Abstract
Background: Small nucleolar RNA host gene 3 (SNHG3), as a novel long non-coding RNA (lncRNA) participates in the oncogenic processes of various cancers. We combined a systematic review and a meta-analysis to assess the potential role of SNHG3 as a pan-cancer marker for cancer diagnosis and prognosis. Methods: Our study comprehensively searched for SNHG3 expression profiling studies from PubMed, Web of Science, Excerpta Medica Database (EMBASE), Cochrane Library, Google Scholar, and The Cancer Genome Atlas (TCGA). The diagnostic capacity of SNHG3 for all cancers in TCGA database was evaluated from the perspective of pooled sensitivity, specificity, diagnostic odds ratio (DOR), area under the curve (AUC) of the summary receiver operating characteristic (sROC) curve. Also, this research studied the correlation of SNHG3 expression and the overall survival to access its prognostic value. Results: A sum total of 11,888 cancer patients and 730 controls from 44 eligible studies were enrolled in this integrated analysis. In TCGA database, SNHG3 was significantly upregulated in most types of cancers (16/33, 48%). The pooled sensitivity, specificity, and DOR with 95% CIs was 0.72 (95% CI: 0.60–0.82), 0.87 (95% CI: 0.84–0.90), and 18 (95% CI: 11–30), respectively. Similarly, the AUC of the sROC curve was 0.89 (95% CI: 0.86–0.92), indicating SNHG3 was highly conserved as a diagnosis biomarker. Additionally, SNHG3 overexpression significantly deteriorated the overall survival of cancer patients (pooled HR = 1.28, 95% CI:1.11–1.48; P < 0.05). Conclusions: These findings suggest that the lncRNA SNHG3 could serve as a promising diagnostic and prognostic biomarker.
Introduction
Currently, cancers cause high mortality and morbidity all over the world. 1 According to the GLOBOCAN estimates, there were approximately 18,078,957 new cases of cancer and 9,555,027 million cancer-associated deaths in 2018. 2 Despite the rapid advances in diagnostic techniques and treatments in recent years, cancers are still the most serious threat to the survival of people around the world. 3
Cancer is a genetically heterogeneous disease.4,5 It is estimated that 30%–50% of cancer cases could be prevented with effective interventions. 6 Thus, the early-stage diagnosis of cancer can significantly improve the prognosis and survival of cancer patients. The emergence of new molecular tumor biomarkers featured with high sensitivity and specificity as well as low false positive rates and complexity have become particularly important, and these biomarkers could be applied to detect cancer progression.
Long non-coding RNAs (lncRNAs) are a set of non-coding RNAs with no protein-coding ability and a length of more than 200 nucleotides. 7 An increasing number of studies have shown that lncRNAs participate in the genesis and development of tumors by regulating the malignant biological behavior of tumor cells at the transcriptional and posttranscriptional levels and the epigenetics of genes.8–10 According to recent research, lncRNAs can serve as diagnostic or prognostic markers in human cancers like BLACAT1, PANDAR, Z38, and so on.11–13
Small nucleolar RNA host gene 3 (SNHG3) is a novel lncRNA, located on chromosome 1p35.3, which is significantly upregulated in several human cancers, including thyroid carcinoma, breast cancer, and lung adenocarcinoma.14,15 Increasing evidence has implied that aberrant SNHG3 expression is directly related to many oncogenic processes, such as tumor cell differentiation, proliferation, metastasis, and invasion. 16 SNHG3 mediates tumorigenesis by transcriptionally regulating p53, AKT/mTOR/ERK, and TGF-β signaling pathway. Aberrantly expressed SNHG3 plays an imperative role in the progression of multiple malignancies; moreover, high expression of SNHG3 significantly predicts the poor prognosis of patients with ovarian cancer, glioma, hepatocellular carcinoma, osteosarcoma, and colorectal cancer.17–21 Based on recent research, SNHG3 serves as a prognostic biomarker in cancers, but few studies reported SNHG3's diagnostic value. More importantly, due to the limited sample sizes and unstable outcomes in previous research, SNHG3's accuracy and reliability are still controversial as a tumor biomarker.
A systematic review and meta-analysis on available online datasets and eligible studies was performed to evaluate and estimate the potential prognostic and diagnostic value of SNHG3 as a pan-cancer biomarker.
