Abstract
BACKGROUND:
MicroRNAs (miRNAs), with noticeable stability and unique expression pattern in plasma of patients with various diseases, are powerful non-invasive biomarkers for cancer detection including endometrial cancer (EC).
OBJECTIVE:
The objective of this study was to identify promising miRNA biomarkers in plasma to assist the clinical screening of EC.
METHODS:
A total of 93 EC and 79 normal control (NC) plasma samples were analyzed using Quantitative Real-time Polymerase Chain Reaction (qRT-PCR) in this four-stage experiment. The receiver operating characteristic curve (ROC) analysis was conducted to evaluate the diagnostic value. Additionally, the expression features of the identified miRNAs were further explored in tissues and plasma exosomes samples.
RESULTS:
The expression of miR-142-3p, miR-146a-5p, and miR-151a-5p was significantly overexpressed in the plasma of EC patients compared with NCs. Areas under the ROC curve of the 3-miRNA signature were 0.729, 0.751, and 0.789 for the training, testing, and external validation phases, respectively. The diagnostic performance of the identified signature proved to be stable in the three public datasets and superior to the other miRNA biomarkers in EC diagnosis. Moreover, the expression of miR-151a-5p was significantly elevated in EC plasma exosomes.
CONCLUSIONS:
A signature consisting of 3 plasma miRNAs was identified and showed potential for the non-invasive diagnosis of EC.
Introduction
Endometrial cancer (EC) is the third most common cancer among women worldwide [1]. However, there is no early detection method for asymptomatic women at the average risk of endometrial cancer [2]. Most guidelines recommend either transvaginal ultrasonography or endometrial biopsy as the initial detection but none of them are completely satisfactory. Diagnostic curettage is a widely used diagnostic method in EC patients. However, diagnostic curettage has several inherent defects, such as invasive and blindness in scraping tissue specimens together with missed diagnosis, and leads to different degrees of endometrial bleeding in some patients. Patients with endometrial cancer may make cancer cells to invade the muscle layer, which may result in tumor metastasis, causing serious consequences [3]. In addition, diagnostic curettage needs to be operated by experienced gynaecologists and possesses risk of missed diagnosis. Thus, identification of accurate and validated biomarkers for EC is needed to improve diagnosis and lead better survival outcome of EC.
MicroRNA (miRNA), a type of non-coding small RNA (18–25 nucleotides), functions as a post-transcrip- tional regulator via the repression of translation or direct degradation of target messenger RNA (mRNA) [4]. Many studies have unraveled its roles in the biological process of various cancers [5, 6, 7]. Meanwhile, circulating miRNAs show great potential of being cancer biomarkers for the stable existence in peripheral serum or plasma. Since blood samples are easy to obtain and the testing procedure is more convenient and cheaper, circulating biomarker can be used alone or combined with other traditional screening methods for preliminary screening before further invasive pathological and imaging examinations. It can help to identify the existence of cancer disease as early as possible, which can be employed for the screening of both undiagnosed suspected cases and healthy population. Several studies have shown that certain miRNAs could serve as highly accurate biomarkers for EC detection [8, 9, 10, 11, 12]. However, these findings lacked consistency between each other because of the varied research methods and study populations. In this study, we performed a four-stage study to determine the diagnostic value of plasma miRNAs using quantitative reverse transcription polymerase chain reaction (qRT-PCR). In addition, we also analyzed miRNA expression in EC tissues and plasma exosomes to promote the understanding of the possible existence form and biological function of the identified miRNAs.
Materials and methods
Information of participants
The study was conducted in accordance with the guidelines of the Hospital Ethics Committee and approved by the Institutional Review Boards of the First Affiliated Hospital of Nanjing Medical University (ID: 2016-SRFA-148). Totally, participants in this study included 93 patients and 79 normal controls (NCs) from the First Affiliated Hospital of Nanjing Medical University and Women’s Hospital of Nanjing Medical University between 2016 and 2017. The cases we collected may come from all over the country, but most of them originated in East China and their ethnic group is basically Han. Patients involved were diagnosed as EC by histopathological examination and didn’t accept any therapeutic intervention such as surgery, radiotherapy or drug treatment before sampling. These healthy volunteers were randomly recruited from the health examine center of the hospital. They were ensured to be EC-free after image examination. In addition, they were familiar with the project and volunteer to participate in this study and had signed the informed consent form. The criteria of sample inclusion have been summarized in Table S1.
