Abstract
OBJECTIVE:
To determine the utility of plasma microRNAs (miRNAs) as biomarkers in cervical cancer (CC).
METHODS:
Some studies were conducted about the specific expression of plasma miRNAs in the diagnosis of CC. Plasma samples of 97 CC patients and 87 normal controls (NCs) were used to identify dysregulation of miRNAs in the training, testing, and external validation phases. Receiver operating characteristic (ROC) curves and area under the ROC curve (AUC) were used to evaluate the sensitivity and specificity of identified individual miRNAs and miRNA panels for the diagnosis of CC. Expression levels of specific miRNAs were also examined in plasma exosomes and tissue samples of CC patients.
RESULTS:
Four plasma miRNAs (miR-146a-5p, miR-151a-3p, miR-2110 and miR-21-5p) which showed up-regulation were identified and validated in CC patients. A panel of the four miRNAs were constructed as potential diagnostic markers for CC. The AUCs of the panel of these four-miRNAs for the training, testing, and external validation phases were 0.911, 0.774, and 0.786, respectively. miR-146a-5p and miR-21-5p levels were all up-regulated in CC tissue specimens, whereas miR-146a-5p, miR-151a-3p, and miR-2110 levels were up-regulated in plasma exosomes.
CONCLUSION:
The signature of the four-miRNAs identified in peripheral plasma is a promising novel biomarker for the diagnosis of CC.
Introduction
Experimental design.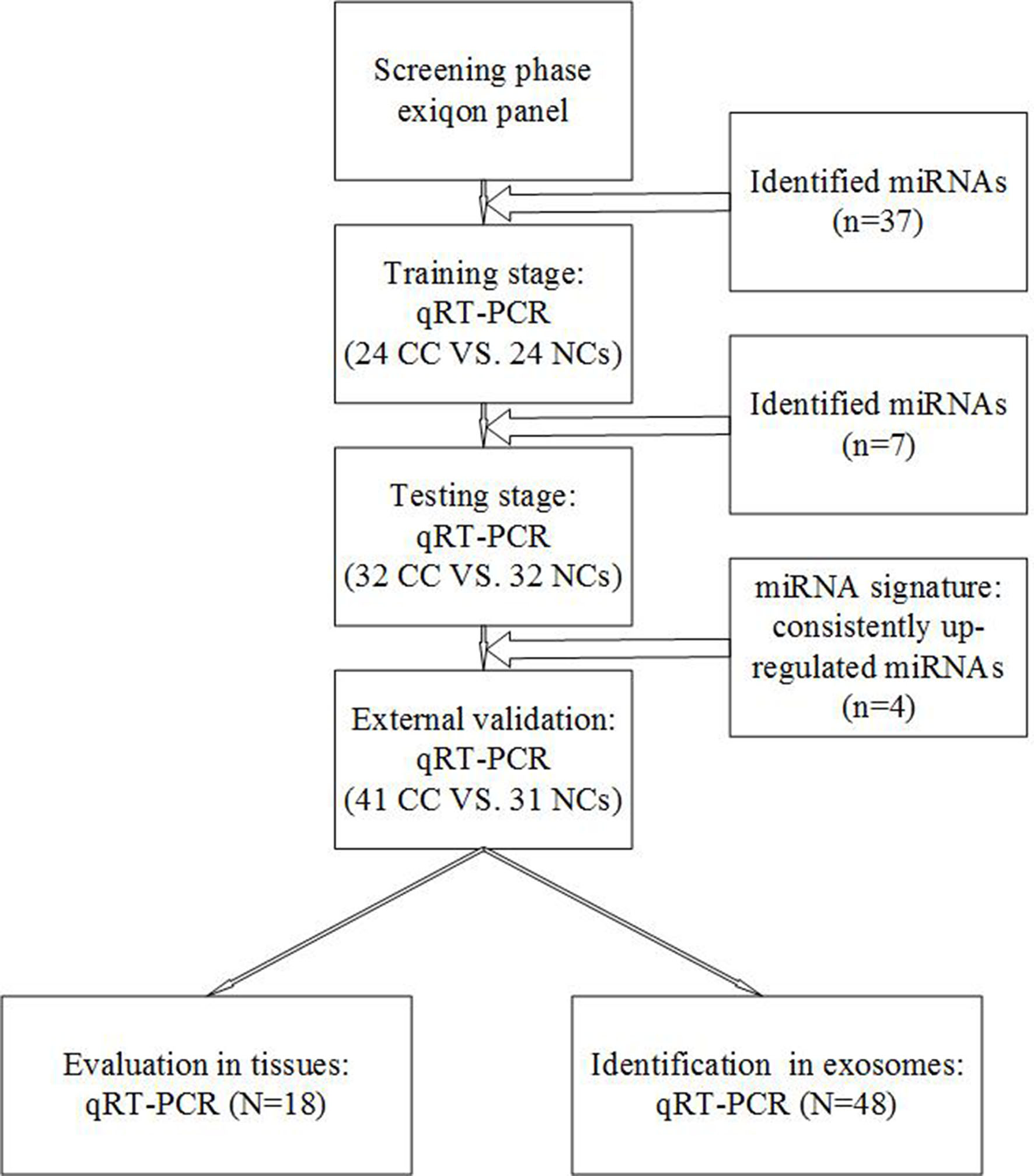
Cervical cancer (CC) remains the second most common cause of cancer-related deaths among females globally [1]. The early diagnosis of CC has been associated with excellent survival; however, some patients in developing countries diagnosed with advanced CC and often at an untreatable stage are associated with very poor survival rates. Although ThinPrep cytological test is an effective method of CC screening, it is limited by its poor sensitivity and specificity [2]. Cervical biopsy is a widely used diagnostic method, in spite of the fact that it has several inherent defects such as invasiveness. Moreover, cervical biopsy requires the availability of an experienced gynecologist. Thus, unprecedented and reliable biomarkers are urgently needed for the early diagnosis and intervention of CC.
