Abstract
Background:
Non-Hodgkin Lymphoma (NHL) is an increasingly prevalent hematological malignancy in Egypt, highlighting the need for a better understanding of its genetic risk factors. The TP53 and MDM4 genes play critical roles in cellular homeostasis and cancer development. This study aimed to assess the frequency of the TP53 (SNP rs1042522) Arg72Pro and MDM4 (SNP rs4245739) A > C polymorphisms as potential risk factors for NHL in adult Egyptian patients.
Methods:
A case-control study was conducted involving 80 adult NHL patients and 100 control age- and sex-matched healthy controls. Genotyping for the TP53 (rs1042522) Arg72Pro and MDM4 (rs4245739) A > C polymorphisms was performed using polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) method.
Results:
A significant association was found between the homozygous TP53 Pro/Pro genotype and increased susceptibility to NHL (47.5% in patients vs 4.0% in controls; P < .001), as well as a higher frequency of the mutant C allele among NHL cases (63.8% vs 28.0%; P < .001). In contrast, no significant association was observed between MDM4 polymorphisms and NHL risk. In addition, analysis of treatment outcomes revealed no statistically significant differences in overall survival or progression-free survival based on TP53 or MDM4 genotypes.
Conclusions:
These findings suggest that the TP53 Arg72Pro polymorphism is a significant genetic marker for NHL susceptibility in the Egyptian population, while MDM4 polymorphisms do not appear to contribute to disease risk. Further studies are warranted to elucidate the genetic mechanisms underlying NHL and to explore their implications for risk stratification and therapeutic strategies.
Keywords
Introduction
Non-Hodgkin Lymphoma (NHL) encompasses a broad category of hematological cancers, characterized by the uncontrolled proliferation of lymphocytes and diverse clinical and biological features. It is the seventh most commonly reported cancer in the United States,1,2 and its incidence has steadily increased over the past few decades. 3
In Egypt, NHL accounts for approximately 10.9% of all cancer diagnoses (around 7305 new cases annually), making it the fourth most common cancer in the country, following liver, breast, and bladder cancers. 4 Diffuse large B-cell lymphoma (DLBCL) is the most prevalent NHL subtype, representing about 53.6% of cases, followed by marginal zone lymphoma (6.75%), small lymphocytic lymphoma (6.7%), follicular lymphoma (2.1%), and the Burkitt lymphoma (1.3%). 5
Tumor suppressor genes play a crucial role in preventing the development and progression of malignancies. They regulate cell growth and maintain genomic stability. Mutations in these genes can lead to uncontrolled cell proliferation and an increased risk of cancer. 6 Among them, the TP53 gene is one of the most critical, often referred to as the “guardian of the genome” due to its roles in regulating the cell cycle, repairing damaged DNA and inducing apoptosis.7,8
More than half of all human cancers harbor genetic alterations in TP53 gene. Mutations and polymorphisms in TP53 can disrupt normal cell regulation, contributing to carcinogenesis. 9 Specifically, single nucleotide polymorphisms (SNPs) in TP53 have been associated with an increased risk of various cancers, including NHL. 10
Most TP53 variants are missense mutations that result in the substitution of a single amino acid. 11 One of the most studied TP53 SNPs is the Arg72Pro variant, which has been linked to elevated cancer risk, particularly among Asian and American individuals carrying the 72 Pro allele.12,13 Furthermore, Chinese individuals with the 72 Pro allele have shown an increased risk of developing NHL. 14
Mouse Double Minute 4 (MDM4) is a negative regulator of P53, binding to its transcriptional activation domain and suppressing the transcription of p53 target genes involved in cell cycle arrest and apoptosis. This inhibition may contribute to lymphomagenesis by promoting the survival and proliferation of genetically abnormal cells. Genetic variants in MDM4, particularly SNPs in its promoter region, have been shown to modulate TP53 activity and may influence cancer susceptibility. 15
Several SNPs in MDM4 have been identified; among them, the rs4245739 A > C polymorphism in the 3’-untranslated region (3’-UTR) of MDM4 creates a binding site for microRNA-191 (miR-191). The C allele enhances miR-191 binding effectively to MDM4 mRNA, leading to reduced MDM4 expression and, consequently, increased p53 activity. This regulatory mechanism may lower the risk of developing NHL.16,17
The interaction between TP53 and MDM4 is crucial for maintaining the balance between cell proliferation and apoptosis, and disruptions in this regulatory axis are frequently observed in lymphomagenesis. 16 Recent studies have underscored the pivotal roles of the tumor suppressor gene TP53 and its negative regulator MDM4, in the pathogenesis of various cancers, including lymphomas.18,19
This study aims to investigate the contribution of TP53 and MDM4 genetic polymorphisms as potential risk factors for NHL among adult Egyptian patients. By elucidating these genetic associations, we hope to enhance understanding of NHL etiology in this population, potentially informing future therapeutic and preventive strategies.
Materials and Methods
Ethical approval
Ethical approval for this study was obtained from the Institutional Review Board of the National Cancer Institute, Cairo University, Egypt (approval number: IRB; 00004025). All participants, including individuals in the control group, provided written informed consent. The study was conducted in accordance with ethical principles outlined in the Declaration of Helsinki.
