Abstract
Background
Individual studies of the association between N-acetyltransferase 1 (NAT1)*10 allele and bladder cancer susceptibility have shown inconclusive results. To derive a more precise estimation of any such relationship, we performed this systemic review and updated meta-analysis based on 17 publications.
Methods
A total of 17 studies were investigated with 4,322 bladder cancer cases and 4,944 controls. The pooled odds ratios (ORs) with 95% confidence intervals (CIs) were used to assess the strength of the association. Subgroup analyses were conducted based on ethnicity, sex, source of controls and detecting methods. Then trial sequential analysis was performed to evaluate whether the evidence of the results was sufficient and reduce the risk of type I error.
Results
There was no association between NAT1*10 allele and bladder cancer risk in a random-effects model (OR = 0.96, 95% CI, 0.84-1.10) or in a fixed-effects model (OR = 0.95, 95% CI, 0.87-1.03). In addition, no significantly increased risk of bladder cancer was found in any other subgroup analysis. Then, trial sequential analyses demonstrated that the results of our study need to be further verified.
Conclusions
Despite its limitations, the results of the present meta-analysis suggested that there was no association between NAT1* 10 allele and bladder cancer risk. More importantly, our findings need to be further validated regarding whether being without the NAT1*10 allele could in the future be shown to be a potential marker for the risk of bladder cancer.
Introduction
Bladder cancer (BCa) is one of the most common cancers of the urinary tract, with an estimated incidence rate of over 380,000 new cases per year and 150,000 deaths worldwide (1). Although the exact cause of BCa remains unknown, a few well-established lifestyle risk factors have been identified, including cigarette smoking and certain occupational exposures such as to aromatic and heterocyclic amines (2). It is now commonly accepted that the etiology of BCa is a complex interaction between environmental factors and genetic susceptibility.
Aromatic amines (e.g., 4-aminobiphenyl) and heterocyclic amines (e.g., PhIP) are 2 of the major carcinogens in BCa. The metabolism of the compounds involves multiple enzymes. Acetylation by N-acetyltransferase (NAT) constitutes an important metabolic step. NAT has 2 isozymes, NAT1 and NAT2 (3). The NAT1 gene is located on chromosome subbands 8p21.3-23.1, and its activity has been detected to be higher in bladder mucosa (4). The enzyme activities of NAT1 are polymorphic. It has been demonstrated that the genetic polymorphisms in NAT1 affect the efficiency of the enzyme for the detoxification of carcinogenic amines (5, 6). There are 4 major NAT1 genotypes – NAT1*4, NAT1*3, NAT1*10, and NAT1*11 – which have been identified frequently. Among these, NAT1*4 is the wild-type allele; NAT1*3 has a single C-A substitution (C1095A); NAT1*11, a rare variant, manifests the deletion of the 9 nucleotides from 1080 to 1088 (7). NAT1*10, with 2 single-base mutations (T1088A and C1095A), has been reported to have higher enzymatic activity (5).
To date, many epidemiological studies have tested the possible association of NAT1*10 allele with the risk of BCa, but the results are still inconsistent or even contradictory. Therefore, we conducted a systemic review and updated meta-analysis of all candidate genetic studies to clarify the real association and identify any statistical evidence. Furthermore, trial sequential analysis (TSA) was for the first time used to clarify whether the evidence for the results was sufficient.
Materials and Methods
Literature search strategy
We performed an exhaustive search of studies that examined the association of NAT1 polymorphisms and susceptibility to BCa. Data were collected from the electronic databases PubMed, Embase and Web of Science available up to July 1, 2016, using the following keywords: “NAT1” or “N-acetyltransferase 1”; “variants” or “polymorphism”; “single nucleotide polymorphism,” and “bladder cancer,” “bladder neoplasm” or “bladder tumor” separately or combined. The research was restricted to human subjects only. All references cited in the retrieved articles were reviewed to identify other relevant publications. Review articles were also inspected to find additional eligible studies. Also, we only considered studies published in English.
Inclusion and exclusion criteria
Studies using an analytic design (case-control, nested case-control or cohort) and employing validated genotype methods to examine the frequency of NAT1 among BCa cases and controls were eligible for inclusion. Studies should include sufficient genotype data for analysis. Case reports, letters, reviews and editorial articles were excluded. Additionally, when duplications or the same patient populations used in several publications existed, only the most recent or complete study was included in this meta-analysis.
