Abstract
While evidence on the impact of the biomechanical environment elasticity on human mesenchymal stem cell (hMSC) behavior is growing, the aspect of micropatterning is still poorly understood. Thus, the present study aimed at investigating the influence of defined environmental micropatterning on hMSC behavior. Following characterization, hMSCs were grown on defined pillar micropatterns of 5, 7, 9, and 11 μm. With respect to cell behavior, primary hMSC adhesion was detected by indirect immunofluorescence (iIF) for paxillin, vinculin, integrin αV, and actin, while proliferation was visualized by histone H3. Morphogenesis was monitored by scanning electron microscopy and the expression of stem cell-specific biomarkers by real-time PCR. Favoritism of primary adhesion of hMSCs on pillar tops occurred at smaller pillar micropatterns, concomitant with cell flattening. While vinculin, integrin αV, and paxillin appeared initially more cytoplasmic, high pillar micropatterns favored a progressive redistribution with polarization to cell tension sites and at cell borders. Accomplishment of morphogenesis at day 3 revealed establishment of fully rotund cell somata at 5 μm, while hMSCs appeared progressively elongated at rising micropatterns. The hMSC proliferation capacity was influenced by pillar micropatterns and gene expression analysis of stem cell- and differentiation-associated biomarkers disclosed clear modulation by distinct pillar micropatterns. In response to environmental biomechanics, our results show that hMSC behavior is governed by pillar micropatterning. In turn, these findings may form the basis to prospectively direct lineage specificity of hMSCs in a customized fashion.
Introduction
Human bone marrow-derived mesenchymal stem cells (hMSCs) are employed for manifold regenerative therapies (21) in pathological conditions as diverse as cardiovascular (25,40) and respiratory diseases (19), diabetes (14), nervous system (33), or hard tissue damage (6,31,45). Ideally fulfilling all prerequisites for engagement in regenerative therapies, hMSCs are easily accessible, broadly available and expandable, and predictably dirigible (3). Among these claims, it is just the cell behavior directing which still remains precarious; however, hMSC in vitro modeling is providing more and more valuable contributions to the understanding of biophysical hMSC fundamentals (9,12,16). Cellular identity is established by a reciprocal interplay of topology, fate, and function, which is coordinated by the fundamental cell competence to perceive and react to environmental cues. The cell surroundings are roughly composed of attributes that can be classified either as soluble factors or physical forces. Both physicochemical features govern the cellular morphogenesis and in consequence the cellular lineage specification by inducing signaling inside cells through specific matrix adhesion receptors to modify cytoskeletal organization and cell shape (7,12). Extracellular matrix adhesion receptors perceive and rapidly respond to external biomechanics by amplifying the original signal intensity inter alia through the initiation of downstream signaling cascades (12). The underlying mechanism involves transmembrane adhesion proteins called integrins that are linked to plasmatic anchor proteins like vinculin and paxillin, which on their part couple actin fibers (7). The clustering of integrins at focal contacts activates tyrosine kinases, that is, focal adhesion kinase (FAK), which phosphorylates numerous targets (17,26) including Rho-GTPase pathways (23,32), hence controlling the actin cytoskeleton remodeling (24). In other words, integrins recruit a number of structural and signaling proteins to the plasma membrane, thereby responding to matrix tension and reciprocally exerting contractility at the cell periphery (12,34). The adhesion dynamics that subsequently navigate the cell shape and consequently the cell fate are tightly linked to environmental anchorage (2), because adhesion is needed for proper metabolism, protein synthesis, and survival (7). These cell–matrix contacts, known as focal adhesions, allow the cells to adhere and to pull on their substratum to which they are bound and in turn mediate the manipulation of the cells by their environment: both mechanical properties of matrix topography (20,23) and elasticity (13,15,28,29) are known regulators of cell differentiation (12,29,38).
With regard to hMSCs, research efforts regarding the impact of biomechanical parameters focused so far on the stiffness of the extracellular environment (10,18). HMSCs were shown to specify lineage and commit to phenotypes with extreme sensitivity to the elasticity of their extracellular substrate (13): soft matrices that mimic brain-operated neurogenic, stiffer matrices that mimic muscle were myogenic, and more rigid matrices that mimic bone proved osteogenic (13). By deforming and stiffening their matrix, hMSCs are able to communicate and align their position and morphology at long distances (41). Noteworthy, one can assume that this feature has a direct influence on the hMSC behavior because the cell–cell dialogue is known to affect the cellular proliferation and differentiation activity (12,23). In line with these results, Chen and colleagues reported that the commitment of hMSCs is predictable by monitoring their contractile status when cultured on microposts of varying elasticity (15). Interestingly, the experimental setup, which employed elastomeric pillar substrates, showed that the density of integrin-rich focal adhesion structure was nearly the same on different substrates (15); hence, it seems that the number of such complexes is the key rather than the substrate elasticity (4). This point of view is supported by the observation that the attachment area size controls the lineage commitment (18): hMSCs preferentially adapt an adipogenic fate when furnished only with small attachment areas but differentiate into osteoblasts on large attachment areas (23). There is evidence that not only the size but also the texture or micropattern of such attachment areas is important: discontinuous attachment areas that are interrupted by longitudinal grooves seem were found to command the cytoskeleton assembly (43) and to affect the transcription of differentiation markers in hMSCs (44).
Taken all together, extracellular biomechanics play an important role in the regulation of many cellular processes (7,8). HMSCs have the ability to sense microscale geometric patterns from their environment, which directly affect the cell shape and hence the cell fate (18,43,44). Nevertheless, data concerning the responses of hMSCs to environmental micropatterns still remain sparse. Thus, our study aimed to shed light on the impact of various anchor point designs on the behavior of bone marrow-derived hMSCs. In detail, we asked whether distinct environmental micropatterns (i) modulate the primary adhesion dynamics, (ii) control the hMSC morphogenesis, and (iii) influence the proliferation and gene transcription of hMSCs. In order to assess the micropattern impact on these hMSC features, we used a model that has previously proven suitable to analyze the mutual inter actions of environmental biomechanics and cells (1,15,37) by employing an array of silicone microposts with varying pillar topography (36). The results of this first-time investigation will form a platform for the future to address and to predictably direct behavior of hMSCs.
