Abstract
BACKGROUND:
Testis-specific genes encoding for long non-coding RNA (lncRNA) have been detected in several cancers; many produce proteins with restricted or aberrant expression patterns in normal or cancer tissues.
OBJECTIVE:
To characterize new lncRNA involved in normal and/or pathological differentiation of testicular cells.
METHODS:
Using bioinformatics analysis, we found that lncRNA LOC100130460 (CAND1.11) is expressed in normal and tumor testis; its expression was assessed in several human cell lines by qRT-PCR. CAND1.11 protein, produced by a single nucleotide mutation, was studied by western blot and immunofluorescence analysis on normal, classic seminoma, and Leydig cell tumor testicular tissues.
RESULTS:
CAND1.11 gene is primate-specific; its expression was low in SH-SY5Y cells and increased when differentiated with retinoic acid treatment. CAND1.11 expression in PC3 cells was higher than in PNT2 cells. CAND1.11 protein is present in the human testis and overexpressed in testicular cancer tissues.
CONCLUSIONS:
This report is one of the few providing evidence that a lncRNA produces a protein expressed in normal human tissues and overexpressed in several testicular cancers, suggesting its involvement in regulating cell proliferation and differentiation. Although further studies are needed to validate the results, our data indicate that CAND1.11 could be a potential new prognostic biomarker to use in proliferation and cancer.
Introduction
A significant increase in the study of long non-coding RNA (lncRNAs) was recorded in the past decades. lncRNAs are considered RNA transcripts longer than 200 nucleotides that do not encode any identifiable peptide product [1, 2]; on the other hand, long-read sequencing technologies promise to improve current annotations, to obtain a complete record of lncRNAs expressed throughout the human lifetime [3]. Furthermore, it has been observed that most of the RNAs transcribed from the human genome are as lncRNAs [4, 5, 6, 7], and the number of those under functional investigation is growing exponentially [8, 9, 10, 11, 12]. lncRNAs have critical maintenance functions, such as transcription, RNA processing and translation, regulation of dosage compensation, and genomic imprinting [13], playing a functional role in various biological processes such as pluripotency, cell development, immune response, differentiation, human disease, etc. [12, 14, 15]. Studies on lncRNAs gene structure, as exons and promoters, demonstrated that they undergo evolutionary purifying selection and provided strong evidence for their functionality [16, 17]. However, they are poorly characterized compared to protein-coding genes. In this context, while lncRNA transcripts appear less conserved than mRNAs, most of their promoter regions are as conserved as those of protein-coding genes [18]. Thanks to new technologies, it has been possible to identify and classify human transcripts based on their spatial expression patterns across organs and tissues, which are relevant for studies of biological processes and diseases [19, 20, 21, 22].
It is well-known that Mammalian testis expresses more lncRNAs as compared to other tissues [17, 23, 24]; moreover, transcriptome studies showed that a dynamic change in lncRNA expression throughout spermatogenesis occurs [25, 26, 27], revealing that some of them possess a significant biological role in testis and, consequently for proper fertility [28, 29]. In fact, in knock-out mouse models of several testis-specific lncRNAs, disturbances in sperm production or quality have been shown to cause marked alterations in spermatogenesis [29, 30, 31, 32].
Notably, several testis-specific genes encoding for lncRNAs, have been detected in various types of cancer [33], leading to their categorization as “cancer-testicle” genes. Interestingly, many of these genes produce proteins with restricted or aberrant expression patterns in normal tissues or different cancer types [33, 34, 35]. Therefore, the study of cancer-testicle genes is of interest to identify new lncRNA targets which might be involved in both normal and/or pathological differentiation of testicular cells. Based on this, we characterized the Homo sapiens LOC100130460 gene and its predicted lncRNA CAND1.11, encoding for a 223 amino acid protein produced by a single nucleotide mutation occurring in the allelic form in the genome. Via Bioinformatics, we evidenced the structure of CAND1.11 gene and protein and the similarity of nucleotides and amino acid sequences with those of other primates. Moreover, by RT-qPCR, CAND1.11 expression was evaluated during differentiation or tumorigenesis in several human cancer cell lines. Finally, using a polyclonal antibody raised against CAND1.11, Western blot (WB), and immunolocalization analyses in human testicular non-pathological and tumor tissues were performed.
