Abstract
Objective:
We aim to identify several microRNAs (miRNAs/miRs)-messenger RNAs (mRNAs) biomarkers correlated to nasopharyngeal carcinoma (NPC) based on an integrated analysis of miRNA and mRNAs microarray expression profiles.
Methods:
The available mRNA and miRNA microarray datasets were retrieved from Gene Expression Omnibus (GEO) database according to pre-determined screening criteria. Differentially expressed miRNA and mRNAs (DEmiRNAs and DEmRNAs) were extracted between NPC and noncancerous nasopharyngeal tissues. The target genes of DEmiRNAs were predicted with miRTarBase followed by the construction of DEmiRNAs-target DEmRNAs network, and functional analyses were performed. The DEmiRNAs expressions were validated and the performance of these DEmiRNAs was assessed by the area under the curve (AUC) values. Finally, the correlations between DEmiRNAs and specific clinical factors were analyzed.
Results:
There were 1140 interaction pairs (including let-7d/f-
Conclusion:
The findings implied let-7d/f-
Introduction
Nasopharyngeal carcinoma (NPC) is a metastatic malignant tumor from the nasopharynx epithelium. 1 The epidemiological evidence suggested that there was a considerable risk for NPC occurrence in Southeast Asia with the incidence rate of approximately 50/100,000 deaths each year. 2 The latest statistics indicated that around 130,000 NPC new cases and 73,000 deaths were estimated across the world during 2018. 3 Although advanced diagnostic tools, effective surgical operation and pharmacological therapy dramatically reduced the occurrence of NPC over past few decades, 4 -7 the corresponding clinical outcomes has been unsatisfied due to delayed diagnosis, distant metastasis, high recurrence and lack of targeted agents. Encouragingly, high-throughput sequencing technology has considerably facilitated molecular genetic analysis of various diseases including cancers. Consequently, increasing investigators have concentrated on uncovering the molecular mechanisms of NPC initiation and development to develop promising therapeutic strategies for NPC treatment. 8,9
MicroRNAs (miRNAs) belong to a subclass of short non-coding RNAs with approximately 22 nucleotides, which can regulate gene expression by binding to the 3’-untranslated region (UTR) of target messenger RNAs (mRNAs).
10
They function as tumor repressors or oncogenes and play imperative roles in tumor biological behaviors including cell proliferation, migration, apoptosis and metastasis.
11
-14
Notably, an increasing body of research has demonstrated that numerous miRNAs were closely associated with NPC occurrence and development according to transcriptome analyses based on microarray expression profiles.
15,16
For example, Chen
Therefore, we searched available miRNA and mRNA microarray datasets from NPC tissues from the National Center for Biotechnology Information Gene Expression Omnibus (GEO) database to perform an integrated bioinformatics analysis. Specifically, the miRNA-mRNA regulatory analysis and functional analysis were performed to identify potential biomarkers related to NPC. Then, the candidate signatures were validated and evaluated. Finally, the correlations between key miRNAs and clinical parameters were predicted. This study will provide a deeper understanding for pathological mechanisms of NPC progression.
Materials and Methods
Data Acquisition
The miRNAs and mRNAs sequencing datasets were retrieved and downloaded from GEO 21 repository using the searching terms of “nasopharyngeal neoplasms” [MeSH Terms] OR “Nasopharyngeal carcinoma” [All Fields] AND “Homo sapiens” [porgn] AND “gse” [Filter]. The available datasets were selected according to the following criteria: 1) the type of studies with eligible datasets was restricted to expression profiling by array or non-coding RNA profiling by array; 2) all data were whole-genome mRNA expression profiles; and 3) these datasets were generated from NPC tissues and noncancerous nasopharyngeal tissues (controls), and all tissues were not treated with stimulant medications. Finally, 5 mRNAs datasets were obtained, including GSE64634 (12 NPC and 4 normal healthy nasopharyngeal tissue specimens), GSE53819 (18 NPC primary tumor tissues and 18 non-cancerous nasopharyngeal tissues), GSE34573 (15 NPC biopsy tissues and 3 normal nasopharyngeal biopsy tissues), GSE13597 (25 undifferentiated NPC biopsy tissues and 3 non-malignant nasopharyngeal biopsy tissues) and GSE12452 (31 NPC tissues and 10 normal nasopharyngeal tissues). Besides, 5 miRNAs microarray datasets were also extracted, including GSE43039 (20 chronic nasopharyngitis tissues and 20 NPC tissues), GSE70970 (246 NPC biopsy tissues and 17 nasopharyngeal epithelium tissues), GSE32906 (16 NPC tissues and 6 normal healthy nasopharyngeal epithelial tissues), GSE32960 (312 NPC tissues and 18 normal nasopharyngeal tissues), and GSE36682 (62 NPC tissues and 18 nasopharyngitis tissues). Table 1 showed the baseline information about mRNA and miRNA datasets.
