Abstract
BACKGROUND:
Aberrant expression of long non-coding RNAs is involved in the progression of ovarian cancer. However, the clinical significance and biological functions of SNHG3 expression was little known in ovarian cancer (OC).
METHODS:
The SNHG3 expression in ovarian cancer tissues and paired adjacent normal tissues was detected using quantitative real time polymerase chain reaction (qRT-PCR). Gain-of function and loss-of function assays were performed in ovarian cancer cells to demonstrate the effects of SNHG3 expression on cell proliferation and invasion. The relative protein expression levels were determined using western blot analyses.
RESULTS:
The expression of SNHG3 was significantly up-regulated in ovarian cancer tissues compared with adjacent normal tissues. Higher SNHG3 expression levels positively associated with FIGO stage, lymph node metastasis, and poor prognosis of ovarian cancer patients. Univariate and multivariate Cox regression analysis implied that FIGO stage, lymph node metastasis, higher SNHG3 expression were independent prognostic factors for overall survival (OS) rate in ovarian cancer patients. Gain-of function and loss-of function assays demonstrated that SNHG3 knockdown inhibited ovarian cancer cell proliferation and invasion abilities. However, SNHG3 overexpression promoted ovarian cancer cell proliferation and invasion abilities. Furthermore, cell proliferation and invasion related protein CyclinD1, CDK1, MMP9 and MMP3 were significantly downregulated after SNHG3 knockdown in ovarian cancer cells, while SNHG3 overexpression had reverse effects. In addition, SNHG3 functioned as an oncogene by regulating GSK3
CONCLUSIONS:
Taken together, our data provide that SNHG3 has potential clinical value of and may serve as target of ovarian cancer treatment.
Background
Ovarian cancer (OC) is one of the most common gynecological malignant tumors in female. About estimated 239,000 women were diagnosed with ovarian cancer in 2012, and 152,000 women died of this disease [1, 2]. Recently, in spite of some advances in therapies including risk-assessment, cytoreductive operation, chemotherapy and molecular-targeted interventions for ovarian cancer, the overall 5-year survival rate at advanced ovarian cancer is less than 30% [3]. Thus, exploring novel potential targets for ovarian cancer treatment is particularly important.
Long non-coding RNAs (lncRNAs) are reported to function as tumor suppressors, oncogenes, or both in human ovarian cancer [4, 5]. Such as, ectopic expressed long non-coding RNA H19 contributes to malignant cell behaviors of ovarian cancer [6]. HOXA11-AS, a novel long non-coding RNA, suppresses the oncogenic phenotype of epithelial ovarian cancer [7]. Long non-coding RNA MALAT1-regulated microRNA 506 and modulates ovarian cancer growth by targeting iASPP [8]. The other lncRNAs including NEAT1 [9], CCAT2 [10], PVT1 [11], and so on, also play crucial regulators in ovarian cancer progression.
Small nucleolar RNA host gene 3 (SNHG3) is an lncRNA whose potential function and mechanism in ovarian cancer remains unknown. In the previous study, Zhang et al. reported that SNHG3 correlates with malignant status and poor prognosis in hepatocellular carcinoma [12]. The long non-coding RNA SNHG3 functions as a competing endogenous RNA (ceRNA) sponging miR-182-5p to promote malignant development of colorectal cancer [13]. In the present study, the expression of SNHG3 was significantly up-regulated in ovarian cancer tissues. Higher SNHG3 expression levels predicted a poor prognosis of ovarian cancer. Moreover, SNHG3 overexpression promoted cell proliferation and invasion capacities by increasing CyclinD1, CDK1, MMP9 and MMP3 expression. SNHG3 functioned as an oncogene by regulating GSK3
Materials and methods
Patient tissue samples
A total of 76 human ovarian cancer tissues and adjacent normal tissues were collected from patients who underwent surgical resection at Department of Gynecology and Obstetrics, Zhujiang Hospital, Southern Medical University from March 2009 to April 2011. Ovarian cancer tissues were confirmed as epithelial ovarian carcinoma by two independent pathologists. None of patients underwent any treatment prior to surgical resection. The study was approved by ethics board of Zhujiang Hospital, Southern Medical University the study and written informed consent was obtained from all patients.