Study design and sample preparation
The present study was divided into four stages (Fig. 1), and in the initial screening phase, 2 peripheral plasma pool samples from 20 EC patients and 1 pooled sample from 10 NCs were selected. Exiqon miRCURY-Ready-to-Use PCR-Human-panel-I+II-V1.M (Exiqon miRNA qPCR panel, Vedbaek, Denmark) was applied to identify dysregulated miRNAs preliminarily. Based on the candidate list obtained in the screening stage, qRT-PCR was used to verify the expression of each candidate miRNA. The external validation phase (27 EC vs. 23 NCs) was designed to finally verify the characteristics of the identified miRNA. In addition, tissue samples (24 EC vs. 24 NCs) and exosomes samples (32 EC vs. 32 NCs) were analyzed to explore the expression of the identified miRNAs.
The flow chart of the experiment design. EC: endometrial cancer; NC: normal control; qRT-PCR: Quantitative real-time polymerase chain reaction.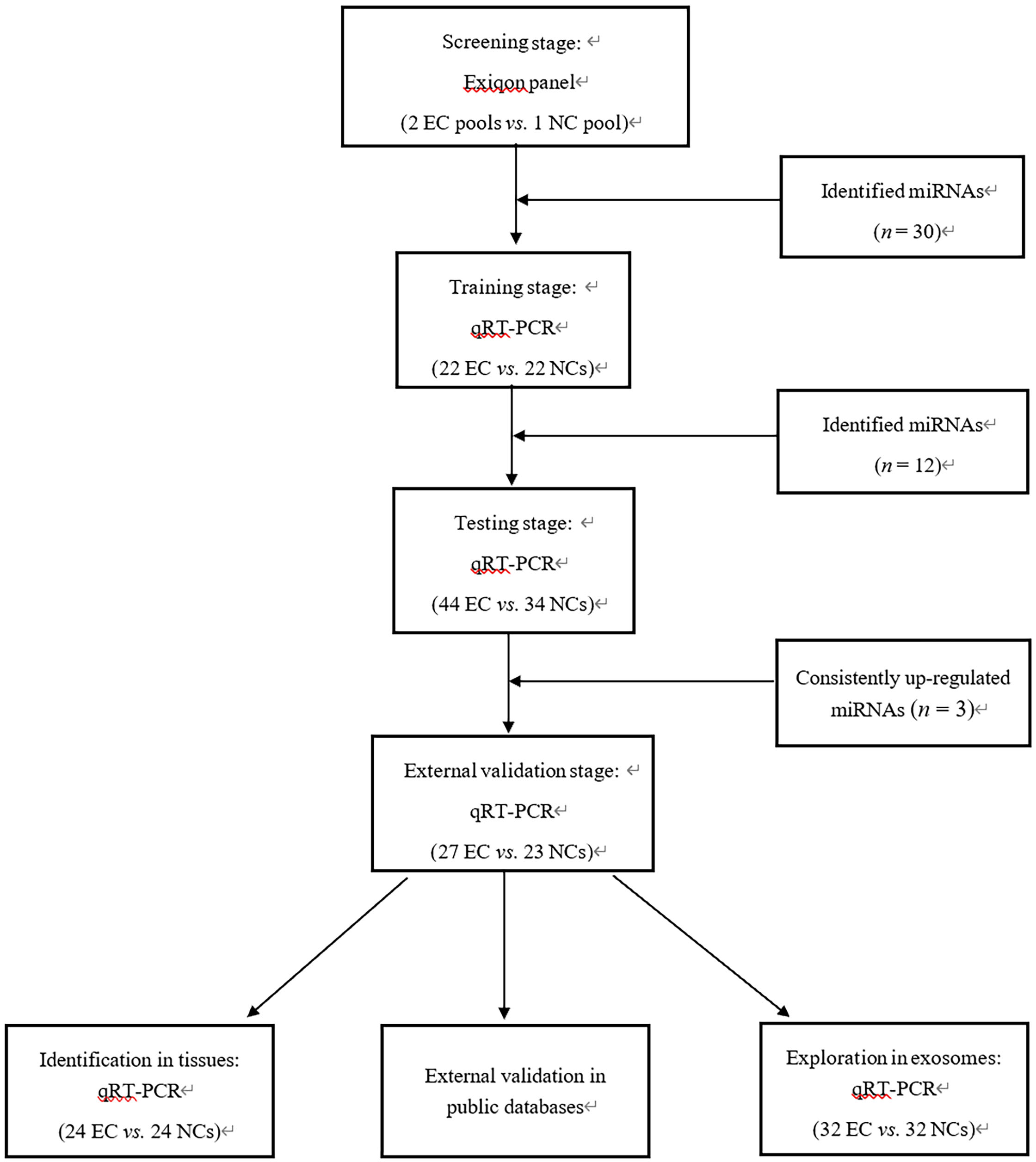
Blood samples from EC patients and healthy controls were collected with ethylenediaminetetraacetic acid (EDTA)-containing tubes (BD, New Jersey, USA). Cell-free plasma was separated from blood samples within 6 hours. The plasma samples were separated using a two-step centrifugation protocol (350 reactive centrifugal force (RCF) for 10 min at 4
Exosomes were isolated from plasma using Exo-Quick Exosome Precipitation Solution (System Biosciences, Mountain View, CA, USA) as described by the manufacturer. Briefly, the exosomes pellets were precipitated from 200
Extraction of RNA
RNA was isolated from 200
Clinical characteristics of 93 EC patients and 79 normal controls
Clinical characteristics of 93 EC patients and 79 normal controls
The specific primers of reverse transcription (RT) and polymerase chain reaction (PCR) from Bulge-Loop
Statistical analysis
Statistical analyses and graph plotting were performed using SPSS 24.0 software (SPSS Inc., Chicago, IL, USA) and GraphPad Prism 7.0 (GraphPad Software, USA). To assess the comparison of miRNA expression between EC patients and healthy controls, nonparametric Mann-Whitney test were conducted. The relationship of miRNAs and clinical characteristics was evaluated by one-way ANOVA or
Bioinformatics analysis