MicroRNAs (miRNAs) are a kind of non-coding RNAs having multi-biological function, such as regulating cell cycle, mediating immune response, inducing apoptosis [3]. The mechanism of miRNA regulating effect for gene expression is combined with the 3’-untranslated region (3’-UTR) of the targeted genes. Consequently, the deregulation and abnormal expression of miRNAs were associated with human pathologies, including cancer. Since the first study by Lawrie et al. who measured miRNAs in the serum, circulating miRNAs had been used for early diagnosis of cancer as reliable noninvasive biomarkers [4]. Several studies exploring the differential expression of circulating miRNAs including CC patients had identified some potential miRNA biomarkers for cancer detection. However, the use of different detection methods and distinct cohorts in these studies underlie the lack of consistency among the findings. Furthermore, most of the studies focused on a small number of specific miRNAs, whereas few studies conducted a comprehensive investigation of the plasma miRNA profiles in CC patients.
In this study, we conducted a multiphase study using quantitative reverse transcription (qRT) polymerase chain reaction (PCR) to identify plasma miRNAs for the diagnosis of CC. miRNAs in exosomes and tissue specimens were further investigated for their potency as circulatory markers for monitoring CC.
Study design and population
Ninety-seven CC patients and 87 normal controls (NCs) were enrolled between 2015 and 2017 at the First Affiliated Hospital of Nanjing Medical University. All CC patients were confirmed by histological biopsy. Collecting blood and tissue samples from the First Affiliated Hospital with Nanjing Medical University after approval by the institutional ethics committee and obtaining written informed consent from the patients. Briefly, blood samples were collected in ethylenediaminetetraacetic acid-containing tubes (BD, NJ, USA), and cell-free plasma was separated within 6 hours after collection. Plasma samples were clarified by spinning at 350
This study was designed to obtain some plasma miRNAs for the diagnosis of CC (Fig. 1). Three plasma pools from 30 CC patients and one pool sample from 10 NCs were detected. All the plasma in this detection belonged to the sample bank of the First Affiliated Hospital of Nanjing Medical University, and not a part of any test phases. miRCURY LNA™ Universal RT microRNA PCR, Serum/plasma Focus microRNA PCR Panel V4 (Qiagen, Germany) was used to screen miRNAs expressing differently between CC samples and NC samples. All the pool samples in initial screening were from biological specimen banks, as previously described [5, 6]. Same as previous studies, the miRNA panel included some literature reported miRNAs associated with specific tumor (CC in the present study). In the training phase, plasma samples from 24 CC patients and 24 NCs were used to confirm the dysregulation of miRNAs identified in the screening phase. Subsequently, in the testing phase, miRNAs were clarified in samples from 32 CC patients and 32 NCs. In the final phase, the diagnostic value of selected miRNAs was validated in an external cohort of 41 CC patients and 31 NCs. Tissue specimens obtained from 18 CC patients belonged to the validation set.