Materials and Methods
All chemicals were purchased from Sigma-Aldrich GmbH, Munich, Germany, unless otherwise indicated. For a detailed description of the pillar assay fabrication and of the experiments performed, see the Supplemental Materials and Methods Section (available at http://proksch-supplements.blogspot.com/).
Pillar Micropattern Device Fabrication and Biofunctionalization
For pillar micropattern device production, the protocols published by Tomakidi and colleagues (27,36) were slightly modified, and the wafer technology was kindly provided by the group of Prof. Dr. Holger Reincke, head of Laboratory of Process Technology of the Department of Microsystem Engineering, Albert-Ludwigs University Freiburg. In brief, the epoxy-based SU-8 10 resin (Microresist Technologies, Germany) was spin-coated onto a clean silicon wafer, the solvent was evaporated, and the wafer was exposed to UV light through a customized negative chromium mask (Masken Lithographie & Consulting GmbH, Jena, Germany). Developing was performed in a propylenglycol monoethylether acetate (PGMEA)-containing developer (mr-Dev.600, Microresist Technologies, Germany). The resulting moulds were covered with polydimethylsiloxane (PDMS; Dow Corning, Midland, MI, USA) vigorously mixed with a hydrosilane cross-linker, baked for 4 h at 65°C, and peeled with a razor blade under sterile conditions. Drops of a fibronectin (FN) solution (10 μg/ml) were pipetted carefully to the pillar arrays (10 s, RT). After withdrawing the drop, the surface was rinsed with phosphate-buffered saline (PBS, Invitrogen, Darmstadt, Germany) and directly used for cell culture. The biofunctionalization success was controlled by indirect immunofluorescence stains (iIF) of the FN immobilized only at the pillar tops. Planar control PDMS devices were likewise biofunctionalized with FN prior to cell seeding.
Cell Culture and Characterization of hMSCs
All experiments have been carried out in accordance to the guidelines of the World Medical Association Declaration of Helsinki and were approved by the Committee of Ethics of the Medical Faculty of the Albert Ludwigs-University Freiburg, Germany (EK-199/09). Human bone marrow-derived mesenchymal stem cells (hMSCs) were obtained from pelvic bone aspirate remnants (bone marrow aspiration pack, Harvest Technologies Corp., Plymouth, MA, USA) of n = 4 healthy patients undergoing hMSC-based sinus floor augmentation. The plastic-adherent cells were cultivated in hMSC proliferation medium (Provitro, Berlin, Germany), passaged up to three to four times (P4) and stored in liquid nitrogen until usage.
Flow Cytometry
Cells were incubated with saturating concentrations of fluorescein isothiocyanate (FITC)-, phycoerythrin (PE)-, peridin–chlorophyll–protein complex cyanin 5 (PerCP-Cy5)-, or allophycocyanin (APC)-conjugated mouse anti-human CD34, CD45, CD10, CD13, CD44, CD73, CD90, CD105, CD166, alkaline phosphatase (ALP) (BD Biosciences, Heidelberg, Germany), mouse anti-human CD146, v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog1 (c-kit), Stro1 (Abcam, Cambridge, UK) antibodies, or isotype-matched controls for 20 min at 4°C in the dark. After intense washing and resuspension in phosphate-buffered saline (PBS), flow cytometric analysis was performed using a FACSCalibur (BD Biosciences) flow cytometer. Data were collected with CellQuest software and analyzed with ModFitLT2.0 software (both BD Biosciences). For each run, 10,000 cell events were gated, and fluorochrome spectral overlap was checked and compensated whenever required.
Multilineage Differentiation Assay
Medium supplements were used to provoke the differentiation of hMSCs into adipocytes (0.5 mM 3-isobutyl-1-methylxanthine, 1 μg/ml insulin, 1 μM dexamethasone, and 100 μM indomethacine), osteoblasts (50 μg/ml ascorbate-2-phosphate, 10 mM β-glycerophosphate, and 10−7 M dexamethasone), and chondrocytes [cell pellets cultivated with 1 ng/ml transforming growth factor-β (TGF-β; R&D Systems, Minneapolis, MN, USA), 50 μg/ml ascorbate-2-phosphate, 1x insulin transferring selenium supplement-x (Invitrogen, Darmstadt, Germany), and 10−7 M dexamethasone] (30). According to established protocols, the adipocytes' lipid vesicles were visualized with the Oil Red O staining technique, the amount of extracellular matrix calcium deposits induced during the osteogenic differentiation was assessed by von Kossa staining, and the chondrogenic differentiation was evaluated by an Alcian Blue staining of the paraffin-embedded cell pellets.
Indirect Immunofluorescence Stainings (iIF)
The adhesion, proliferation capacity, and the cytoskeleton organization of hMSCs on pillar micropatterns were checked by iIF using mouse anti-human vinculin, integrin αV, histone H3, and β-actin (all Abcam, Cambridge, UK) or paxillin antibody (R&D Systems, Minneapolis, MN, USA), which were detected by a goat anti-mouse IgG1 Alexa Fluor 488 antibody (Invitrogen, Darmstadt, Germany). After actin fiber staining using Texas Red-labeled phalloidin and nuclei counterstaining with DAPI, the cells were mounted (Fluoromount G, Southern Biotech, Birmingham, AL, USA) and photographed with a Biozero BZ-8000 fluorescence microscope equipped with a CCD camera (Keyence Corp., Neu-Isenburg, Germany).
Scanning Electron Microscopy
In order to assess the cell morphology, cells were fixed with 8% formaldehyde and stored at +4°C until usage. After rinsing with PBS, specimens were dehydrated in ascending alcohol series (ranging from 30% to 100% ethanol, three times each for 20 min at RT), critical point dried (CPD 030 Critical Point Dryer, Bal-Tec AG, Balzers, Liechtenstein), and immediately sputter coated with gold palladium for 60 s at 60 mA (SCD 050, Balzers, Liechtenstein). The cells were examined using a LEO 435 VP scanning electron microscope (Zeiss, Oberkochen, Germany).
