Abstract
Background
Cancer is one of the leading causes of death in the world. Several observational studies have suggested a significant association of the MDM4 SNP34091 A>C polymorphism with cancers. However, the results of the published studies are inconsistent.
Materials and methods
PubMed, Embase/Ovid and the Chinese National Knowledge Infrastructure were searched for relevant studies with a time limit of April 20, 2016. Pooled odds ratios (ORs) with 95% confidence intervals (CIs) were used to evaluate the strength of the association between MDM4 polymorphism and cancer risk. Publication bias was estimated using Begg's funnel plots and Egger's regression test.
Results
A total of 19,328 patients and 51,058 controls were included in the analysis. Overall, a significantly decreased risk of cancer was associated with MDM4 SNP34091 polymorphism for the allele model (C vs. A, OR = 0.715, 95% CI: 0.622-0.821, p = 0.000), dominant model (CC + AC vs. AA, OR = 0.684, 95% CI: 0.563-0.831, p = 0.000), recessive model (CC vs. AC + AA, OR = 1.139, 95% CI = 1.055-1.230, p = 0.001) and heterozygote model (AC vs. AA, OR = 0.687, 95% CI = 0.568-0.832). In the subgroup analysis by cancer type, no significant association was found in the breast cancer subgroup. In the subgroup analysis by geographical region, 2 genetic models, the allele and heterozygote models, showed a significant association in Chinese populations.
Conclusions
The results of our meta-analysis showed that the MDM4 SNP34091 A>C polymorphism may function as a protective factor against cancer risk.
Introduction
Cancer is a leading cause of death and a serious disease threatening public health in developed and developing countries worldwide. Based on the GLOBOCAN worldwide estimates for 2012, newly diagnosed cancers amount to 14.1 million and cancer-related deaths to 8.2 million (1). It is well known that the etiology and development of cancer are complex, and can be affected by environmental and genetic factors (2). In addition to exogenous factors, genetic variations are crucial in the development of cancer. Single nucleotide polymorphisms (SNPs) are the most common genetic variation contributing to the individual susceptibility to cancer (3). Following the finding of this relationship, a large number of candidate genes were studied in case-control studies (4).
The tumor suppressor gene TP53 plays a pivotal role in many physiological processes including apoptosis, cell cycle and metabolism (5). TP53 mutation has proved to be among the most common mutations in various cancer types. Maintaining the correct level of p53 protein is therefore imperative. Murine double minute 4 (MDM4), also known as MDMX or HDMX, is a structurally homologous protein of MDM2 (6). MDM4 and MDM2 are the major negative regulators of TP53. Both MDM2 and MDM4 can inhibit TP53 by directly binding and masking its transactivation domain, which results in p53 protein degradation via the ubiquitination pathway. However, MDM4 alone is unable to cause degradation (7–9); it must form the MDM2/MDM4 heterodimer. Interestingly, the MDM2/MDM4 heterodimer showed a more potent degrading effect than the MDM2 homodimer (10). MDM4 overexpression has been observed in many types of cancer and was found to contribute to cancer initiation, maintenance and progression (7, 11).
MDM4 polymorphisms and mutations have been frequently studied because of their impact on TP53 in some types of cancers. Most studies focused on MDM4 SNP34091 A>C (rs4245739). The MDM4 SNP34091 A>C polymorphism in the 3′-untranslated region of MDM4 creates a putative target site for miR-191-5p and miR-887-3p, which might lead to less expression of MDM4 (12, 13). However, individual studies yielded inconsistent or conflicting findings because of heterogeneous data collection and inadequate sample sizes. To eliminate this inconsistency, we conducted a meta-analysis of all eligible case-control studies published to date and estimated the cancer risk of MDM4 polymorphisms.
Materials and methods
Publication search
The databases PubMed, Embase/Ovid and the Chinese National Knowledge Infrastructure (CNKI) were searched using the following search terms: “murine double minute 4”, “MDM4”, “polymorphism”, “variants” and “cancer” to identify relevant articles on the subject. The last retrieval date was April 20, 2016. In addition, the references of the included studies were searched to identify additional potentially relevant studies. The language was restricted to English and Chinese. The included studies met the following inclusion criteria: 1) evaluation of the association between MDM4 SNP34091 A>C polymorphism and cancer susceptibility; 2) case-control study or cohort study; 3) the genotype distribution of the polymorphism in cases and controls was sufficient to estimate the odds ratios (ORs) with a 95% confidence interval (CI) and a p value. The main exclusion criteria were as follows: 1) case reports, review articles, and editorials; 2) no control population; 3) duplication of a previous publication; 4) insufficient available data.