Cell lines culture and cell transfection
Human ovarian cancer (OC) cell lines (SKOV3, OVCAR3, A2780, and ES2) and one normal ovarian cell line, HOSE were purchased from Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences (Shanghai, China). Cells were cultured in RPMI-1640 supplemented with 10% fetal bovine serum (FBS) and added with 100 U/mLpenicillin and 100
RNA isolation and quantitative real-time PCR (qRT-PCR)
The Trizol reagent (Invitrogen) was used to extract RNA from tissues and cells. RNA was reversed to First-strand cDNA using the Reverse Transcriptase Kit (TAKALA, Dalian, China). The reaction was at 37
Cell proliferation assay
Cell Counting Kit-8 (CCK-8) kit (Dojindo, Tokyo, Japan) was used to assess cell viability after cell were transfected with si-NC and si-SNHG3-2 or pcDNA3.0-vector or pcDNA3.0-SNHG3. After cells transfection, the cells were seeded into 96 well plates at a concentration of 2000 cells/well. The CCK-8 reagent was added into the well at 24, 48, 72 and 96 h and incubated at 37
SNHG3 expression is higher in ovarian cancer and associates with poor prognosis. (A) Expression of SNHG3 was measured in 76 pairs of ovarian cancer tissues and adjacent normal tissues using qRT-PCR assay. (B) Expression of SNHG3 was measured in human ovarian cancer (OC) cell lines (SKOV3, OVCAR3, A2780, and ES2) and one normal ovarian cell line HOSE using qRT-PCR. (C) Kaplan-Meier survival curve and log-rank test were performed to determine the association between SNHG3 expression levels with overall survival rate of ovarian cancer patients. Data are shown as mean 
Cells were transfected with si-NC and si-SNHG3- 2 or pcDNA3.0-vector or pcDNA3.0-SNHG3 and then were seeded into 6 cm dishes (500 cells/wells). After cells were cultured for 7 days in a humidified atmosphere containing 5% CO
Cell invasion assay
Transwell chamber (50 mg/L BD Matrigel 1: 8 diluent) was used in the study. The cells at a concentration of 1
Western blot analysis
Cells were lysed using RIPA buffer (Roche, Indianapolis, IN, USA) that was added proteinase inhibitor cocktail (Roche, Indianapolis, IN, USA). Total cell protein (45
Statistical analysis
Statistical analyses were performed using SPSS 20.0 software and results were shown as mean
Relationship between clinicopathological feather and SNHG3 expression in 76 ovarian cancer patients was shown