RNA extraction
RNA was isolated from 200
Isolation of exosomes
Exosomes were extracted from plasma depending on manufacturer’s instructions of Exo-Quick exosome precipitation solution (System Biosciences, Mountain View, CA, USA). All samples used for exosome analysis belonged to validation set. Briefly, exosome pellets precipitated from 200
Quantitative real-time polymerase chain reaction
Specific primers (RiboBio, China) were used for reverse transcription (RT) and PCR amplification of miRNAs, and RT-PCR were performed as described previously [6, 7]. miRNA amplification and detection were achieved using the SYBR
Statistical analysis
The miRNA screened in the miRCURY LNA™ Universal RT microRNA PCR, Serum/plasma Focus microRNA PCR Panel V4 was selected according to the order of fold change value. The stability of plasma miRNA expression levels was determined by calculating the stability value (M) using the geNorm software (version 3.5). Nonparametric Mann-Whitney test was used to assess the statistical significance of differential expression of miRNAs in CC patients in three phases. The relationship between identified miRNAs and clinical characteristics was evaluated using one-way analysis of variance or chi-square test, and the correlation of expression patterns of miRNAs in plasma was analyzed using Pearson’s correlation coefficient. We established the miRNA panel by logistic regression analysis and evaluated the sensitivity and specificity of CC by receiver operating characteristic (ROC) curves and area under the ROC curve (AUCs).
SPSS 20.0 software (SPSS, Chicago, IL, USA) and GraphPad Prism 7.0 (GraphPad Software, USA) were used to analysis the relative data in this study.
Demographic and clinical characteristics of individuals contributing to the 184 plasma samples and 18 tissue samples in the study
Demographic and clinical characteristics of individuals contributing to the 184 plasma samples and 18 tissue samples in the study
Expression levels of the four plasma miRNAs in the training and testing phases (presented as the mean
Characteristics of subjects
