Abstract
BACKGROUND:
Circulating microRNAs (miRNAs) prove to be potential non-invasive indicators of cancers. The purpose of this study is to profile serum miRNA expression in breast cancer (BC) patients to find potential biomarkers for BC diagnosis.
METHODS:
The miRNA expression patterns of serum samples from 216 BC patients and 214 normal control subjects were compared. A four-phase validation was conducted for biomarker identification. In the screening phase, the Exiqon miRNA qPCR panel was employed to select candidates, which were further analyzed by quantitative reverse transcriptase PCR in the following training, testing, and external validation phases.
RESULTS:
A 12-miRNA (let-7b-5p, miR-106a-5p, miR-19a-3p, miR-19b-3p, miR-20a-5p, miR-223-3p, miR-25-3p, miR-425-5p, miR-451a, miR-92a-3p, miR-93-5p, and miR-16-5p) panel in serum was constructed. The diagnostic performance of the panel was assessed using ROC curve analyses. The area under the curves (AUCs) were 0.952, 0.956, 0.941 and 0.950 for the four separate phases, respectively. Additionally, the expression features of the 12 miRNAs were further explored in 32 pairs of BC tumor and para-tumor tissues, and 32 pairs of serum exosomes samples from patients and healthy subjects. miR-16-5p, miR-106a-5p, miR-25-3p, miR-425-5p, and miR-93-5p were highly overexpressed and let-7b-5p was conversely downregulated in tumor tissues. Excluding miR-20a-5p and miR-223-3p, the 10 other miRNAs were all significantly upregulated in BC serum-derived exosomes.
CONCLUSION:
A signature consisting of 12 serum miRNAs was identified and showed potential for use in non-invasive diagnosis of BC.
Introduction
Clinical characteristics of 216 BC patients and 214 NCs in the study
Clinical characteristics of 216 BC patients and 214 NCs in the study
BC: breast cancer; NC: normal control.
Breast cancer (BC) has caused a heavy health burden for females in both developed and developing countries [1, 2]. According to the 2018 Global Cancer Statistics, BC is the most frequently diagnosed cancers and remains one of the leading causes of cancer-related deaths for females worldwide [3]. In China, it is now the most frequently diagnosed cancer among women, and has shown a consistent increase in global morbidity and mortality rates in recent decades [4]. Despite effective treatments such as surgical operation in early stages, many patients remain undiagnosed until the disease develops distal metastases [5]. Thus, in some ways, early diagnosis, rather than delayed treatment, can provide a greater benefit for BC patients. Conventional diagnostic strategies often show high rates of false positives or over-diagnosis; this may cause direct or indirect damage to patients, greatly limiting their range of applications [6, 7, 8, 9]. Commonly used tumor biomarkers like cancer antigen 15-3 also have some limitations, including low sensitivity and specificity for BC diagnosis [10]. Therefore, it is very important to discover novel non-invasive biomarkers.
MicroRNAs (miRNAs) are families of small, single-stranded RNA molecules (containing about 22 nucleotides) that regulate gene expression by binding target mRNA sequences [11]. They participate in many cancer-related processes including apoptosis, differentiation, and cell cycle regulation [12, 13, 14]. The dysregulation of specific miRNAs is reported to be majorly important in tumor development and progression [15]. Importantly, the stable presence of miRNAs in the blood enables them to be potent signatures for cancer detection, including BC [16, 17]. Different studies have reported various circulating miRNAs (e.g. miR-21, miR-145, miR-373, and miR-199a) as potential biomarkers for BC [18, 19, 20, 21]. However, these results were often inconsistent, possibly because they analyzed different subtypes and stages of the disease, or the studies had different designs and subjects. Furthermore, many studies only included a small number of cases and controls [17]. Systematic research with a larger study cohort and a more precise study method needs to be performed for the discovery of more reliable biomarkers.
Here, we conducted a four-phase experiment to profile serum miRNA expression using the miRCURY Ready-to-Use PCR, Human panel I
Study subjects
Totally, 248 histopathologically diagnosed BC patients (all previously untreated) and 214 healthy normal controls (NC) were included in the study. All participants were recruited from the Jiangsu Provincial People Hospital during 2016 and 2017. All experiments were conducted under the approval of the Institutional Ethical Committee. Written informed consent was obtained from each participant.
Serum samples (216 BC vs. 214 NCs) obtained from the participants were divided into four sets. Table 1 shows the clinical features of the study subjects in each set. The distribution of patients’ age, TNM stage, grade, or molecular subtypes was unbiased across the four phases with
Study design
The experiment was performed in four phases: the screening, training, testing, and external validation phases. The flow chart of experiment design is shown in Fig. 1.