The enrichment of overlapping target gene signaling pathways of these miRNAs was performed using the online tool of DIANA-miRPath v3.0 [14]. The potential biological processes and pathways involved by miRNAs were evaluated using the assembly of Gene Ontology (GO) biological process and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway databases.
Results
Characteristics of participants
A total of 172 plasma samples (93 EC vs. 79 NCs) were analyzed in this four-stage experiment. The schematic diagram of the experimental procedure was shown in Fig. 1 and the clinical characteristics of the study subjects were listed in Table 1. There was no significant difference in the distribution of age between EC patients and NCs during the four stages (
Endometrial cancer diagnostic capability of the signature and previously identified biomarkers (presented as the results of ROC curve analysis using the data from GEO dataset)
Endometrial cancer diagnostic capability of the signature and previously identified biomarkers (presented as the results of ROC curve analysis using the data from GEO dataset)
Expression levels of the three identified miRNAs in the plasma of 93 EC patients and 79 female NCs. Horizontal line: mean with 95% confidence interval (CI). A: miR-142-3p; B: miR-146a-5p; C: miR-151a-5p. N: normal controls; T: tumor.
In the screening phase, 174 miRNAs were measured by the Exiqon miRCURY-Ready-to-Use PCR-Human-panel-I+II-V1.M based on the qRT-PCR platform in 2 peripheral plasma pools from 20 EC patients; 1 pooled sample came from 10 female NCs. The criteria of candidate miRNAs for further analysis were listed: (a) a Ct-value
Evaluation of serum miRNAs by qRT-PCR
Firstly, in the training stage, 12 differentially expressed miRNAs (has-miR-142-3p, has-miR-24-3p, has-miR-151-5p, has-miR-155-5p, has-miR-423-3p, has-miR-146a-5p, has-miR-154-5p, has-miR-197-3p, has-miR-181a-5p, has-miR-29c-3p, has-miR-125b-5p, has-miR-143-3p) were identified. In the following testing stage, the plasma samples of 44 EC patients and 34 female NCs were used for validation, and 3 miRNAs were consistently upregulated. To verify the diagnostic capacity of the three miRNAs for EC patients, the expression levels of miRNAs were further assessed among an additional independent cohort of 27 EC patients and 23 NCs. Combined results showed that miR-142-3p, miR-146a-5p, and miR-151a-5p were significantly up-regulated in the plasma of EC patients (Fig. 2).
