Abstract
BACKGROUND:
Metastasis often leads to poor prognosis in nasopharyngeal carcinoma (NPC) patients. Evidence has indicated the important roles of microRNA (miRNA) in cancer metastasis. The aim of this study was to identify and verify the key miRNAs that might be involved in the development of NPC metastasis.
METHODS:
Microarray data were obtained and analyzed to screen the differentially expressed miRNAs (DEMs) between NPC tissues with metastasis and those without metastasis. The target genes of the DEMs were predicted and their functions were annotated. Then, candidate hub genes were screened out through protein-protein interaction analysis, and the key miRNAs were identified. Afterwards, the expression levels of the key miRNAs were assessed by qRT-PCR based on an in vitro model.
RESULTS:
A total of 22 DEMs were screened out, and 616 target genes were predicted. Gene Ontology (GO) and pathway enrichment analysis showed that the target genes may be enriched in a diversity of GO terms and signaling pathways. Among them, eleven hub genes were identified, such as PTEN, KAT2B, CCND1, STAT3, and MAP3K5. Moreover, a five-miRNA profile (miR-106b, miR-17, miR-20b, miR-18a and miR-93) was identified and their expression levels were tested to be up-regulated in high-metastatic NPC cells relative to low-metastatic ones.
CONCLUSION:
The present study revealed that five miRNAs (miR-106b, miR-17, miR-20b, miR-18a and miR-93) and several hub genes such as PTEN, KAT2B, CCND1, STAT3, and MAP3K5, might play critical roles in the development of NPC metastasis. Future investigations are needed to confirm the results.
Introduction
Nasopharyngeal carcinoma (NPC) is a squamous cell malignancy arising from the epithelial cells of the nasopharynx, and its incidence is high in Southeast Asia. Approximately 70% of patients present with regional lymph node metastasis and 10% with distant metastasis when diagnosis [18]. Metastasis is an important influence factor for the tumor-node-metastasis staging (TNM) that has been regarded as the most important prognostic factor for NPC [10].
Multiple factors might contribute to NPC metastasis. For instance, up-regulated expression of cyclooxy- genase-2 might have a correlation with lymph node metastasis and disease progression in NPC [20]. ARHGAP42 promotes migration and invasion abilities of NPC cells in vitro, and its high expression is associated with poor metastasis-free survival of NPC patients [6]. Likewise, over-expression of NgR3 may promote NPC cell migration by suppressing E-cadherin, thus resulting in poor prognosis in NPC patients [5]. However, little has been known about the molecular mechanisms involved in this process.
Recently, microRNAs (miRNAs) have attracted much attention because they have been introduced to play a role in a variety of disorders. miRNAs are a class of short non-coding RNA (ncRNA) sequences that can bind to mRNAs and inhibit their expression, thus modulating various biological processes, such as cell proliferation, inflammation, and even the development of cancer [12]. A single miRNA may target multiple genes, and conversely, a single gene may be targeted by multiple miRNAs. The miRNAs and their target genes constitute a complex regulatory network and thus ensure the balance of various biological processes. Reports have shown that miRNAs play different roles in regulating the metastatic abilities of cancer cells. For example, miR-652 may promote endometrial cancer metastasis by targeting RORA [13]. Conversely, miR-302c-3p may suppress the migration ability of hepatocellular cancer cells by targeting TRAF4 [21]. Hence, the roles of various miRNAs were different in the regulation of cancer metastasis.
In the present study, we aimed to screen key miRNAs that play crucial roles in the development of NPC metastasis and assess their possible functions. First, differentially expressed miRNAs (DEMs) between tissues from NPC with metastasis and those without metastasis were screened out by analyzing the microarray-based data. Then, the target genes of the candidate DEMs were predicted and the hub genes were further screened out and assessed. The DEMs that might regulate the hub genes have been regarded as the key miRNAs. Afterward, the expression levels of the key miRNAs were verified in cell models.