Flow chart of the experimental design. BC: breast cancer; NC: normal control; qRT-PCR: quantitative reverse transcription PCR.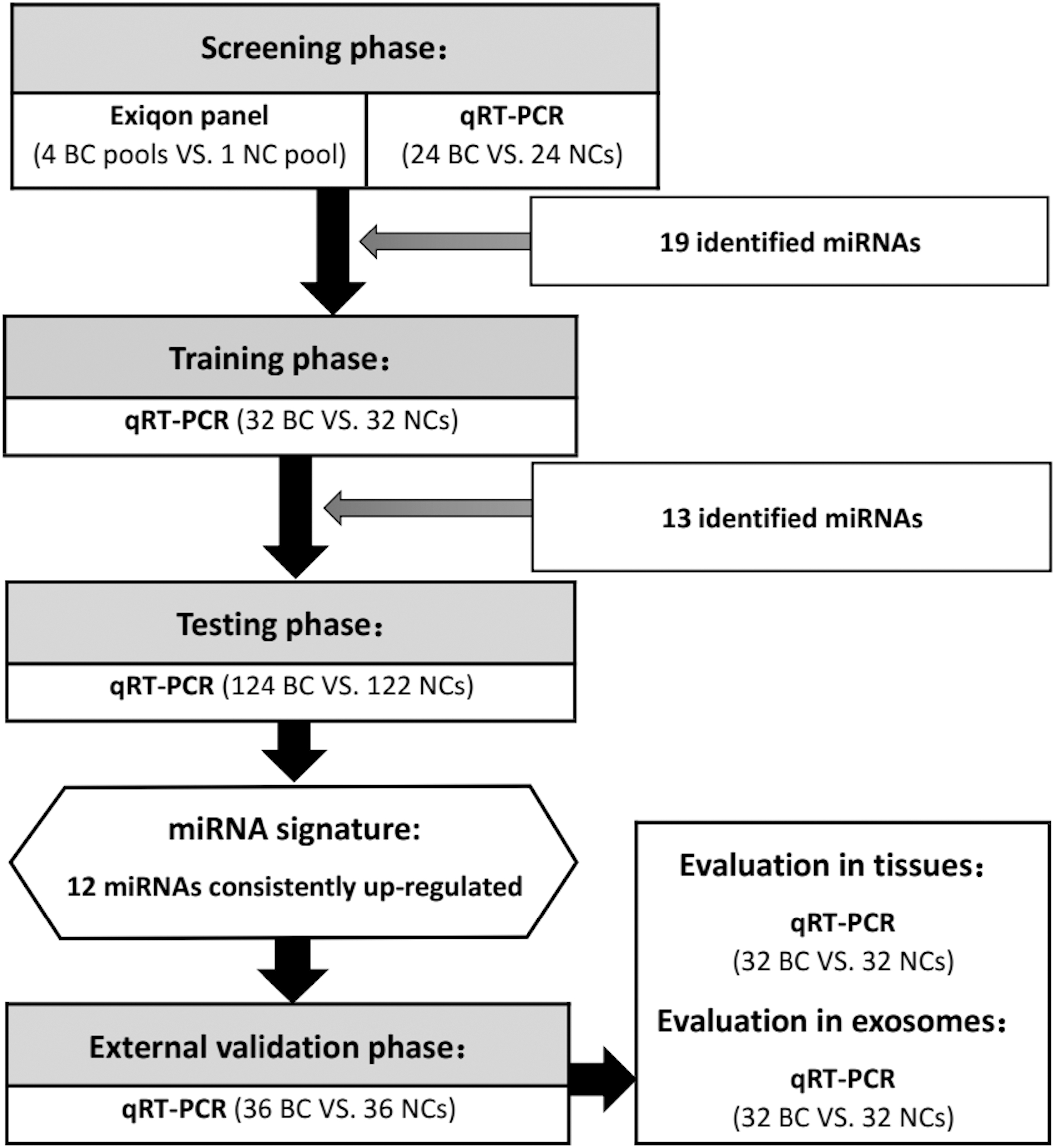
The initial screening phase was performed in two steps: first, possible candidate miRNAs were identified using the commercial plasma/serum miRNA profiling panel (Exiqon miRCURY-Ready-to-Use PCR-Human-panel-I
The training (32 BC vs. 32 NCs) and testing phases (124 BC vs. 122 NCs) were designed for precise confirmation of promising serum miRNA biomarkers. Based on the candidate list obtained in the screening stage, the expression of each candidate miRNA was validated using qRT-PCR. The external validation phase (36 BC vs. 36 NC) was designed for the final verification of the identified miRNA signatures. Moreover, 32 pairs of BC tumor and para-tumor tissue samples, as well as serum exosomes samples (32 BC vs. 32 NCs) were analyzed to explore the expression of the identified miRNAs.