A List of mRNA and miRNA Datasets Information for NPC.

mRNA, messenger mRNA; miRNA, microRNA; NPC, nasopharyngeal carcinoma; N, the number of normal nasopharyngeal tissue specimens; P, the number of nasopharyngeal carcinomas tissue specimens
Data Pre-Processing and Identification of Differentially Expressed RNAs (DERs)
For data pre-processing, the annotated mRNAs and miRNAs in each sequencing platform were obtained and over-lapped. Herein, a total of 12180 over-lapping genes among mRNA microarray datasets were identified while there were 196 over-lapping miRNAs across miRNA expression profiles. Then, gene or miRNA distribution matrix were obtained. If the data does not conform to a normal distribution, log2 transformation will be conducted to standardize data. Subsequently, we used the metaDE package in R to extract differentially expressed miRNA and mRNAs (DEmiRNAs and DEmRNAs) between NPC and control samples using false discovery rate (FDR) < 0.05 as the screening cutoff. 22 Finally, the hierarchical clustering analysis was conducted with the pheatmap package (https://cran.r-project.org/package=pheatmap) in R language.
Prediction of Target Genes for DEmiRNAs and Functional Analyses
Admittedly, miRNAs exert significant roles at post-transcription processes via regulating mRNA levels. The updated miRTarBase (version 7.0; http://miRTarBase.mbc.nctu.edu.tw/) database contains thousands of experimentally verified miRNA-target interactions (MTIs) involving 29 species. More specifically, it collects 2599 miRNAs and 15064 corresponding targets for human beings. Herein, this database was employed to predict the target genes for screened DEmiRNAs and the Cytoscape software 23 was utilized to construct the DEmiRNA-target DEmRNA regulatory network. We would regard those target DEmRNAs as candidate genes for the following analyses.
Afterward, we further explored the underlying biological roles of these genes, the Gene ontology (GO) analysis was firstly performed on the basis of 3 categories molecular function (MF), cellular component (CC) and biological process (BP) using BiNGO plugin in cytoscape software. Notably, the
Validation of DEmiRNAs in Plasma Samples of NPC Patients
To examine whether DEmiRNAs in NPC tissues can be detected in datasets from plasma samples of patients undergoing NPC, the relevant microarray datasets were searched from GEO database. Finally, 2 datasets (GSE43329 and GSE48442) from plasma samples from NPC patients were obtained. Specifically, there were 31 NPC cases and 19 healthy control plasma samples in GSE43329 dataset while GSE48442 comprised of 20 NPC cases and 10 control samples. Following this, the miRNA expression levels were computed by t-test and the receiver operating characteristic (ROC) analysis was implemented to evaluate the performance of DEmiRNAs using the pROC package (https://cran.r-project.org/web/packages/pROC/index.html) in R language.
Association Analysis Between DEmiRNAs and Clinical Parameters
To further explore whether there are correlations between DEmiRNAs and some clinical factors, the association analysis was carried out. The clinical information was respectively collected from mRNA and miRNA datasets. As indicated in Table S1, the clinical data in GSE64634 and GSE13597 were unavailable. GSE53819 contained the information about age and gender. Two datasets (GSE13597 and GSE12452) included NPC stage data. Therefore, the clinical data was incomplete in mRNA datasets. For miRNA datasets, GSE32906 and GSE32960 contained NPC stage information (GSE32906: 4 patients in stage I-II, III and IV, respectively; GSE32960: 12, 86, 91 123 patients in stage I, II, III and IV, respectively; Table S1
Results
Screening of DEmiRNA and DEmRNAs
A total of 28 DEmiRNAs were identified between NPC tissues and controls based on the pre-determined criteria, which consisted of 13 up-regulated miRNAs and 15 down-regulated miRNAs (Table 2). Additionally, 2593 DEmRNAs, including 1399 up-regulated genes and 1194 down-regulated genes, were also obtained after microarray data mining and standardization (Table S2). The Figure S1 displayed the differential expression levels of all genes in NPC tissues among all datasets and the top 20 DEmRNAs were exhibited in Table 3.