Material and methods
Data source
To obtain DEMs between tissues from NPC patients with metastasis and those without metastasis, we downloaded the public gene expression profile (GSE36682) from the GEO database (
This dataset was deposited by Wei et al. in 2012, containing 17 samples from NPC patients with metastasis and 45 samples from NPC patients without metastasis. The experiment type was a ncRNA profiling based on the Human miRNA 1K platform.
Screening of DEMS
The data of GSE36682 were analyzed by GEO2R that is a web tool on the basis of limma R packages from the Bioconductor project. The results were downloaded in text format, in which the miRNAs that met the cut-off criteria of adjusted
MIRNA target gene prediction and functional annotation
To know the possible target mRNAs of the candidate DEMs, several computational microRNA-target prediction tools have been considered. mirDIP database was selected for use because it includes the integration of different tools that have been established for minimizing the shortcomings of individual databases [16].
Candidate mRNA with an integrated score of more than 0.9 was selected as a miRNA target in this database. To annotate the functions of the predicted target genes, Gene Ontology (GO) and KEGG pathway enrichment analysis were carried out through the Kobas tool [19]. A
Protein-protein interaction (PPI) network construction
To screen the possible hub genes/proteins in the development of metastasis, the target genes were submitted to STRING [15] and Cytoscape tools to predict the interaction relationship among them.
In the STRING tool, a combined score of not less than 0.40 (medium confidence score) was set as the cut-off criterion.
The hub genes/proteins were selected based on their relations with other proteins, which were sorted by the degree value in the network.
Cell culture
The clones of the human nasopharyngeal carcinoma CNE2 cell line, S18 (high-metastatic ability) and S26 (low-metastatic ability) [9], were conserved in our laboratory. Cells were cultured in RPMI-1640 media (Gibco, Carlsbad, CA, USA) containing 10% fetal bovine serum in a humidified atmosphere containing 5% CO
RNA extraction and MIRNA quantitative PCR
Total RNA was extracted from cultured cells with Trizol reagent (Invitrogen) according to the manufacturer’s protocol. The RNA concentration was quantified using a NanoDrop ND-1000 spectrophotometer (Nanodrop, Waltham, MA, USA). Only samples, with an OD A260/A280 ratio close to 2.0, were subsequently analyzed because it implies that the RNA was pure.
Reverse transcription (RT) was performed using a TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA) based on the manufacturer’s instructions. The human TaqMan MicroRNA Assay Kit (Applied Biosystems, Foster City, CA, USA) was used to quantify the amount of miRNAs. Quantification of miRNA was conducted by qPCR using an Applied Biosystems 7900HT thermocycler (Applied Biosystems). The PCR amplification conditions were presented as follows: 95
Statistical analysis
The expression levels of miRNAs between S18 and S26 were compared by the
Results
Identification of DEMS between NPC tissues with metastasis and those without metastasis
The dataset GSE36682 was retrieved from the GEO database, which contained a total of 62 NPC tissue specimens. These specimens were divided into two groups according to the metastasis status (17 metas- tasis and 45 non-metastasis specimens, respectively). The DEMs were screened out from the comparison of the two groups, respectively. There were 22 DEMs identified (12 up-regulated and 10 down-regulated miRNAs) (Table 1).
The DEMs screened from GSE36682
The DEMs screened from GSE36682
#A positive FC value indicates that the miRNA expression level was down-regulated in the metastasis group, whereas a negative FC value indicates that the miRNA expression level was up-regulated in the metastasis group, Note: Thresholds used in the screening analysis: adjusted
To predict the target genes of the identified DEMs, they were submitted to the mirDIP database for analysis. As a result, a total of 616 predicted targets of these miRNAs were identified according to the screening criteria.
To annotate the functions of these target genes, we submitted these gene IDs to the Kobas tool for analysis. As a result, these genes were enriched in 1481 GO terms. The top terms were binding, cell part, regulation of biological process, regulation of metabolic process, and regulation of cellular process (Fig. 1a).