Quantitative Real-Time PCR Arrays
Total cellular RNA was pooled from biological replicates (n = 11–16 single pillar devices of each planar, 5-, 7-, 9-, and 11-μm micropattern) after accurately washing the cells with PBS. The RNA was purified using a guanidium–thiocyanate method (RNeasy Mini kit, Qiagen, Hilden, Germany) and stored at −80°C. The RNA integrity and quantity were verified using the Experion RNA StdSens chip microfluidic technology according to the manufacturer's instructions (Bio-Rad Laboratories, Munich, Germany). cDNA was synthesized from 500 ng of total RNA, and genomic DNA contamination was eliminated by using the First Strand cDNA Synthesis Kit (SA Biosciences, Frederick, MD, USA) in a C1000 Thermal Cycler (Bio-Rad, Munich, Germany). For real-time PCR, cDNA samples were diluted and amplified in duplicate using the precoated Human Mesenchymal Stem Cell RT2 Profiler™ PCR Array (SABiosciences, Frederick, MD, USA) with an RT2 SYBR Green qPCR Master Mix in a CFX96 cycler (Bio-Rad, Munich, Germany). The products' specificity of each amplicon was checked by examining the melting temperatures (heating at 0.05°C/s to 95°C). Data were collected using the CFX96 Manager Software version 1.0 (Bio-Rad, Munich, Germany) and analyzed using the PCR Array Data Analysis Software (SA Biosciences, Frederick, MD, USA). Herein, the fold change (2-ΔΔCt) was calculated as the normalized gene expression (2-ΔCt) in the test sample divided the normalized gene expression (2-ΔCt) in the control sample.
Results
Human Mesenchymal Stem Cells (hMSCs) Fulfill the Standard Characterization Criteria
Dealing with MSCs, the current terminology is confusingly heterogeneous, and a number of terms have so far been employed to describe progenitor cell populations derived from different tissues with different levels of inherent stem cell properties (3). We employed the standard hMSC characterization criteria of the International Society for Cellular Therapy (ISCT)—MSCs must be (i) adherent to plastic in vitro; (ii) positive for CD73, CD90, and CD105, while lacking CD45 and CD34; and (iii) able to differentiate at least to osteoblasts, adipocytes, and chondroblasts in vitro—in order to define the nature of the human MSCs (21) that were subsequently seeded onto our micropatterned pillar devices.
Plastic Adherence
In order to purify the hMSC fraction from bone marrow aspirates, we eliminated red blood cells and selected the mononuclear cells that readily adhered to their culture device. The latter did not hesitate to adapt a polymorphous and elongated polygonal shape and proliferated well while emanating from cell colonies.
Expression of Surface Markers
Flow cytometry screening of four to six times passaged cells derived from n = 4 patients revealed that the cells were reasonably homogeneous in size and granularity and were not labeled for CD34 (mean ± SEM: 0.25 ± 0.14%) and CD45 (0.03 ± 0.02%), but robustly expressed CD73 (80.45 ± 6.133%), CD90 (96.76 ± 1.09%), and CD105 (94.57 ± 4.15%). In addition, we analyzed the presence of further hMSC- or hard tissue mesenchymal cell-associated markers including CD10 (18.83 ± 9.71%), CD13 (86.90 ± 8.70%), CD44 (37.39 ± 10.04%), and CD166 (17.06 ± 1.20%). Although the typical hMSC surface marker spectrum was detected, the expression rate of alkaline phosphatase (4.73 ± 2.77%), c-kit (0.03 ± 0.02%), Stro1 (0.13 ± 0.09%), and CD146 (2.79 ± 1.46%) was fairly low (Fig. 1A), which may in part be a consequence of the trypsinization process that preceded the flow cytometric analysis. Taken together, the tiny number of bone marrow-derived, double-labeled CD34/CD45 population (0.02 ± 0.01%) likely allows for excluding that the isolated cells were of hematopoietic origin.

Fundamental human mesenchymal stem cell (hMSC) characterization. (A) Flow cytometric analyses of proliferated cells. Representative dot and histogram plots show that the cells were reasonably homogeneous in size and granularity and express a panel of surface markers that is requested to classify them as hMSCs including CD73, CD90, and CD105 while double-lacking CD34 and CD45, a fact that distinguishes them from eventually coisolated hematopoietic stem cells. In addition, hMSCs express further hMSC- and hard tissue mesenchymal cell-associated surface markers including CD13 and CD44. Selection gate R1: representative set of value limits, black dots: all events, Red dots: gated events. Green curves: negative controls, purple curve areas: positive samples. (B) Cytological stains and light microscopy images showed that hMSCs displayed both a multilineage differentiation potential and tendency to emerge from colonies (Oil Red O: adipogenic, von Kossa: osteogenic, and Alcian Blue: chondrogenic induced hMSCs). Scale bar: 50 μm. C-kit, v-kit Hardy–Zuckerman 4 feline sarcoma viral oncogene homolog; ALP, alkaline phosphatase.
Multilineage Differentiation
The hMSCs were successfully transformed in adipocytes, osteoblasts, or chondrocytes. After 21 days of adipogenic differentiation induction, the abundant number of hMSC-derived adipocytes was easy to identify by Oil Red O stains of their huge lipid droplets (Fig. 1B). Von Kossa staining allowed the visualization of calcium deposits in the extracellular matrix of osteogenic-induced hMSCs (Fig. 1B). After a 21-day culture period in micromass pellets induced by TGF-β, the extracellular matrix of hMSCs comprised acid mucopolysaccharides as revealed by Alcian Blue-staining (Fig. 1B), indicating a chondrogenic differentiation.
Colony Formation
Freshly isolated hMSCs started to grow in a colony-like pattern upon adherence to their cell culture device. Since this colony formation property of the cells was only observed at very early stages when the cell number was still limited, we documented the discrete number of arising colonies photographically without performing a colony-forming unit assay (Fig. 1B).
Fibronectin-Coated Pillar Devices Exhibit Defined Micropatterns
Consistent with previous studies (27,36), polymerization of PDMS in the silicon wafer yielded pillar devices with posts of constant height (15 μm) and diameter (5 μm) as revealed by scanning electron microscopy (SEM; Supplemental Fig. 1A, available at: http://proksch-supplements.blogspot.com/). The pillar arrays were configured with distinct micropatterns: 5, 7, 9, and 11 μm (Supplemental Fig. 1B1, B2, available at: http://proksch-supplements.blogspot.com/). Since the naked pillars per se render an unfavorable environment for cell adhesion due to their silicone-based nature, it was of importance to bioactivate the pillars such that they gained an extracellular substrate appealing cell adhesion. To ensure a defined adhesion of the hMSCs only on the pillar tops, we exerted biofunctionalization of the pillars with FN exclusively on the pillar tops. Indirect immunofluorescence stains of the fibronectintouched pillar posts revealed the successful coating of the pillar heads (Supplemental Fig. 1C, available at: http://proksch-supplements.blogspot.com/). Since the red fluorescence was exclusively seen on the pillar heads and the interspaced regions were devoid of fluorescence, it can be insured that only the pillar tops provide a basis for adherence of the seeded cells (Supplemental Fig. 1C, available at: http://proksch-supplements.blogspot.com/).