Patient selection
This case-control study included 80 consecutive patients diagnosed with NHL who presented to the Adult Oncology Department of the National Cancer Institute (NCI), Cairo University, between 2018 and 2022. The control group comprised 100 healthy bone marrow donors, matched for age and sex, recruited from the Bone Marrow Transplantation Unit. These individuals underwent standard procedures, including bone marrow aspiration, to exclude any occult hematological malignancies.
The reporting of this case-control study conforms to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines (see Supplementary File for the completed STROBE checklist).
Exclusion criteria include prior cancer treatment and co-occurrence with other neoplastic conditions. The NHL diagnoses were confirmed using standard clinical and pathological criteria. Peripheral blood samples were collected from patients prior to the initiation of any therapeutic intervention. Demographic and clinical data, including age, sex, diagnosis, treatment history, and disease status, were extracted from patient medical records.
Patients were followed for a minimum of 24 months post-treatment to evaluate treatment outcomes including resistance and relapse and to determine overall survival (OS) and disease-free survival (DFS). All patients received treatment in accordance with standardized protocols at NCI, Cairo University.
Comprehensive clinical assessments, including medical history, physical examination, computed tomography (CT) scans, and laboratory investigations (complete blood count [CBC], lactate dehydrogenase [LDH], beta-2 microglobulin, bone marrow aspirate [BMA], and bone marrow biopsy [BMB]), were performed for all subjects.
Sample collection and DNA extraction
Peripheral blood samples (3-5 mL) were collected from both patients and controls into EDTA-coated tubes. Samples were either processed immediately for DNA extraction or stored at −20°C for later use. Genomic DNA was extracted from peripheral blood leukocytes using the QIAamp DNA Blood Mini Kit (QIAGEN, Hilden, Germany; Cat No. 51104), following the manufacturer’s standard protocol.
Genotyping of polymorphisms
Genotyping of the TP53 rs1042522 (Arg72Pro) and MDM4 rs4245739 A > C polymorphisms was performed using polymerase chain reaction (PCR) followed by restriction fragment length polymorphism (RFLP) analysis. Each PCR reaction was carried out in a total volume of 25 µL, consisting of 12.5 µL of Dream Taq PCR Master Mix (2X; Thermo Scientific, Lithuania; Cat No. K1071), 1 µL of each primer, 5 µL of template DNA, and 5.5 µL of distilled water.
The thermal cycling conditions for the MDM4 gene included an initial denaturation at 95°C for 2 minutes, followed by 35 cycles of denaturation at 94°C for 30 seconds, annealing at 58°C for 30 seconds, and extension at 72°C for 30 seconds, with a final extension step at 72°C for 10 minutes. For the TP53 gene, after an initial denaturation at 94°C for 2 minutes, 30 cycles were performed at 94°C for 30 seconds (denaturation), 60°C for 30 seconds (annealing), and 72°C for 30 seconds (extension), followed by a final extension at 72°C for 10 minutes.
Primers used were as follows:
TP53 forward: 5’-TTGCCGTCCCCAAGCAATGGATGA-3’; reverse: 5’-TCTGGGAAGGGACAGAAGATGAC-3’.
MDM4 forward: 5’-AAGACTAAAGAAGGCTGGGG-3’; reverse: 5’-TTCAAATAATGTGGTAAGTGACC-3’.
For TP53 gene genotyping, 10 µL of the PCR product was digested with 0.5 to 2 µL of BstUI (AccII) restriction enzyme (Thermo Scientific; Cat No. FER.921), along with 2 µL of 10× buffer and 16 µL of nuclease-free water. The mixture was incubated overnight at 37°C. The presence of the polymorphic allele abolishes the BstUI restriction site, producing distinct band patterns:
Homozygous mutant (CC): 113 bp and 86 bp bands
Heterozygous mutant (CG): 199 bp, 113 bp, and 86 bp bands.
Wild-type (GG): a single 199 bp band (Figure 1A).

Identification of MDM4 A > C and TP53 Arg72Pro polymorphism genotypes by gel electrophoresis.
For MDM4 genotyping, 10 µL of PCR product was digested with 1 µL of FastDigest MspI enzyme (Thermo Scientific; Cat No. FD0554), 2 µL of 10× buffer, and 17 µL of nuclease-free water. Digestion was performed at 37°C for 5 minutes. The MspI site produces the following patterns:
Homozygous wild-type (AA): 134 bp band.
Heterozygous (AC): 134 bp, 111 bp, and 23 bp bands.
Homozygous (CC): 111 bp and 23 bp bands (Figure 1B).
The PCR products were resolved on 2% agarose gels stained with ethidium bromide and visualized under UV light.
Statistical analysis
Sample size was calculated using Epi Info software, based on detecting an odds ratio (OR) of 2.18 with 90% confidence and a 1:1 case-to-control ratio. The expected prevalence in the control group was set at 38%, representing the minor allele frequency (MAF) of the TP53 rs1042522 polymorphism in the general population. 10 A relative precision of 40% was considered acceptable for this exploratory study. Based on these parameters, the required sample size for the control group was 87 subjects, yielding a total required study population of 174 participants.