Eleven hub genes sorted by degree and their relevant miRNAs
Eleven hub genes sorted by degree and their relevant miRNAs
Go analysis (a) and KEGG enrichment analysis (b) of the target genes.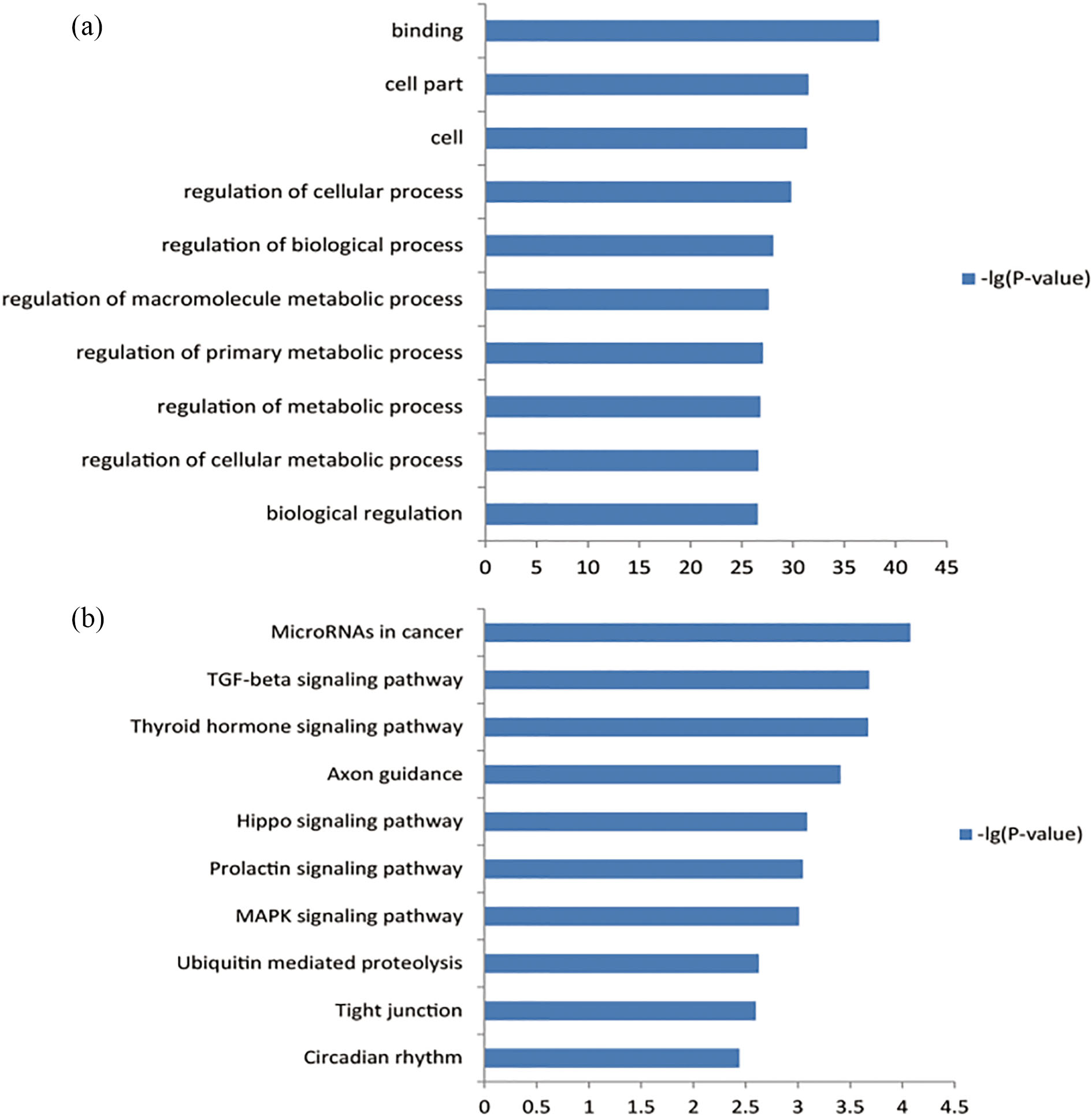
The pathway enrichment analysis showed that these target genes were enriched in 41 KEGG terms. The most significant dysfunctional pathways were varied, such as MicroRNAs in cancer, TGF-beta signaling pathway, Thyroid hormone signaling pathway, Axon guidance, and Hippo signaling pathway (Fig. 1b).