The Behavior of hMSCs Is Governed by the Biomechanical Cue of Micropatterning
Behavioral cell reactions to environmental stimuli appear in multiple ways. In the context of biomechanics, we addressed the following basic cell reactions to distinct micropatterns: (i) adhesion, (ii) morphogenesis, (iii) proliferation, and (iv) gene transcription.
The Dynamics and Molecule Distribution of hMSC Adhesion Depends on the Pillar Micropattern
Characterized by the feature of adherence, the very first mechanism observable in hMSCs in vitro is their primary adhesion to the respective growth surface. In order to monitor the adhesion dynamics of hMSCs established on pillar devices with 5-, 7-, 9-, and 11-μm micropatterns, the cells were allowed to adhere to the FN-coated pillars for 3 and 6 h after seeding.
An intriguing feature of hMSCs on the variopatterned devices concerned the shape of cells visible on the biofunctionalized pillars. Together with the cell shape, the β-actin organization varied tremendously dependent on the micropattern: on planar PDMS surfaces and 5-μm pillar micropatterns, hMSCs adhered to the micropillars displaying a flattened morphology and a well-structured β-actin organization already 3 h after cell seeding (Fig. 2A1–2). When seeded onto 7-μm pillar micropatterns, the hMSCs covered a smaller area and displayed a more polygonal shape with loosely organized globular β-actin molecules (Fig. 2A3). Awt 9- and 11-μm pillar micropatterns, the freshly seeded hMSCs showed an intensifying spherical profile with a completely disorganized β-actin cytoskeleton (Fig. 2A4–5). These findings were even emphasized by the finding that besides the cytoskeleton organization dynamics also the number of successfully adhered cells was micropattern related: the higher the pillar distance, the smaller amount of cells was adhered (Fig. 2B1–5). After 3 h, hMSCs had already flattened, being about to trace the pillar hole pattern with their actin fibers (Fig. 2B1) and resting carpet-like on 5-μm micropatterns reflected by a relatively low fluorescence signal at overview magnifications (Fig. 2B2). At 7-μm micropatterns, the hMSCs were similarly couched in an early outspread manner but the cell layer featured some gaps (Fig. 2B3). At 9 and even more at 11-μm micropatterns, a decreasing number of cells was adhered and flattened in shape (Fig. 2B4,5). In other words, despite a homogenous FN immobilization only on the pillar tops (Supplemental Fig. 1C, available at: http://proksch-supplements.blogspot.com/), the number of properly adhered and flattened hMSCs obviously decreased at micropatterns with increasingly distant pillars.
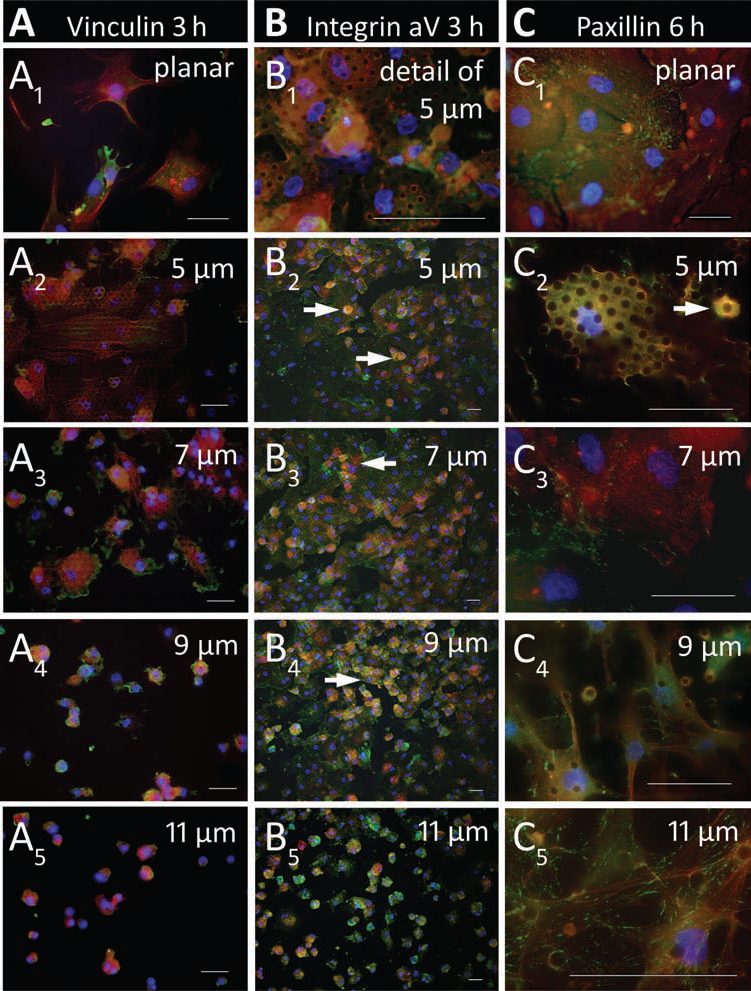
Micropattern-dependent adhesion dynamics. Green: adhesion molecules, red: actin cytoskeleton, yellow-orange: merge of green-colored adhesion molecules and red-colored actin, blue: cell nuclei (DAPI). Scale bar: 50 μm. (A1,2, B1,2) Three hours after seeding, the cells adhered readily to planar surfaces (A1) and small-scale pillar micropatterns of 5 μm (A2, B1,2), where they formed an uniform layer, adopted a flattened shape, and exhibited cytoplasmic adhesion molecules appearing as an equal green-colored haze that covered the cell body (A: vinculin, B: integrin αV) and an already partially organized actin cytoskeleton. The spatial colocalization of adhesion molecules and actin cytoskeleton resulting in a yellow-orange merged color highlighted the random intracellular adhesion molecule distribution. At 7-μm micropatterns, (A3) the adhesion molecules polarized at the cell borders and (B3) the hMSC layer was less homogenesous. (A4, B4) At the 9-μm pillar micropattern, the adhesion process was just implemented: the still spherical cells already displayed the pillar “hole” micropattern. (A5, B5) This effect was found to a lower extent on 11-μm micropatterns at which the hMSCs had not yet stuck after 3 h. At high-scale micropatterns, both the adhesion and the actin molecules appeared cloudy within still spherical, nonadherent cells (A4,5, B4,5). Cells that were still in the act of settling were round-shaped and displayed a brighter fluorescence signal in part above the microscopic focusing level (white arrows). (C) Six hours after seeding, the hMSCs accomplished to adhere to each micropattern (C1–5). At planar and small-scale pillar micropatterns, paxillin molecules (green) were randomly distributed over the cell body (C1–3), while they progressively accumulated at cell tension sites with rising pillar miropatterns (white arrows, C5). At high-scale micropatterns, the anchor point deficit was echoed by a progressively elongated and branched cell shape with cell extensions that contained clearly visible stress fibers (red, C4).