Data management and statistical analysis were conducted using SPSS (version 20). Categorical variables were summarized using percentages, while numerical variables were expressed as means ± standard deviations or medians with ranges, as appropriate. The Chi-square test was employed to assess associations between clinical variables and the TP53 (rs1042522) and MDM4 (rs4245739) polymorphisms.
Overall survival was defined as the time from diagnosis to death from any cause, with patient still alive at last follow-up considered censored.
Disease-free survival (DFS) was defined as the time from complete remission (CR) to documented relapse or death, with patients without relapse censored at the date of the last follow-up. The OS and DFS were estimated using the Kaplan-Meier survival analysis, and survival curves were compared using the log-rank test. Statistical significance was defined as a 2-sided alpha level of .05, with 95% confidence intervals. 20
Results
Clinical and laboratory characteristics of non-Hodgkin lymphoma patients
This study included 80 patients with NHL, aged between 18 and 79 years, with a mean age of 47.2 ± 15.1 years. The patient group was nearly evenly distributed, comprising 39 males (48.8%) and 41 females (51.2%), yielding a male-to-female ratio of approximately 1:1.05. The control group consisted of 100 individuals—60 males (60.0%) and 40 females (40.0%), aged between 18 and 79 years, with a mean age of 43.8 ± 13.6 years. Patients and controls were matched by age and sex. Detailed patient characteristics are presented in (Table 1).
Clinical and laboratory characteristics of 80 NHL patients.
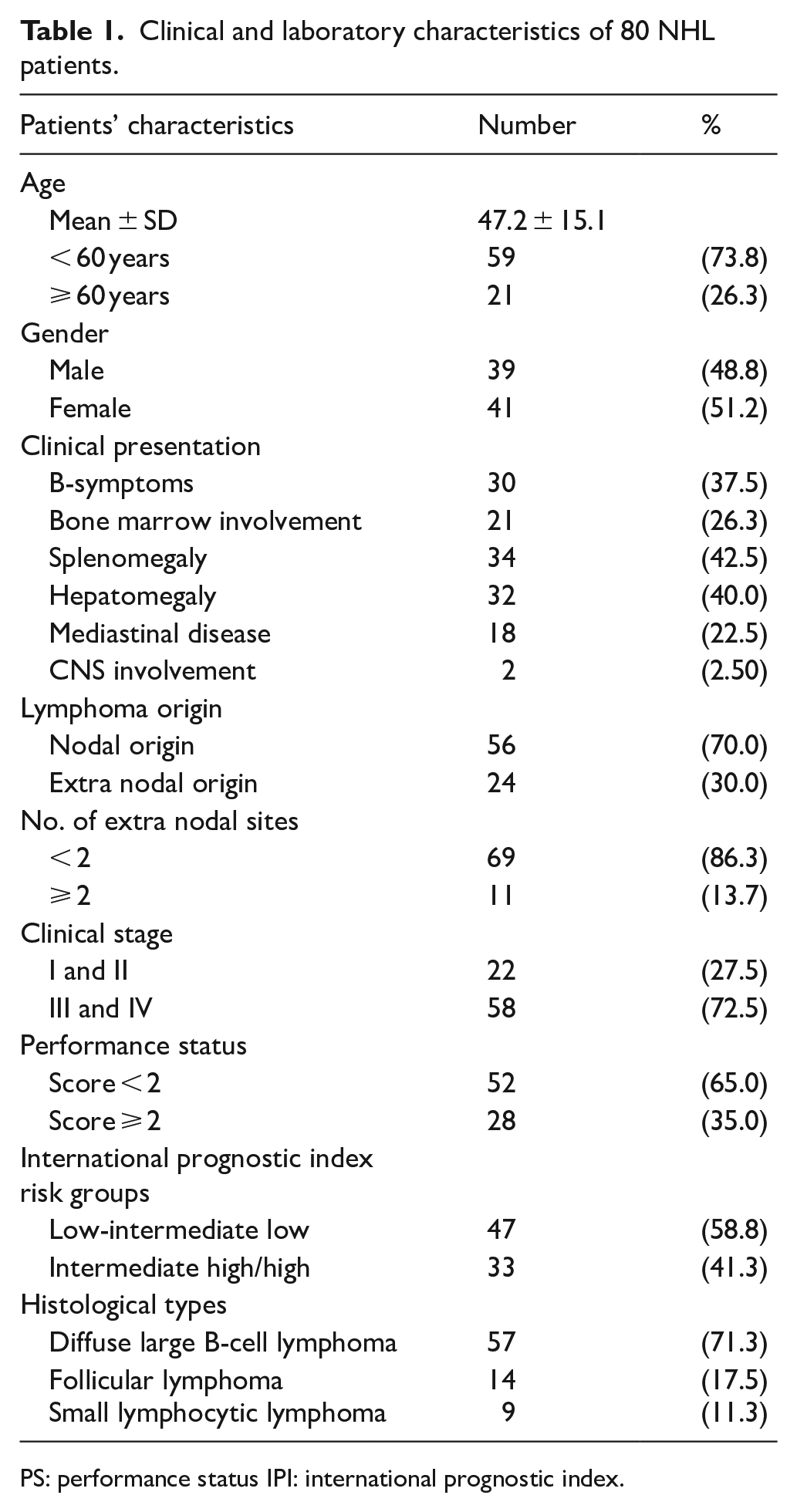
PS: performance status IPI: international prognostic index.