Relationship between clinicopathological feather and SNHG3 expression in 76 ovarian cancer patients was shown
Univariate and multivariate Cox analysis for the overall survival (OS) time in 76 ovarian cancer patients
HR: hazard ratio, CI: confidence interval,
SNHG3 expression is higher in ovarian cancer tissues and associates with clinicopathological features
First, qRT-PCR analyses were performed to assess SNHG3 in 76 ovarian cancer tissues and adjacent normal tissues collected from patients who underwent surgical resection. As shown in Fig. 1A, the relative SNHG3 expression was higher in ovarian cancer tissues than that in the corresponding adjacent normal tissues (
Higher SNHG3 expression associates with poor prognosis of ovarian cancer patients
Furthermore, Kaplan-Meier survival curve and log-rank test were performed to determine the association between SNHG3 expression levels with overall survival (OS) rate of ovarian cancer patients. Our results demonstrated that higher SNHG3 expression significantly associated with poor OS rate compared with lower SNHG3 expression in ovarian cancer patients (Fig. 1C). Univariate and multivariate Cox regression model was performed to determine whether SNHG3 expression and other clinicopathologic feathers were independent prognostic markers for OS in ovarian cancer patients. Our results demonstrated that SNHG3 expression (HR
Upregulation of SNHG3 promotes cell proliferation and invasion of ovarian cancer
SNHG3 promotes cell growth of ovarian cancer in vitro. (A) and (B) The relative expression of SNHG3 was reduced using two siRNAs targeting SNHG3 in SKOV3 and A2780 cells. (C) The relative expression of SNHG3 was over-expressed using pcDNA3.0-SNHG3 in OVCAR3 cells. (D) and (E) The cell growth rates were determined by performing CCK-8 assay after knockdown of SNHG3 in SKOV3 and A2780 cells. (F) The cell growth rates were determined by performing CCK-8 assay after overexpression of SNHG3 in OVCAR3 cells, compared to relative control groups, respectively. Data are shown as mean 
SNHG3 promotes cell colony formation of ovarian cancer in vitro. (A) and (B) Colony forming assays were shown after knockdown of SNHG3 in SKOV3 and A2780 cells. (C) and (D) Colony forming assays were shown after overexpression of SNHG3 in OVCAR3 cells, compared to relative control groups, respectively. Data are shown as mean 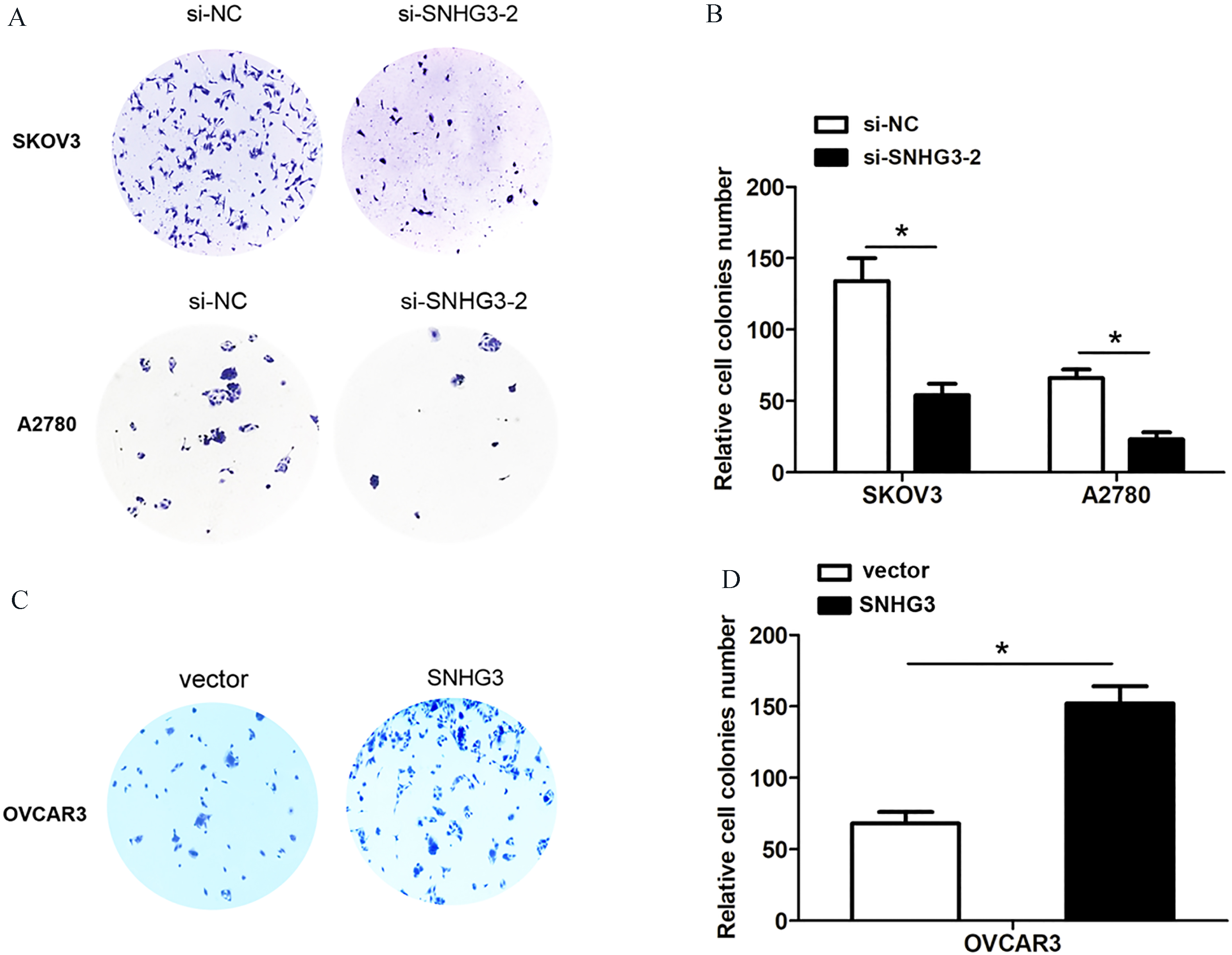