Serum samples were separated from whole blood (5 ml) samples collected with SST Advance tubes after clotting for 30 min (BD Co., Franklin Lakes, NJ, USA), using a centrifugation process (3000
Exosomes samples were isolated from serum using the Exo-Quick Exosome Precipitation Solution (System Biosciences, Mountain View, CA, USA). According to the manufacturer’s protocol, 50
RNA extraction
The mirVana PARIS Kit (Ambion, Austin, TX, USA) was used to extract total RNA from serum and exosome samples following the manufacturer’s protocol. For tissue specimens, Trizol (Invitrogen, Carlsbad, CA, USA) was used for RNA extraction. 5
Quantitative reverse transcription PCR (qRT-PCR)
The extracted miRNAs were amplified using the Bulge-Loop miRNA qRT-PCR Primer Set (RiboBio, Guangzhou, China) on the LightCycler 480 Real-Time PCR System (Roche Diagnostics, Basel, Switzerland). The procedure of RT reaction was 42
Statistical analysis
Differences in miRNA expression was assessed using the Wilcoxon rank-sum test for serum and exosomes samples. The paired nonparametric test was performed to analyze paired tissue samples. The one-way analysis of variance or chi-square test were conducted to analyze the association between the identified miRNAs and the demographic and clinical features of the BC patients. A logistic regression model was then built to construct a BC-related predictive panel. A receiver-operating characteristic (ROC) curve was constructed, and the area under the ROC curve (AUC) was calculated to assess