Evaluation of diagnostic value
We used ROC analysis to evaluate the performance of the 3 miRNAs in identifying EC patients from NCs. Firstly, we separately analyzed the three miRNAs in a combined cohort of 93 EC patients and 79 NCs. The AUCs were 0.689 (95% confidence interval (CI): 0.611–0.767) for miR-142-3p, 0.694 (95% CI: 0.616–0.772) for miR-146a-5p, and 0.680 (95% CI: 0.601–0.759) for miR-151a-5p, respectively (Fig. S1). Though single miRNA could distinguish EC from healthy controls, a combination of several miRNAs might provide stronger differentiation power than individual miRNA. The three miRNAs were combined as a panel using logistic regression analysis and the equation to calculate the probability of EC was: Logit (P)
Receiver-operating characteristic (ROC) curves for the three-miRNA signature to discriminate EC patients from NCs. (A) the combined data of training, testing and external validation stages (93 EC vs. 79 NC): AUC 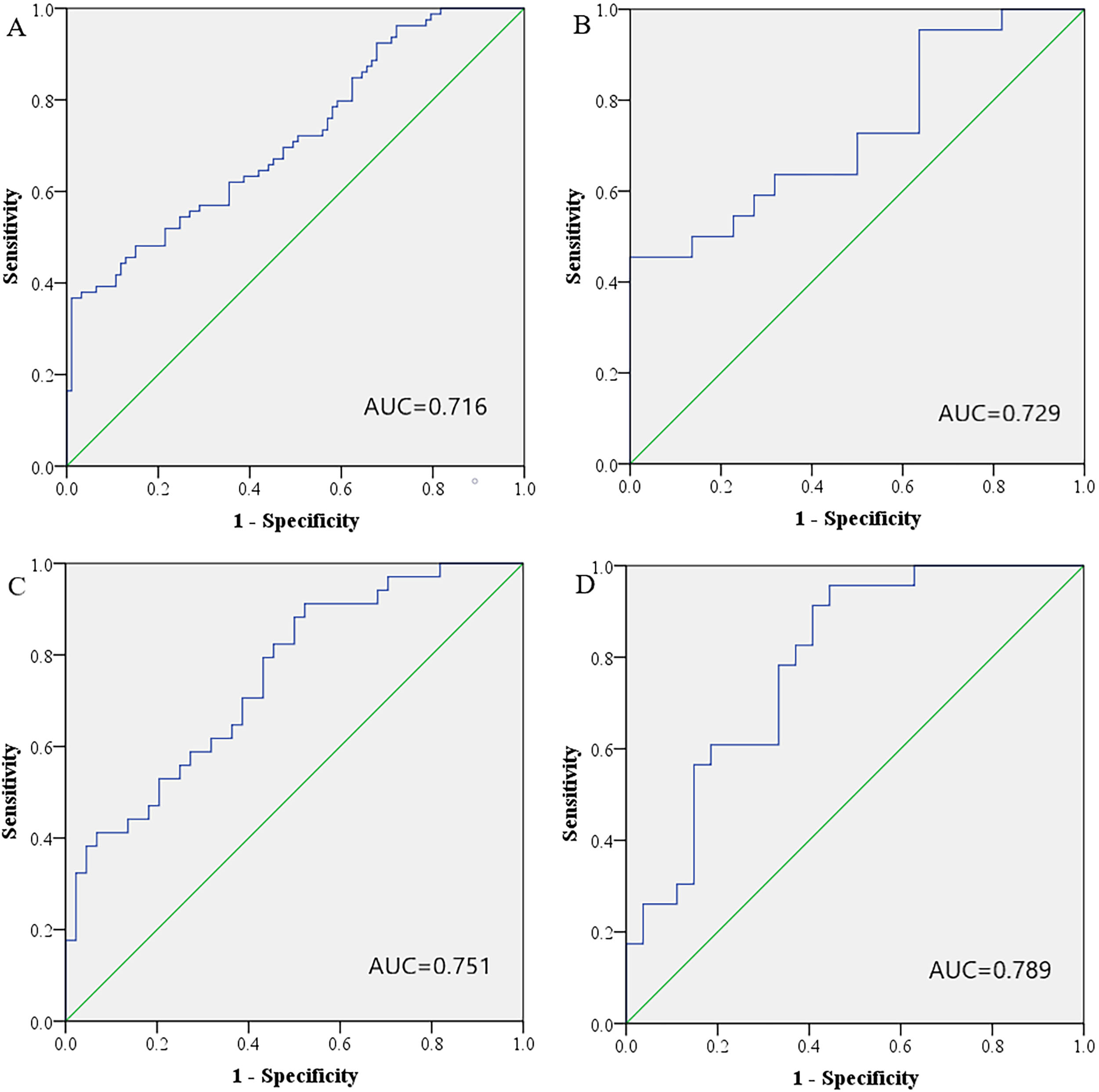
In addition, this study analyzed the correlation between the identified miRNAs and clinicopathological parameters of 93 EC patients. Based on the analysis of FIGO stages, there was no statistically significant difference in the expression of the three miRNAs in the early stage (I) and advanced stage (II+III+IV) (Fig. S3). Subgroup analyses based on histological grade were also conducted and miR-142-3p was significantly overexpressed in well-differentiated EC patients (Fig. S4). The expression of miR-142-3p and miR-146a-5p in EC patients in pre-menopausal status was significantly higher than that of post-menopausal status (Fig. S5). The ROC curves of early stage (I) and advanced stage (II+III+IV) compared with NC