Data extraction
Data extraction was carried out independently by 2 investigators. Disagreements were resolved by discussion. For each included study, the following information was collected: first author's name, year of publication, tumor type, country of study population, source of control, genotyping method, allele or genotype frequencies, and p value for Hardy-Weinberg equilibrium (HWE) of controls. The patient ethnicities were categorized as Caucasian, Asian or African. Publications that included more than 1 polymorphism were considered as separate studies. When more than 1 article was published by an author, the study with the largest number of participants was included.
Statistical analysis
In this meta-analysis, ORs with 95% CIs were calculated to assess the strength of the MDM4 SNPs and cancer susceptibility. The pooled ORs were calculated for allele comparison, dominant and recessive models, homozygote comparison and heterozygote comparison. The Z test was performed to estimate the significance of the pooled ORs, and p<0.05 was considered statistically significant. The HWE of each study describing control subjects was evaluated by the chi-square test at a significance level of p<0.05. The chi–square-based I2 test and the p value of the Q test were used to evaluate the heterogeneity of the studies. As a guide, p>0.05 indicated a lack of heterogeneity, I2 of <25% indicated low heterogeneity, I2 ranging from 25% to 75% indicated moderate heterogeneity, and I2 of >75% indicated high heterogeneity. If p>0.05, the pooled ORs were calculated using a fixed-effects model. Otherwise, a random-effects model was used. Furthermore, sensitivity analyses were performed by sequentially removing each eligible study. Publication bias of the literature was assessed with Begg's funnel plots. P<0.05 was considered representative of statistically significant publication bias. All p values were 2-sided, and statistical analyses were conducted using the STATA software version 12.0 (StataCorp).
Results
Characteristics of studies
Following the search strategy, 75 articles in PubMed, 26 articles in Ovid, and 22 articles in CNKI were identified. After removal of duplicates, 97 potentially relevant studies were retrieved. Based on the inclusion and exclusion criteria, 90 studies were excluded and 7 were included in this meta-analysis (14–20). Three case-control studies involved 2 different geographical regions (15, 18, 19), so they were considered as separate studies. The flow diagram is shown in Figure 1. A total of 19,328 patients and 51,058 controls were included. The characteristics of the included studies are summarized in Table I.

Flowchart of study identification.
Main characteristics of included studies in the meta-analysis
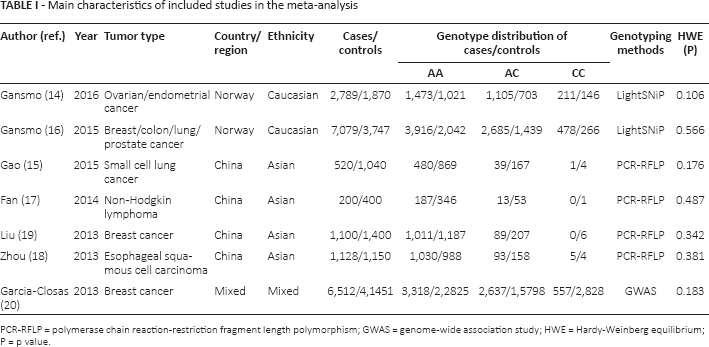
PCR-RFLP = polymerase chain reaction-restriction fragment length polymorphism; GWAS = genome-wide association study; HWE = Hardy-Weinberg equilibrium; P = p value.
Quantitative synthesis
Five different genetic models (allele, dominant, recessive, homozygote and heterozygote) were assumed to observe the association between the MDM4 rs4245739 polymorphism and cancer risk. The results are presented in Table II and Figure 2. Significant associations were found in the allele model (C vs. A, OR = 0.715, 95% CI: 0.622-0.821, p = 0.000), dominant model (CC+AC vs. AA, OR = 0.684, 95% CI: 0.563-0.831, p = 0.000), recessive model (CC vs. AC+AA, OR = 1.139, 95% CI = 1.055-1.230, p = 0.001) and heterozygote model (AC vs. AA, OR = 0.687, 95% CI = 0.568-0.832, p = 0.000). No significant associations were found in the homozygote comparison (CC vs. AA: OR = 1.063, 95% CI: 0.842-1.342, p = 0.608). Considering that geographical region and cancer type might bias the overall results, subgroup analyses were conducted according to these covariates in order to enhance the reliability and stability of our meta-analysis. With regard to cancer types, breast cancer failed to show any significant association. With regard to geographical region, 2 genetic models (allele and heterozygote) showed significant associations in Chinese populations.
