Abstract
BACKGROUND:
We have previously characterized esophageal carcinoma-related gene 4 (ECRG4) as a novel tumor suppressor gene, which is frequently inactivated in nasopharyngeal carcinoma and breast cancer. Nevertheless, the expression status and prognostic significance of ECRG4 maintain elusive in human gastric cancer. Herein, we examined ECRG4 expression profile in gastric cancer and assessed its association with clinicopathological characteristics and patient survival.
METHODS:
Online data mining, real-time RT-PCR and immunohistochemistry were employed to determined ECRG4 expression at transcriptional and protein levels in tumors vs. noncancerous tissues. Statistical analyses including the Kaplan-Meier survival analysis and the Cox hazard model were utilized to detect the impact on clinical outcome. Moreover, ECRG4 expression was silenced in gastric cancer SGC7901 cells, and cell proliferation, colony formation and invasion assays were carried out.
RESULTS:
ECRG4 mRNA and protein levels were obviously downregulated in cancer tissues than noncancerous tissues. Statistical analyses demonstrated that low ECRG4 expression was found in 34.5% (58/168) of primary gastric cancer tissues, which was associated with higher histological grade (
CONCLUSIONS:
Our observations reveal that ECRG4 expression is involved in gastric cancer pathogenesis and progression, and may serve as a candidate prognostic biomarker for this disease.
Introduction
Gastric cancer is the fourth most common type of cancer worldwide in terms of incidence and second in terms of mortality [1, 2]. The prognosis is heavily influenced by recurrence and metastasis. Surgery has so far been the only curative treatment choice for gastric cancer, and it must be conducted promptly. Currently, the 5-year overall survival rate of gastric cancer remains no more than 25% [2]. The molecular mechanisms underlying gastric tumorigenesis are mostly unknown. Consequently, developing novel molecular biomarkers during carcinogenesis and progression is urgently required to more accurately predict patient outcomes and individualize treatments for gastric cancer.
Esophageal carcinoma-related gene 4 (ECRG4), also called augurin or C2ORF40, was originally characterized via comparing gene expression differentiation between human normal esophageal epithelium and cancer tissues [3]. ECRG4 has been identified as a candidate tumor suppressor gene which is frequently inactivated in various cancer types, including human esophageal cancer, thyroid carcinoma, prostate cancer, breast cancer, gastric cancer, hepatocellular carcinoma, renal cell cancer, colorectal carcinomas and glioma, mainly due to its gene promoter methylation [4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15]. Consistently, we previously stated that promoter methylation-mediated silencing of ECRG4 in nasopharyngeal carcinoma and its overexpression inhibits the malignant phenotypes of cancer cells [16]. In addition, we observed that decreased ECRG4 expression is significantly correlated with unfavorable patient survival and is an independent prognostic indicator in breast cancer [17]. Our results suggest that ECRG4 might not only offer novel tailor-made therapeutic targets for cancers, but may also contribute to prediction of patient outcomes. Therefore, in the current study we first examined ECRG4 expression profiles in primary gastric cancer tissues. Following that, we analyzed these results in combination with clinicopathologic parameters and survival data. Finally, we elucidated the biological functions of silencing ECRG4 regarding cancer cell proliferation and invasion.
Materials and methods
Patients and tissue samples
20 gastric cancer tissues and paired adjacent noncancerous mucosa were collected from an independent cohort of gastric cancer patients undergoing endoscopy in 2020. These specimens were obtained and immediately processed using an RNALater reagent (Invitrogen; Thermo Fisher Scientific, Inc., Waltham, MA, USA) and kept at
Correlation between ECRG4 expression and clinicopathological variables
Correlation between ECRG4 expression and clinicopathological variables
ECRG4 mRNA expression was examined in various cancer types and cancer cell lines using the ONCOMINE database (
RNA extraction and reverse transcription- quantitative polymerase chain reaction (RT-qPCR)
Total RNA was extracted from cells transfected with ECRG4 siRNAs or scrambled siRNA using a TRIzol reagent (Invitrogen) and was reversely transcribed using an M-MLV First Strand kit (Invitrogen) according to the manufacturer’s protocols. The cDNA was subjected to qPCR with the following gene-specific primers: ECRG4 forward 5
Western blot analyses
This procedure has been described before [16, 17, 20, 26, 27, 28, 29, 30]. In brief, equivalent amounts of proteins extracted from cell lysates were separated using 10% SDS-PAGE and then transferred to PVDF membranes (Millipore, Billerica, MA, USA). After blocked in 5% skim milk in TBS containing 0.1% Tween-20 (TBST) buffer, the membranes were were incubated with the indicated primary antibodies overnight at 4
Immunohistochemistry and evaluation
Immunohistochemistry (IHC) was performed using a standard EnVision complex method described previously [17, 20, 23, 29, 30]. Briefly, after undergoing deparaffinization and rehydration, endogenous peroxidase blocking and antigen retrieval, 4-
The staining evaluation was conducted using the following standard: ten random 400
Cell culture and transfection
One established gastric cancer cell line SGC-7901 were cultured as described before [30]. A pre-designed validated siRNA against 5
Cell proliferation and colony formation assays
Cell proliferative and clonogenic capacities were determined using a 3-(4,5-dimethylthiazol-2-yl)-2 5-diphenyl-2H-tetrazolium bromide (MTT) colorimetric assay and a colony formation (soft agar culture) assay, respectively, as described previously [16, 26, 27, 28].