Materials and methods
Databases searches and bioinformatics analysis
Searches of new lncRNAs expressed in the testis and in testicular cancers were carried out in the National Center for Biotechnology Information (NCBI) nucleotide databases (
Cell culture
The SH-SY5Y (human neuroblastoma, ATCC), HEK293T (human embryonic kidney, ATCC), HT-29 (human colon cancer, kindly granted by Prof. Marina De Rosa, University of Naples Federico II, Italy), U2OS (human bone osteosarcoma, kindly granted by Prof. Mimmo Turano, University of Naples Federico II, Italy), RKO (colon carcinoma, kindly granted by Prof. Mimmo Turano, University of Naples Federico II, Italy), PNT2 (human prostate epithelium, ATCC) and PC3 (human prostate cancer, ATCC) cell lines were grown and propagated in Dulbecco’s Modified Eagle Medium (DMEM, EuroClone, Milan, Italy) supplemented with 2 mM L-glutamine (EuroClone), 1% penicillin/streptomycin (EuroClone) and 15% fetal bovine serum (FBS, EuroClone) for SH-SY5Y and 10% for the other cell lines.
RNA isolation, retrotranscription, RT-PCR, and RT-qPCR
Total cellular RNA was isolated using TRI Reagent
The melting curve and the gel electrophoresis confirmed that only one amplicon of the expected size, was produced under our conditions. The specificity of RT-qPCR reactions was further validated by sequencing analysis of the amplification product by the Sanger method performed by an external service (Bio-Fab, Rome, Italy).
Tissue samples
Three human placental samples were obtained from patients undergoing a cesarean section. Each specimen was quickly immersed in liquid nitrogen and stored at
Testicular biopsy specimens were obtained from the suspected testicular nodule, and an experienced pathologist confirmed the presence of cancer by extemporaneous evaluation. The number of collected samples was 6 non-pathological (NP), 10 classic seminoma (CS), and 6 Leydig cell tumor (LCT). Written informed consent, from all subjects involved in the study, was obtained. Each tissue sample was divided into two halves: one half was quickly immersed in liquid nitrogen and stored at
Production of antibody
Polyclonal antisera were raised in rabbits using a synthetic peptide for immunization. The optimal peptide sequence was retrieved using the CAND1.11 protein of 223 amino acids in length. The peptide with the following sequence TGSCSVAWLECSGANMT, was synthesized by an external service (Proteogenix, Schiltigheim, France). After cross-linking with albumin by formaldehyde treatment, the peptide was injected subcutaneously with complete Freund’s adjuvant. The injection was repeated after 2 weeks. Then, two additional injections were performed with incomplete Freund’s adjuvant. Antibody was isolated from serum by ammonium sulfate precipitation. The specificity of CAND1.11 antiserum was checked by pre-adsorbing the primary antiserum with five-fold excess of the corresponding epitope and assessed via WB analysis.
Total protein extraction and WB analysis
The placental and testicular tissue samples were lysed in radioimmunoprecipitation assay (RIPA) lysis buffer (#TCL131; Hi Media Laboratories GmbH; Einhausen, Germany) supplemented with 10
Immunofluorescence (IF) analysis
The fixed tissues were dehydrated in increasing alcohol concentrations before paraffin embedding. Histological evaluation of the samples was determined in our previous papers [53, 54]. For IF staining, 5
Statistical analysis
The results from independent biological replicates in triplicate are expressed as mean
Results and discussion
lncRNAs are known for contributing to the regulation of many physiological processes and are also significant controllers in the biology of tumors [15, 40]. Testis is the mammalian tissue expressing the majority of lncRNAs [41], and many of them have been identified in physiological and pathological spermatogenesis, but, to date, just a few have been validated and functionally characterized [42].