the predictive performance of the identified miRNA signature for BC diagnosis. SPSS 20.0 (SPSS Inc., Chicago, IL, USA) and GraphPad Prism 7.0 (GraphPad Software, San Diego, CA, USA) were employed for analyzing data and constructing graphs. A two-sided value of
Bioinformatics analysis
The potential targets and involved pathways of the identified miRNAs were predicted using DIANA-miRPath v3.0, a tool for the analysis of miRNA-involved pathways. Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) analyses were conducted on the basis of experimentally verified miRNA-pathway interactions from the DIANA-TarBase7.0.
Results
MiRNA quantification in screening phase
In the screening phase, a total of 28 dysregulated miRNAs were identified using the Exiqon miRNA quantification platform, with 22 upregulated and 6 downregulated. Then the 28 miRNAs were re-confirmed by qRT-PCR using 24 BC and 24 NC serum samples. Finally, a total of 19 miRNAs with consistent aberrant expression in BC serum samples were identified as candidate miRNAs (Supplementary Table S1).
Identification of miRNA biomarkers in serum by qRT-PCR
In the training phase, 13 (let-7b-5p, miR-106a-5p, miR-19b-3p, miR-19a-3p, miR-20a-5p, miR-223-3p, miR-25-3p, miR-425-5p, miR-451a, miR-92a-3p, miR-93-5p, miR-16-5p, and miR-24-3p) out of the 19 miRNAs were consistently upregulated in BC serum samples compared with NCs (
Expression levels of the 12 serum miRNAs in the training and testing phases (presented as mean
SD;
CT, relative to combination of cel-miR-39 and miR-1228)