This trend may reflect the progressive difficulty for the cells to find anchor points for adhesion, which was also observed with regard to the adhesion molecule distribution. Labeled with green fluorochromes, distinct molecules of cellular focal adhesion sites became visible in a micropattern-dependent gradual accumulated grain pattern, preferentially localized around the pillar tops: already 3 h after seeding, in hMSCs on 5-μm pillar micropatterns, the actin-binding protein vinculin and the transmembrane anchoring molecule integrin αV were expressed equally distributed throughout the cell body discretely tracing the pillar arrangement (Fig. 2A2, B1–2). The distribution of all adhesion-associated proteins obviously followed the pillar micropattern: the more distant the pillars, the more circumferential the vinculin and integrin staining; in other words, the adhesion molecules were polarizing at cell borders (Fig. 2A3–5, B3–5). This observation again suggests that an extracellular regime with distant anchor points seriously hampers the proper cell adhesion. Paxillin, a focal adhesion-associated signal transduction adaptor protein, was found to cluster in freshly adhered hMSCs (Fig. 21–5), thereby indicating that the environmental biomechanics rapidly affect their cell functions. In cells seeded to planar control surfaces, the adhesion-associated molecules were stochastically distributed after 6 h (Fig. 2C1). Compared to planar surfaces, the adhesion-associated molecules were clustered much more articulately after cell installation on micropillars, where they clearly sketched (Fig. 2C2) and surrounded the apical micropost sites (Fig. 2C3). The polarization of the adhesion molecules to cell tension sites in hMSCs seeded to high-scale micropatterns suggests adhesion sites other than the pillar tops at micropatterns with remote pillars already 6 h after cell seeding (Fig. 2C5). Interestingly, at high-scale pillar micropatterns, the cells displayed huge filopodia (Fig. 2C4) containing well-organized stress fiber meshes (Fig. 2C5).
Generally, the micropattern-related adhesion dynamics were also reflected by the distribution dynamics of the focal adhesion-associated molecules. Although these explicitly aggregated in cells seeded onto each micropattern, they were redistributed more rapidly on small-scale micropatterns, where a period of 3 h was sufficient for the adhesion molecules to aggregate (Fig. 3A1, B1) but not in cells at high-scale micropatterns that were currently not yet completely adhered (Fig. 3A2, B2). The adhesion molecule aggregates were already more pronounced or even redistributed in cells on small-scale micropatterns (Fig. 3A3, B3), when the aggregation process took place in cells on high-scale micropatterns, then at least 6 h was required (Fig. 3A4, B4).

Adhesion molecule distribution relative to time and micropattern exemplified for paxillin (A, green) and vinculin (B, green). Red: actin cytoskeleton, yellow-orange: merge of green-colored adhesion molecules and red-colored actin, blue: cell nuclei (DAPI). Scale bar: 50 μm. (A1) Already 3 h after seeding, paxillin formed articulate aggregates around the pillar tops in adherent cells at 5-μm micropatterns. (A2) In contrast, cells had not yet accomplished to adhere at 11-μm micropatterns after 3 h but still remained round-shaped and exhibited a cloudily structured actin cytoskeleton (white arrows). The green cell-covering haze corresponded to paxillin molecules that were not aggregated but rather equally distributed in cells that proceeded to adhere. (A3) After 6 h, cells at 5-μm micropatterns displayed a plane shape and evenly redistributed paxillin molecules. (A4) At 11-μm micropatterns, cells were outstretched adopting a polygonal shape and exhibited clearly visible paxillin aggregates at the cell tension sites and around the pillar tops. Of note, cells on high-scale micropatterns dangled between the pillars such that parts of their somata went out of microscopic focus resulting in a partly fuzzy appearance. (B1) Again, cells seeded at 5-μm micropatterns were already adhered after 3 h and illustrated a flattened shape with accentuated actin molecules. Vinculin was visible in a randomly distributed manner both in adhered cells and in settling cells located above the microscopic focus level (white arrow). (B2) Even more of the latter were detectable at 11-μm micropatterns (white arrows), where the cells had not yet achieved to flatten and to build an organized actin cytoskeleton. (B3) After 6 h, cells at 5-μm micropatterns still exhibited a plane shape and even more explicitly structured actin fibers, while vinculin was polarized at the pillar tops. (B4) In contrast, cells at 11-μm micropatterns revealed a branched shape and more scattered vinculin molecules that retraced the pillar “hole” pattern.