This multiphase study of 184 subjects including 97 CC patients and 87 NCs, assessed differentially expressed circulating miRNAs. Detailed characteristics of study participants whose plasma samples were allocated to the four study phases (screening, training, testing, and external validation; Fig. 1) are summarized in Table 1 and Supplement 1. There were no significant differences in age distribution between CC patients and NCs (
Selection of endogenous reference
Alternative reference miRNAs (miR-16-5p, miR-103a-3p, RNUB6, and miR-1228-5p) included in the present study were selected because they had relatively stable expression levels based on previous studies [9, 10]. In geNorm, high variations in expression are indicated as high M values and reflects low stability, whereas low M values indicate high expression stability [11]. In plasma samples, the lowest M values belonged to miR-16-5p and miR-103a-3p (Supplement 2); therefore, miR-103a-3p was selected as the internal reference control in the present study because of its smaller standard deviation.
Four indicated miRNAs expression in plasma which were from CC patients (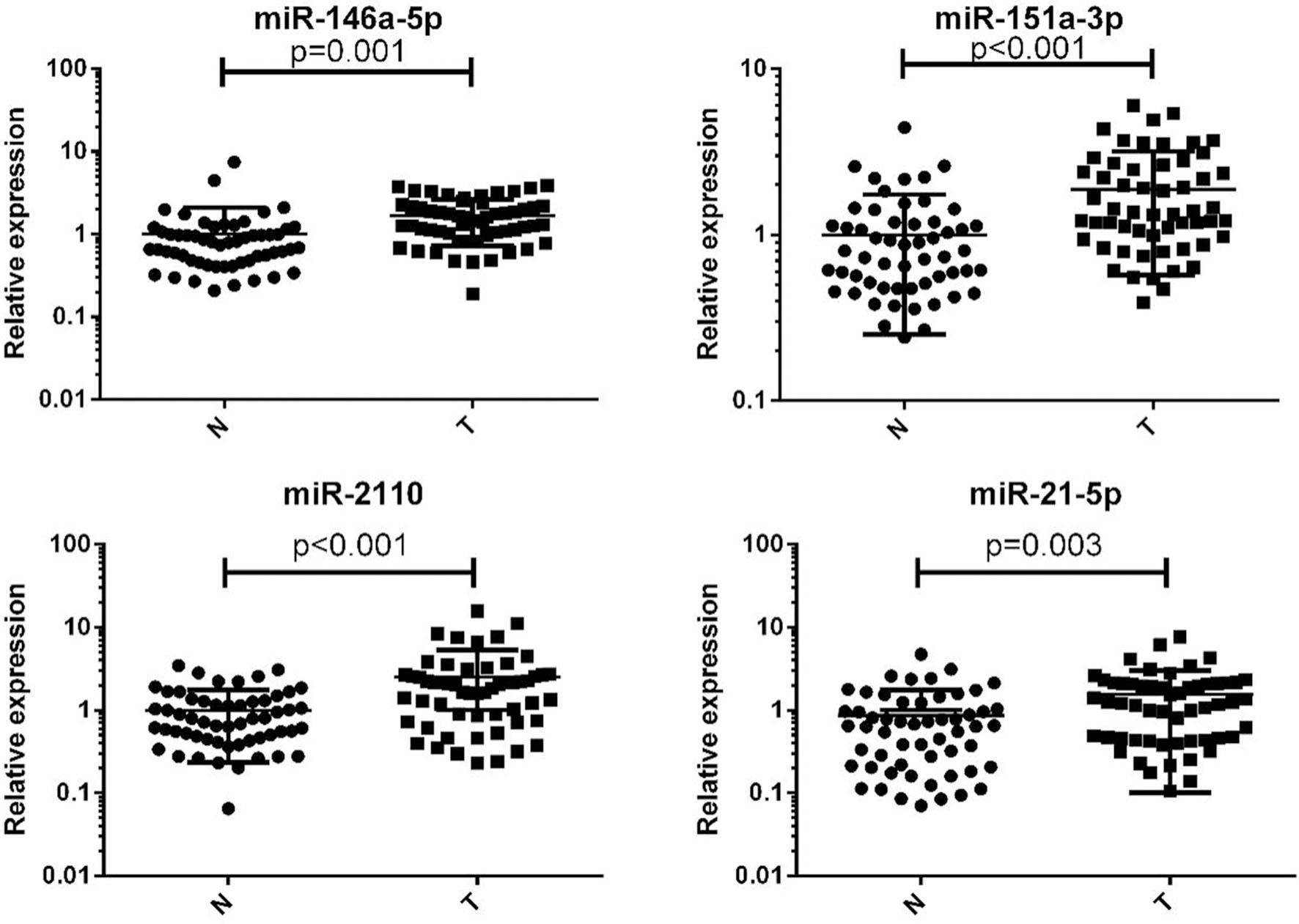
We used the miRCURY LNA™ Universal RT microRNA PCR, Serum/plasma Focus microRNA PCR Panel V4 to measure 179 miRNAs of interest in three peripheral plasma pools from 30 CC patients and one pool sample from 10 NCs. Candidate miRNAs were selected based on the following criteria: (a) a Ct value of
Confirmation of candidate miRNAs by qRT-PCR
The 37 differentially expressed candidate miRNAs were initially tested in 24 CC and 24 NC samples in the training phase (Supplement 3). A total of 7 miRNAs demonstrated aberrant expression and were validated in the testing phase. After the testing stage, 56 CC and 56 NC samples were analyzed together. The overall combined results of the training and testing phases showed that miR-146a-5p, miR-151a-3p, miR-2110, and miR-21-5p were significantly up-regulated in CC patients (Table 2, Fig. 2; additional miRNAs are presented in Supplement 4). In addition, to ascertain whether these four miRNAs could be used as a potential panel for the diagnosis of CC, an additional analysis including independent cohort of 41 CC patients and 31 NCs was conducted.