group are 0.720 (95% CI: 0.642–0.797) and 0.713 (95% CI: 0.547–0.878) respectively, which proves that 3-miRNA marker can distinguish EC patients at various stages from NCs (Fig. 4). The ROC analysis was further conducted for histological grade and menopausal status. As is shown in the Fig. 4, the 3-miRNA signature performs well in discriminating EC cases of different histological grade and menopausal status from healthy controls, with AUCs of 0.768 (95% CI: 0.666–0.870) for grade I, 0.733 (95% CI: 0.649–0.816) for grade II, 0.672 (95% CI: 0.547–0.797) for grade III, 0.792 (95% CI: 0. 673–0.911) for pre-menopause status and 0.678 (95% CI: 0. 587–0.768) for post-menopause status.
ROC curves of the 3-miRNA panel for discriminating EC patients at different clinicopathological parameters from NCs. (A) Stage I: AUC 
External validation of the 3-miRNA signature in TCGA-UCEC (A), GSE35794 (B), and GSE25405 (C) datasets. A: TCGA-UCEC: AUC 
The diagnostic value of the identified 3-miRNA signatures was further verified in three public datasets: The Cancer Genome Atlas Uterine Corpus Endometrial Carcinoma (TCGA-UCEC) dataset, GSE35794 dataset (18 tumor samples and 4 normal controls), and GSE25405 dataset (41 tumor samples and 7 normal controls) (Fig. 5). In the TCGA-UCEC dataset, the identified signatures still performed well in discriminating EC tumor tissues (
Comparison with previously identified signatures
Comparing the diagnostic capabilities of the signature with the five previously reported signatures could similarly distinguish EC patients from healthy people. As is shown in Supplementary Table 2, the 3-miRNA signature in this study is superior to the other miRNA biomarkers in EC diagnosis.