To search for new lncRNAs expressed in the normal testis and testicular cancers, we carried out exploration in NCBI nucleotide databases with the following keywords: lncRNAs, gametogenesis, testis, and cancer. Among the retrieved gene/transcript sequences, we identified an mRNA (AI652043.1) for the CAND1.11 gene. This gene has also been described as a member of candidate clusters corresponding to novel Cancer/Testis (CT) genes [34]. In that paper, the authors reported that, probably due to partial sequences for CAND1.11 transcript, no open reading frames (ORF) longer than 110aa were identified, leading to the hypothesis that this transcript might belong to the growing class of non-coding regulatory RNAs [34]. This hypothesis seems corroborated by the NCBI reference sequence for CAND1.11 (NR_103765.1), which is associated with a transcribed lncRNA of 1019 bases in length.
Expression level of CAND1.11 transcript in different cell lines. (A) The gene expression level was calculated by the 2-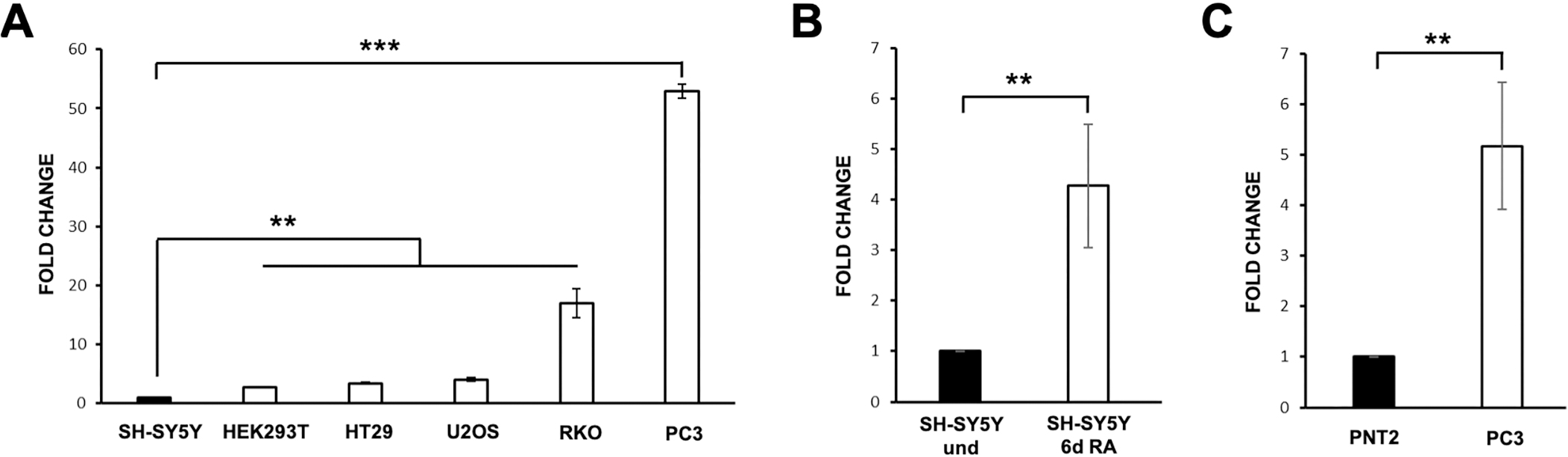
To evaluate if the CAND1.11 gene expression is cell-specific, we designed a pair of primers for use in RT-qPCR assays. We analyzed its transcript level in different human cell lines in their proliferative state: SH-SY5Y (neuroblastoma), HEK293 (human embryonic kidney), HT29 (colorectal adenocarcinoma), U2OS (osteosarcoma), RKO (colon carcinoma) and PC3 (prostate cancer). The results showed a different expression level among the various cell lines, with the highest level in PC3 cells as compared to all the others (Fig. 1A). Afterwards, we evaluated if CAND1.11 expression level may change in diverse conditions, to this end, we used SH-SY5Y cells that can be differentiated by RA stimulation, changing from an undifferentiated to mature-like state after six days of treatment. As shown in Fig. 1B, CAND1.11 transcript level increased following RA treatment compared to control undifferentiated SH-SY5Y (