Cell invasion assay
Cell invasiveness was evaluated using a Matrigel invasion chamber assay as described before [6, 16, 26, 27, 28]. Briefly, cells were plated into 24-well Matrigel-coated Transwell inserts (EMD Millipore, Billerica, MA, USA) at the density of 1
Statistical analyses
Statistical analyses were conducted using the SPSS 26.0 statistical software package (SPSS Inc., Chicago, IL, USA). The difference of ECRG4 expression between tumors and nancancerous tissues was calculated by Student’s
ECRG4 is down-regulated in gastric cancer tissues. (A) The mRNA levels of ECRG4 in different cancer types. A graphic obtained from Oncomine indicates the numbers of datasets with significant over-expression (Red) or under-expression (Blue) at the transcriptional level in cancer tissues relative to normal tissues. Cell color was determined by the best gene rank percentile for the analyses within the cell, and the gene rank was analyzed by percentile of target genes in the top of all genes measured in each analysis. (B) ECRG4 was distinctively high expressed in gastric cancer cell lines according to CCLE analysis. (C) ECRG4 mRNA levels in gastric cancer were analyzed using the UALCAN database. NG, normal gastric (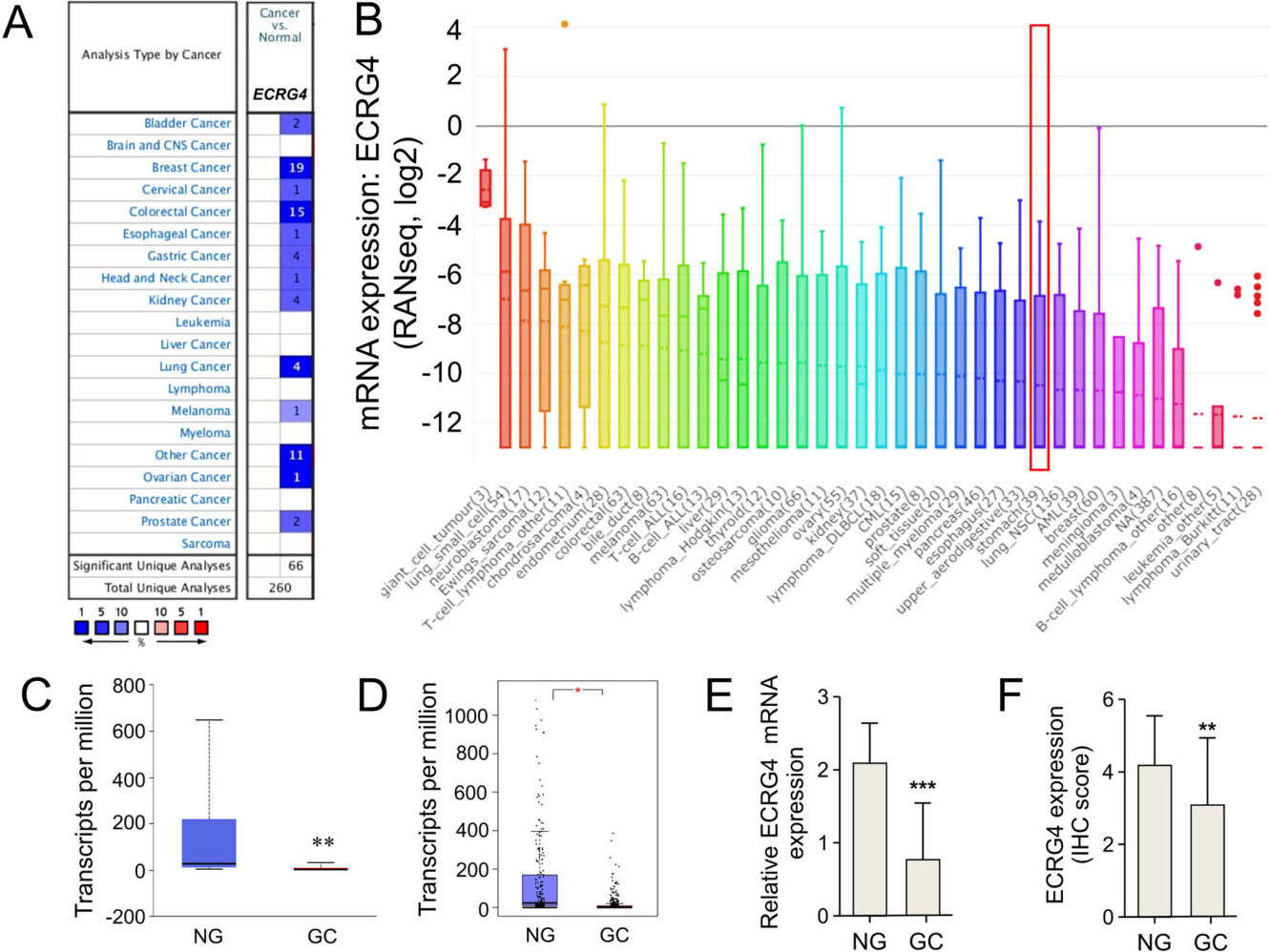