The Differentially Expressed miRNAs in NPC.

miRNA, microRNA; NPC, nasopharyngeal carcinoma; FDR, false discovery rate
The Top 40 Differentially Expressed mRNAs in NPC.

mRNA, messenger mRNA; NPC, nasopharyngeal carcinoma
Prediction of DEmiRNA Targets and Functional Analyses
Firstly, we determined 751 target genes for DEmiRNAs based on the miRTarBase 7.0. Then, the DEmiRNAs-target DEmRNAs network involving 1140 interaction pairs was established, of which, there were 280 up-regulated DEmiRNAs-down-regulated DEmRNAs interactions such as miR-452-
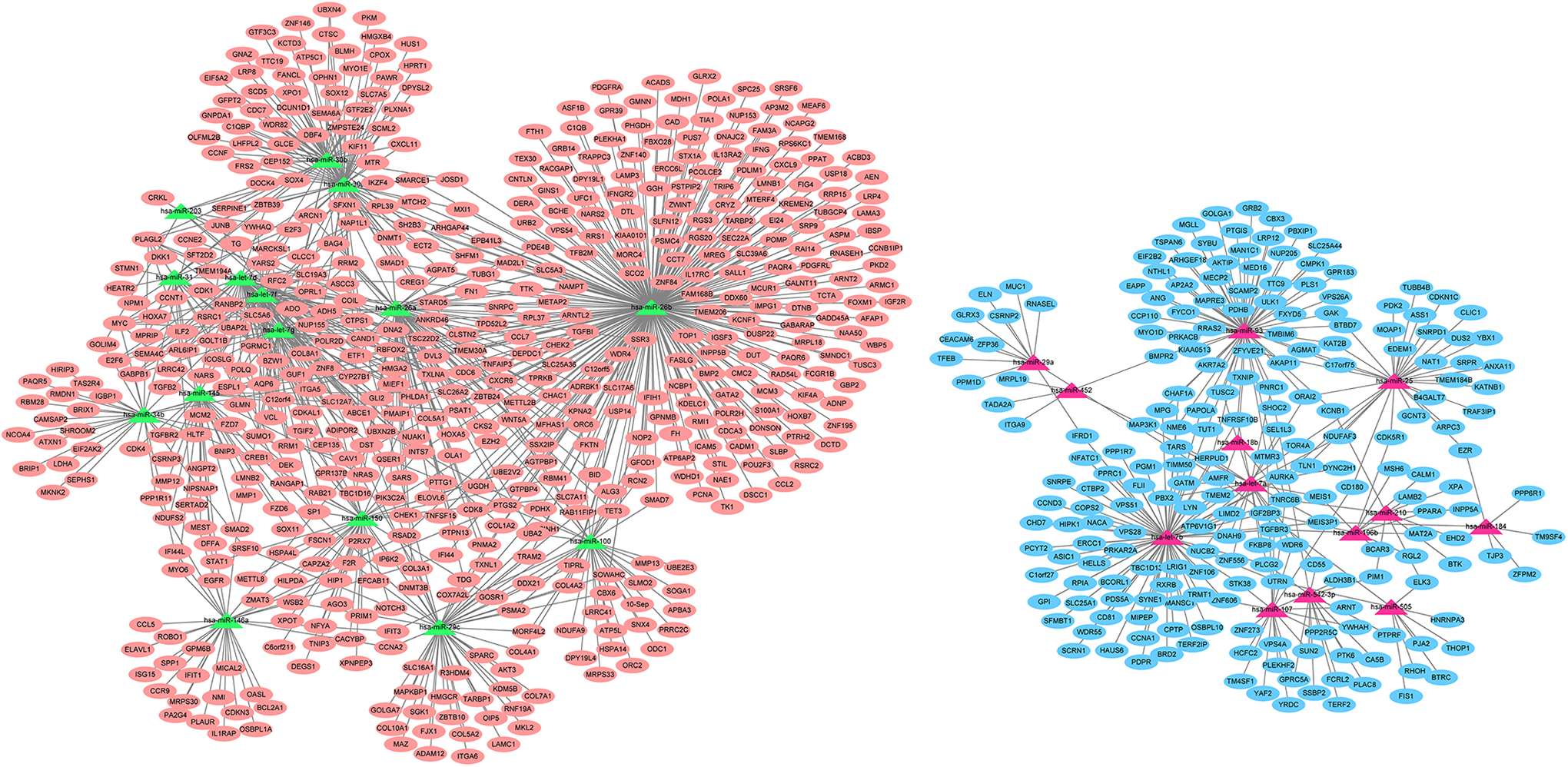
The DEmiRNAs-target DEmRNAs network. The orange ovals show the up-regulated target genes for DEmiRNAs and blue ovals indicates the down-regulated target genes for DEmiRNAs in nasopharyngeal carcinoma tissues. The red triangles indicate up-regulated miRNAs while green triangles denote down-regulated miRNAs in nasopharyngeal carcinoma tissues. DEmiRNAs: differentially expressed microRNAs.
The Functional Analyses of Target Genes for Differentially Expressed miRNAs in NPC.

GO, Gene ontology; BP, biological process; MF, molecular function; CC, cellular component; NPC, nasopharyngeal carcinoma
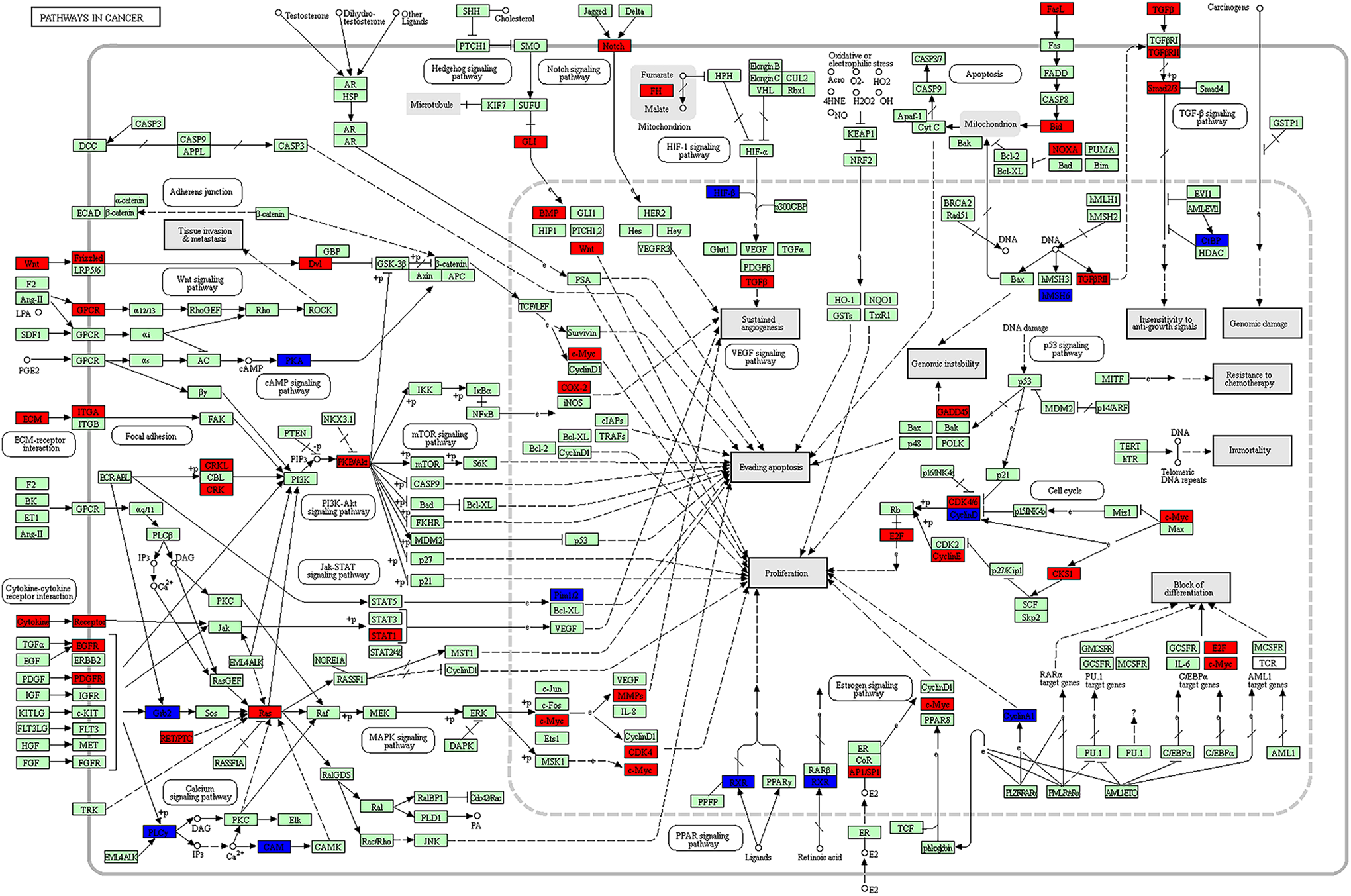