The results indicated that the molecular mechanisms underlying the development of metastasis were complicated. Multiple signaling pathways might be involved in this process.
Eleven hub genes sorted by degree from protein-protein interaction analysis.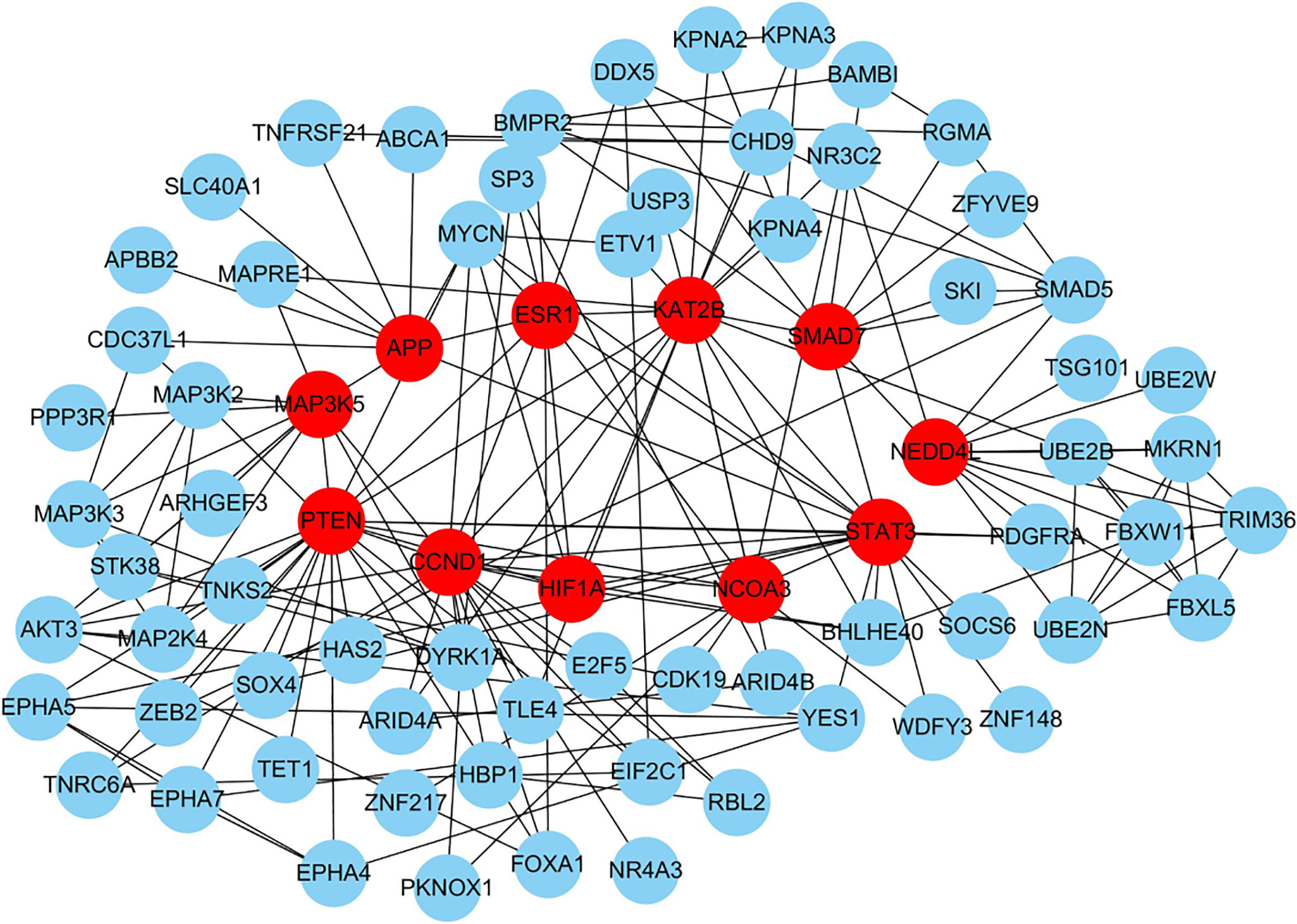
To screen the hub genes/proteins of the target genes, the genes were submitted to the STRING website for analysis. The genes with a confidence of 0.40 were selected for establishing PPI networks.