SNHG3 promotes cell invasion in ovarian cancer cells. (A) Cell invasion assay was shown after knockdown of SNHG3 in SKOV3 and A2780 cells compared to control group. (B) Cell invasion assay was shown after overexpression of SNHG3 in OVCAR3 cells, compared to control group. (C) and (D) Western blot analysis of CyclinD1, CDK1, MMP9 and MMP3 expression after knockdown of SNHG3 in SKOV3 cells or overexpression of SNHG3 in OVCAR3 cells. Data are shown as mean 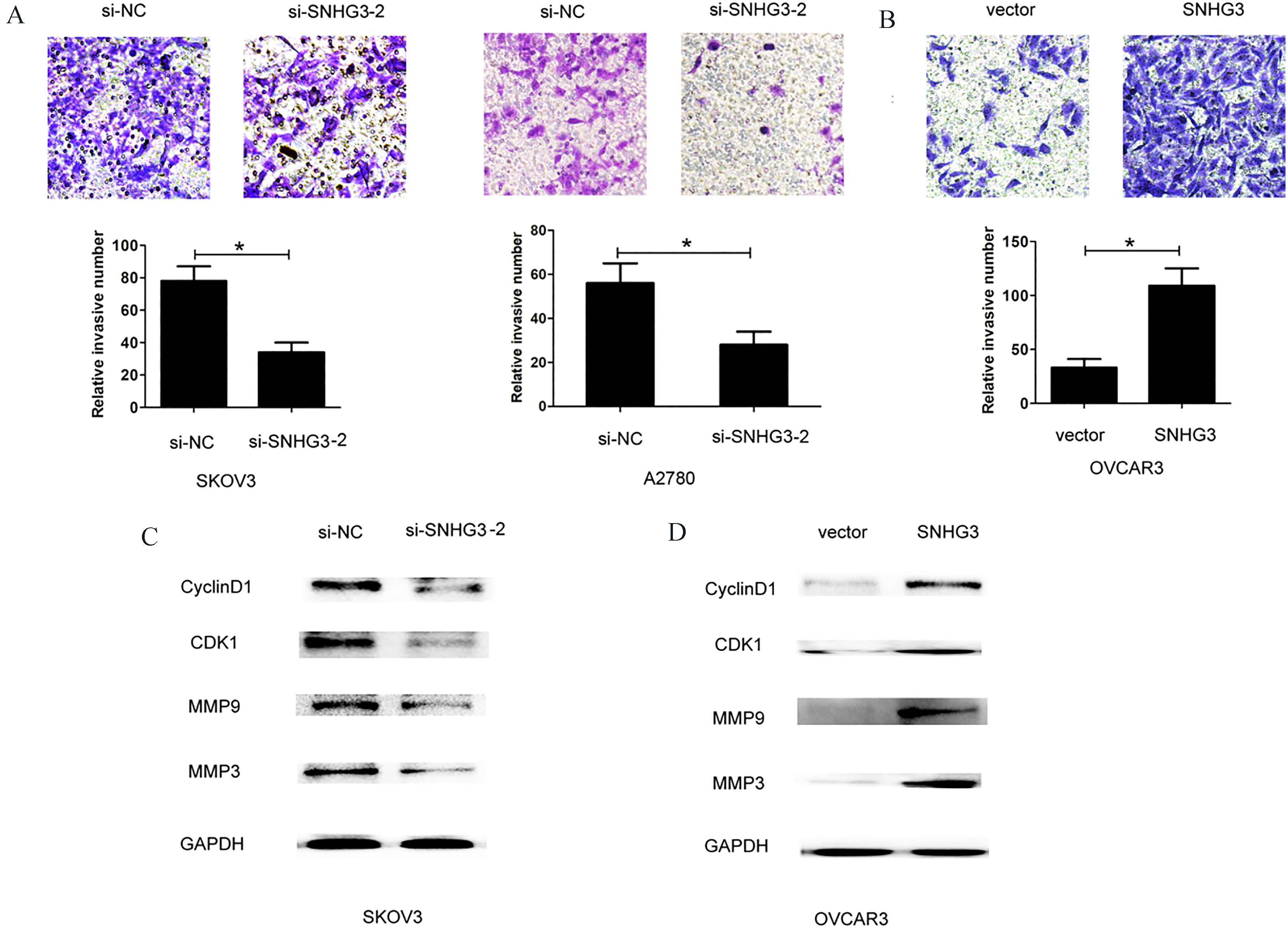
To investigate whether SNHG3 expression could affect cell proliferation and invasion of ovarian cancer, we performed lose-function and gain-function assays by transfecting two siRNA oligos targeting SNHG3 or si-negative control into SKOV3 and A2780 cells, or transfecting pcDNA3.0-vector or pcDNA3.0-SNHG3 into OVCAR3 cells (Fig. 2A and B), the siRNA-SNHG3-2 was used for following experiments due to higher silencing efficiency of SNHG3 in SKOV3 and A2780 cells. The CCK8 cell viability assay demonstrated that knockdown of SNHG3 using siRNA-SNHG3-2 inhibited cell growth rate in SKOV3 and A2780 cells, while SNHG3 over-expression by pcDNA3.0-SNHG3 enhanced cell growth rate in OVCAR3 cells (Fig. 2C and D). Cell colony forming assays showed that knockdown of SNHG3 using siRNA-SNHG3-2 reduced cell colonies number in SKOV3 and A2780 cells, while SNHG3 over-expression using pcDNA3.0-SNHG3 enhanced colonies number in OVCAR3 cells (Fig. 3A–D). To determine whether SNHG3 expression affected tumor cell invasion, we performed transwell invasion assays by knocking down SNHG3 in SKOV3 and A2780 cells or SNHG3 over-expression in OVCAR3 cells. The results demonstrated that SNHG3 knockdown suppressed cell invasion ability in SKOV3 and A2780 cells, whereas, SNHG3 over-expression enhanced cell invasion ability in OVCAR3 cells (Fig. 4A and B).
Additionally, we also explore the effects of SNHG3 expression on cell proliferation and invasion related protein CyclinD1, CDK1, MMP9 and MMP3. The results showed that CyclinD1, CDK1, MMP9 and MMP3 expression levels were significantly downregulated after SNHG3 knockdown in SKOV3 cells. However, SNHG3 over-expression enhanced their expression in OVCAR3 cells (Fig. 4C and D). Thus, these results demonstrated that SNHG3 promoted cell proliferation and invasion of ovarian cancer.