Among the NHL patients, 63 (78.7%) achieved CR following treatment; however, 49.2% of them experienced relapse. Seventeen patients (21.2%) were resistance to treatment. By the end of the study, 60.0% of the patients had died due to the disease, while 40.0% were still alive (Supplementary Table 1).
Genotyping of MDM4 and TP53 genes in non-Hodgkin lymphoma patients and controls
We examined the frequencies of the TP53 (rs1042522, Arg72Pro) and MDM4 (rs4245739, A > C) polymorphisms in NHL patients and compared them with those in the control group. The rs4245739 SNP in the MDM4 gene involves the A and C alleles, with A being the ancestral variant. This polymorphism is located on chromosome 1 (1q32.1) and represent a non-coding transcript variant (Supplementary Table 2 and Figure 1A).
The rs1042522 SNP in the TP53 gene involves the C and G alleles, with G being ancestral. It is located on chromosome 17 (17p13.1) and results in a codon change from CGC to CCC, leading to a substitution of arginine with proline (Supplementary Table 2 and Figure 1B).
Genotype distribution and adjusted ORs are presented in Table 2. The homozygous genotype of TP53 was significantly more frequent in NHL patients (47.5%) than in the controls (4.0%) (P < .001, odds ratio [OR] = 7.27, 95% CI = 3.91-13.53). Mutant genotypes (CG and CC) was observed in 80.0% of patients compared with 52.0% of controls (P < .001, OR = 2.24, 95% CI = 1.49-3.35) (Table 2).
Genotype distribution of MDM4 rs4245739 A > C and TP53 rs1042522 Arg72Pro polymorphism in 80 NHL patients and 100 controls.

Significant P-value ⩽ .05.
Bold values represent significant values.
Furthermore, the mutant C allele of TP53 was significantly more prevalent in patients (63.8%) than in controls (28%) (P < .001, OR = 4.52, 95% CI = 2.89-7.06). However, no statistically significant differences were found in the allele or genotype frequencies of MDM4 (rs4245739, A > C) between the patient and controls (Table 2).
The TP53 CC genotype was significantly more frequent in patients with diffuse large B-cell lymphoma (DLBCL; 49.1%), follicular lymphoma (FL; 42.9%), and small lymphocytic lymphoma (SLL; 44.4%) compared with controls (4.0% in each comparison; P < .001, P < .001, and P = .001, respectively). The mutant C allele of TP53 was also significantly more frequent in patients with DLBCL (64.0%), FL (64.3%), and SLL (61.1%) compared with controls (28.0%) (P < .001, P = .020, and P = .006 respectively; Table 3).
Relationship between MDM4 rs4245739 A > C and TP53 rs1042522 Arg72Pro polymorphism and NHL histological subtypes.

No significant associations were observed between MDM4 polymorphisms and NHL subtypes, except in FL patients, where the homozygous CC genotype was more prevalent (14.3%) compared with controls (1.0%) (P = .010). However, this may be a chance finding due to small sample size (Table 3).
Association of MDM4 and TP53 polymorphisms with patient characteristics
The heterozygous (AC) and homozygous (CC) genotypes of the MDM4 gene were significantly associated with increased frequencies of splenomegaly and lower hemoglobin levels compared with the wild-type (AA) (P = .048 and P = .040, respectively). However, we cannot rely on these findings due to small sample size. No other significant associations were found between MDM4 genotypes and clinical characteristics (Table 4).
Characteristics of 80 NHL patients according to genotype distribution of MDM4 rs4245739 A > C and TP53 rs1042522 Arg72Pro polymorphism.

Association of MDM4 and TP53 polymorphisms with treatment response and outcomes
Using rs4245739 AA and rs1042522 GG as reference genotypes, no significant differences were observed in the distribution of MDM4 or TP53 genotypes or alleles between patients who achieved CR and those who did not, indicating no apparent influence on treatment response (Table 5). Similarly, no significant differences were detected between relapsed patients and those who remained in remission (Supplementary Table 3).
Response to therapy according to genotype distribution of MDM4 rs4245739 A > C and TP53 rs1042522 Arg72Pro polymorphism in 80 NHL patients.

OR, odds ratio; CI, confidence interval.
Overall survival and progression-free survival in relation to MDM4 and TP53 polymorphisms
Survival analyses revealed a 3-year OS rate of 39.2%, with a mean OS of 20.6 months (95% CI = 17.5-23.6) among all NHL patients. The 3-year progression-free survival (PFS) rate was 22.2%, with a mean PFS of 13.6 months (95% CI = 10.2-17.0) (Supplementary Table 4).
For TP53 genotypes, the mean 3-year OS was 50 months for wild-type, 30.8 months for heterozygous, and 40.5 months for homozygous genotypes (P = .282). Corresponding PFS values were 28.6, 29.2, and 18.1 months, respectively (P = .87) (Figure 2).