Stratified analyses of the MDM4 SNP34091 A>C polymorphism on cancer risk
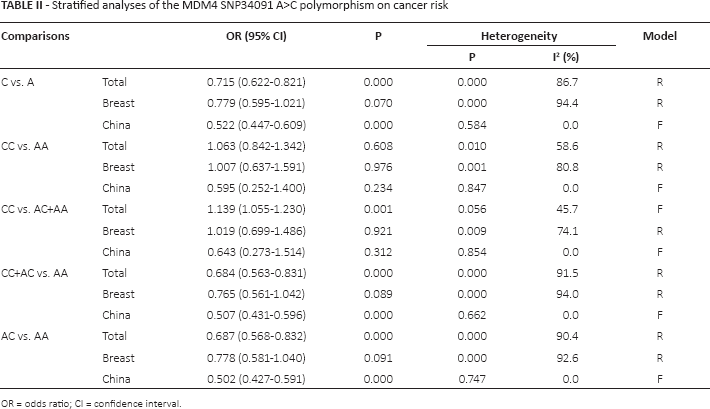
OR = odds ratio; CI = confidence interval.
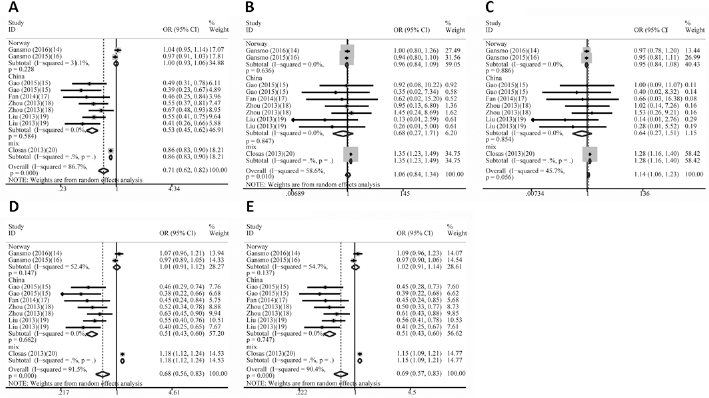
Forest plot for MDM4SNP34091 A>C polymorphism and cancer susceptibility. (A) = allele model; (B) = homozygote model; (C) = recessive model; (D) = dominant model; (E) = heterozygote model.
Heterogeneity and sensitivity analyses
Heterogeneity was evaluated using the chi–square-based I2 test; the results showed that heterogeneity existed in most of the overall comparisons and specific subgroup analyses (Tab. II). However, in the subgroup analysis by region, no significant heterogeneity was found in the China group. Sensitivity analysis was conducted to verify the effect of each study on the overall OR. Studies were sequentially omitted from the equation. The results showed that the pooled ORs of the 6 polymorphisms were not materially altered by the contribution of any individual study. These results showed that the meta-analysis was statistically robust (Fig. 3).
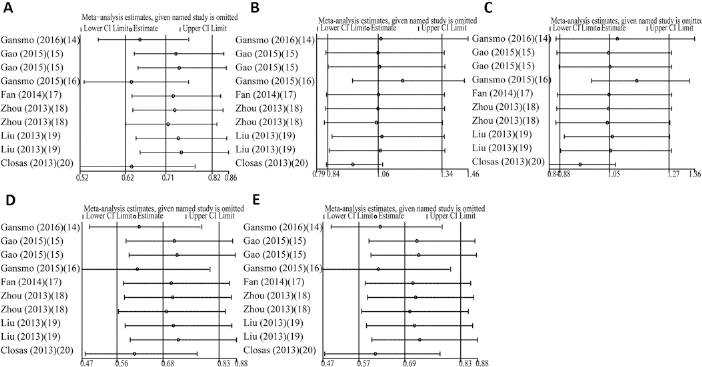
Sensitivity analyses in overall cancers. (A) = allele model; (B) = homozygote model; (C) = recessive model; (D) = dominant model; (E) = heterozygote model.