As the highest level of its transcript occurred in PC3 cells, we also investigated CAND1.11 transcript level in a corresponding normal cell line, the PNT2 (prostate epithelium). The results showed that its transcript level in the PNT2 cells was lower than in PC3 cells (
CAND1.11 nucleotide and protein sequences. (A) CAND1.11 nucleotide sequence (NR_103765.1) and corresponding predicted protein. (B) nucleotide alignment of different EST retrieved in the NCBI database. The accession number is reported on the left of each sequence. The corresponding amino acid sequences for the nucleotide sequence with the internal stop codon (red square) and the additional EST with the TGC codon encoding for a cysteine (red square) are reported under the nucleotide sequence. (C) Alignment of putative amino acid sequences of primate CAND1.11 sequences. Hs, Homo sapiens (NR_103765.1, TGA codon was manually changed in TGC codon to obtain the longer protein sequence), Pp, Pan paniscus (XM_003818154.1), Gg, Gorilla gorilla (XM_004050695.2).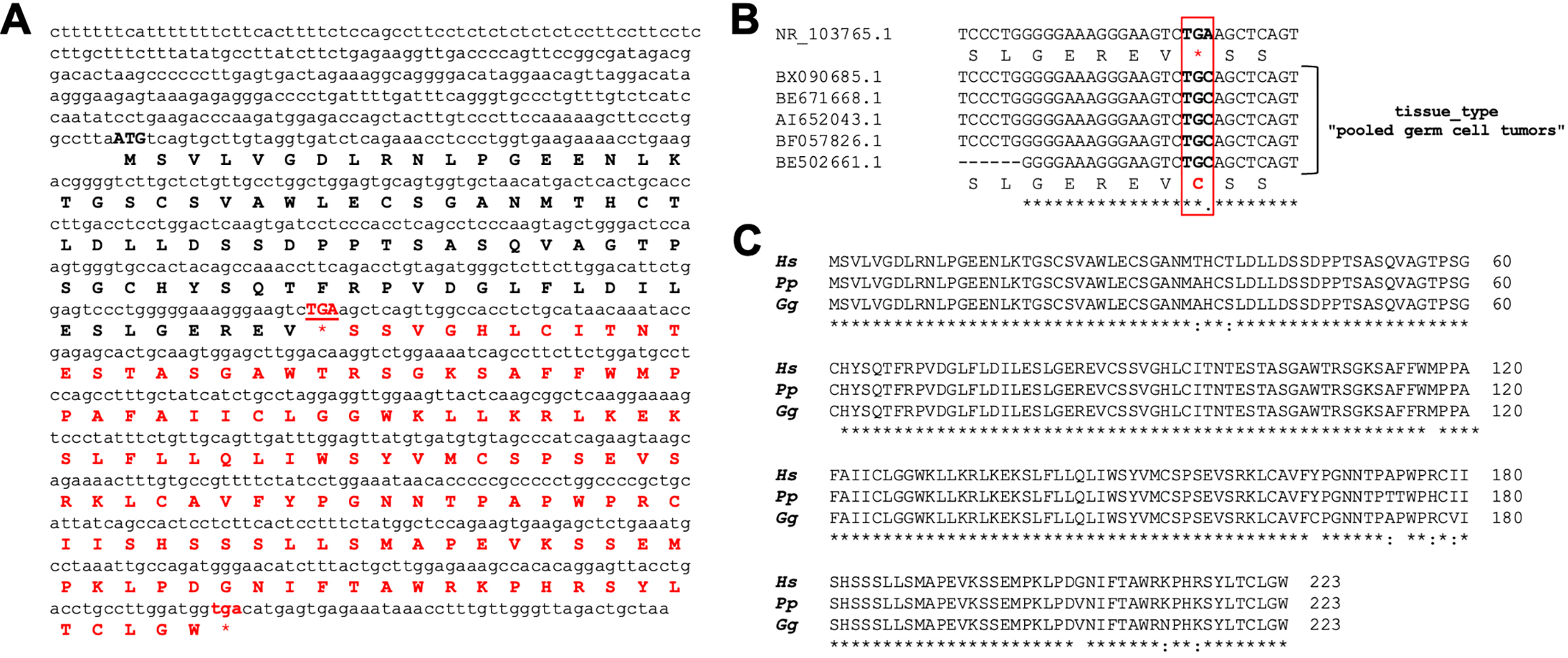
CAND1.11 gene structure. Exon-intron organization of CAND1.11 human (Hs) and gorilla (Gg) gene based on the transcript nucleotide sequence (Hs, NR_103765.1; Gg, XM_004050695.2). TGA/C indicates the internal stop codon (TGA) and the alternative TGC codon found in the same ESTs.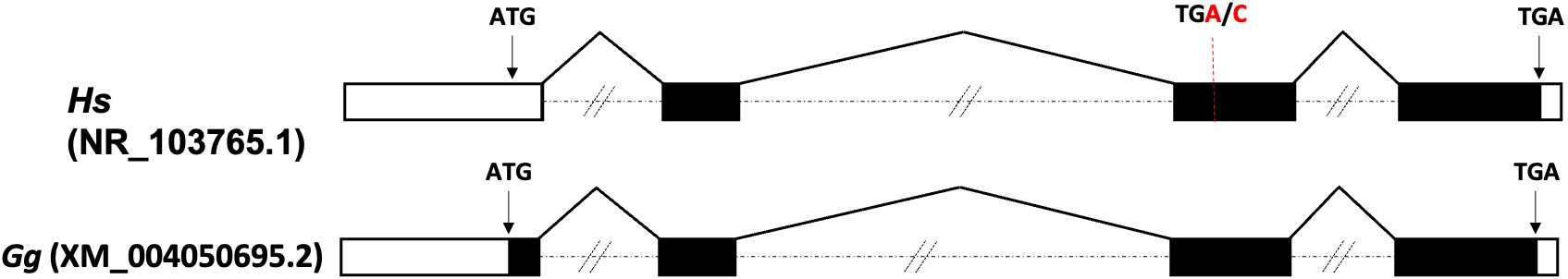
Interestingly, although initially classified as lncRNAs, many of these transcripts contain ORF and then may code for functional peptides or small proteins that play an important role in the pathogenesis of many diseases, including cancer. For example, CASIMO1 and SMIM30, two lncRNA-encoded microproteins, promote cell proliferation in breast and liver cancer, respectively [see for review 46]. Here, the bioinformatically translated nucleotide sequence of CAND1.11 also showed a putative ORF of 258 bases, encoding for a protein of 86aa in length (amino acid sequence in bold black in Fig. 2A).