Genetic alterations of the ECRG4 gene and its association with patient survival in gastric cancer via the cBioPortal database analysis. (A) Oncoprint in cBioPortal represented the proportion and distribution of samples with ECRG4 gene alterations. The figure was cropped on the right to exclude samples without alterations. (B) Alterations frequency of the ECRG4 gene (mutation and amplification) in subtypes of gastric cancer. (C-F) The impacts of genetic alterations of ECRG4 on overall survival (C), disease-free survival (D), progression-free survival (E) and disease-specific survival (F) in patients with gastric cancer. (G) Analysis of the top 10 genes with concomitant alteration of ECRG4.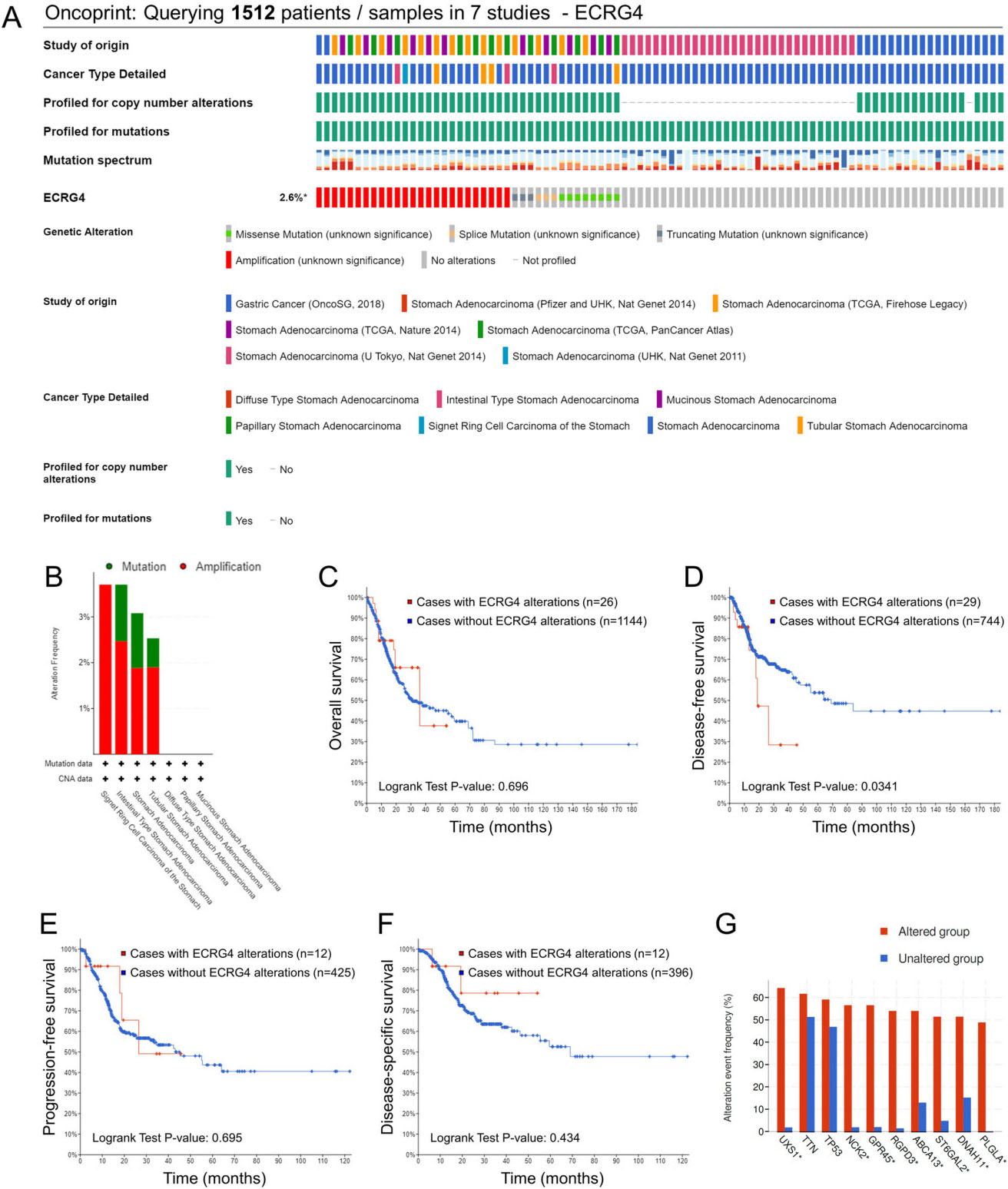
Copy number variance (CNV) analysis of the ECRG4 gene in gastric cancer as well as their impacts on overall survival. (A) Gene copy number of ECRG4 in GC was obtained from the Oncomine database. (B) CNV distribution in stomach adenocarcinoma (STAD) specimens. (C) Spearman correlation analysis between CNV and mRNA expression of ECRG4 in STAD. (D) Relationship between ECRG4 CNV and overall survival in STAD patients.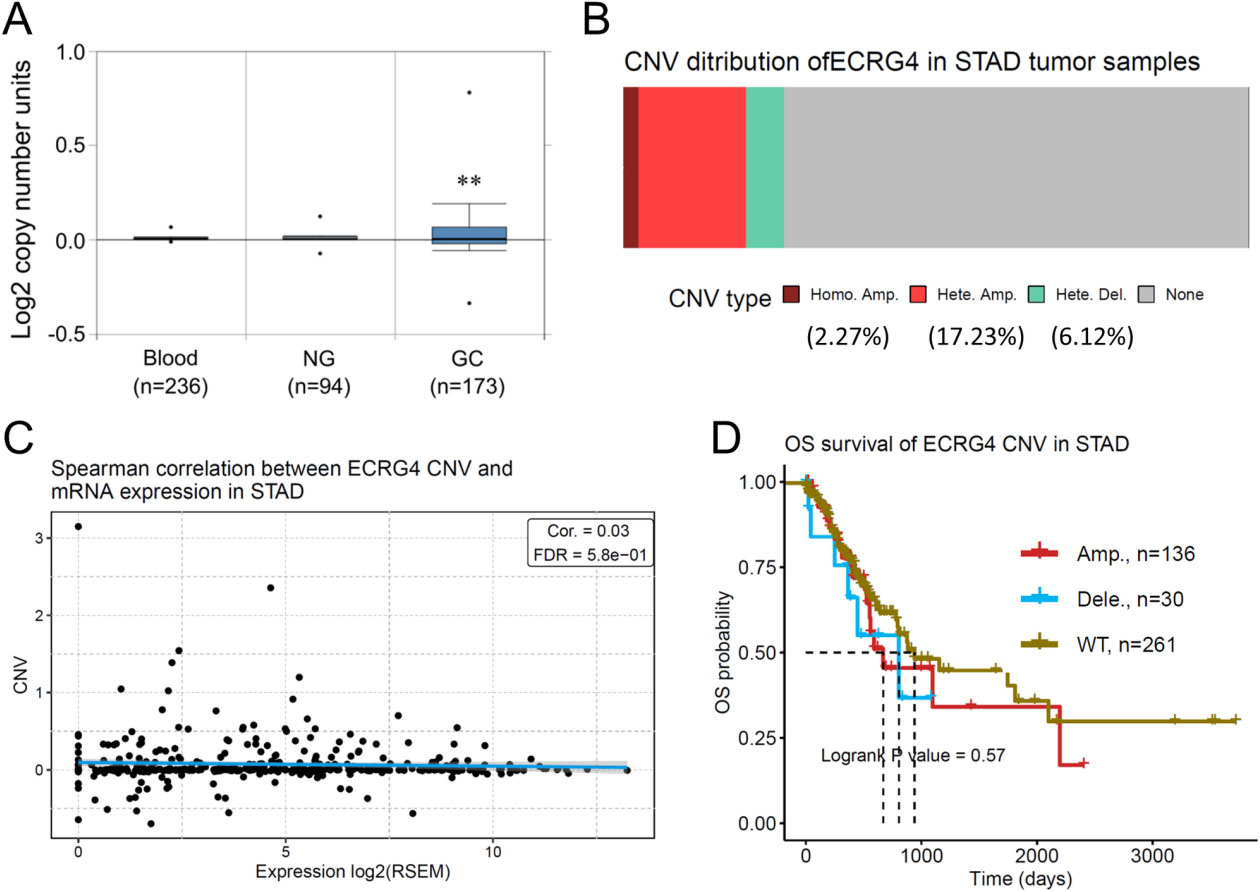