Data extraction
Two investigators (Z. X. and X. L.) independently extracted the data with a standard protocol, and the result was reviewed by a third investigator (B. Y.). We extracted the following information: first author's surname, year of publication, country of origin, ethnicity of subjects, sex of subjects, source of controls, genotype method and source of controls, and subdivided the subjects into 2 groups (without *10 and with *10) by numbers of NAT1*10 alleles in control and case groups, respectively.
Statistical analysis
In this meta-analysis, the following variations for NAT1 polymorphism were evaluated: without NAT1*10 and with NAT1*10. All associations were presented as the pooled odds ratios (ORs) with the corresponding 95% confidence intervals (CIs). After that, subgroup analyses were conducted on the basis of ethnicity, sex, source of controls and detecting methods. Between-study heterogeneity was tested using the chi-square-based Q-test. In addition, we separately used random-effects and fixed-effects models to analyze the data for the stability of the results (8, 9). Sensitivity analysis was conducted by excluding a single study 1 by 1, to check for stability and reliability on the pooled ORs by repeating the meta-analysis. In addition, publication bias was investigated by Begg's funnel plot, and the funnel plot was assessed by Egger's linear regression test (10). Statistical significance was considered when the p value of Egger's test was <0.05. All statistical analyses were carried out using STATA software (version 12.0; StataCorp LP, College Station, TX, USA). To ensure the reliability and accuracy of the results, 2 reviewers (Z. X. and X. L.) inputted the data in the statistical software programs independently and obtained the same results. The study group has substantial experience in conducting oncological research and has published similar studies elsewhere.
Trial sequential analysis
The outcomes of a meta-analysis might result in type I errors due to repetitive testing of accumulated data and are prone to systematic or random errors with sparse data collected (11, 12). TSA was introduced to estimate the required information size by adjusting the threshold for significance level with dispersed data and to confirm greater statistical reliability of the data than with a traditional meta-analysis (11, 13-15). In the current meta-analysis, TSA was performed by maintaining a 95% confidence interval, a 20% relative risk reduction, an overall type I error of 5% and a statistical test power of 80%, from which the required information size was calculated and the trial sequential monitoring boundaries were constructed. If the cumulative Z-curve (the blue line) crosses the trial sequential monitoring boundary (sloping red line) or exceeds the required information size (vertical red line), a significant result has been reached, and no further studies are needed. Otherwise, if the cumulative Z-curve does not cross the boundary or the information size required has not been reached, we need additional clinical trials to reach the adequate information size required for obtaining sufficient evidence (11, 12, 16-18). The TSA software (TSA, version 0.9, 2011; Copenhagen Trial Unit, Copenhagen, Denmark) was used in this study.
Results
Study characteristics
Based on the inclusion and exclusion criteria, 17 studies were evaluable in the current meta-analysis (19-35). The first one was published in 1997 (22) and the last in 2012 (32). The detailed characteristics of the selected studies are presented in Table I. The process of literature search and exclusion is shown in Figure 1. A total of 17 studies were investigated with 4,322 BCa cases and 4,944 controls. All studies were case-control studies that evaluated the association between NAT1 polymorphism and BCa susceptibility. Among these previous studies, 11 studies were conducted in white populations, 1 was in a non-Asian population, 2 were in mixed race populations and the remaining 3 studies were conducted in Asian populations. In addition, restriction fragment length polymorphism (RFLP) analysis was reported in 12 publications, and TaqMan analysis was performed in the remaining 5 studies.