Expression levels of the 12 serum miRNAs in the training and testing phases (presented as mean
BC: breast cancer; NC: normal control; FC: fold change.
Expression levels of twelve identified serum miRNAs in the combined training and testing phases. N: normal control; T: tumor; Horizontal line: mean with 95% CI.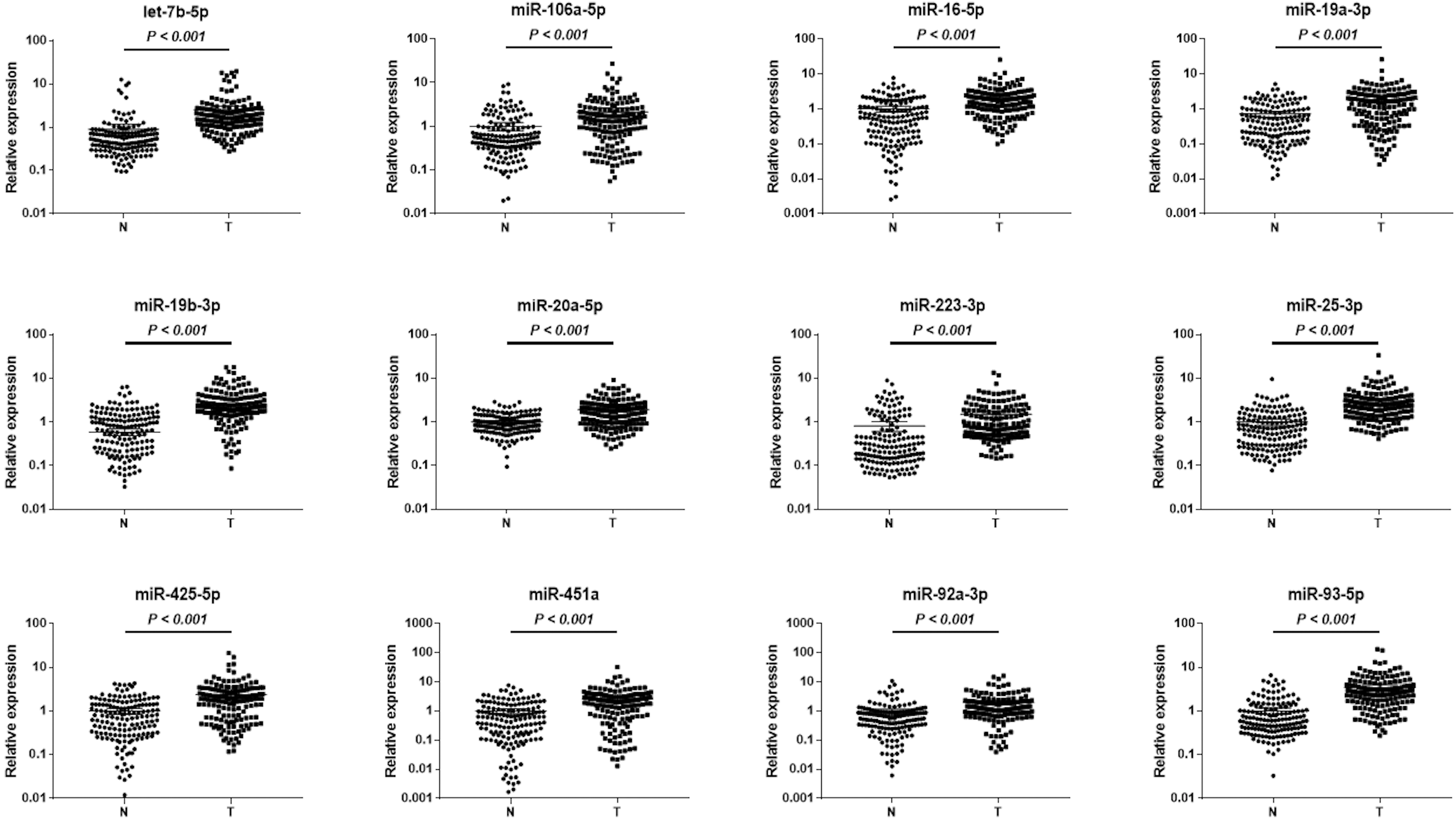
Receiver operative characteristic (ROC) curve analysis of the 12-miRNA panel for breast cancer (BC) detection. A: Combined training and testing phase cohorts; B: Training phase; C: Testing phase; D: External validation phase. AUC: area under the ROC curve.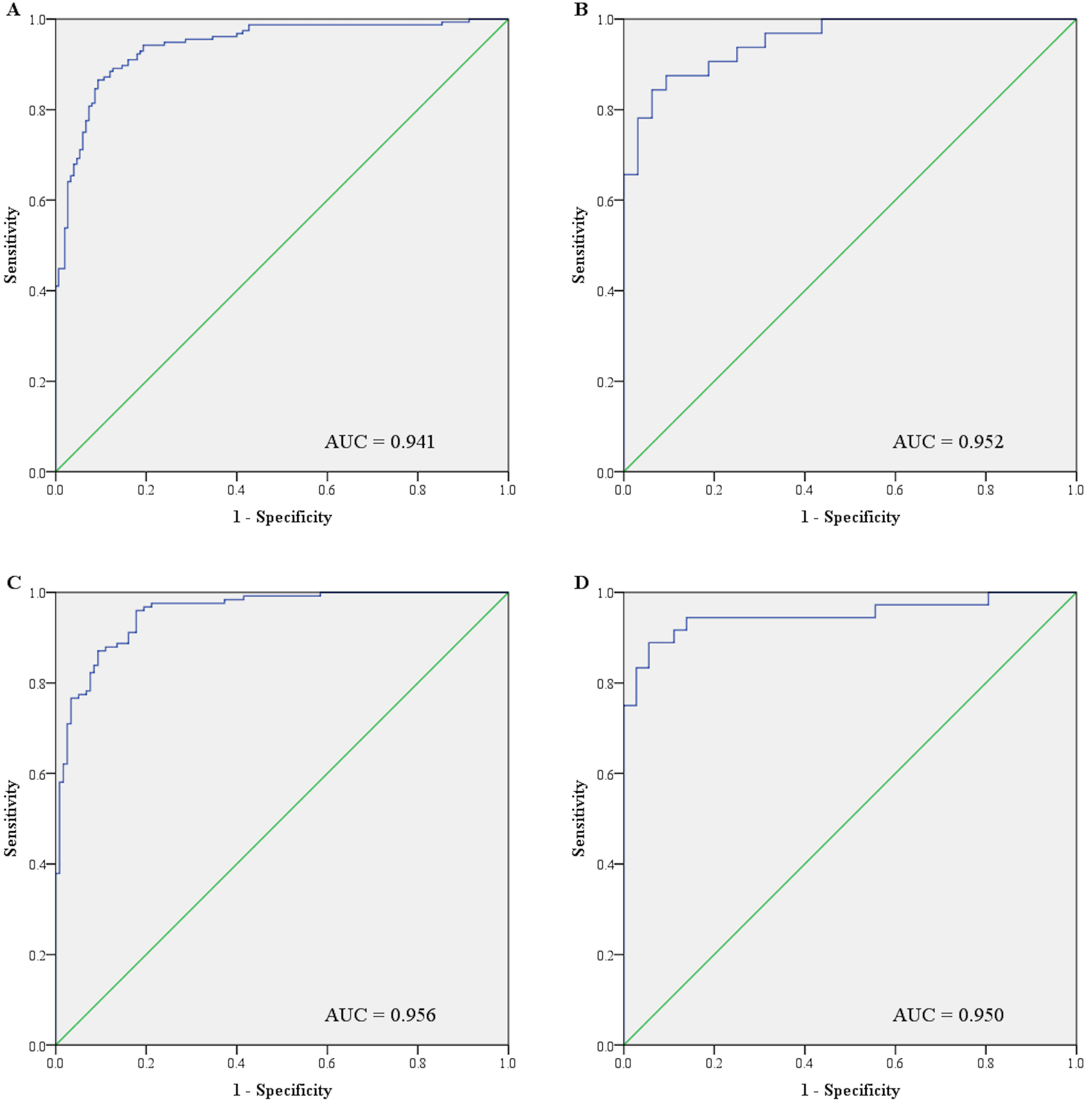
Expression levels of the twelve miRNAs in paired tissue samples. N: normal control; T: tumor; Horizontal line: mean with SEM.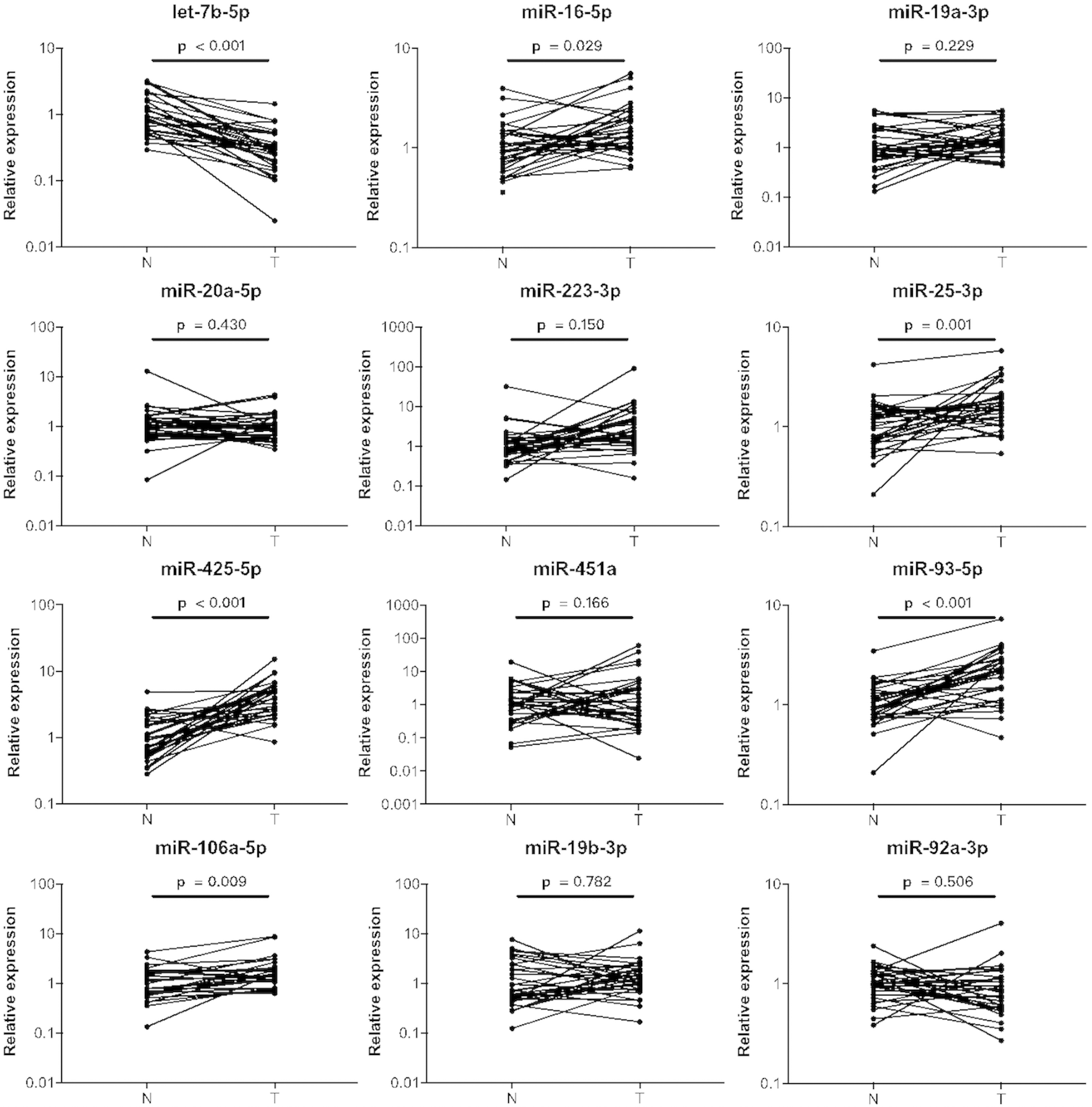
Expression levels of the twelve miRNAs in serum-derived exosomes. N: normal control; T: tumor; Horizontal line: mean with SEM.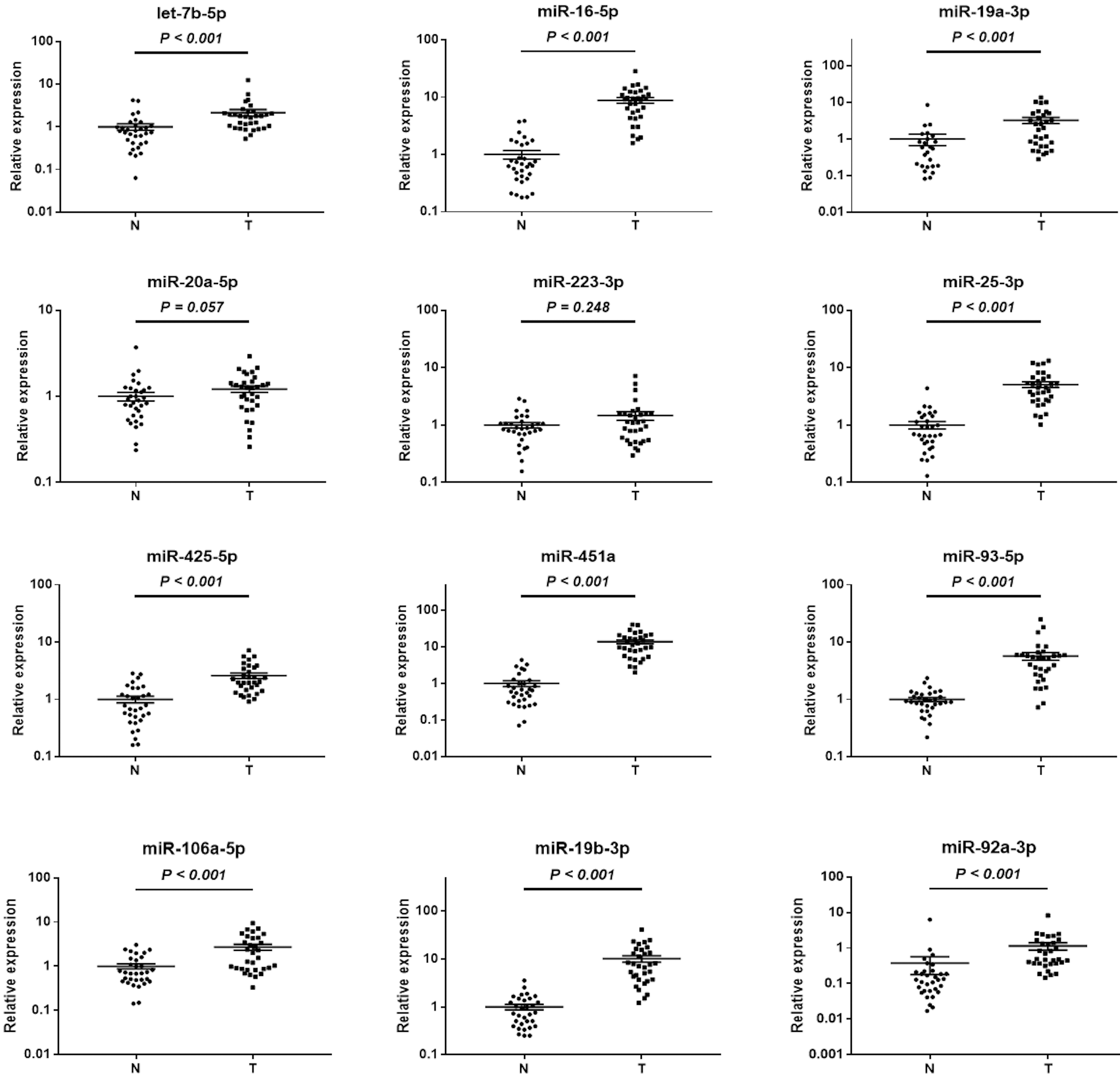
To assess the diagnostic performance of the 12 serum miRNAs, we built ROC curves for each of the miRNAs using the combined data of training and testing phases (Supplementary Fig. S1). We then combined the 12 serum miRNAs and established a 12-miRNA panel for BC detection using logistic regression analysis. The probability of BC prediction using the panel was determined by the following formula: Logit (P)