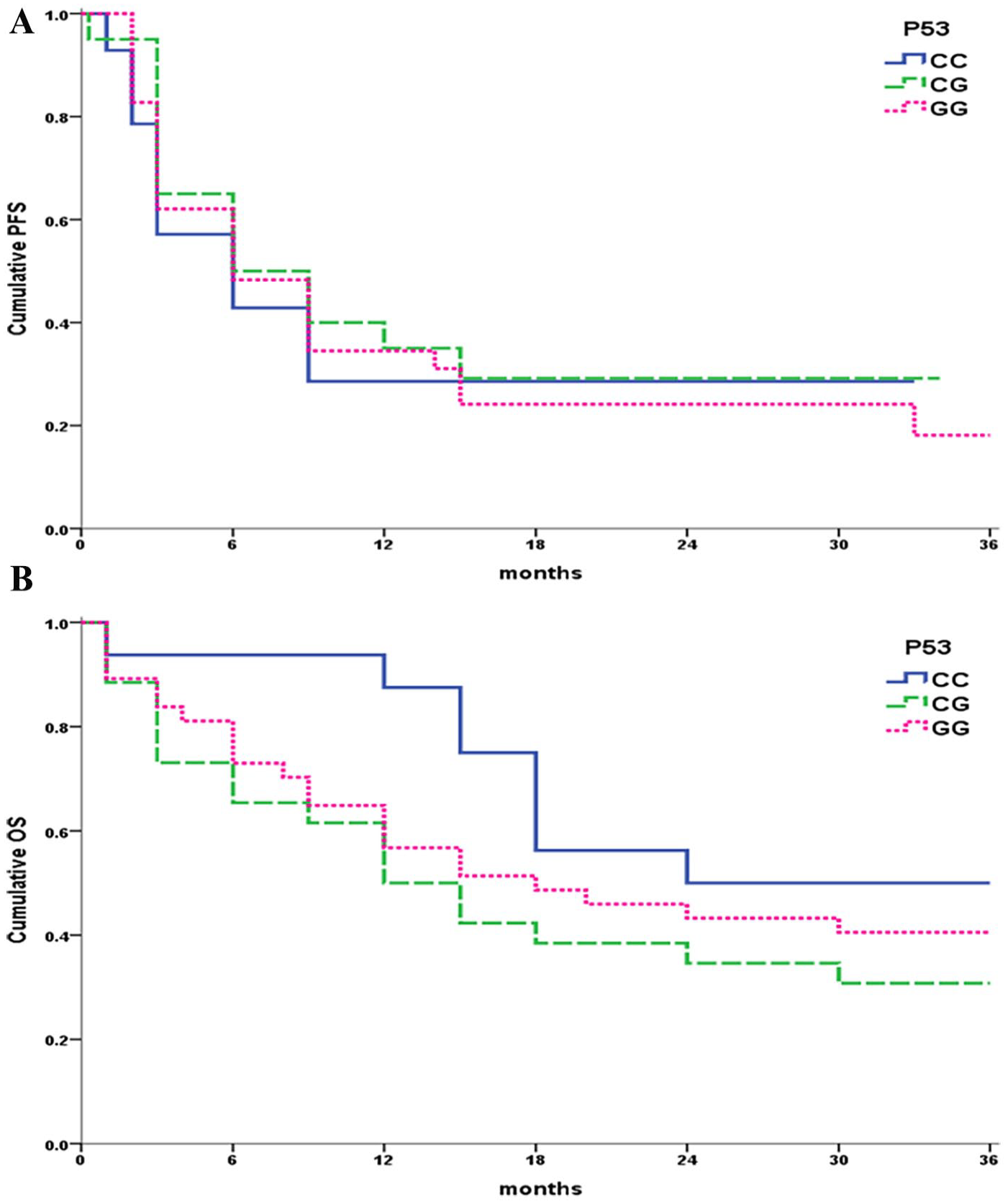
OS and PFS according to TP53 genotypes in 80 NHL cases.
For MDM4 genotypes, the mean 3-year OS was 42.5 months for the wild-type, 38.2 months for heterozygous, and 20 months for homozygous genotypes (P = .312). The PFS values were 32.1, 13.3, and 25 months, respectively (P = .636) (Figure 3). No statistically significant differences were found in OS or PFS between patients with wild-type and mutant genotypes of TP53 and MDM4.

OS and PFS according to MDM4 genotypes in 80 NHL cases.
Logistic regression analysis
Logistic regression analysis demonstrated that the TP53 CG + CC genotypes were significantly associated with an increased risk of NHL occurrence (P < .001; Table 6). No other genotypes, including those of MDM4, were significant predictors of treatment response or survival (Table 6).
Regression analysis for prediction of NHL occurrence, remission, and relapse using multiple confounders.