Characteristics of studies investigating the association between NAT1 polymorphisms and risk of bladder cancer
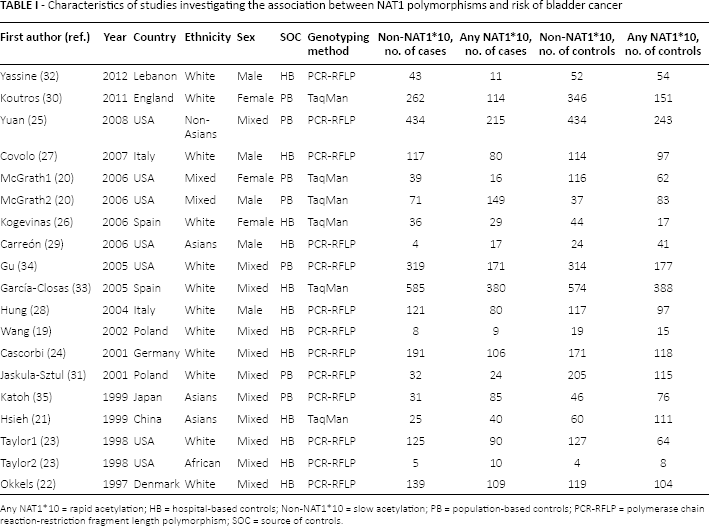
Any NAT1*10 = rapid acetylation; HB = hospital-based controls; Non-NAT1*10 = slow acetylation; PB = population-based controls; PCR-RFLP = polymerase chain reaction-restriction fragment length polymorphism; SOC = source of controls.
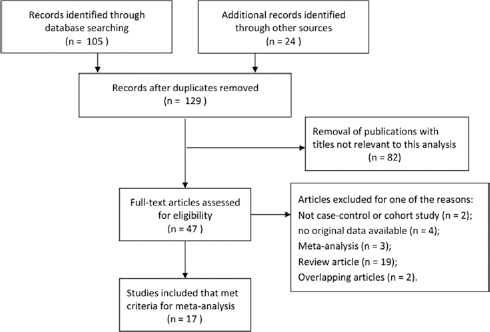
Flow diagram of literature search and selection process.
Quantitative synthesis results
A summary of the meta-analysis findings of the association between NAT1*10 allele and the risk of BCa is listed in Table II. The results of the meta-analysis showed that there was no association between NAT1*10 allele and BCa risk in the comparisons of individuals without NAT1*10 and with NAT1*10, in the random-effects model (OR = 0.96, 95% CI, 0.84-1.10) or in the fixed-effects model (OR = 0.95, 95% CI, 0.87-1.03) (Fig. 2).
Main results of the meta-analysis of NAT1 polymorphism and bladder carcinoma risk
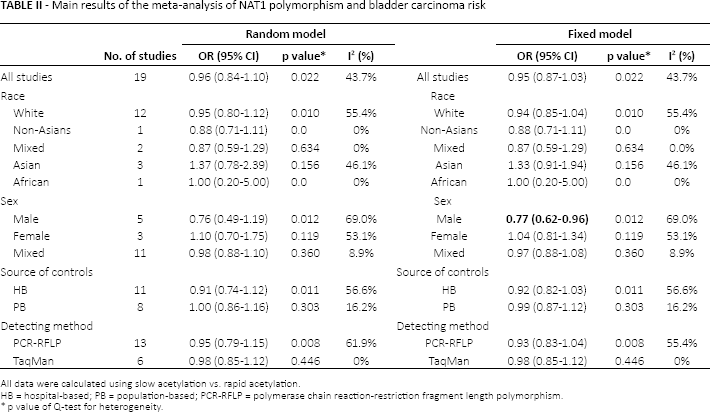
All data were calculated using slow acetylation vs. rapid acetylation.
HB = hospital-based; PB = population-based; PCR-RFLP = polymerase chain reaction-restriction fragment length polymorphism.
p value of Q-test for heterogeneity.
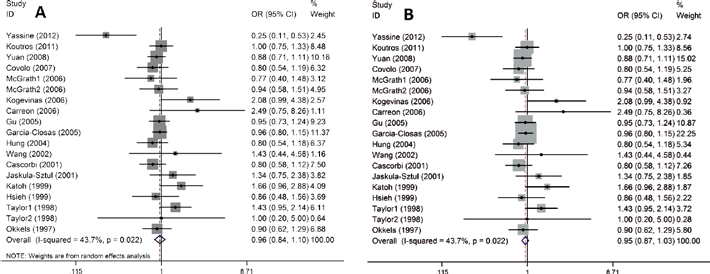
Forest plots of bladder cancer risk associated with the NAT1*10 allele. A. random-effects model, B. fixed-effects model.
For the stratified analyses by ethnicity, sex, source of control and detecting methods, the results are shown in Figure 3. There were no obvious associations between NAT1*10 allele and the risk of BCa in any other subgroup analysis.