However, this panel may still be unsuitable for clinical use due to the high testing cost caused by the large number of tested items. Therefore, we tried to reduce the number of miRNAs to be analyzed to construct a smaller panel with a similar diagnostic ability. Through evaluation among multiple combinations, we found the combination of serum miR-25-3p, miR-451a, and miR-93-5p (the three miRNAs with the highest variations between BC and NCs) also exhibited relatively high diagnostic performance while requiring fewer miRNAs to be detected. The equation for the logistic regression model from this 3-miRNA panel was Logit (P)
External validation in public databases
The diagnostic value of the identified 12-miRNA and 3-miRNA signatures was further verified in two public datasets: The Cancer Genome Atlas Breast Invasive Carcinoma (TCGA-BRCA) dataset and GSE73002 dataset (Supplementary Fig. S3). In the TCGA-BRCA dataset, the identified signatures still performed well in discriminating BC tumor tissues (
Comparison with previously identified signatures
The diagnostic capability of the two signatures was compared with seven previously reported signatures that could similarly differentiate BC patients from healthy people using the data from TCGA-BRCA dataset [18, 21, 26, 27, 28, 29, 30]. As shown in Supplementary Table S4, the 12-miRNA signature had the highest AUC value compared with the other markers, and the performance of the 3-miRNA panel was only behind the signature proposed by Fermin et al. The two signatures identified in this study are superior to the other serum miRNA biomarkers in BC diagnosis.