Immunohistochemical staining of ECRG4 in human gastric cancer tissues and adjacent noncancerous tissue. (A) Strong staining of ECRG4 in gastric cancer tissue. (B) Moderate ECRG4 staining in gastric cancer tissue. (C) Negative ECRG4 staining in gastric cancer tissue. (D) Positive ECRG4 staining in adjacent noncancerous tissue. Original magnification, 400 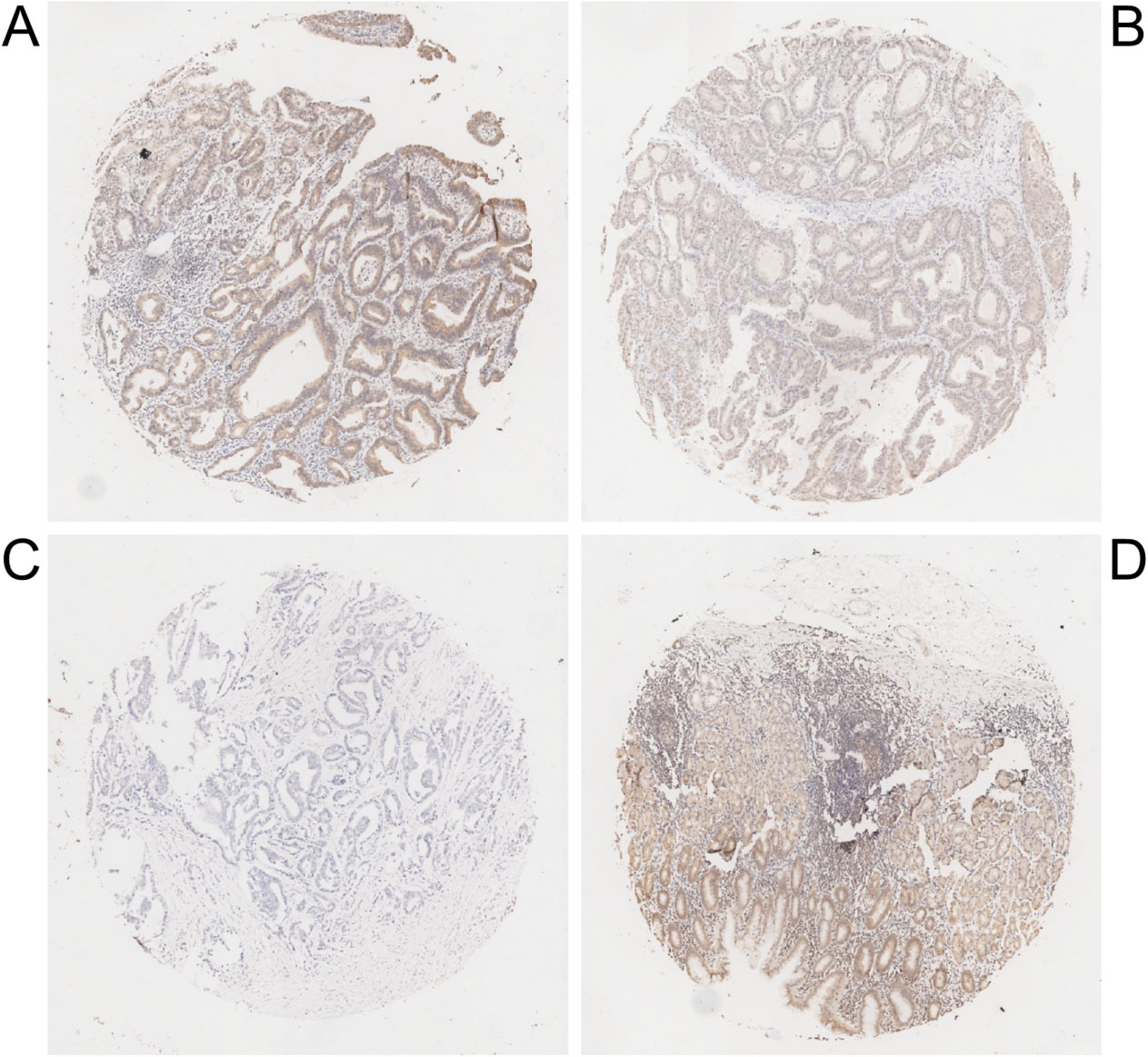
Differential expression analysis of ECRG4 in gastric cancer
We first performed differential expression analysis of ECRG4 in primary gastric cancer and adjacent noncanerous tissues using several online databases. As displayed in Fig. 1A, the Oncomine database revealed lower ECRG4 mRNA level in 24.6% of (64/260) unique analyses. Regarding gastric cancer, all four unique analyses indicated its down-regulation. The CCLE database analysis revealed that ECRG4 mRNA expression level in gastric cancer cell lines are listed in the 29
Impact of the ECRG4 gene alterations on patient survival
We next examined the gene alteration status of ECRG4 and its impact on patient survival in gastric cancer using the GSCA online database. The genetic alteration rate for the ECRG4 gene was found to be 2.6% in seven studies containing 1512 gastric cancer patient (Fig. 2A). Interestingly, ECRG4 gene amplification occurred most frequently in single ring cell carcinoma of the stomach, while ECRG4 gene mutations appeared most frequently in stomach adenocarcinoma, and especially in intestinal and tubular stomach adenocarcinoma (Fig. 