The aim of the study
The aim of the study was to investigate the expression of SNHG3 in human cancers, the diagnostic value of SNHG3 in human cancers, the relationship between SNHG3 and the overall survival (OS) of cancer patients.
Materials and methods
This study was conducted based on the Cochrane Collaboration by following the PRISMA statement guidelines. No ethical approval and patient consent were required.
Literature search and study design
This study was conducted in accordance with the standard guidelines for a meta-analysis of original studies. The PubMed, Web of Science, Excerpta Medica Database (EMBASE), Cochrane Library, Google Scholar, and TCGA were searched. The latest search time was February 28, 2021, and the detailed search strategy is shown in Supplementary Table 1.
Inclusion and exclusion criteria
In this meta-analysis, the inclusion criteria are listed here: (a) the original data were reported and a complete peer review article was published; (b) the expression of SNHG3 was detected by reverse transcription polymerase chain reaction (RT-PCR); (c) the patient was diagnosed with cancer; and (d) studies with enough original data were used for statistical analysis of survival information, such as providing or calculating 95% CI and the hazard ratio (HR).
The exclusion criteria were as follows: (a) repetitive articles; (b) case reports, letters, expert opinions, comments, editorials, and reviews; (c) studies without available or unqualified data, such as 95% CI or HRs; and (d) non-human studies.
Data extraction and processing based on TCGA database
We constructed a biological network at the systems level based on available data from TCGA database (http://genome-cancer.ucsc.edu/). Based on TCGA data portal, 33 different types of cancer datasets including tumor samples and normal samples from TCGA database were accessed. RNA expression matrices and clinical information were obtained by using R for subsequent analysis. Totally, 11,093 samples were analyzed in our study, including 730 specimens of normal tissue samples, and 10,363 specimens of tumor tissue samples (Supplementary Table 2). An unpaired Student t test was used to compare the differences between tumor tissues and adjacent normal tissues regarding gene expression; P < 0.05 was statistically significant.
Diagnostic meta-analysis
Based on the SNHG3 expression levels of the different cancer samples collected from the TCGA database aforementioned, IBM SPSS ver. 20.0 (Stata Corporation, College Station, TX, USA) was used to calculate the area under the curve (AUC), sensitivity, and specificity of each kind of cancer. Also, the sensitivity, specificity, diagnostic odds ratio (DOR), AUC of the summary receiver operating characteristic (sROC) curve were applied to assess the diagnostic accuracy of SNHG3 via Stata ver. 15.0 software. Potential publication bias was determined by Deeks’ funnel plot asymmetrical test in this diagnostic meta-analysis.
Data extraction and quality assessment
Data from the selected original literature were screened and extracted independently by two researchers (Dingting Wang and Longfei Zou) in accordance with the criteria. For any differences that arose, the two investigators decided whether to include the study through joint discussion, and, if necessary, a third party (Conghong Zhang) made the final decision. In addition, we contacted the original author for more information if the data were incomplete.
Key components of all qualified studies were included as follows: cancer type, publication year, number of cases, first author, country origin, cut-off value, 95% CI of the HR, and the HR value. If the abovementioned data were not cited in the original studies or no replies were received from authors via email, then these items were deemed as “not reported” (NR).
The quality of each study was evaluated independently by three authors using the Newcastle–Ottawa Scale (NOS). The NOS contains eight items within three domains and the total maximum score is 9. Using the tool, each study is judged on eight items, categorized into three groups: the selection of the study groups; the comparability of the groups; and ascertaining of either the exposure or outcome of interest for case-control or cohort studies, respectively. A study with a score of 7–9 has high quality; 4–6, high risk; and 0–3, very high risk of bias. According to the NOS, only the eligible high-quality studies achieving a score of 6 or higher were included.