The Pillar Micropattern Determines the Morphogenesis of hMSCs
Morphogenesis, that is, the development of a regular cell shape, effectively depends on proper cell adhesion. In consequence, the hMSC morphology was found strongly to depend on the pillar micropattern: the above-mentioned difficulty for hMSCs to adhere to distant pillars was reflected by the phenomenon of increasing cell branching when anchoring to remote pillars (Fig. 2C4). One step further, we determined the cell morphology after completion of initial adhesion and migration processes, and after 3 days, the hMSC tendency to adapt their shape to the pillar micropattern became even more evident. Cells that were grown on 5-μm pillar micropatterns assumed a flatly widespread shape, which turned out to be the closest match to hMSCs' regular shape (Fig. 4A, B). Their short and broad pseudopodia accomplished to anchor at side-by-side pillars (Fig. 4C). When grown on 7-μm pillar micropatterns, the globally rotund cell shape did not change significantly (Fig. 4D), but the cells presented manifold thin filopodia-like structures (Fig. 4E) with which they reached for consecutively arranged pillars (Fig. 4F). In comparison, hMSCs that have been cultivated on 9-μm pillar micropatterns displayed a more elongated cell body with tendency to adhere adjacent to the pillars (Fig. 4G, H), thereby practically mantling the pillars (Fig. 4I). This trend was continued in hMSCs grown on 11-μm pillar micropatterns, where the cells had an enormously stretched aspect (Fig. 4J), and exhibited vast pseudopodia-like extensions that clasped the pillars and anchored aside of them (Fig. 4K, L). Completely in contrast, cells grown on planar PDMS surfaces adopted an absolutely flattened plane and polygonal shape (Fig. 4M, N) and reached with broad and short finger-like cell extensions for their plain adhesion area (Fig. 4O). These versatile irregularities in hMSC morphology may be another hint that an environment with remote anchor points is unfavorable for true cell adhesion. This point of view becomes even more emphasized by the fact that after 3 days not only the external cell shape but also the internal cytoskeleton structure was diversified by the pillar micropattern (Fig. 5A). The β-actin fibers of cells cultivated on planar control surfaces resembled well-regulated longitudinal stripes, which paralleled the cells' lengthwise axis (Fig. 5A1). When grown on 5-μm micropatterns, the β-actin fibers also appeared well organized but rather horizontally striped (Fig. 5A2). In hMSC grown on 7-μm micropatterns, the β-actin fibers were less organized but accentuated the cell borders, hereby highlighting the wavelike adaption of the cell body to the pillars (Fig. 5A3). In hMSCs cultivated on high-scale micropatterns, the cells appeared less outspread but rather slender and outstretched with cell extensions that stepped out of the microscopic focus setting (Fig. 5A4,5). Both in the somata and in the extensions of hMSCs cultivated on high-scale micropatterns, the β-actin fibers materialized in a frowzy manner without clear self-organization (Fig. 5A4,5).

Cell morphogenesis after 3 days. (A–C) Representative scanning electron microscopy images of hMSCs grown on 5-μm pillar micropatterns revealed a rotund cell shape and pillar anchoring with short and broad pseudopodia that spanned the pillar gap. (D–F) At 7-μm pillar micropatterns, hMSCs displayed a fairly rotund morphology tagged with manifold thin and filopodia-like extensions. (H–I) Cells grown on 9-μm pillar micropatterns exhibited an elongated shape and tended to adhere aside of the pillar tops, thereby mantling the pillars. (J–L) The 11-μm pillar micropattern provoked an enormously stretched cell morphology with vast pseudopodia-like cell extensions that adhered on top and aside of the pillars. (M–O) In contrast, the cells adopted a completely flattened and polygonal shape and adhered with broad and short finger-like cell extensions to planar PDMS surfaces.

Micropattern-dependent cytoskeleton aggregation and mRNA expression. (A) After 3 days, the actin fibers (green) appeared (A1) longitudinal on planar surfaces, (A2) horizontally banded on 5 μm, (A3) accentuated at the cell borders on 7 μm, and (A4,5) increasingly less organized on high-scale micropatterns. The in part cloudy actin fiber display results from the phenomenon of sagged cells reaching out of microscopic focus being spanned between the pillars (blue: DAPI). Scale bar: 50 μm. (B) Clustergram of micropattern-dependent stem cell- and differentiation-associated gene expression normalized each to a panel of housekeeping genes (details: text) shows that distinct micropatterns differentially regulate gene expression. Colors indicate the magnitude of gene expression (green: minimal, black: average, and red: maximal gene expression levels). Clustergram columns: micropatterns (from left to right: 5, 7, 9, and 11 μm, planar), clustergram lines: investigated genes. (C) Bar graph of genes with significantly micropattern-regulated expression (cut-off level: 1.4-fold), among which the majority is associated with hMSC lineage specification. y-axis: fold change expression (2-ΔΔCt) calculated as normalized gene expression (2-ΔCt) in test samples to normalized gene expression (2-ΔCt) in planar surface samples (n = 11–16 each).
Taken together, the pronounced biaxial shape of hMSCs visible at the 5-μm pillar micropattern is nearly completely lost when hMSCs were exposed to a biomechanical environment composed of an 11-μm configuration. This default in regular shape formation is substantiated by long cell stretches, frequently observed at the 11-μm micropattern, and initiation of these stretches can be already noticed at intermediate pillar micropatterns of 7 and 9 μm. Hence, pillar devices with large-scale micropatterns may point to inappropriate biomechanical conditions for the performance of hMSC-innate cell behavior. Proper adhesion is indispensable for the development of regular hMSC morphology, and the increase of cell extensions correlated with increasingly distant pillar micropatterns may likely be a consequence of the cells' efforts to adapt to an environment inconvenient for proper adhesion. This situation strongly suggests that regular hMSC morphology is governed by environmental micropatterns.