Expression of identified miRNAs in tissue and plasma exosomes samples
In present study, the expression patterns of the three miRNAs were examined in 24 pairs of tissue samples of EC patients. However, none of them were significantly up-regulated in tumor tissues compared to their matched controls (Fig. 6). In addition, the expression of the three miRNAs was also examined in plasma exosomes from 32 EC patients and 32 female NCs. As is shown in Fig. 6, the significant dysregulation of exosomal miR-151a-5p was consistent with the dysregulation in plasma. No significant difference was found in miR-142-3p and miR-146a-5p expression.
Expression of the three miRNAs in the tumor tissues and plasma exosomes. Horizontal line: mean with 95% CI. A: The expression of miR-142-3p in the tumor tissues; B: The expression of miR-146a-5p in the tumor tissues; C: The expression of miR-151a-5p in the tumor tissues; D: The expression of miR-142-3p in the plasma exosomes; E: The expression of miR-146a-5p in the plasma exosomes; F: The expression of miR-151a-5p in the plasma exosomes; N: normal controls; T: tumor.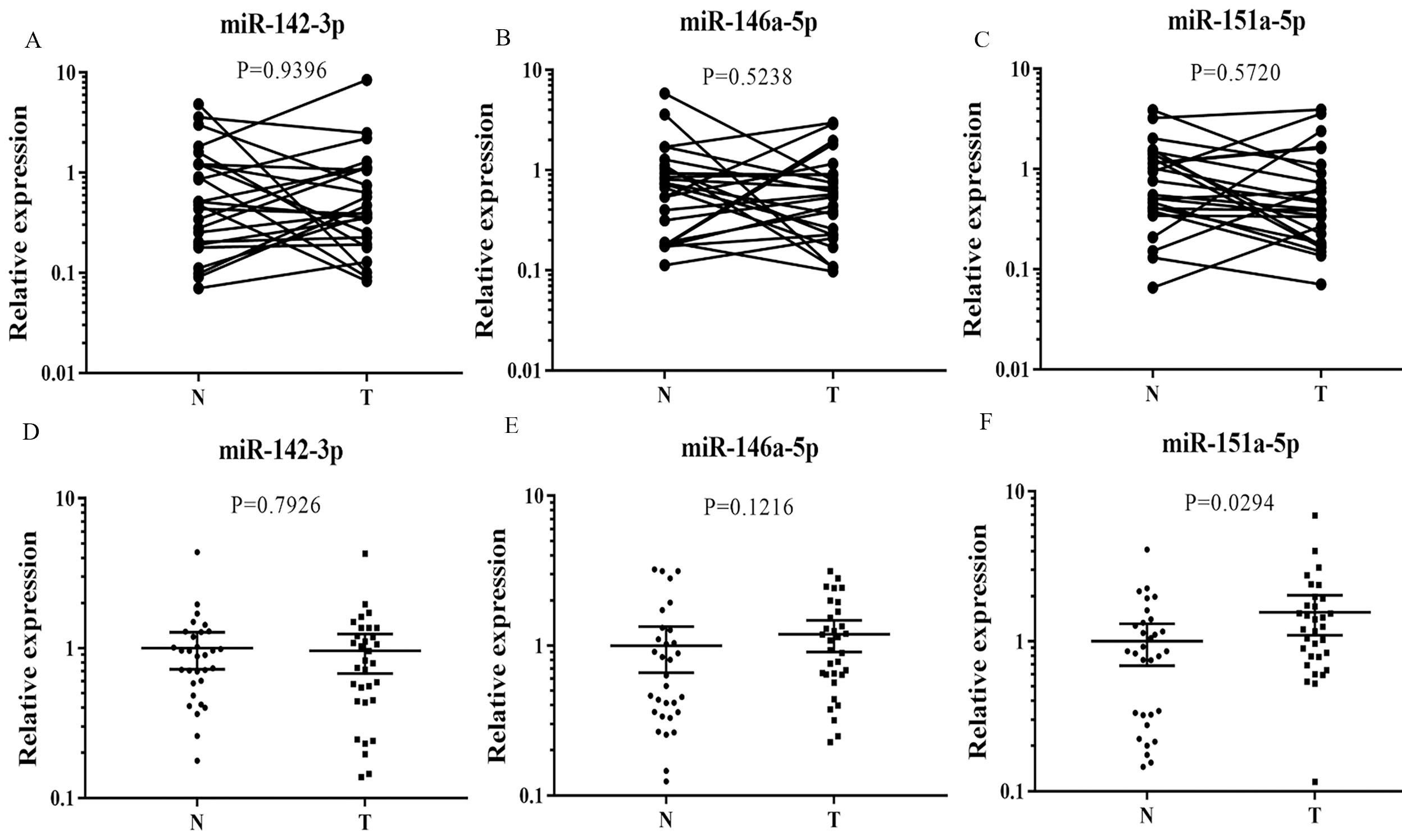
We utilized DIANA-mirPath v3.0 (
Discussion
EC is one of the most common malignant tumors in females. Although quantities of cases have been diagnosed at early stages, there are still a proportion of patients who show high-grade malignancy or recurrence and have shorter overall survival.