2B). Following analyses by Kaplan-Meier plot with a log-rank test, no significant association was found between ECRG4 gene alteration and patient survival regarding overall survival, progression-free survival and disease-specific survival (Fig. 2C, E and F). Nevertheless, ECRG4 gene alteration was observed to be significantly associated with disease-free survival for gastric cancer (
Kaplan-Meier survival curves with univariate analyses (log-rank) according to the expression statues of ECRG4 in patients with gastric cancer. (A) The survival curve of ECRG4 expression in all gastric cancer patients. (B and C) The survival curves of ECRG4 expression in N0 and N1–N3 patients. (D and E) The survival analysis of ECRG4 expression in M0 patients. (F) The survival curve of ECRG4 in stage I/II and stage III/IV patients.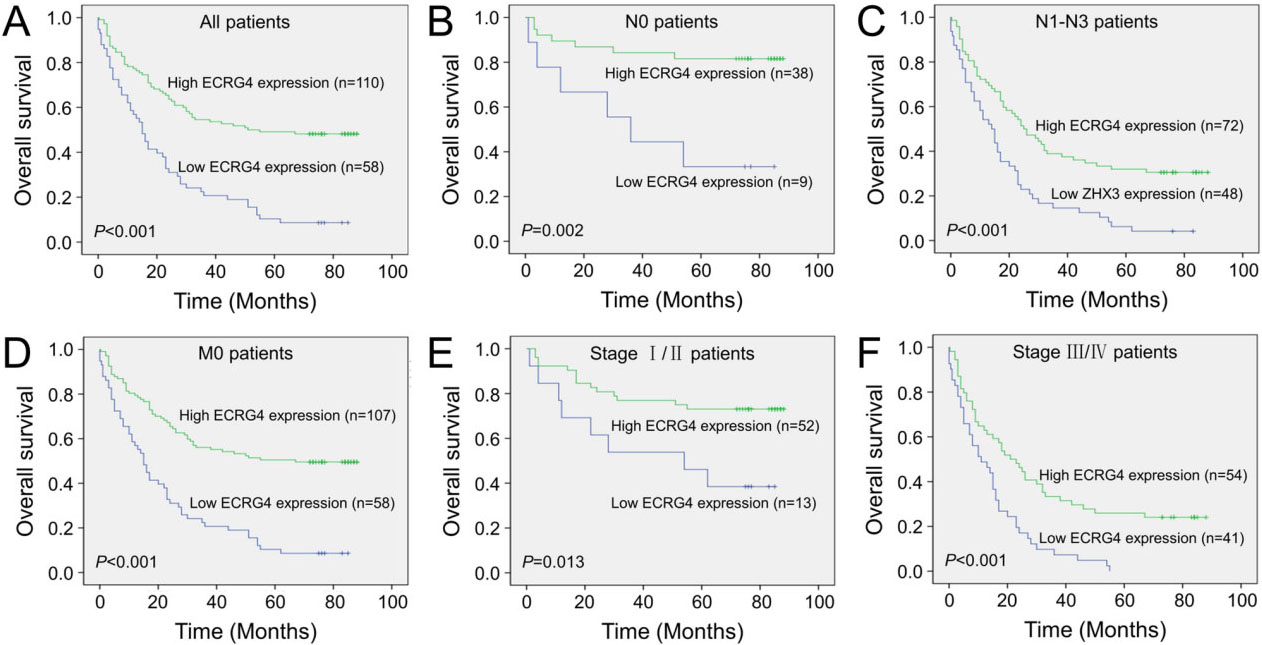
To corroborate the above findings, we characterized the CNV status of the ECRG4 gene in gastric cancer. The amplification of the ECRG4 gene in gastric cancer was observed by analyzing DNA copy numbers using the Oncomine database (Fig. 3A). In addition, the GSCA database analysis revealed that the CNV distribution rates for homo-amplification, hetero-amplification and hetero-deletion were 2.27%, 17.23% and 6.12%, respectively (Fig. 3B). Spearman correlation analysis revealed a negligible correlation between mRNA expression and CNV status of ECRG4 in gastric cancer (
We further assessed the correlation between ECRG4 expression and various clinicopathological characteristics in gastric cancer, using two tissue microarray chips containing total 180 primary gastric cancer specimens. However, 12 gastric cancer tissue samples in tissue microarray chips were lost during IHC staining. Positive ECRG4 immunostaining was found primarily in the cytoplasm of cancer cells (Fig. 4). In a cohort of 168 informative gastric cancer specimens, high ECRG4 expression was observed in 65.5% (110/168) of primary tumors, whereas low expression was found in 34.5% (58/168) of primary tumors. Low ECRG4 expression was significantly associated with older patient age (
To evaluate the prognostic impacts of ECRG4 expression on patient outcomes, Kaplan-Meier survival analyses was conducted. As demonstrated in Fig. 5A, high ECRG4 expression was observed to be correlated with a better overall survival rate in patients with gastric cancer (
Univariate and multivariate Cox proportional hazards model for overall survival in gastric cancer patients
Univariate and multivariate Cox proportional hazards model for overall survival in gastric cancer patients
HR, hazard ratio; CI, confidence interval.