The Gene Expression Profile of hMSCs Is Determined by the Pillar Micropattern
To contribute to the identification of putative target genes modulated by the environmental biomechanics, we compared the signal intensity of cDNA arrays derived from control hMSCs grown on planar PDMS surfaces with hMSCs cultivated on pillar arrays with 5-, 7-, 9-, and 11-μm micropatterns, respectively (Fig. 5B). To this end, cDNA was pooled from 11 to 16 biological replicates of each pillar or planar micropattern configuration, and the expression of 57 key genes involved in maintaining pluripotency and self-renewal status was profiled. In general, the transcription of these genes tended to be manipulated by the environmental micropattern, and this globally discrete modification in gene expression may be attributed to the relatively short cultivation period of 3 days. However, clear micropattern-dependent differences became apparent in the transcription profiles (Fig. 5B): an intriguing finding was that mesenchymal stem cell-associated genes were expressed at higher levels in cells grown on micropatterns with remote pillar distances (Fig. 5B). Among these, the transcription of leukemia inhibitory factor (LIF), basic fibroblast growth factor (bFGF), nucleoside diphosphate-linked moiety X motif 6 (NUDT6), and nestin were upregulated exclusively in cells grown on 11-μm pillar micropatterns (Fig. 5B). In comparison, the expression of the respective genes was downregulated in cells grown on 5-, 7-, and 9-μm pillar micropatterns, except for nestin, of which the expression was upregulated by the 9-μm pillar micropattern as well (Fig. 5B). Even though the transcription of single hMSC-related genes like KIT ligand was found to be mostly upregulated upon culture on other pillar micropatterns, the vast majority of stem cell-related genes was found to be up regulated by the 11-μm pillar micropattern (Fig. 5B). This observation suggests that a biomechanical environment with remote anchor points favors the preservation of an immature footprint of hMSCs. Not surprisingly, we detected no exclusive micropattern preference in terms of differentiation markers. Although the latter were globally expressed in an oscillating manner, it was possible to compile some trends with respect to the distinct lineage specifications: each direction was associated with certain environmental micropattern favorites. Genes that are attributed to an adipogenic differentiation like the peroxisome proliferator-activated receptor γ were favorably expressed upon culture on 5-μm micropatterns (Fig. 5B). Compared to planar surfaces, a chondrogenic differentiation was nevertheless mostly hindered by the 5- and 9-μm micropatterns: the 5-μm micropattern most prominently downregulated the transcription of sex-determining region Y box 9 (SOX9) and transforming growth factor-β1 (TGF-β1) (Fig. 5B), while the 7- and, in part, the 11-μm micropattern favored the transcription of bone morphogenic protein 6 (BMP6), growth differentiation factor 6 (GDF6), and K(lysine)acetyltransferase 2B (KAT2B) (Fig. 5B). The expression of osteogenic differentiation molecules was quite heterogeneous; nevertheless, the most well-known osteogenic markers were found to be upregulated by the 9-μm-micropattern: BMP2, collagen 1α1 (COL1α1), and FGF10. Although the osteoblastspecific runt-related transcription factor 2 (RUNX2) was in general slightly downregulated compared to hMSCs cultured on planar control substrates, the 9-μm micropattern best prevented this downregulation (Fig. 5B). In the context of environmental biomechanics, we additionally assessed the transcription of the adhesion site molecules integrin α6 and αV as well as the mediator molecule of cell shape-dependent lineage specification, Rho-GTPase A. In comparison to planar surfaces, the expression of both integrin genes was generally downregulated (Fig. 5B). However, in particular the 5-μm micropattern prevented the downregulation of integrin α6, a receptor that is highly selective for laminin. In contrast, the transcription of the fibronectin-binding molecule integrin αV was downregulated at the 5-μm micropattern to the greatest extent, such that it is suspected to play a more significant role in primary adhesion (Fig. 2B). Despite this downregulation in integrin transcription, the 5-μm micropattern augmented the expression of the downstream integrinsignaling molecule RhoA (Fig. 5B) compared to planar control hMSCs, such that two different conclusions may be drawn: firstly, the micropattern-related mechanotransduction employs RhoA and secondly, integrin αV may not be the main player in coupling of established cell adhesion and lineage specification. Although the shift in transcription was in general discrete for the vast majority of genes assayed, a significant regulation (cut-off level 1.4-fold) was found for a total of 13 genes that are depicted in Figure 5C. Among these, 12 are attributed to a certain lineage specification while only the brain-derived neurotropic factor (BDNF) is considered as a mesenchymal stem cell marker. Taken together, these data suggest that the environmental micropatterning of anchor points influences the hMSC behavior via modulation of gene transcription.
Pillar Micropatterns Influence the Detection of the Proliferation Marker Histone H3 in hMSCs
Keeping in mind that the extracellular micropattern directs the hMSC adhesion dynamics and morphogenesis, we next analyzed whether the distinct micropatterns influenced the cell proliferation capacity after 3 days. In this context, cells grown on each micropattern exhibited histone H3 localized inside the cell nuclei (Fig. 6A–E). However, clear micropattern-related differences came to light with regard to the signal frequency: on 5-μm micropatterns, almost each nucleus contained granular histone H3 molecules (green, Fig. 6A1,2), while considerably less cells showed histone H3-comprising nuclei on 7-μm micropattern (Fig. 6B1,2). This trend was continued on 9- (Fig. 6C1,2) and 11-μm micropatterns (Fig. 6D) where the fluorescence signal intensity generally decreased. Cells cultivated on planar control surfaces only in part exhibited histone H3 expression within their nuclei (Fig. 6E). These findings suggest that the basic cell function of proliferation capacity was in evidence influenced by the environmental micropattern.

Micropattern-dependent proliferation marker expression. Green: proliferation marker histone H3, red: actin fibers, blue: cell nuclei (DAPI). Scale bar: 50 μm. After 3 days, histone H3 was detected in nearly all cells grown on (A1, 2) 5 μm, but to a lower extent in cells grown on 7 μm (B1, 2). Even less histone H3 was detected in hMSCs cultivated on 9-μm (C1, 2) and 11-μm micropattern (D). Also cells grown on planar control surfaces in part displayed histone H3 expression (E). The visualization of the actin cytoskeleton (red) illustrated the micropattern-dependent cell morphology with outspread cell shapes and nuclei bedded on pillars of small-scale micropatterns (A1, 2, B1,2) and elongated cell somata with nuclei trapped between the pillars of high-scale micropatterns (C1,2, D).
Again, the micropattern effect on cell shape became obvious, while flattened cells cultivated on small-scale micropatterns displayed nuclei that clearly covered the pillar tops (Fig. 6A1,2, B1,2), elongated hMSCs on large-scale micropatterns disclosed nuclei that were rather trapped between than underpinned by the pillars (Fig. 6C1,2, D). Further, the high-scale micropattern scenario revealed that the nuclei were located at a considerably lower microscopic focusing level than the β-actin fibers. Hence, it appeared obvious that cells were buckled and bent over remote pillars, while they came across a comfortable bed of proximate pillars at the small-scale micropatterns.
Discussion
Here we show that the environmental micropattern directly affects the performance of hMSCs. This issue is captivating with regard to two key aspects of regenerative therapies: in vitro, our data open the road for a selective manipulation of hMSCs and in vivo, our observations serve to be prospectively implemented in a more precise prediction of hMSC behavior in the manifoldness of their applications (21) and last but not least with special regard to developing appropriate hMSC transplantation carriers (35,39,45). The pillar arrays used in our study provide exquisite control over the cellular anchor point micropattern, thereby enabling evant observations regarding the impact of biomechanical cues on stem cell performance. HMSCs adhered to the pillars at varying dynamics with clear preferences for small interpillar spacings and their shape profoundly developed as a function of the anchor point micropattern. These findings were reflected by a differential detectability in histone H3 proliferation marker and modulated gene transcription in contingency with the various environmental micropatterns.