Diagnostic value of plasma miRNAs for CC
To evaluate the diagnostic value of identified plasma miRNAs in distinguishing between CC patients and NCs, ROC curves were determined and AUCs were calculated. AUCs of miR-146a-5p, miR-151a-3p, miR-2110, and miR-21-5p in plasma were 0.770 [95% confidence interval (CI) 0.681–0.858], 0.752 (95% CI 0.663–0.841), 0.721 (95% CI 0.625–0.816), and 0.677 (95% CI 0.577–0.776), respectively, in the combined cohorts of the training and testing phases (Supplement 5).
Moreover, the equation for the panel of these four miRNAs were as follows: Logit(P)
Discriminating CC patients from NCs by ROC curves for 4-miRNA panels in plasma. Area under the Curve (AUC). A. A combined two cohorts from the training and testing phases (A: 97 CC patients’ and 87 NCs’ plasma); B. Training Phase (24 CC patients and 24 NCs’ plasma); C. Testing Phase (32 CC patients’ and 31 NCs’ plasma); D. External validation phase (41 CC patients’ and 31 NCs’ plasma).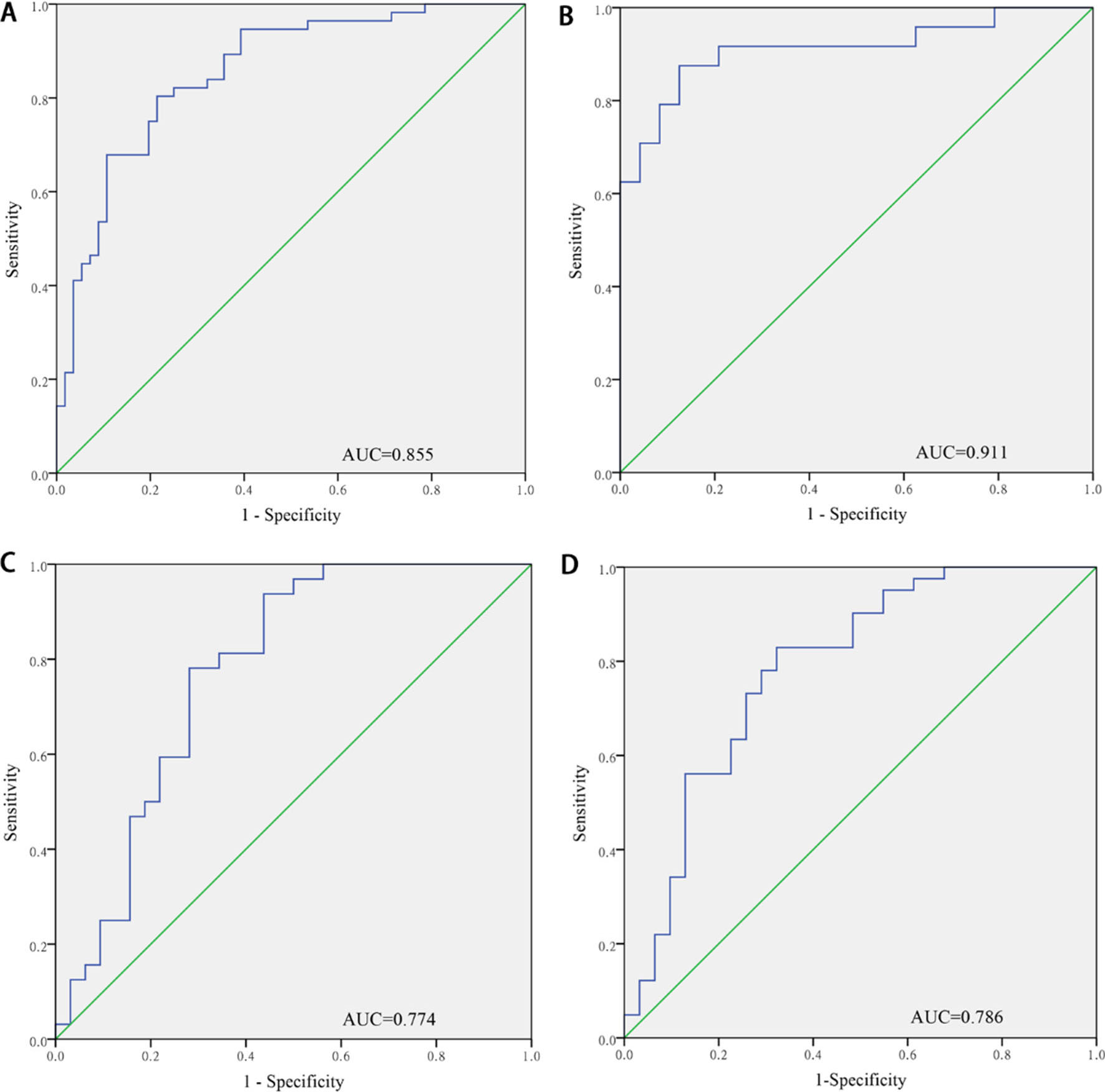
The relationship between identified miRNAs and clinicopathological parameters, including histological grade, FIGO stage, and pathological type, were analyzed for the 97 plasma samples from CC patients (Supplement 7). Plasma miR-151a-3p levels were significantly higher in patients with grade I disease than in those with grade II or III disease as well as in patients with stage I disease than in those with stage II disease. Plasma miR-21-5p levels were increased in patients with grade II or III disease than in those with grade I disease. Finally, the expression of miR-146a-5p was higher in adenocarcinoma than in squamous cell carcinoma.
Evaluation of miRNAs in tissue samples
Four miRNAs expression levels were evaluated in 18 tissue samples from CC patients. Among the four miRNAs, only miR-146a-5p and miR-21-5p showed significantly higher level of expression in tumor tissue samples than the matched controls (Fig. 4).