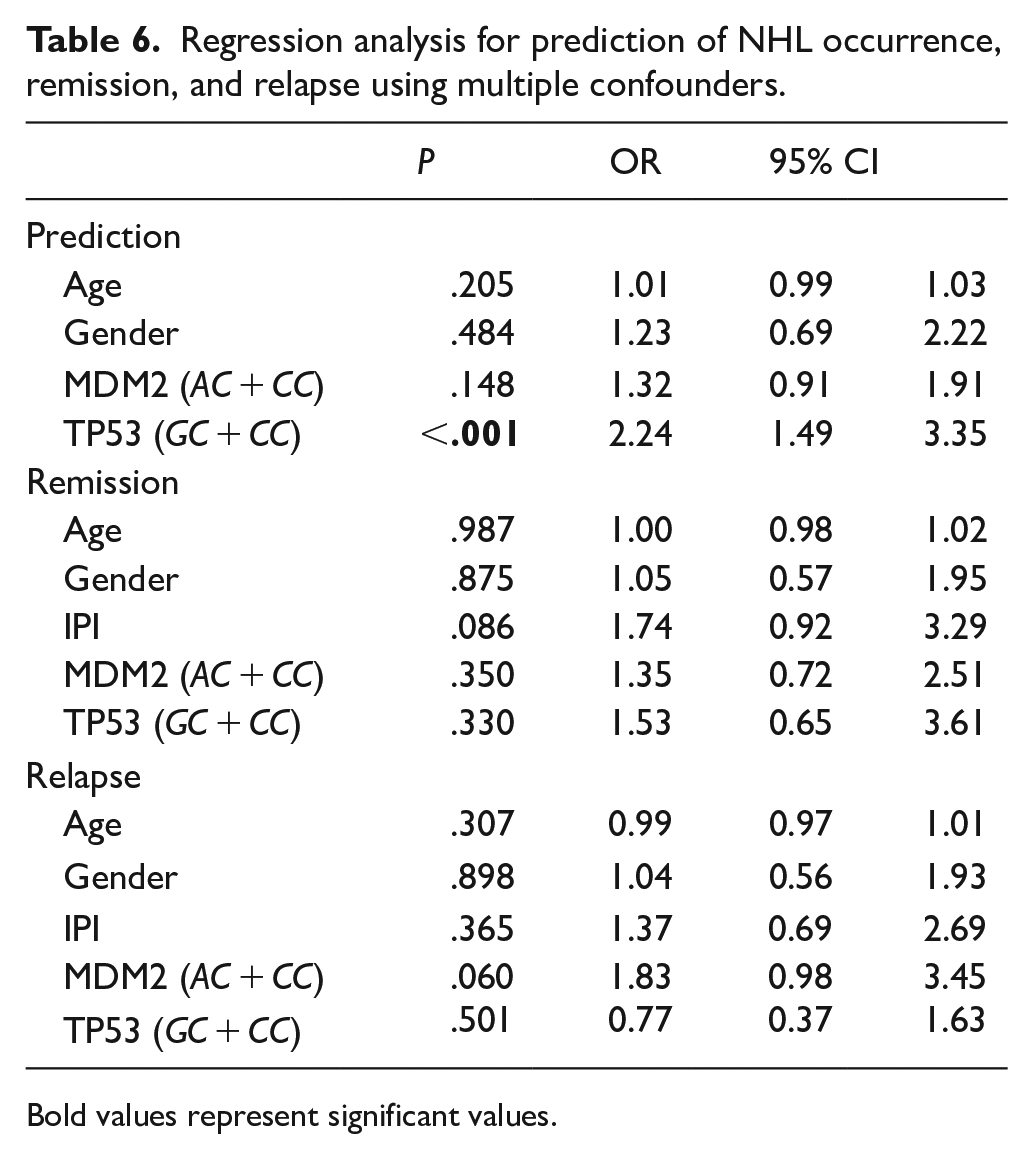
Bold values represent significant values.
Discussion
The TP53 gene is a tumor suppressor gene that plays a critical role in preventing the development and progression of cancer. Polymorphism in TP53 can affect its function and may have implications for various types of cancer, including lymphoma. One of the most common TP53 polymorphisms is rs1042522, also known as the codon 72 polymorphism or TP53 Arg72Pro. 21
The current study provides insights into the genetic and clinical landscape of NHL in an Egyptian population, with a focus on polymorphisms in TP53 (rs1042522, Arg72Pro) and MDM4 (rs4245739, A > C). Our findings highlight a significant association between these polymorphisms and susceptibility to NHL, although their clinical relevance appears limited. To our knowledge, this is the first study to investigate these polymorphisms in the Egyptian population.
The TP53 gene, known for its tumor suppressor functions, is crucial for regulating apoptosis, maintaining genomic stability, and controlling cell cycle progression. In this study, the TP53 rs1042522 polymorphism, particularly the homozygous CC genotype, was significantly associated with an increased risk of developing NHL (P < .001, OR = 7.27). Moreover, mutant genotypes (CG and CC) were present in 80.0% of patients compared with 52.0% of controls (P < .001), supporting the hypothesis that TP53 mutations contribute to NHL pathogenesis. These findings are consistent with earlier studies that have demonstrated an association between TP53 polymorphisms, especially the Arg72 variant, and increased susceptibility to cancer, particularly hematological malignancies.22,23
Often referred to as the “guardian of the genome,” the TP53 gene plays a vital role in regulating the cell cycle and preserving genomic integrity. The Arg72Pro polymorphism is known to affect protein function, influencing both apoptosis and DNA repair mechanisms. 24 The increased frequency of the Pro/Pro genotype in our NHL cohort suggests a potential predisposition to lymphomagenesis, likely due to an impaired apoptotic response and increased genomic instability. Previous studies have linked the Pro variant to reduced apoptosis in response to stress, 25 which may contribute to the malignant transformation of lymphoid cells and the associated risk of developing NHL.