The first issue addressed by our experimental setting, namely adhesion, is a meaningful feature of hMSCs that are known to be capable of homing to different tissues, a process that requires the competence to adhere both to the endothelium and the destination tissues (5,42). In fact, the hMSCs clearly discriminated the various micropatterns: already after short seeding periods, the hMSCs had readily adhered to the small-scale pillar micropattern devices showing an increasingly organized actin-fiber assembly. In contrast, remote pillar micropatterns seriously hampered hMSC adhesion and cytoskeleton organization. In addition, micropatterns with increasingly distant pillars fairly correlated with an increase of cell extensions, peculiar at 11-μm configurations where the cells were literally branched. This finding appears likely to be connected with the hMSCs' endeavour to adapt to a biomechanical environment inconvenient for proper adhesion. The number of adhesion molecules expressed by hMSCs (11) suggest that the perception and transduction of environmental biomechanics follow the same standards—that is, focal adhesion signaling—as in other solid tissue cells (2,7,12,17,34). Since the dynamics and molecule distribution of hMSC adhesion clearly attuned to the different micropatterns, it seems a logical consequence that the environmental anchor point micropattern may substantially control the hMSC behavior. The adhesive, together with the morphological preference for small-scaled pillar interfaces, suggests that this spatial organization provides a suitable environment for hMSC adhesion, while large pillar micropatterns may represent unfavorable biomechanical conditions for the realization of cell-specific morphology. In other words, adhesion is the most basic prerequisite of morphogenesis, and disrupting of regular morphology has been shown in hMSCs to be critical for fundamental cell functions including DNA synthesis (11,20,23).
To aid in understanding the role that adhesion-mediated micropatterns play in regulating hMSC differentiation, we performed gene expression analysis using DNA microarrays. After cultivating hMSCs on variomicropatterned pillar devices, we have seen a clear lineage preference for individual pillar micropatterns. Since iIF and SEM images revealed that the adhesion molecule distribution and morphology of cells that were grown on the corresponding pillar micropatterns were manifestly different, one can assume that cell adhesion and thus cell morphology correlate with lineage specification. To be more precise, cell morphology is indispensable for the display of adequate cell physiology, the basis of proliferation, and adopting a certain fate thereby supporting tissue regeneration. This consideration leads us to reflect on possible explanations of how adhesion, morphology, and cell function in terms of gene expression and proliferation may be interlinked. Since our results perfectly match reports on the impact of further biomechanical parameters—like cell shape, spreading or matrix stiffness—on hMSC behavior (13,15,20,23), we speculate that environmental micropatterns act via the same mechanotransduction pathways. Although the mechanism by which hMSCs adopt a biomechanical-conditioned fate is not yet fully understood, evidence is accumulating that the degree of cytoskeletal tension—that is tightly linked to cell adhesion and in consequence to cell morphology—is directly correlated to the cell differentiation status (13,15,20). In this light, it is likely that the observed effects of micropattern-dependent cell morphology and cytoskeletal organization in hMSCs play a key role: the more high scale the environmental micropatterns, the more stretched and hence the more tensioned the cell morphology. It is obvious that cell shape relies on the inherent cytoskeleton structure (12), and the fact that remote pillars do not comfortably underpin but rather stretch the cells suggests that actin fibers are pretty tensioned herein. Disrupting the cytoskeleton (20) or inhibiting RhoA (23) favors an adipogenic specification, but enhancement of the cytoskeleton contractility (20,23), cell adhesion capability (20), and extracellular matrix (ECM) stiffness (13) eases osteogenic differentiation. In our experimental setup, the culture on 5-μm micropatterns yielded a higher degree of adipogenic transcript expression, whereas the expression of osteogenic markers opted for the 9-μm pillar micropattern. Interestingly, we found that stem cell marker genes were mainly attributed to 11-μm micropatterns, such that we hypothesize that extreme conditions rather conserve an immature imprint. Matrix elasticity, cell spreading, and environmental micropattern each result ultimately in cell shape modulation. It was reported that these shape cues regulate hMSC destiny in a manner that involves mitogen-activated protein kinase and Wnt signaling cascades (20,23). One of the simplest examples of the influence of cell shape on differentiation has emerged from chondrogenic specification approaches that are usually performed in 3D pellet cultures (18). The most puzzling finding regarding the anchorage dependency of hMSCs was that the chondrogenic differentiation-associated gene transcripts were expressed fairly oscillating. We hypothesize that this peculiarity may be attributed to the fact that 2D culture in flattened shape may lead to an inhomogeneous chondrogenic specification (18,22). It is tempting to assume that cells differentiate in vitro in response to shape cues that are similar to native tissues and precisely cell shape is determined by adhesive interactions and matrix biomechanics. Although it remains unclear whether the micropattern-dependent cell morphology of individual hMSCs is predictive of differentiation, we observed a strong correlation between the environmental micropattern and the cell fate. This notion is supported by our finding that not only the adhesion- and cytoskeleton assembly-associated gene transcripts but also the proliferation-associated molecule histone H3 is differentially regulated by various pillar micropatterns.
Taken together, we conclude that the hMSC shape and thus the hMSC fate is biomechanically not only governed by matrix stiffness (4,10,13,15) and consecutive cell geometry (20) and spreading (23) but also by the adhesion-mediating microtopography as we have unequivocally shown in this paper. Such knowledge will be precious for the future identification and characterization of hMSC-tailored biomechanical conditions that can prospectively be implemented in adjuvant therapy concepts or the creation of new materials in regenerative medicine.
Footnotes
Acknowledgments
The authors are grateful to Prof. Dr. Holger Reinecke, head of the Laboratory of Process Technology of the Department of Microsystem Engineering, Albert-Ludwigs University Freiburg, for manufacturing the silicon wafers, necessary for PDMS pillar micropattern production. The authors acknowledge the excellent technical assistance of Yrgalem Abreha, Brunhild Saaler, and Heike Jahnke, who supported the cell culturing and conducted the preparation and documenting of the scanning electron microscopic images. The study was funded by the Research Committee of the Medical Faculty of the University Freiburg Medical Center (PRO723/09) and by the Deutsche Gesellschaft für Zahn-, Mund- und Kieferheilkunde (DGZMK, 11/2010). The authors declare no conflict of interest.