Publication bias
Publication bias in all of the studies was assessed by Begg's funnel plot and Egger's test. Publication bias was found in 3 models (p = 0.025 for C allele vs. A allele; p = 0.000 for AC vs. AA; p = 0.000 for CC+AC vs. AA). For the remaining 2 models the graphical funnel plots were symmetrical and no publication bias was present (p = 0.161 for CC vs. AA; p = 0.168 for CC vs. AC+AA; Fig. 4). In the subgroup analysis based on geographical region, no publication bias was found.
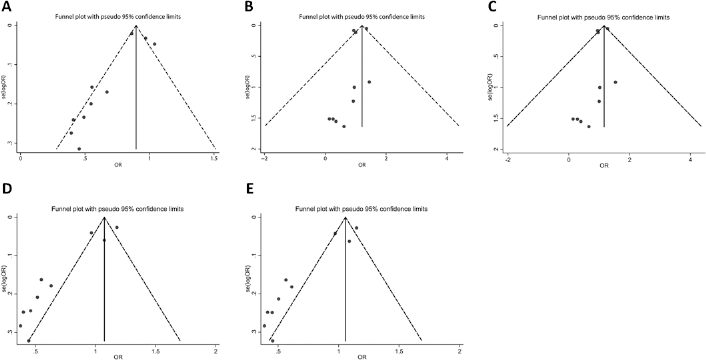
Funnel plot for publication bias test. (A) = allele model; (B) = homozygote model; (C) = recessive model; (D) = dominant model; (E) = heterozygote model.
Discussion
The TP53 tumor suppressor gene is a commonly mutated gene in human cancers (21). This gene encodes a DNA-binding transcription factor that regulates sequence-specific target genes involved in cell growth inhibition, senescence and apoptosis, providing a powerful intrinsic defense against cancer (22). MDM4 is a transcriptional regulator that binds to and inhibits p53 (6). Gansmo et al (16) found that the MDM4 SNP34091 (rs4245739) polymorphism was not associated with risks for colon, lung and prostate cancer. However, Fan et al (17) found a significantly decreased non-Hodgkin lymphoma risk among carriers of the MDM4 rs4245739 C allele in Chinese. This research pooled these studies together. This is the first meta-analysis to detect the association between MDM4 rs4245739 polymorphism and cancer risk. In this meta-analysis, the association between MDM4 polymorphism and cancer was observed in the overall comparison and in specific subgroup analyses based on geographical region and cancer site.
The reason why the MDM4 rs4245739 polymorphism affects the cancer risk may be its influence on the miRNA binding site. This polymorphism creates a putative target site for miR-191-5p and miR-887-3p, such that MDM4 expression is affected by both miRNAs. miR-191-5p is overexpressed in osteosarcoma, colon cancer, and pancreatic cancer (23–25), but its expression is relatively lower in prostate cancer (26). The tissue-specific expression of miRNA may explain the different roles of the MDM4 rs4245739 polymorphism in cancer.
Considering the complexity of tumor etiology, we conducted subgroup analyses in this study. However, only the breast cancer subgroup and the China subgroup were analyzed because of the limited number of samples. A significant association was observed in the overall comparison except for the breast cancer group, where no significant associations were observed. This trend may be attributed to the different genetic backgrounds and risk factors of different cancers.
Heterogeneities between studies were found in this meta-analysis. Ethnicity is one of the most important factors affecting heterogeneity. In this gene polymorphism site, completely different genotype frequencies in Chinese and Norwegians were observed. In the subgroup analysis of China, no significant heterogeneities were found. In addition, tumor sites and characteristics also contribute to the heterogeneity.
This meta-analysis has some limitations. First, we must note that a publication bias was found in 3 model comparisons. So the results from these 3 models must be treated with caution. Second, the analysis did not consider gene environment interactions due to insufficient data. Finally, the data for each type of cancer were not sufficient, so future studies comprising larger subgroup populations are required.
In conclusion, the results of our meta-analysis show that the MDM4 SNP34091 A>C polymorphism may function as a protective factor against the cancer risk. Further studies with different ethnicities and large population sizes should be conducted to reach a comprehensive conclusion.
Footnotes
Financial support: None.
Conflict of interest: The authors declare that there is no conflict of interest regarding the publication of this paper.