The presence of the mutant C allele in 63.8% of patients compared with 28.0% of controls (P < .001, OR = 4.52) further supports the role of TP53 in NHL development. This allele has previously been implicated in cancer progression due to its potential to impair apoptosis and weaken tumor suppression. Our findings are consistent with previous studies that have reported similar associations between TP53 mutations and aggressive forms of NHL.22,26
Our results align with prior literature suggesting that alterations in the TP53 gene contribute to the risk of various malignancies, including lymphomas. 27 Notably, the high prevalence of the C allele in our NHL patients indicates that this polymorphism may serve as a genetic marker for NHL susceptibility, warranting further investigation into its underlying mechanisms.
Consistent with our observation, a case-control study conducted by Kim et al 13 involving 945 cases and 1700 controls reported a significantly higher risk of NHL associated with the homozygous TP53 Pro/Pro genotype in the Korean population (P = .040). Similarly, Fan et al 11 found that both Pro/Pro and Arg/Pro genotype were associated with a markedly higher risk of NHL compared with the Arg/Arg wild genotype (P = .002 and P = .007, respectively).
Furthermore, a meta-analysis by Weng et al 28 assessed the contribution of the TP53 Arg72Pro polymorphism to the development of hematological malignancies. They observed a significantly increased risk of NHL associated with the heterozygous genotype (Arg/Pro vs Arg/Arg: OR = 1.18, 95% CI = 1.02-1.35) and the dominant genotype (Arg/Pro + Pro/Pro vs Arg/Arg: OR = 1.19, 95% CI = 1.03-1.34). These results are in line with our findings, suggesting that Arg72Pro polymorphism may contribute to NHL susceptibility. In contrast, studies conducted in European Caucasian populations reported no significant association between the Arg72Pro polymorphism and NHL risk,29,30 possibly due to ethnic variations influencing genetic susceptibility.
Regarding the effect of Pro variant in other cancer types, Ounalli et al 31 reported an increased risk of chronic lymphocytic leukemia (CLL) associated with this variant. It has also been implicated in heightened risk for osteosarcoma, melanoma, thyroid, and prostate cancer. However, no significant association has been found between the Pro variant and ovarian or glioma tumors. 26
In our study, no significant association was detected between TP53 Arg72Pro polymorphism and clinical parameters, laboratory findings, treatment response, or survival outcomes. Consistent with our findings regarding survival, Bittenbring et al 30 in a study on European Caucasians reported that the TP53 Arg72Pro had no effect on survival outcomes in patients with DLBCL. Conversely, Chiappella et al 32 found that DLBCL patients harboring TP53 mutation had lower response rate to chemo-immunotherapy and poorer outcomes in terms of OS and PFS.
In contrast, Liu et al 33 reported that among patients treated with CHOP or CHOP-like therapy, those with the Arg/Arg and Arg/Pro genotypes had a significantly higher 5-year OS rate compared with those with the Pro/Pro genotype (P = .001). In addition, the Arg/Arg and Arg/Pro genotypes were associated with a better complete response (CR) rate (P = .007) than the Pro/Pro genotype. Ethnicity may influence the penetrance of the TP53 Arg72Pro polymorphism in cancer susceptibility, which could explain these observed disparities.12,13
The MDM4 gene negatively regulates the TP53 protein. Single nucleotide polymorphisms in the MDM4 gene can affect its activity and consequently influence tumor susceptibility and prognosis. 34 In this study, we also investigated the association between the MDM4 rs4245739 A > C polymorphism and the risk of developing NHL.
Three genotypes―wild-type AA, heterozygous AC, and homozygous CC—were observed in both the patient and control groups. Our analysis revealed no statistically significant differences in genotype frequencies between NHL patients and controls. In this context, numerous studies have reported varying and sometimes conflicting results.
Contrary to our findings, Fan et al 11 demonstrated that carriers of the MDM4 rs4245739C allele had a significantly lower risk of developing NHL compared to those with the A allele. A meta-analysis by Chen et al 34 also reported that the rs4245739 polymorphism is associated with both NHL and ovarian cancer and may confer a reduced risk of cancer, particularly in Asian populations. Moreover, among various cancer types, only esophageal squamous cell carcinoma was significantly associated with the rs4245739 polymorphism.
Several studies have found that the rs4245739 A > C polymorphism is significantly associated with reduced cancer risk.11,35 -37 However, other studies have reported minimal or no effect of this polymorphism on cancer susceptibility.38 -40 Xu et al 17 noted that no pooled analysis had been conducted specially for NHL, gastric, colon, or prostate cancers, which may hinder comprehensive evaluation of this polymorphism’s clinical significance.
MDM4 is known to inhibit TP53 function, and its dysregulation has been implicated in tumorigenesis. 41 However, the lack of correlation observed in our study suggests that MDM4 may not play a crucial role in NHL susceptibility within our sample population. This finding aligns with previous studies that have reported inconsistent associations between MDM4 polymorphisms and various cancer types, indicating a complex interplay of genetic factors in lymphoma development.