To determine the feasibility that ECRG4 expression may act as a prognostic predictor for gastric cancer, we employed univariate and multivariate Cox-regression models for further analysis. On univariate analysis, we found that patient age (
We finally investigated the biological consequences of silencing ECRG4 expression in gastric cancer cells. The efficiency of ECRG4 knockdown using a pre-design validated siRNA was confirmed by western blot analysis (Fig. 6A). We found that SGC-7901 cells with silenced ECRG4 expression represented a significant increase in cell proliferation and colony formation than those transfected with non-specific control siRNAs (Fig. 6B and C). In addition, ECRG4 knockdown impact on cell invasiveness was assessed using a Matrigel assay, in which SGC-7901 cells with ECRG4 knockdown indicated a significant increase in invasiveness than their control cells (Fig. 6D). Western blot analysis revealed that ECRG4 knockdown led to markedl downregulation of representative junction proteins such as E-cadherin and
Knockdown of ECRG4 promotes gastric cancer cell growth and invasion. (A) Knockdown of ECRG4 in SGC-7901 cells as indicated western blot. Knockdown of ECRG4 promotes SGC-7901 cell proliferation assessed by MTT method (B) and colony formation assay (C). (D) Knockdown of ECRG4 promotes the invasive ability of SGC-7901 cells using a Matrigel assay. All experiments were repeated at least three times. siNS, non-specific control siRNA. **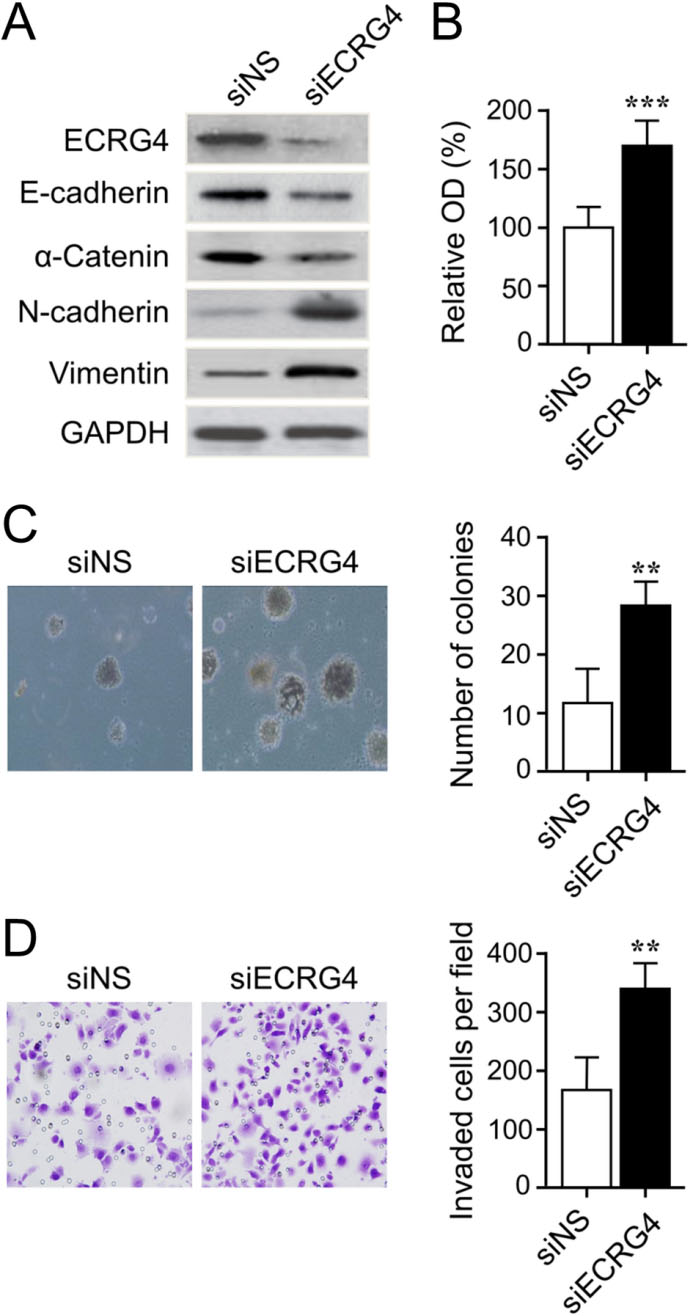
This study builds on our previous work to decipher the relationship between gene expression profiles and cancer outcomes, as well as to identify novel prognostic biomarkers and molecular targets. We were prompted to study ECRG4 because it frequently appeared on the top list of genes whose expression correlates with cancer prognosis. Based on our previous findings that exhibit evidenced functions of ECRG4 as a potential tumor suppressor [16, 17], we evaluated ECRG4 expression profile utilizing multiple approaches, e.g. in silico data-mining and IHC analysis. In this regard, this is the first report to examine ECRG4 as a molecular biomarker by means of combining these two avenues. The above results not only indicated downregulated ECRG4 expression in gastric cancer tissues vs. noncancerous tissues, but also demonstrated its strong correlation with metastasis and tumor advancement. Out data suggest that ECRG4 may serve as an independent prognostic indicator in gastric cancer patients, consistent with our recent report in breast cancer [17].