Association of identified miRNAs with pathological features
The associations of the 12 miRNAs with clinical characteristics such as TNM stage and molecular subtype were further analyzed in the 216 BC patients and 214 NCs. In BC patients at different TNM stages, all the 12 serum miRNAs were consistently significantly upregulated when compared to those in NCs (
Subgroup analyses based on molecular subtypes were also conducted. The 216 BC cases were divided into four subtypes: Luminal A (
Tissue miRNA expression
In addition, the expression patterns of the identified miRNAs were explored in 32 pairs of BC tumor and para-tumor normal tissue samples (Fig. 4). Among the 12 miRNAs, miR-16-5p, miR-106a-5p, miR-25-3p, miR-425-5p, and miR-93-5p were significantly overexpressed in both serum and tissue samples of BC patients; however, the expression level of let-7b-5p was significantly lowered in BC tissues compared to that in para-tumor tissues (
Serum exosome miRNA expression
We analyzed the expression of the 12 miRNAs in serum exosomes samples (32 BC vs. 32 NCs) by qRT-PCR for further exploration. All the identified miRNAs, except miR-20a-5p and miR-223-3p, were consistently upregulated in serum exosomes in BC patients compared to NCs (
Functional exploration using bioinformatics analysis
We conducted Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) analyses for functional exploration of the 12- and 3-miRNA signatures, and the constructed heatmaps and summary sheets are described in Supplementary Fig. S8 (12-miRNA), Supplementary Fig. S9 (3-miRNA), and Supplementary Table S7. We concluded that many of the 12 identified miRNAs were commonly involved in biological processes relating to various cancers, such as colorectal cancer (10/12 miRNAs), prostate cancer (9/12 miRNAs), and thyroid cancer (9/12 miRNAs). Several typical cancer-related pathways also showed a close association with the 12 identified miRNAs, including the p53 signaling pathway (8/12), Hippo signaling pathway (9/12), EGFR signaling pathway (11/12), cell death (11/12), and others. The common pathways (i.e. DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest, G1/S transition of mitotic cell cycle) probably involved by the 3-miRNA signature was also summerized (Supplementary Fig. S9). Functional annotation results indicated that the 12-miRNA and 3-miRNA panels may have a critical role in tumor initiation and development, and indirectly demonstrated the reliability of the miRNA expression differences discovered in this study between BC patients and NCs. These results show potential for further large-scale verification and an investigation of the related mechanisms in future studies.
Discussion
BC is the leading cause of cancer deaths in females worldwide. The accurate diagnosis of BC is of critical importance to improve clinical outcomes. However, common screening and diagnostic strategies such as mammograms have several limitations [6]. Tumor-derived miRNAs in blood have been established as potent markers for cancer diagnosis and prognosis prediction [25]. Until now, serum miRNA biomarkers such as miR-21, miR-484, miR-1246, and miR-199a have been reported to be useful for BC diagnosis in various studies [17, 21, 26, 29, 31, 32, 33]. However, these studies often have conflicting results due to the use of diverse miRNA quantification platforms, study subjects, or analytical methods.
In the present study, miRNA expression in serum was profiled using qRT-PCR for 216 BC patients and 214 NCs. A total of 12 serum miRNAs (let-7b-5p, miR-106a-5p, miR-19a-3p, miR-19b-3p, miR-20a-5p, miR-223-3p, miR-25-3p, miR-425-5p, miR-451a, miR-92a-3p, miR-93-5p, and miR-16-5p) were significantly overexpressed in BC patients compared to NCs. A 12-miRNA panel with steady diagnostic performance for BC detection was further established. Moreover, we found that the 12-miRNA panel successfully discriminated BC patients in different stages or with different molecular subtypes from NCs, which further confirmed its diagnostic value. In tissues, among the 12 miRNAs, miR-16-5p, miR-106a-5p, miR-25-3p, miR-425-5p and miR-93-5p were consistently overexpressed, and let-7b-5p was conversely downregulated, in BC tissues compared to those in NCs.