GSK3
SNHG3 functions as an oncogene by affecting GSK3
Emerging evidences have shown that lncRNAs play critical roles in several cancers. In ovarian cancer development and progression [14]. Qiu et al. reported that overexpression of long non-coding RNA HOTAIR predicts poor patient prognosis and promotes tumor metastasis in epithelial ovarian cancer [15]. Depletion of UCA1 was involved in the downregulation of matrix metallopeptidase 14 (MMP14) expression, a target gene of miR-485-5p in epithelial ovarian cancer [16]. Hua revealed that long noncoding RNA CCAT2 knockdown suppresses tumor progression by sponging miR-424 in epithelial ovarian cancer [17]. LncRNA MALAT1 promotes proliferation and metastasis in epithelial ovarian cancer via the PI3K-AKT pathway [18]. These evidences indicated that lncRNAs function as a regulator in ovarian cancer progression.
The long non-coding RNA SNHG3 was found to function as a competing endogenous RNA to promote malignant development of colorectal cancer [13]. In the present study, we demonstrated that the expression levels of SNHG3 were significantly up-regulated in ovarian cancer tissues compared with adjacent normal tissues. Higher SNHG3 expression levels predicted a poor prognosis of ovarian cancer patients. Furthermore, Gain-of function and loss-of function assays demonstrated that SNHG3 overexpression promoted cell proliferation and cell invasion. However, SNHG3 knockdown inhibited cell proliferation and cell invasion. Moreover, cell proliferation and invasion related protein CyclinD1, CDK1, MMP9 and MMP3 were significantly downregulated after SNHG3 knockdown in ovarian cancer cells. These findings showed that SNHG3 could affect ovarian cancer progression.
Additionally, SNHG3 knockdown increased levels of glycogen synthase kinase 3
In conclusion, our results demonstrated that SNHG3 was higher expression in ovarian cancer tissues and cells. Higher SNHG3 predicted a poor prognosis of ovarian cancer. Moreover, increased SNHG3 expression enhanced cell proliferation and invasion and activated GSK3
Footnotes
Acknowledgments
We thank the financial support from the Hainan Key Science and Technology Project (ZDXM2015069), the Hainan Natural Science Foundation-funded Project (813209), and the Hainan Health Department Research Project Approval (2012PT-09).
Conflict of interest
None.