Four identified miRNAs (including miR-146a-5p, miR-151-3p, miR-2110, miR-21-5p) expression levels in CC patients’ tumor tissues. The relative expression (2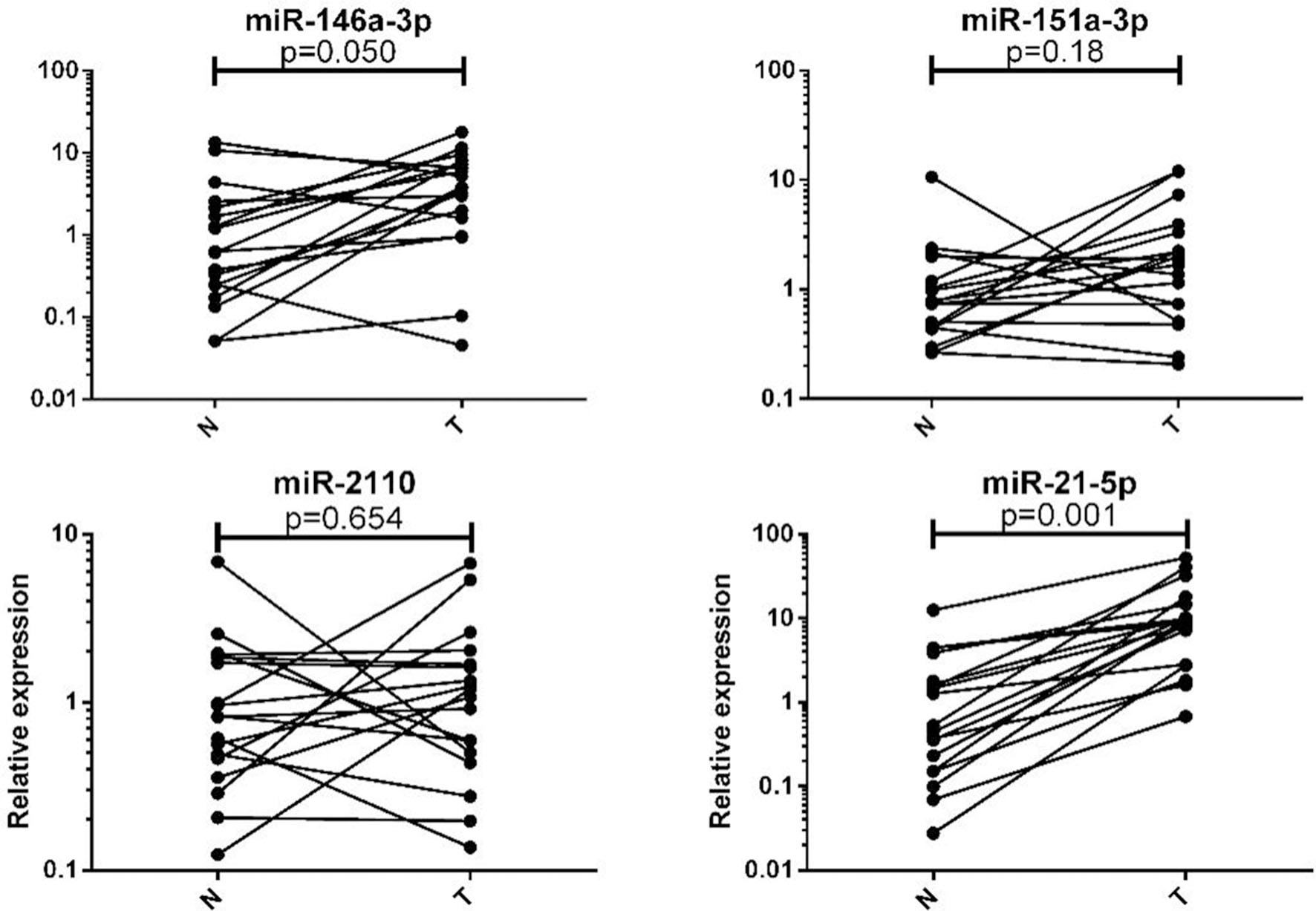
The identified miRNAs gene expression levels in 24 CC patients’ and 24 NCs’ plasma exosomes. The relative expression (2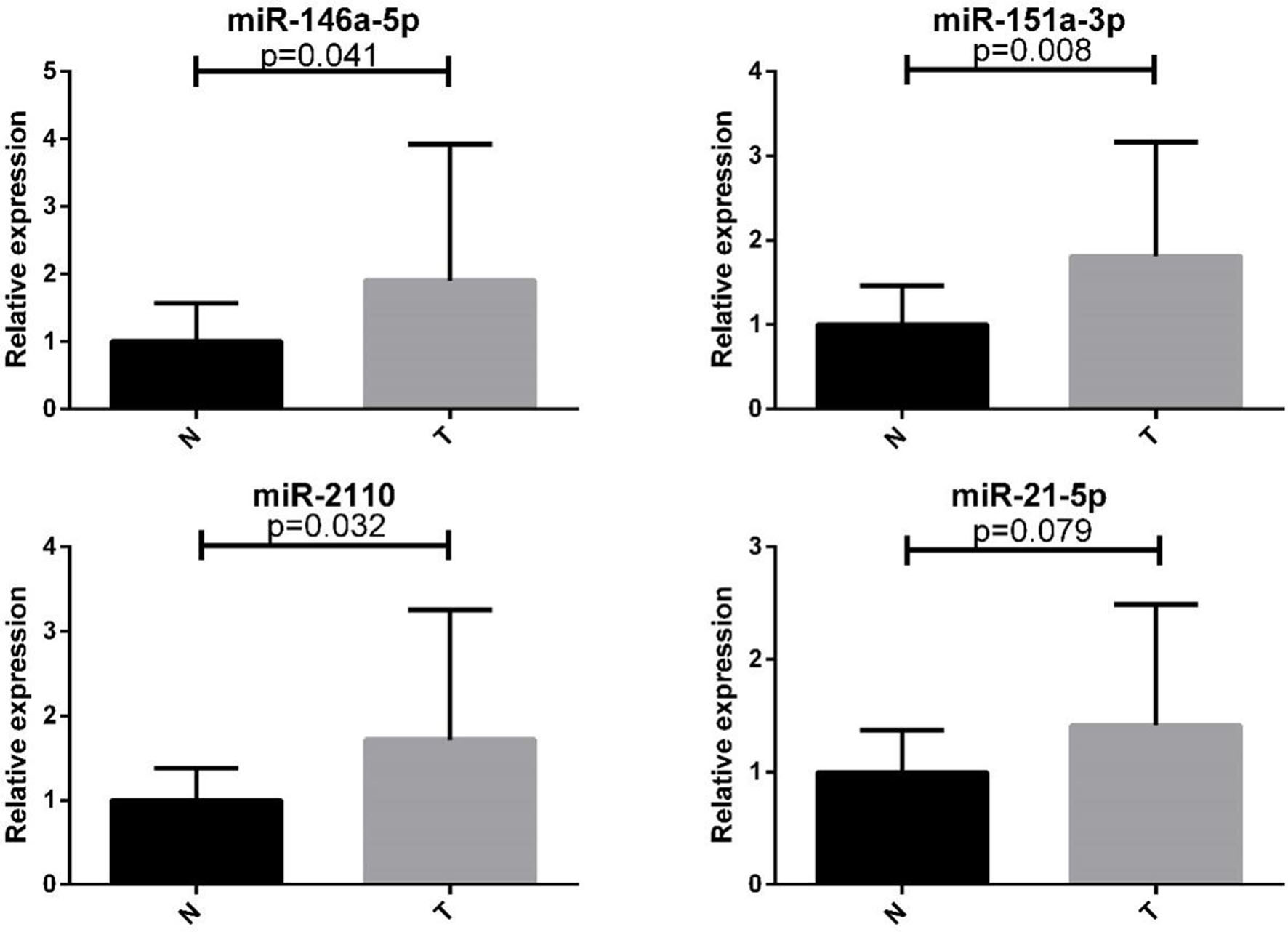
Four identified miRNAs expression levels were examined in plasma exosomes from 24 CC patients and 24 NCs to evaluate their potential presence in plasma exosomes in CC patients. As shown in Fig. 5, changes in levels of exosomal miR-146a-5p, miR-151a-3p, and miR-2110 were consistent with those in plasma. The expression of miR-21-5p was also increased in plasma exosomes of CC patients compared with those in NCs, albeit not reaching statistical significance.
Analysis of candidate miRNAs with a bioinformatics database
In this study, we applied an online analysis tool (DIANA-mirPath v3.0) to predict the targets of identified miRNAs derived from IANA-TarBase v7.0. Thus, this database was used to decipher potential functions of the four candidate miRNAs identified in the present study (miR-146a-5p, miR-151a-3p, miR-2110, and miR-21-5p). These programs were used for KEGG pathway analysis and gene ontology (GO) category analysis to investigate relevant pathways. Subsequently, 14 biological processes associated with the four identified miRNAs were identified in the GO category analysis, including the biological process and cellular nitrogen compound metabolic process. Among the four miRNAs, miR-21-5p appeared to be more closely related to the pathological mechanisms of cancer, including an interaction with cell cycle and the p53 signaling pathway in the KEGG pathway analysis. Heatmaps of target pathways are shown in Supplement 8. These results indicated the possible roles of these four candidate miRNAs in the pathogenesis of CC.