Pathway in cancer from the Kyoto Encyclopedia of Genes and Genomes enrichment analysis. The red color shows the up-regulated genes while blue color represents the down-regulated genes in nasopharyngeal carcinoma tissues. The green color indicates genes which had no significant difference between nasopharyngeal carcinoma tissues and control tissues.

MicroRNAs in cancer from the Kyoto Encyclopedia of Genes and Genomes enrichment analysis. The red color shows the up-regulated genes while blue color represents the down-regulated genes in nasopharyngeal carcinoma tissues. The green color indicates genes which had no significant difference between nasopharyngeal carcinoma tissues and control tissues.
Confirmation of DEmiRNAs in NPC Plasma Samples
For GSE43329 dataset, we found that let-7b, let-7f, let-7 g, miR-107 and miR-29a were dramatically under-expressed in NPC plasma compared to normal controls (

The ROC curves of key miRNAs in GSE43329 and GSE48442 datasets. The X-axis indicated 1-specificity and Y-axis represented sensitivity. (A) ROC curve of let-7f in GSE43329 dataset; (B) ROC curve of let-7 g in GSE43329 dataset; (C)ROC curve of miR-29a in GSE43329 dataset; (D) ROC curve of miR-452 in GSE48442 dataset. ROC: receiver operating characteristic.
Correlation Analysis between DEmiRNAs and Clinical Parameters
The relationships between DEmiRNAs and 2 clinical parameters (stage and overall survival) were analyzed (Figure S2-4). The results showed that miR-150 and miR-29a were associated with NPC stage (miR-150:
Discussion
NPC has been recognized as a serious public health issue around the globe due to the high prevalence and mortality.
24
A growing number of researchers have concentrated on identifying potential miRNA signatures related to NPC in the past decade. For example, Li
Extensive studies have demonstrated that miRNAs might serve as oncogenes and contribute to the initiation and progression of NPC through regulating their target genes that participate in numerous cellular processes.
27,28
We carried out an integrated analysis of miRNAs and mRNAs microarray datasets and our findings indicated that down-regulated let-7d, let-7f and their targets genes (up-regulated MYC and HMGA2) played essential roles in carcinogenesis. Besides, let-7d and let-7f were also validated to be under-expressed in NPC plasma samples in comparison with healthy controls from other datasets, which suggested that plasma let-7d and let-7f level may be potential biomarkers for NPC detection. Overwhelming evidence has demonstrated that miRNA let-7 family (-a, -b, -c, -d, -e, -f, -g and –i) exerted regulatory roles at the transcriptional and post-transcriptional levels among species and their aberrant expression might be closely linked with the pathogenesis of cancers such as NPC.