Of interest, this amino acid sequence is different from that obtained by translation of the reference nucleotide sequence (AI652043), reported by Bettoni et al. [34] because of a nucleotide substitution that changes the “TGC” triplet, encoding for a cytosine amino acid, into a “TGA” stop codon (indicated in red uppercase in the Fig. 2A). This evidence led us to deeper investigate about the presence of this stop codon in different Expression Sequence Tags (ESTs) reported in the NCBI database. We found a series of ESTs generated by pooled germ cell tumors possessing the TGC triplet instead of the stop codon (Fig. 2B), as for the sequence reported by Bettoni et al. [34]. The analysis of the NCBI Single Nucleotide Polymorphisms (SNPs) database confirmed the presence of this A
Result of the CPAT software analysis for the NR_103765.1 nucleotide sequence with TGA or TGC codon
To gain further evidence in line with the hypothesis that CAND1.11 gene may encode for a protein, we exploited using the Coding Potential Assessment Tool (CPAT) software. We submitted the entire nucleotide sequence of the 1019 bases in length RNA, both with “TGC” and “TGA” triplets, to the analysis of the CPAT. As shown in Table 1, while the version of the RNA with the stop codon retrieved no coding potential, on the contrary, the RNA sequence with “TGC” triplet showed a coding probability value above the cut-off (0.364).
In addition, we should mention that few lncRNAs encode for peptides [48, 49, 50]; an example is that of ANRIL, a small peptide encoded by a lncRNA that is a new biomarker in human cancer [51].