Discussion
As an early event in the tumor development, aberrant miRNA expression and circulating miRNAs might have utility as noninvasive biomarkers in CC. Circulating miRNAs could potentially detect CC in the early stage. Previous studies has proposed a serum miRNA panel including four miRNAs (miR-16-2*, miR-195, miR-2861 and miR-497) to identify cervical intraepithelial neoplasia [12]. RNAs, including miRNAs, may be released from the blood cells into the serum during coagulation [13]. Plasma miRNAs, which might differ from serum miRNAs, were closer to the physiological levels in CC patients. In the present study, the microarray (serum/plasma focus microRNA PCR panel) was used to screen out a number of miRNAs which have potential diagnostic value. In addition, we reviewed previous studies to make up the possible deficiency of the microarray (such as small sample size). Eventually, five miRNAs (miR-196a, miR-205a-5p, miR-20a-5p, miR-21-5p and miR-218) reported in the literature were screened in the first phase. All the miRNA proved to be closely associated with CC in vitro of in vivo [14, 15, 16, 17, 18]. Therefore, five miRNAs were selected as a supplement in CC biomarkers specifically. However, only miR-21-5p was verified to have important significance in differential diagnosis of CC. The main reason is the differences of miRNA expression pattern between serum and plasma. In view of this question, in a previous study where miRNA expression profiles were compared between plasma and serum samples, only plasma-derived miRNAs were associated with clinical parameters in breast cancer [6]. From the present study report of the assessed plasma samples from 97 CC patients and 87 NCs, our results demonstrated that four plasma miRNAs (miR-146a-5p, miR-151a-3p, miR-2110, and miR-21-5p) were significantly up-regulated in CC compared with those in NCs. The four-miRNAs panel was constructed to diagnose CC with high sensitivity and specificity. In the training phase, the dysregulation of miR-142-3p, miR-584, and miR-296 was observed in CC patient samples. However, a similar outcome was not found in the testing phase, including a larger cohort. The external validation phase further expounded and proved the validity of diagnostic efficiency of the four miRNAs. Similar level change of miRNA in CC tissue samples indicated that these miRNAs play a crucial role in tumorigenesis and progression.
In the four increasing miRNAs, miR-146a-5p was previously reported to be involved in the pathogenesis of CC, whereas miR-146a was demonstrated to promote CC cell viability via targeting IRAK1 and TRAF6 [19]. Furthermore, miR-151a was identified as an oncomiRNA in non-small cell lung cancer and was shown to target E-cadherin mRNA and induce proliferation, migration, and partial EMT [20]. However, very few studies explored the functions of miR-2110. One study has showed that miR-2110 played a clinically relevant role in maintaining the expression levels of MYCN and miRNA in neuroblastoma [21]. miRNA-21-5p, another oncomiRNA, involved in tumorigenesis, has been reported to be up-regulated in various cancers including breast cancer [22], ESCC [23], and clear cell renal cell carcinoma [24]. In addition, miRNA-21-5p had been reported to be a predictor of recurrence in young GC patients [25]. The diagnostic value of circulating miRNA-21-5p levels had also been evaluated in thymic epithelial tumors [26] and non-small-cell lung cancer [27].
Given that the majority of circulating miRNAs originate from tumor tissues [28], expression levels of the four miRNAs were also assessed in CC tissue samples. Only two identified miRNAs (miR-146a-5p and miR-21-5p) were consistently and significantly up regulated in CC tissues samples. However, the dysregulation of miR-151a-3p and miR-2110 expression were not found in CC tissue samples in the present study, which might be due to the limited number of available samples. Moreover, the release and transportation of miRNAs in tumor could led the differences between plasma samples and tissue samples. The biological roles of these two miRNAs in CC required further investigation. Although the tissue miRNAs directly reflected the signature of tumor, the plasma miRNAs panel could reflect the signature of CC in body fluid, therefore the plasma panel might have higher value in early diagnosis.
Recent studies had reported that miRNA-containing exosomes can be detected in blood [29]. This was a newly discovered mechanism in which a primary tumor might influence the gene expression in target cells. Compared with NCs, cancer patients have elevated levels of tumor-derived exosomes in the plasma [30], the present study further investigated whether the four identified miRNAs were present in the exosomes of CC patients. Our analysis revealed that three out of the four signature miRNAs were increased in the plasma-derived exosomes of CC patients, whereby the small sample size might account for the lack of a change in miR-21-5p in the exosomal fraction of the plasma. Another possibility involved the potential cofractionation of miR-21-5p with protein complexes rather than exosome. As a kind of late-eluting miRNA, miRNA-21-5p could cofractionated with argonaute 2, which was the key effector protein component of the miRNA-induced-mediated silencing complex that directly binds with miRNAs [31]. In brief, the functions and mechanisms of these exosomal miRNAs required further investigation.
Overall, this multiphase study identified and validated a panel of four-miRNAs in the plasma of CC patients as a potential noninvasive biomarker for the diagnosis of CC. The combination of this plasma-based miRNA biomarker panel and current early diagnostic tools could be applied to the screening of CC patients.
Footnotes
Acknowledgments
This work was supported by the National Natural Science Foundation of China [Grant number: 81672400; 81702364], the Natural Science Foundation of Jiangsu Province [Grant number: BK20171085] and Postgraduate Research and Practice Innovation Program of Jiangsu Province [Grant number: KYCX19_ 1158].
Conflict of interest
The authors declare that they have no competing interests.
Supplementary data
The supplementary files are available to download from