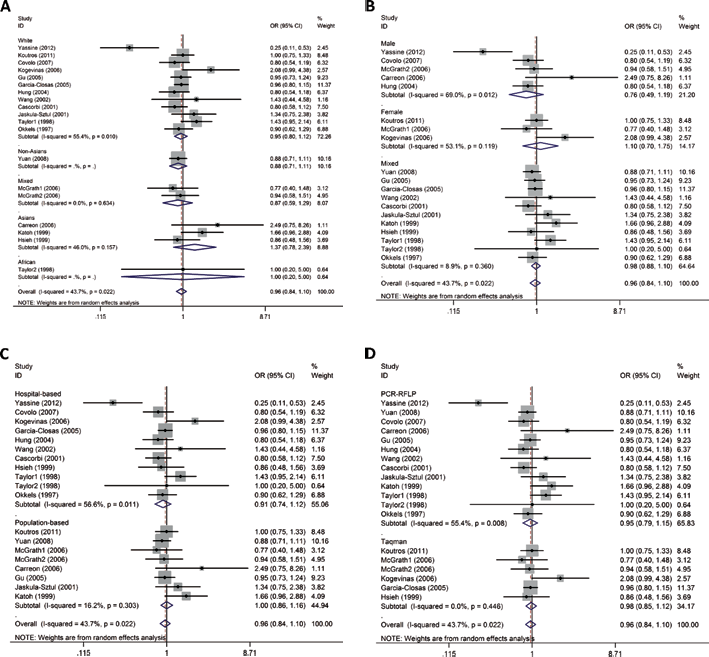
Forest plots of subgroup analysis of bladder cancer risk associated with the NAT1*10 the random model. (A) stratified by ethnicity; (B) stratified by sex; (C) stratified by source of control; (D) stratified by genotyping method.
Sensitivity analysis
Sensitivity analysis was performed by omitting 1 included study at a time, to distinguish the influence of each individual study on the pooled ORs, repeating the meta-analysis. The sensitivity analysis of the association between the NAT1*10 allele and the risk of BCa is shown in Supplementary Figure 1 (available online at www.biological-markers.com – Sensitivity analysis under the random model), demonstrating that the recalculated ORs were not substantially influenced. Therefore, the sensitivity analysis indicated that our results were robust and stable.
Publication bias
The Begg's funnel plot and Egger's test were performed to assess the publication bias of the literature in this meta-analysis. The shape of the funnel plot did not reveal any evidence of obvious asymmetry, and the outcome of Egger's test provided statistical evidence for the symmetry of Begg's funnel plot (p = 0.487). Thus, there was no obvious publication bias of the literature included in this meta-analysis (see Supplementary Figure 2, available online at www.biological-markers.com – Begg's funnel plot of publication bias test in the random model).
TSA results
In our present study, Figure 4 shows that not only did the cumulative Z-curve not cross the trial sequential monitoring boundary, but also the total number of cases and controls were not more than the required information size, showing the results to need to be further verified.
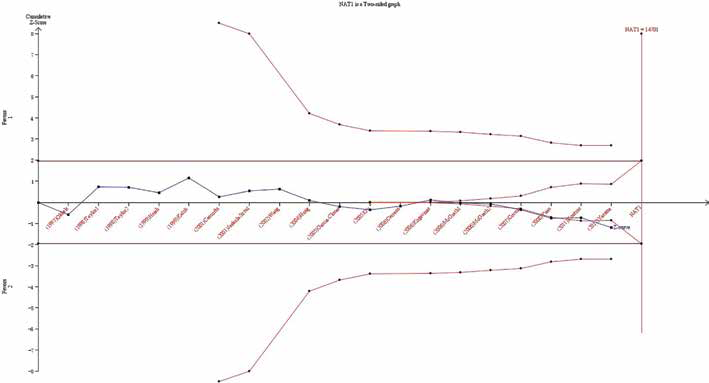
Trial sequential analysis of the association between NAT1 polymorphism and the risk of bladder cancer. The required information size was calculated based on a 2-sided α = 5%, β = 15% (power 85%), and a relative risk reduction of 20%.