MiR-106a-5p, miR-19b-3p, and miR-92a-3p are derived from the miR-106a-363 cluster. The tumor promoting function of the miR-106a-363 cluster has been demonstrated in several reports [34, 35, 36, 37]. In this experiment, we discovered that the expression level of miR-106a-5p was significantly higher in both the serum and tissue of BC patients. According to previous reports, the aberrant expression of serum miR-106a-5p has been observed in many tumors, such as prostate cancer, astrocytoma, osteosarcoma, and lung squamous cell carcinoma, but was rarely reported in BC [39, 40, 41]. Jonsdottir et al. discovered that increased miR-106a-5p and miR-25 expression in BC tissues had some associations with high proliferation, cytokeratin 5/6 positivity, and ER-negativity status, indicating a poor prognosis for patients [42]. In contrast, it was reported by Jin et al. that miR-19b-3p inhibited the proliferation of BC cells [43]. Circulating miR-19b-3p was found to be a potential biomarker for gastric cancer [44, 45]. However, there is little information about its relationship with BC and requires further investigation. MiR-92a-3p has been defined as a circulating BC marker in several researches, indicating its underlying correlation with the disease state and biological process of BC [18, 46, 47].
MiR-25-3p, along with miR-93-5p, belongs to the miR-106b-25 cluster which locates on chromosome 7. MiR-19a-3p and miR-20a-5p, identified here, belongs to the miR-17-92 cluster on chromosome 13. These two clusters are paralogs of the miR-106a-363 cluster, which play important roles in cancer-related processes [48, 49, 50, 51, 52]. In our study, miR-25-3p and miR-93-5p were upregulated in both serum and tissue samples from BC patients. We proposed that these miRNAs might be released by BC tumor cells into circulation and have a major association with BC development. Similar results, as well as their determined functions in tumor progression via different mechanisms in BC, have also been reported by several studies [53, 54, 55]. For instance, Chen et al. once reported the overexpression of miR-25-3p in triple-negative BC tissue promoted tumor proliferation by negatively regulating the tumor suppressor BTG2 [56]. Li et al. found that miR-93 can regulate cell cycle progression in BC by suppressing PTEN via the PI3K/Akt pathway [57]. Bai et al. recently reported that miR-20a-5p can promote tumor cell growth by targeting RUNX3 in triple-negative BC [58]. Serum miR-25-3p and miR-93-5p have been proposed as markers for cancer diagnosis in prostate cancer, lung cancer, and osteosarcoma, but were rarely reported in BC [38, 39, 40]. Similar results were observed for miR-20a-5p and miR-19a-3p [39, 40, 59, 60].
In this study, miR-425-5p and miR-16-5p were significantly upregulated in both serum and tissue in BC, while miR-451a and miR-223-3p was upregulated in serum but had no expressing differences in tissue. However, according to previous studies, miR-451a was downregulated in BC tumor tissues, which could promote cell growth, proliferation, migration, and invasion by weakening its tumor-suppressive function [61, 62, 63]. The dysregulation of miR-223-3p in blood circulation has been discovered in colorectal adenocarcinoma, esophageal squamous cell carcinoma, and lung cancer [64, 65, 66]. MiR-223-3p may also inhibit tumor progression in a variety of other cancer types [67, 68, 69]. However, there is scarce evidence of its role in BC [45, 60, 70, 71, 72, 73]. More research is required to decipher the exact roles of these identified miRNAs in BC.
In addition to serum and tissue samples, miRNA expression was also evaluated in serum-derived exosomes. Exosomes are membrane-derived vesicles released by various cell types that carry various molecular cargos and function in cell-to-cell communication [74, 75]. Exosomes are a major potential existing form of circulating miRNAs in addition to argonaute2 complexes [76, 77]. The dysregulation of exosomal tumor-related miRNAs may hint to their potential functions. In this study, 10 of the 12 identified miRNAs (excluding miR-20a-5p and miR-223-3p) were upregulated in serum-derived exosomes. These results confirmed our hypothesis that these miRNAs and their exosomal forms are not only potential promising biomarkers for BC detection, but also might play important roles in disease progression.
In summary, we identified a 12-miRNA signature in serum (let-7b-5p, miR-106a-5p, miR-19a-3p, miR-19b-3p, miR-20a-5p, miR-223-3p, miR-25-3p, miR-425-5p, miR-451a, miR-92a-3p, miR-93-5p, and miR-16-5p) which may serve as a promising signature for the accurate diagnosis of BC after four-phase screening. However, more research is required to further verify of our results and explore the functions of these miRNAs in BC-related biological processes. While there is still a long way to go for these miRNA signatures to be widely applied in clinical practice, this may be useful in combination with other standard detection methods.
Footnotes
Acknowledgments
This work was supported by the National Natural Science Foundation of China [Grant numbers: 81672400, 81672788, 81702364].
Conflict of interest
The authors declare that they have no conflict of interest.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-201547.
