco-Moraes et al., 1996).
In conclusion, the current data have demonstrated the expression of many genes in parallel after focal ischemia, and there appear to be quite discrete and region specific changes in the regulation of the genes. Families of genes were found to be up-regulated and confirmed many of the previous findings using standard techniques (Koistinaho and Hokfelt, 1997). This was a proof of concept study for utilizing high density oligonucleotide probe arrays to monitor gene expression in a diseased state. This methodology provides a novel way to study changes in gene expression in different animal models of cerebral ischemia and to identify genes that may be commonly expressed in areas of brain resistant to the neuronal damage. The challenges that lie ahead are in attempting to characterize these genes and to identify a function for them. With regards to ischemia, questions arose whether these changes were just an adaptive response of the brain to stress or were some of these genes potentially future drug targets because of their expression in tissue that withstood the ischemic insult. This study is just the beginning of a wealth of information that can be obtained about disease states that would refine our therapeutic strategies for these diseases. With the help of informatics and multiple disease models we should be able to narrow down the number of genes as potential new therapeutic targets for the treatment of neurodegenerative disorders such as stroke.
Footnotes
Acknowledgments
The authors thank Dr. R. Heller, Dr. P. Cannon, and Dr. P. Klonowski for the design of the first prototypic rat oligonucleotide probe array. The authors also thank Dr. J.A. Kemp and Dr. J. Mous for their enthusiasm and support for this project; Dr. J.A. Kemp for critical reading of the manuscript; Adrian Roth for helping with analysis of the data from the probe arrays; and Dr. G. Richards for helping with the in situ hybridization technique.