Circulating miRNAs, including miRNAs in serum, plasma, or circulating microcarriers such as peripheral exosomes, have particular advantages as diagnostic markers for cancers. Firstly, circulating miRNAs are quite easy to obtain without damage which permits the dynamic monitoring of patients and the routine examination of healthy people. Secondly, PCR, the critical step of miRNA detection, can be more sensitive since it maximums the difference signal between cases and controls. Compared with other conventional screening methods, miRNA detecting method is also cheaper, quicker, and more objective. Last but not least, the aberrant expression of certain miRNAs in blood circulation not only reflects the existence of disease status but also indicates the potential mechanisms of tumor development, progression, invasion, and metastasis. Circulating miRNA can be both a biomarker and a therapeutic target [17, 18]. Although the sensitivity and specificity of different miRNA biomarkers may fluctuate greatly, stable circulating miRNA signatures, combined with other traditional methods, can play a potent role in assisting clinical diagnosis.
We reviewed the current methodologies of detecting circulating miRNAs and analyzed several key factors affecting miRNA quantification. On the one hand, considering that not only race and population differences but also variety of samples will affect the outcome of the research, multicenter data is needed for further verification. On the other hand, there are multiple standardized methods for the quantitative analysis of circulating miRNA, including adding endogenous or exogenous reference. At present, there is no generally accepted endogenous reference for circulating miRNA analysis [19, 20]. It was reported that correcting sample processing differences by adding exogenous miRNAs may be a better method for quantitative analysis of circulating miRNAs [19, 21, 22]. In addition, inconsistent platforms usually show different results in the initial screening stage. Exiqon miRNA qPCR panel was proved to have better linearity and sensitivity in the quantification of miRNAs at relatively lower abundance than the TaqMan platform [23, 24, 25].
In recent years, the diagnostic potential of circulating miRNAs has been gradually discovered in EC. Potential biomarkers such as circulating miR-222, miR-223, miR-186, miR-204, miR-484, and miR-23a were continuously discovered, but these results lacked consistency within these studies. Through four-stage screening, we investigated the application of plasma miRNAs for EC diagnosis among 93 EC patients and 79 NCs in this study, and we found that a biomarker combining over-expressed miR-142-3p, miR-146a-5p, and miR-151a-5p in plasma could be utilized to diagnose EC.
During the screening stage, we used the Exiqon miRNA qPCR panel to initially screen differentially expressed plasma miRNAs in 2 EC and 1 NC pooled samples. In the testing and the training stage, we further used qRT-PCR to validate the candidate miRNAs from the screening stage. In this study, three plasma miRNAs (miR-142-3p, miR-146a-5p and miR-151a-5p) with significantly higher expression levels that could accurately discriminate EC patients from healthy women. A panel combing these three miRNAs was then built using logistic regression analysis for EC detection. The AUCs of the panel turned out to be 0.729, 0.751, and 0.789 for the training, testing, and external validation stage, respectively. Moreover, the expression levels of the three miRNAs were also examined in tissues and plasma-derived exosomes samples. However, none of them were significantly up-regulated in tumor tissues compared to their matched controls. MiR-151a-5p were up-regulated in the plasma exosomes of EC patients.