29,30
Luan
MiR-452, another vital regulator, was over-expressed in NPC tissues compared with normal controls, which was further confirmed in NPC plasma samples. Moreover, a lower expression of miR-452 represented a higher survival rate. Existing studies suggested that miR-452 played key roles in pathogenic mechanisms of a wide variety of cancers, including non-small cell lung cancer, prostate cancer and hepatocellular carcinoma.
35
-37
However, the potential effects of miR-452 on NPC initiation have not been elaborated. Herein, a close correlation between miR-452 and down-regulated ITGA9 was detected and functional analyses implied that this gene was primarily responsible for cellular processes. ITGA9 was located at 3p21A region and a linkage analysis showed that the mutations in this segment might induce NPC formation.
38
Later on, Ng
Although several prominent gene and miRNAs biomarkers have been identified using bioinformatics methods, there are still limitations in our current analysis. Firstly, a comprehensive bioinformatics analysis based on a larger sample size needs to be carried out to confirm our results. Secondly, the corresponding experiments such as qPCR array and NPC cell experiments require to verify several potential RNA transcripts and corresponding signaling pathway. Finally, the extensive clinical information will also be requisite to integrate into following analysis to evaluate prognosis.
Conclusion
In summary, we performed an integrated analysis of miRNAs and mRNAs to identify potential biomarkers for NPC treatment. The results showed that let-7d-MYC/HMGA, let-7f-MYC/HMGA2 and miR-452/ITGA9 axis were possibly associated with NPC progression, which will provide novel insights into developing novel therapeutic strategies for diagnosis and treatment of NPC. However, a detailed bioinformatics analysis and experimental validation also need to be undertaken in following investigation.
Supplemental Material
Supplemental Material, Figure_S1 - An Integrated Analysis of mRNAs and miRNAs Microarray Profiles to Screen miRNA Signatures Involved in Nasopharyngeal Carcinoma
Supplemental Material, Figure_S1 for An Integrated Analysis of mRNAs and miRNAs Microarray Profiles to Screen miRNA Signatures Involved in Nasopharyngeal Carcinoma by Lei Liu, Hailing Wang, Chaohui Yan and Shudong Tao in Technology in Cancer Research & Treatment
Supplemental Material
Supplemental Material, Figure_S2 - An Integrated Analysis of mRNAs and miRNAs Microarray Profiles to Screen miRNA Signatures Involved in Nasopharyngeal Carcinoma
Supplemental Material, Figure_S2 for An Integrated Analysis of mRNAs and miRNAs Microarray Profiles to Screen miRNA Signatures Involved in Nasopharyngeal Carcinoma by Lei Liu, Hailing Wang, Chaohui Yan and Shudong Tao in Technology in Cancer Research & Treatment
Supplemental Material
Supplemental Material, Figure_S3 - An Integrated Analysis of mRNAs and miRNAs Microarray Profiles to Screen miRNA Signatures Involved in Nasopharyngeal Carcinoma
Supplemental Material, Figure_S3 for An Integrated Analysis of mRNAs and miRNAs Microarray Profiles to Screen miRNA Signatures Involved in Nasopharyngeal Carcinoma by Lei Liu, Hailing Wang, Chaohui Yan and Shudong Tao in Technology in Cancer Research & Treatment
Supplemental Material
Supplemental Material, Figure_S4 - An Integrated Analysis of mRNAs and miRNAs Microarray Profiles to Screen miRNA Signatures Involved in Nasopharyngeal Carcinoma
Supplemental Material, Figure_S4 for An Integrated Analysis of mRNAs and miRNAs Microarray Profiles to Screen miRNA Signatures Involved in Nasopharyngeal Carcinoma by Lei Liu, Hailing Wang, Chaohui Yan and Shudong Tao in Technology in Cancer Research & Treatment
Supplemental Material
Supplementary_tables - An Integrated Analysis of mRNAs and miRNAs Microarray Profiles to Screen miRNA Signatures Involved in Nasopharyngeal Carcinoma
Supplementary_tables for An Integrated Analysis of mRNAs and miRNAs Microarray Profiles to Screen miRNA Signatures Involved in Nasopharyngeal Carcinoma by Lei Liu, Hailing Wang, Chaohui Yan and Shudong Tao in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Authors’ Note
Lei Liu and Hailing Wang contributed equally to this work.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