In the network, the top 11 node proteins, including PTEN, KAT2B, CCND1, STAT3, MAP3K5, NEDD4L, ESR1, HIF1A, APP, SMAD7, and NCOA3, showed a strong association with other proteins (with a degree value of not less than 10), suggesting that they have higher hub degrees than other genes (Table 2, and Fig. 2). Therefore, the hub genes/proteins may play important roles in the development of NPC cell metastasis.
To identify the DEMs that target the hub genes, the miRNAs were also listed in Table 2. As shown in this table, miR-106b, miR-17, miR-20b, miR-18a and miR-93 were selected as the key miRNAs.
Verification of key miRNAs by qRT-PCR
The high-metastatic nasopharyngeal carcinoma cell line S18 and the low-metastatic cell line S26 were cultured. The expression levels of the key miRNAs were tested by qRT-PCR.
The results showed that the expressions of these five miRNAs (miR-106b, miR-17, miR-20b, miR-18a, and miR-93) were up-regulated in S18 cells compared with those in S26 cells, respectively, in accordance with the microarray-based data (Fig. 3).
The expression levels of the 5 key miRNAs in S18 and S26 cells, as determined by qRT-PCR. 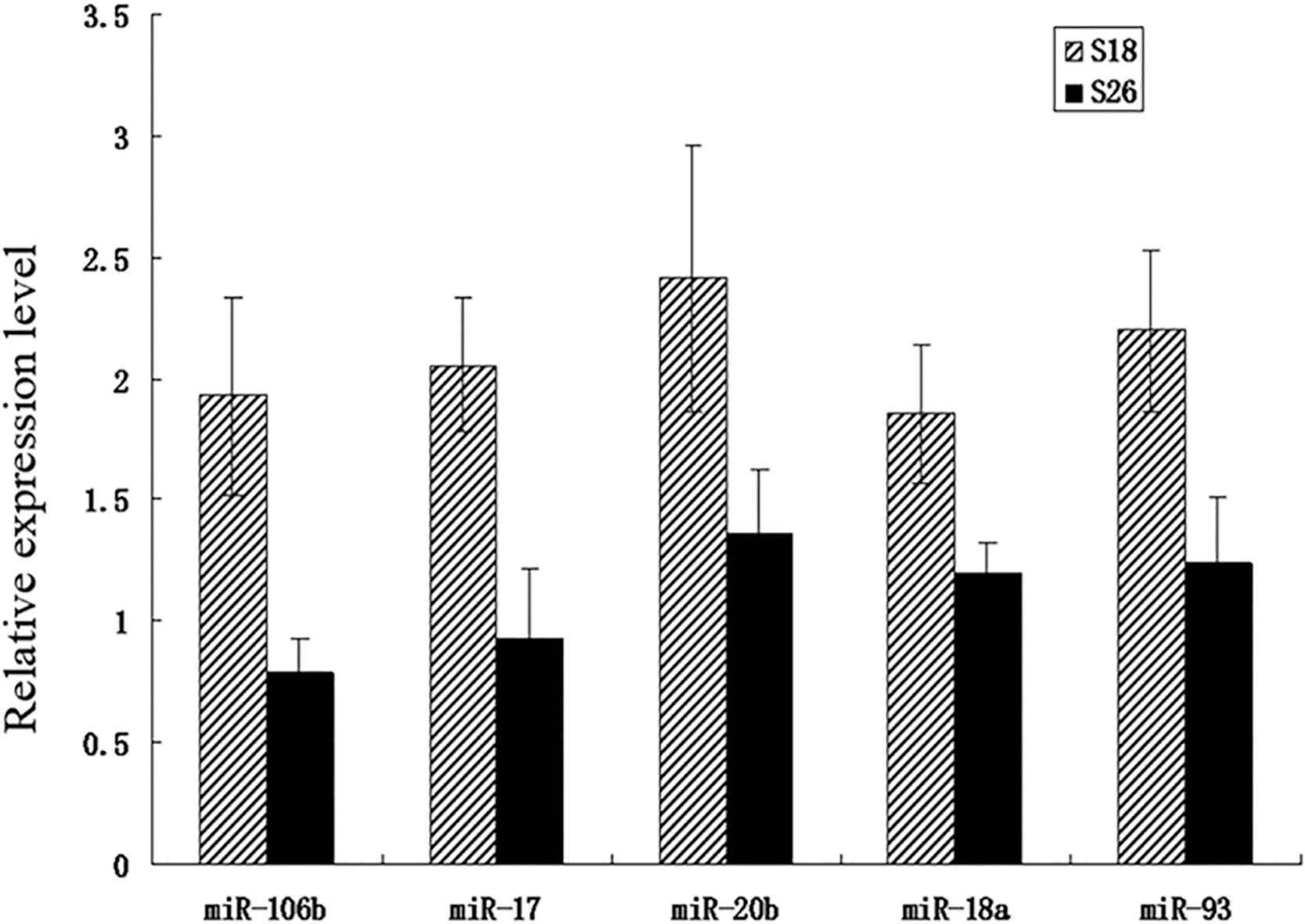
The occurrence of metastasis in NPC patients is a serious problem. The molecular mechanisms of metastasis have not been fully understood. In the present study, we screened out a number of miRNAs and their target genes that might play a role in the development of metastasis by using bioinformatics approaches. As a