Statistical methods
Pooled sensitivity, pooled specificity, negative likelihood ratio (NLR), positive likelihood ratio (PLR), and DOR were calculated with corresponding 95% CIs to evaluate the diagnostic value of SNHG3. The diagnostic accuracy of the included studies was presented on the sROC curve. Heterogeneity among the included studies was determined using the I-squared statistic, with I2 values greater than 50% suggesting that substantial heterogeneity was present. A fixed-effects model was used to analyze the pooled results when the included studies showed moderate heterogeneity (I2 < 50%). Otherwise, a random effects model was employed (I2 > 50%). HRs and 95% CIs were further calculated, and GraphPad Prism 8.0 software was used to correlate survival data. Further, Engauge Digitizer 10.0 was applied to obtain survival data based on Kaplan–Meier curves. The Tierney method was used to calculate HRs and 95% CI if data were not available in published articles. The current meta-analysis was performed via Stata ver. 15.0 (Stata Inc., University Station, TX, USA). The pooled HR was of statistical significance when 95% CI excluded 1. HR > 1 indicated that patients with high SNHG3 expression had a poor prognosis. The funnel plot and Begg’s test were further adopted to access potential publication bias. A P-value < 0.05 indicated potential publication bias.
All the reported P-values were two-sided, and a P < 0.05 had statistical significance.
Results
Search results and description of studies
A schematic PRISMA flowchart describing the screening, inclusion and exclusion criteria is shown in Supplementary Figure 1 and Supplementary Table 1. The literature search yielded 229 potentially relevant publications, which were related to the topic of cancer biomarkers and included SNHG3. Of these, 111 studies eligible for inclusion were confirmed after excluding duplicate studies (118). Subsequently, 23 studies with unrelated content were excluded. The remaining full-text articles of 88 studies were assessed for suitability. However, 60 studies were dismissed because of apparent irrelevance, and 17 studies were removed because of insufficient data. Finally, 11 published studies15,17–26 and 33 datasets from TCGA were included in this meta-analysis. The SNHG3 expression values were determined before any related treatment.
SNHG3 expression in human cancers based on TCGA database
To address the role of SNHG3 in carcinogenesis, we first evaluated SNHG3 expression in TCGA datasets. The bioinformatics analysis results suggested that SNHG3 showed relatively high expression in tumor tissues versus normal tissues (Figure 1).