Considering that the NCBI database reports that the expression of the CAND1.11 gene is at a high level in a many normal human tissues (i.e. placenta, testis, fat, breast, lung and trachea), while it is at a low level or not expressed in other tissues, to further investigate on its protein, we raised a polyclonal antibody against CAND1.11 protein to use it to analyze the expression and localization in normal and pathological human testicular tissues. Testicular cancer is a commonly used definition of a particular heterogeneous pathology, including several types of cancer, which are classified and divided into different subgroups by the International Agency for Research in Cancer of the World Health Organization [52]. Among them, CS and LCT are two of the most common solid tumors in the adolescent and young adult male population between 20 and 40 years old [53, 54].
W B and IF analysis of CAND1.11 in NP, CS and LCT testicular tissue samples. (A) WB analysis was carried out on total protein extracts from each sample using CAND1.11 antiserum pre-adsorbed 5-fold excess with the epitope. (B) WB analysis showing the protein level of CAND1.11 (25 kDa) and 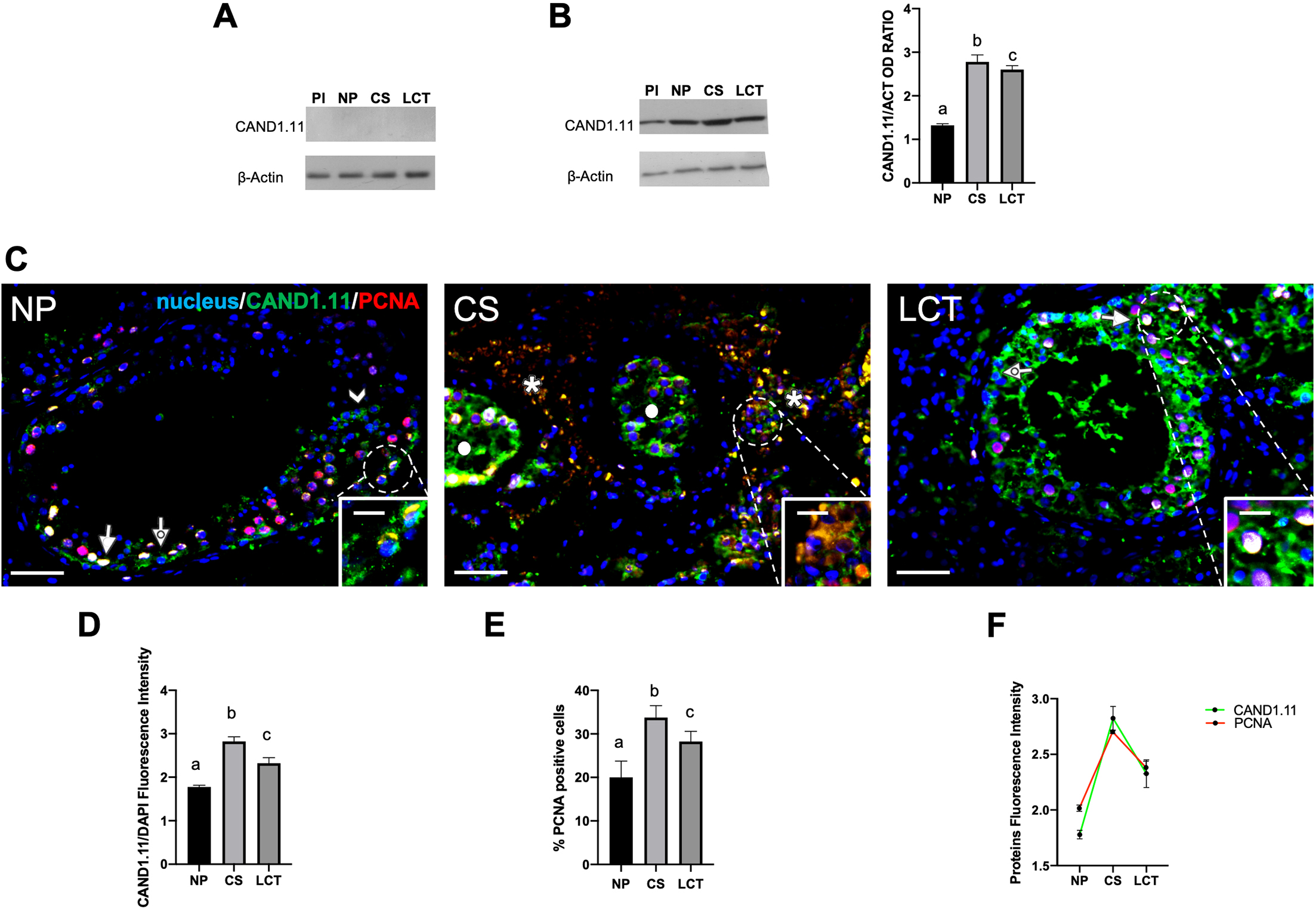
Firstly, we verified the antibody specificity by incubating protein samples with the CAND1.11 antiserum preabsorbed with the corresponding epitope, and the WB result gave no signal, confirming the specificity of the antibody through competitive peptide blocking (Fig. 4A); secondly, the results showed that CAND1.11, with a molecular weight of about 25 kDa, was expressed in human placenta (used as a positive control), as well as in human testicular tissues, as reported in Fig. 4B. Interestingly, CAND1.11 protein level increased in the cancer tissues, and particularly of 110% in CS (
CAND1.11 IF analysis, performed along with PCNA, a commonly used proliferation marker, confirmed the trend observed by WB. In fact, in NP, CAND1.11 localized at cytoplasmic and nuclear levels in both mitotic spermatogonia (SPG; dotted arrow; Fig. 4C and inset) and meiotic spermatocytes (SPC; arrow; Fig. 4C), with a more intense signal in the latter, as well as in differentiating spermatids (SPT; arrowhead; Fig. 4C). Moreover, a clear CAND1.11 and PCNA co-localization, evidenced by the intermediate yellow-orange dye in the nucleus of SPC, was observed. Thus, the specific localization of CAND1.11 in the nucleus and in the cytoplasm of SPG and SPC in the human testis may suggest the hypothesis of its involvement in the proliferative and differentiative phases of spermatogenesis. Other studies are in progress to better clarify this point.