result, a total of 22 DEMs were identified. These miRNAs and their target genes might be enriched in a number of GO terms, such as binding, cell part, regulation of biological process, regulation of metabolic process, and regulation of cellular process. Moreover, these genes might be enriched in a diversity of signaling pathways, such as MicroRNAs in cancer, TGF-beta signaling pathway, Thyroid hormone signaling pathway, Axon guidance, and Hippo signaling pathway. Among these target genes, eleven hub genes were identified, such as PTEN, KAT2B, CCND1, STAT3, and MAP3K5. Additionally, a five-miRNA profile (miR-106b, miR-17, miR-20b, miR-18a and miR-93) that may target the hub genes were identified as the key miRNAs and their expression levels were further verified by qRT-PCR.
Evidence indicates that miRNAs might have an effect on the invasive and metastatic abilities of NPC cells. For instance, low miR-342 expression was associated with distant metastasis in NPC patients, and elevated expression of miR-342 may inhibit NPC cell growth and invasion by targeting ZEB1 [23]. miR-328 can suppress NPC cell migration and inhibit the epithelial-mesenchymal transition (EMT) process by targeting CD44 [8]. Likewise, miR-203a-3p inhibits metastasis via targeting LASP1 in NPC [7]. Nevertheless, not all the miRNAs and their target genes can play a central role in mediating cancer cell metastasis. To find the key miRNAs that might target the hub genes, we used bioinformatics methods for identifying them.