In this study, we also assessed treatment outcomes in a cohort of patients with different genotypes of TP53 and MDM4, focusing on their association with CR, OS, and PFS. Our results revealed a high CR rate of 78.7%, with 21.3% of patients exhibiting resistance to treatment. While the overall response was favorable, we found no statistically significant associations between TP53 and MDM4 genotypes and treatment outcomes, raising important questions regarding the influence of genetic variants on therapeutic response.
Previous studies have suggested that TP53 mutations are often associated with poor clinical outcomes in various malignancies, particularly hematologic cancers.42,43 However, our findings demonstrated no significant differences in treatment response across TP53 genotypes. This suggests that, although TP53 mutations may influence tumor biology, they do not appear to be decisive in determining treatment efficacy within our cohort. Specifically, the OS for patients with wild-type, heterozygous, and homozygous TP53 genotype was 50, 30.8, and 40.5 months, respectively (P = .282), indicating no clear prognostic distinction.
Similarly, the MDM4 gene, despite its known role in regulating p53 activity, did not show a significant impact on patient outcomes. The OS for wild-type, heterozygous, and homozygous MDM4 genotypes was 42.5, 38.2, and 20 months, respectively (P = .312), mirroring the trend seen with TP53. These findings are consistent with prior research suggesting that MDM4 alterations do not uniformly predict clinical outcomes in cancer treatment. 44
Limitations
The primary limitation of this study is the relatively small sample size, particularly in the subgroup analyses of clinical outcomes, which was constrained by limited financial resources. In addition, although we examined the role of TP53 and MDM4 polymorphisms, the genetic architecture of NHL is likely to involve a broader range of genetic and epigenetic factors. Future investigations should aim to integrate genomic data with transcriptomic and proteomic profiles to comprehensively explore the molecular determinants of treatment resistance and survival. Such an approach could enhance our understanding of the complex biological mechanisms underlying NHL pathogenesis and progression.
Conclusion
This study underscores a significant association between TP53 polymorphisms and increased NHL risk in an Egyptian population, with the mutant C allele conferring higher susceptibility to the disease. While MDM4 polymorphisms were not associated with NHL risk, they may influence certain clinical parameters; however, their overall prognostic relevance remains uncertain. Further research is warranted to elucidate the molecular mechanisms through which these genetic variants impact NHL and to identify potential therapeutic targets that may improve clinical outcomes. Larger, multi-ethnic cohort studies are essential to validate these findings and to investigate possible gene-environment interactions affecting NHL susceptibility and prognosis.
Supplemental Material
sj-docx-1-onc-10.1177_11795549251352047 – Supplemental material for Contribution of TP53 and MDM4 Genetic Polymorphisms as a Risk Factor in Non-Hodgkin Lymphoma in Adult Egyptian Patients
Supplemental material, sj-docx-1-onc-10.1177_11795549251352047 for Contribution of TP53 and MDM4 Genetic Polymorphisms as a Risk Factor in Non-Hodgkin Lymphoma in Adult Egyptian Patients by Eman A Helal, Naglaa M Hassan, Mahmoud M Kamel, Mahmoud A Amer and Roxan E Shafik in Clinical Medicine Insights: Oncology
Supplemental Material
sj-docx-2-onc-10.1177_11795549251352047 – Supplemental material for Contribution of TP53 and MDM4 Genetic Polymorphisms as a Risk Factor in Non-Hodgkin Lymphoma in Adult Egyptian Patients
Supplemental material, sj-docx-2-onc-10.1177_11795549251352047 for Contribution of TP53 and MDM4 Genetic Polymorphisms as a Risk Factor in Non-Hodgkin Lymphoma in Adult Egyptian Patients by Eman A Helal, Naglaa M Hassan, Mahmoud M Kamel, Mahmoud A Amer and Roxan E Shafik in Clinical Medicine Insights: Oncology
Footnotes
Acknowledgements
The authors extend their sincere gratitude to the patients and control participants, whose cooperation made it possible to carry out this project.
Ethical considerations
This study received ethical clearance from the Institutional Review Board of the National Cancer Institute, Cairo University, Egypt (Ethical Approval Number: IRB; 00004025). All procedures were conducted in accordance with the ethical standards of the Declaration of Helsinki for research involving human participants.
Consent to participate
Written informed consent was obtained from all participants (patient and control) after the study objectives were explained and prior to blood sample collection. Confidentiality of all participant data was strictly maintained.
Consent for publication
NA.
Author contributions
EAH, NMH, and RES contributed to project preparation, study design, and drafting of the manuscript. EAH, NMH, MMK, and RES were responsible for managing purchasing tasks and project schedules. NMH coordinated specimen collection and transport and implemented quality assurance procedures throughout the laboratory workflow. EAH, NMH, MMK, and RES conducted the laboratory analysis. Patient selection and data collection were performed and supervised by MAA and MMK. All authors have read and approved the final manuscript.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
All data and materials generated or analyzed during this study are available upon request. The corresponding author is the designated contact person for data access inquiries.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