Remarkably, in CS, peculiar CAND1.11, and PCNA signals were seen in the cytoplasm and the nucleus of some seminoma cell clusters (asterisks; Fig. 4C and inset), showing a significant increase in CAND1.11 intensity (
Finally, in LCT, although the fluorescent signal intensity was stronger than that observed in NP (
Here we showed that the primate-specific lncRNA CAND1.11 is expressed in normal and cancer cell lines determining that its expression changes during differentiation and/or tumorigenesis in tumor cell lines. Furthermore, this report is one of the few showings that a lncRNA, CAND1.11, produces a protein of 223 amino acids expressed in normal human tissues during life and over-expressed in several human testicular cancers.
While this study is preliminary, mainly due to the reduced number of human testis samples used, it supports the putative role of CAND1.11 in regulating cell cycle progression that occurs before, during, and after normal and pathological cell differentiation. However, although more studies are needed to validate all the above findings, these data indicate that CAND1.11 could be used as a potential novel biomarker both prognostic of proliferation and cancer.
Funding
This work was supported by the Italian Ministry of University and Research (Grant No. PRIN 2020 to FA).
Competing interests
The authors declare that they have no competing interests.
Author contributions
Conception: Sergio Minucci, Francesco Aniello.
Interpretation or analysis of data: Aldo Donizetti, Massimo Venditti, Davide Arcaniolo, Vincenza Aliperti, Anna Maria Carrese.
Preparation of the manuscript: Aldo Donizetti, Massimo Venditti.
Revision for important intellectual content: Marco De Sio, Michele Caraglia, Sergio Minucci, Francesco Aniello.
Supervision: Sergio Minucci, Francesco Aniello.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Ethics approval
The study was conducted in accordance with the Declaration of Helsinki and approved by the Institutional Ethics Committee of University of Campania “Luigi Vanvitelli” (protocol code 206 approved on 15 April 2019).
Consent to participate
A written informed consent was obtained from all subjects involved in the study.