MiRNAs can bind to the 3’-untranslated region of the target mRNAs and thus suppress their translation. Hence, screening the target genes and exploring their possible functions may help us better understand the roles of miRNAs in the development of metastasis. A total of 616 targeted genes of the 22 DEMs were predicted. Of these genes, eleven ones were calculated to be the hub genes, namely, PTEN, KAT2B, CCND1, STAT3, MAP3K5, NEDD4L, ESR1, HIF1A, APP, SMAD7, and NCOA3. These hub genes have been suggested to play a role in the metastasis of cancers. For instance, the down-regulation of PTEN might promote ovarian cancer cell migration [14]. KAT2B may regulate miR-200c/141 and influence tumor metastasis in breast cancer [22]. CCND1 expression correlates with occult nodal metastases in oral cancer [11]. Collectively, these hub genes might play crucial roles in the development of NPC metastasis.
Since the hub genes may act as central factors in the metastasis of NPC, the DEMs that target them might be regarded as the key miRNA in this process. As shown in Table 2, a total of five miRNAs (miR-106b, miR-17, miR-20b, miR-18a, and miR-93) were identified as the key miRNAs. Evidence showed that these five miRNAs may be involved in cancer metastasis. Reports showed that miR-106b promotes the EMT process by targeting Smad 7 and thus facilitates the migration of esophageal cancer cells [2]. miR-17 can promote tumor metastasis in colorectal cancer by targeting PTEN [3]. miR-20b significantly induces the colony formation and invasiveness of breast cancer cells and contributes to brain metastasis [1]. miR-18a can promote breast cancer metastasis, while down-regulation of miR-18a by ginsenoside Rd may attenuate breast cancer metastasis [17]. miR-93 can also promote gastric cancer metastasis by targeting TIMP2 [4]. Taken together, these key miRNAs may play critical roles in cancer metastasis. In the present study, the in vitro experiment verified that the expression levels of these key miRNAs were up-regulated in high-metastatic NPC cells relative to low-metastatic NPC cells, respectively, suggesting that these five key miRNAs may also play a crucial role in the development of NPC metastasis.
Several limitations might be involved in the present study. First, the present study only concerned miRNA, one type of ncRNAs. Future studies are required to explore the roles of other ncRNAs, such as LncRNAs and ceRNAs, in the metastasis of NPC. Second, circulating miRNAs, as non-invasive biomarkers for disease detection, have attracted much attention. Future studies concerning circulating miRNAs for cancer stage diagnosis and prognosis would be of great value for NPC research. Third, only the expression levels of the key miRNAs were tested in a pair of cell lines. Given that validation in limited kinds of cell lines may overestimate the differential expressions of the miRNAs, future studies using more cell lines are needed to overcome the shortcomings. Moreover, in vivo experiments are also required because in vitro experiments cannot simulate the situations in vivo.
Despite the limitations, the present study provides a very useful clue for us to study the metastatic mechanisms in NPC. The discovery of the present study might have several impacts on clinical practice. First, the identified miRNAs might be used as biomarkers predicting the metastatic potential of NPC. Second, these miRNAs might have prognostic values. Since they may play crucial roles in the metastasis of NPC, they may also act as prognostic indexes for the patients. Third, these miRNAs might be used as potential treatment targets. However, further investigations are warranted to confirm these hypotheses.
In summary, five miRNAs (miR-106b, miR-17, miR-20b, miR-18a and miR-93) were identified as key miRNAs, and several genes, such as PTEN, KAT2B, CCND1, STAT3, and MAP3K5, were identified as hub genes in the initiation and development of NPC metastasis. The key miRNAs were verified by a cell model to be up-regulated in high-metastatic NPC cells compared with low-metastatic ones. Future studies are needed to confirm the conclusions.
Footnotes
Acknowledgments
This study was partly supported by the Special funding from China science foundation for postdoctors (2015T80962).
Conflict of interest
The authors declare that they have no conflict of interest.