We utilized multiple avenues to reveal the first comprehensive analysis of ECRG4 in a cohort of 168 specimens from patients with primary resected gastric cancer and relevant online databases. We found that ECRG4 expression, at both mRNA and protein levels, was significantly down-regulated in tumor tissues than paired noncancerous tissues, which was supported by data mining, RT-qPCR and IHC analyses. The immunostaining patterns of ECRG4 observed in gastric cancer are similar to previous reports in esophageal cancer, hepatocellular carcinoma, renal cell cancer, colorectal cancer, and breast cancer [4, 12, 13, 14, 17, 31]. These findings suggest the notion that loss of ECRG4 expression may play an important role in pathogenesis and progression of gastric cancer. Our analyses discovered that decreased ECRG4 expression was associated with several clinicopathologic indicators, including higher histological grade, lymph node metastasis, invasive depth and advanced disease stage. This result was supported by similar reports in colorectal cancer and hepatocellular carcinoma [12, 14]. Yet, we did not observe a significant relationship between ECRG4 expression and distant metastasis, disagreeable with findings studied in renal cell cancer [13]. One possibility may be due to the actual composition of samples in which M1 samples were barely 1.8% (3/168). Further research with a larger sample scale of distant metastasis tumors may be more informative to clarify this issue.
Although a recent study has provided that decreased ECRG4 expression determined by IHC analysis was closely related with lymph node metastasis in gastric cancer, the prognostic implications of ECRG4 expression in gastric cancer remain missing [9]. In this study, decreased ECRG4 expression was observed to be correlated with unfavorable overall survival of patients with gastric cancer. The prognostic value of ECRG4 expression observed in gastric cancer is consistent with its essential role as an independent prognostic predictor for patient survival reported in esophageal cancer, breast cancer, renal cell cancer and nasopharyngeal carcinoma [4, 6, 13, 17, 32]. Through IHC analysis and in vitro cell experiments, we identified that ECRG4 expression may serve as a prognostic biomarker that also holds great therapeutic promise for gastric cancer.
Because gastric cancer patients usually present with locally advanced disease, biomarkers other than lymph node metastasis are required to forecast patient outcomes [33]. In this study, high ECRG4 expression was found to be associated with better overall survival rates in patients without lymph node metastasis or distant metastasis, and in stage I/II patients. These observations indicate that ECRG4 expression may be valuable in predicting the prognosis of patients with early-stage disease.
The alterations of DNA could contribute to the cancer initiation, progress, diagnosis, prognosis and therapy [20]. Through online database analysis, we found a negligible correlation between mRNA expression and CNV status of the ECRG4 gene in gastric cancer. As a recognized integrator and an indispensable node of major growth factor signaling pathways, ECRG4 is modulated by multiple layers of mechanisms such as transcriptional repression, gene promoter hypermethylation, genetic alteration and RNA splicing. The intrinsic differences in each type of tumors might be another possible explanation. Further, an unobvious FDR was not statistically supported the association and more strict setting may be needed to avoid excessive false positive.
In conclusion, our results reveal that ECRG4 expression is downregulated in gastric cancer and this alteration was strongly correlated with metastasis and tumor advancement. ECRG4 expression is an independent prognostic factor that holds therapeutic promise for gastric cancer. Clearly, further examination of ECRG4-modulated signaling pathways may promote our ability to assess the prognosis in patient with gastric cancer.
Availability of data and materials
The dataset used and/or analyzed in the present study is available from the corresponding authors on reasonable request.
Author contributions
Conception: Yanjie You.
Interpretation or analysis of data: Yanjie You, Shengjuan Hu.
Preparation of the manuscript: Yanjie You, Shengjuan Hu.
Revision for important intellectual content: Yanjie You, Shengjuan Hu.
Supervision: Yanjie You, Shengjuan Hu.
Footnotes
Acknowledgments
This work was supported by the Foundation of Ningxia Medical University (grant no. XZ2019011) and in part by the National Natural Science Foundation of China (grant no. 81860426), the Natural Science Foundation of Ningxia, China (grant no. 2021AAC03318), the Construction Project of Healthy Ningxia (grant no. 2021-NW-091) and the Key Research and Development Program of Ningxia, China (grant no. 2019BEG03005).
Conflict of interest
No potential conflicts of interest were disclosed.