Discussion
This systematic review provides the most recent and comprehensive assessment of the NAT1*10 allele in BCa. Many previous studies in the past 2 decades have investigated such an association, but the results have remained inconclusive. Among these, Carreón et al (29) found individuals carrying NAT1wt/*10 and NAT1*10/*10 showed a higher relative risk of BCa. However, Gu et al (34) found there was no significant association between risk of BCa and the NAT1*10 allele. Evidence suggested that the polymorphisms in activating and detoxifying enzymes might interact to affect the level of DNA damage sustained by a specific tissue and ultimately influence disease risk (36). Therefore, the imbalances between activation and detoxification processes might result in an increase in the susceptibility of BCa due to the accumulation of carcinogen metabolites, such as from cigarette smoking or exposure to carcinogenic amines.
As a statistical analysis tool, meta-analyses have a greater power than a single study especially in analyzing unexplained studies and can provide more substantial results. In the present study, there was a much stronger advantage toward showing any association between the NAT1*10 allele and susceptibility to BCa: for example, the sample size in this meta-analysis was larger than that of any individual study, which might make our results more reliable and robust. The current meta-analysis suggested that the NAT1*10 allele could not influence the risk of BCa.
NAT1*10 was suspected to lead to higher enzymatic activity than NAT1*4 in the bladder mucosa (5). However, based on the 17 studies in our meta-analysis, there was no association between the NAT1*10 allele and BCa risk. We further conducted a subgroup analysis and found no obvious association between the NAT1*10 allele and the risk of BCa.
In addition, because of the limited data, it could be that this meta-analysis was underpowered to explore any real association. Therefore, TSA was adopted in this meta-analysis, for the first time to reduce the risk of type I error and verify whether the evidence of our results was sufficient.
In conclusion, we need a better method to analyze and understand the association between the NAT1*10 allele and the risk of BCa. Therefore, to guarantee the reliability of our meta-analysis, more research should focus on the influence of different factors in the future.
TSA is a powerful and useful approach toward the goal of summarizing evidence and determining the required information size in meta-analyses (12, 37). To reduce the risk of a type I error and estimate whether further trials are needed, TSA is introduced to calculate the required information size for the meta-analysis with the adaptation of monitoring boundaries (38). However, the meta-analyses not reaching the required sample size are analyzed with trial sequential monitoring boundaries, which is similar to interim monitoring boundaries in a single trial (12, 17, 18, 39). Compared with the traditional meta-analysis techniques, TSA shows the potential to be more reliable. If the sufficient information size is not reached, false positive results are eliminated early due to random errors and repeated significance testing in meta-analyses (11, 12, 39). Furthermore, when reliable evidence is obtained, as demonstrated by TSA, researchers can stop implementation of further studies. Otherwise, it is necessary to reestimate the additional number of patients required to obtain reliable results in the meta-analyses, thereby guiding researchers in subsequent studies (40, 41). In the present meta-analysis, the number of cases and controls included were not more than the required information size, which meant that our results might not be firm evidence of effect.
Notably, there were several limitations to this meta-analysis. Firstly, the controls were not uniformly defined, because some controls were hospital-based. Hence, nondifferential misclassification bias is possible. Secondly, all published studies were written in English. It is possible that some related, published or unpublished studies that might meet the inclusion criteria were missed. Thus, some inevitable publication bias might exist in the results of the meta-analyses, though the funnel plots as well as Egger's linear regression tests indicated no remarkable publication biases. What is more, in the subgroup analysis by ethnicity, the population of whites came from 3 continents – North America, Europe and the Middle East – which might cause a selection bias. Furthermore, we excluded any interactions among genes and environment such as NAT2, the influence of smoking and exposure to benzidine or any other carcinogens, and lacking the original data of the studies included limited our further evaluation of potential interactions. Additionally, additional studies about exploring the risk effects of this polymorphism with susceptibility to BCa need to be further validated in subsequent studies.
Conclusions
In summary, despite its limitations, the results of the present meta-analysis suggested that there was no association between the NAT1 polymorphism and the risk of BCa in the comparisons of individuals without NAT1*10 and with NAT1*10. Moreover, in the subgroup analysis, there were no obvious associations between the NAT1*10 allele and susceptibility to BCa. More importantly, our findings need to be further validated regarding whether being without the NAT1*10 allele could in the future be shown to be a potential etiology and marker for the risk of BCa.
Footnotes
Financial support: No grants or funding have been received for this study.
Conflict of interest: We declare that we have no conflict of interest.