Deng et al. demonstrated that miR-142-3p was detected to be down-regulated in cervical cancer cells, and over expression of miR-142-3p inhibited proliferation and invasion of cervical cancer cells by targeting FZD7 [26]. In addition, Wang et al. validated miR-142-3p and miR-146a-5p as the part of miRNA signature that could serve as an independent predictor for overall survival (OS) of EC patients [27]. We found that the expression of miR-142-3p and miR-146a-5p in premenopausal EC patients was significantly higher than that in postmenopausal patients. It was reported that menopause could be an independent prognostic factor for EC and premenopausal EC patients would have a better prognosis. This further indicates that the mechanism and relationship between menopause and these two miRNAs are worthy of further study.
Zhou et al. once reported overexpression of miR-146a attenuates the effect of NIFK-AS1 on M2 polarization inhibition of macrophages and estrogen-induced endometrial cancer cell proliferation, migration and invasion [28]. It was reported that miR-146a-5p is involved in the pathogenesis of CC, and miR-146a has been shown to promote CC cell viability via targeting IRAK1 and TRAF6 [29]. In addition, miR-151a was identified as an oncomiRNA in non-small cell lung cancer. It has been shown to target E-cadherin mRNA and induce migration of proliferation, and partial EMTEMT [30]. However, there is little evidence of its role in EC. More research is needed to decipher the exact role of these identified miRNAs in EC.
The menstrual phase of the endometrium includes inflammation as a physiologic component [31]. menstruation involves the synthesis and release of inflammatory factors, including cytokines, growth factors, COX-2, and prostaglandins, as well as activation of NF-
In order to further verify the results of the study, we also explored plasma miRNAs for EC detection in tissues. However, in our study, the expression of the three miRNAs in tissue samples is not consistent with the miRNA plasma profile. We speculated that discrepancy between EC plasma and tissue samples might be due to the release and transportation of miRNAs in tumor and its microenvironment. Other biological factors might also affect the result, such as different biological states of tumors, influence of other non-tumor cells or some immunological factors. Circulating miRNA signals may reflect the changes in body fluids of patients with diseases, and the tissue miRNA profile may mainly show local organ diseases. Anyway, a larger number of tissue samples were in demand for further validation.
Exosomes are small cell-derived vesicles (30–120 nm) existing in almost all kinds of body fluids and can carry a variety of molecules including miRNAs. Exosomal miRNAs are associated with various cancers which shows their great potential of being diagnostic tools [38, 39, 40, 41]. Therefore, to explore the potential form and function of the three miRNAs, we further analyzed miRNA expression in plasma-derived exosomes. Exosomal miR-151a-5p was increased in EC compared to NCs. The findings might give some hints about cell-to-cell communication and cell-to-microenvironment communication mediated by the two exosomal miRNAs in the development and progression of EC.
Limitations of our study still remained. First of all, since the samples in this study were all from Chinese patients, the clinical outcomes of different races and biological behaviors were not considered. Secondly, the sample size in our study was relatively small and the diagnostic value in patients with advanced stage could not be well evaluated. Last but not least, the precise mechanisms by which these three miRNAs facilitate EC progression remained largely obscure. Our study was still in preclinical stage, and there would be a long way to go for future clinical application.
Conclusion
In summary, our study has identified and validated a three-miRNA signature (miR-142-3p, miR-146a-5p and miR-151a-5p) in plasma as a potential non-invasive biomarker for EC diagnosis.
Experimental subject statement
Approval for the present study was obtained from the Institutional Ethical Committee and participants have given written informed consent (ID: 2016-SRFA-148). The experiments were undertaken with the understanding and written consent of each subject, and that the study conforms with The Code of Ethics of the World Medical Association (Declaration of Helsinki), printed in the British Medical Journal (18 July 1964).
Authors’ contributions
Conception: Z.W and Z.H; Interpretation or analysis of data: Z.S.Y and C.W.F; Preparation of the manuscript: F.X.C; revision for important intellectual content: C.M.M, F.X.C, and L.C; Supervision: Z.C and L.C.Y.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-200972.
sj-docx-1-cbm-10.3233_CBM-200972.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-200972.docx
Footnotes
Acknowledgments
This work was supported by the Natural Science Foundation of China [Grant number: 81672400].