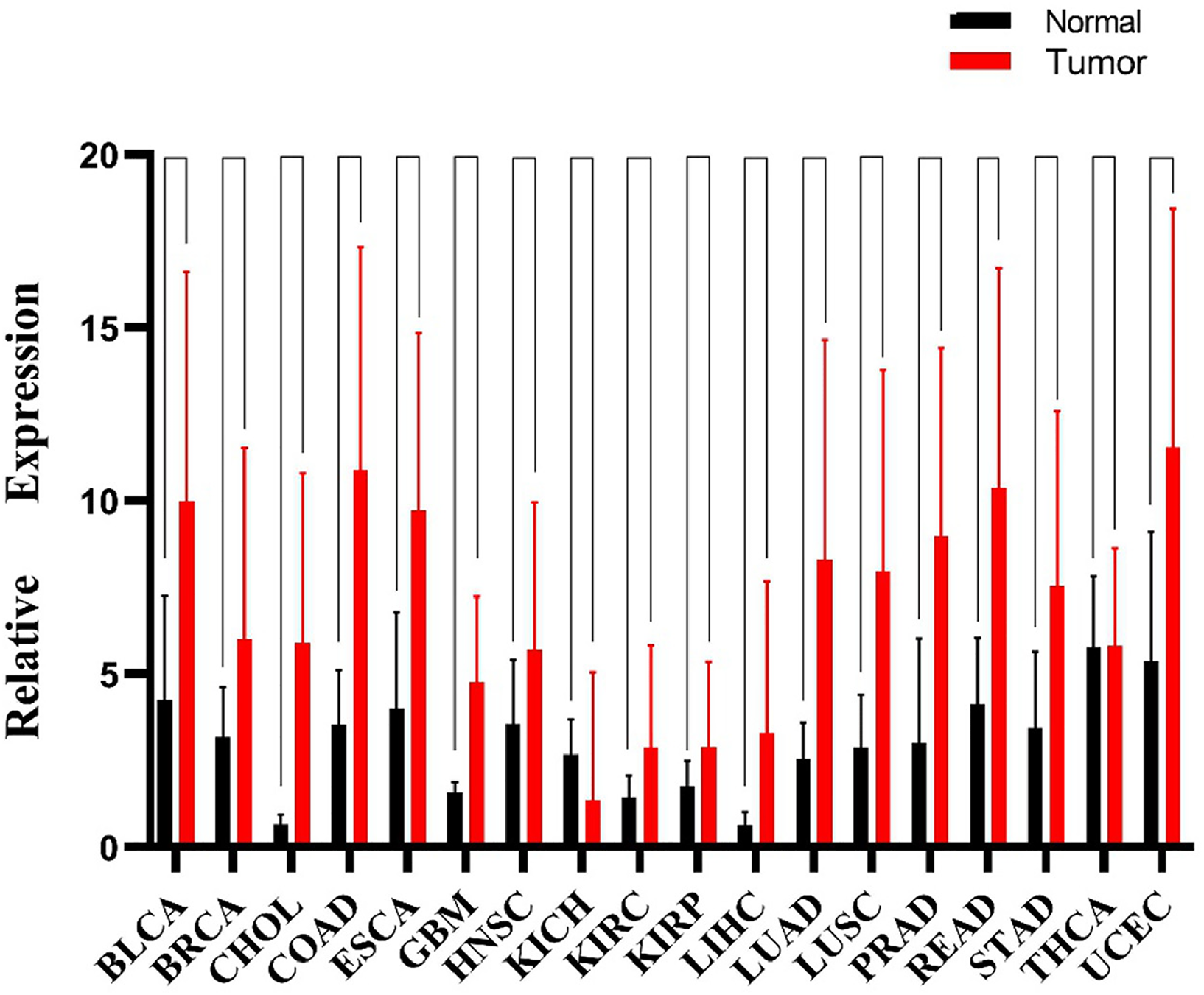
Expression of SNHG3 in human cancers based on the TCGA database.
The data relevant to gene expression of 33 human cancers were obtained from TCGA database. The RNA–Seq count matrices of 18 malignancies were gathered based on R by excluding datasets without sufficient tumor tissues or normal tissue samples. Our results clearly show that SNHG3 was considerably upregulated in 16 of 18 tumor tissue samples (Figure 1), which included bladder urothelial carcinoma (BLCA), breast invasive carcinoma (BRCA), cholangiocarcinoma (CHOL), colon adenocarcinoma (COAD), esophageal carcinoma (ESCA), glioblastoma multiforme (GBM), head and neck squamous cell carcinoma (HNSC), kidney renal clear cell carcinoma (KIRC), kidney renal papillary cell carcinoma (KIRP), liver hepatocellular carcinoma (LIHC), lung adenocarcinoma (LUAD), lung squamous cell carcinoma (LUSC), prostate adenocarcinoma (PRAD), rectum adenocarcinoma (READ), stomach adenocarcinoma (STAD), and uterine corpus endometrial carcinoma (UCEC). Nevertheless, alterations of SNHG3 expression were not observed from tumor-tissue samples of kidney chromophobe (KICH) and thyroid carcinoma (THCA), which may be due to limited data and incomprehensive investigation.
Diagnostic value of SNHG3 in human cancers
The obtained AUC, specificity, and sensitivity implied that SNHG3 was of significance in diagnosing BLCA (AUC 0.84), BRCA (0.69), COAD (0.92), ESCA (0.86), HNSC (0.64), KIRC (0.70), KIRP (0.66), LIHC (0.93), LUAD (0.89), LISC (0.86), PRAD (0.91), READ (0.86), STAD (0.81), UCEC (0.82), but not THCA (0.48) (Supplementary Table 2). The forest plots of pooled sensitivity and specificity with their 95% CIs for individual studies are shown in Figure 2(a). The overall pooled results for sensitivity, specificity, PLR, NLR, and DOR with their 95% CIs were 72.00% (95% CI: 60.00–82.00%, Figure 2(a)), 87.00% (95% CI: 84.00–90.00%, Figure 2(a)), 5.80% (95% CI: 4.60–7.30), 0.31 (95% CI: 0.22–0.46), and 18 (95% CI: 11–30) respectively; which introduced the SNHG3 as accurate indicator for the diagnosis of different cancers. Subsequently, it could be concluded that sROC curve analysis based on the methods used in a previously published study32 was adopted to further identify the diagnostic value of SNHG3 in human cancers. The pooled sensitivity and specificity, along with AUC of sROC curve, was 0.72, 0.87, and 0.89, respectively (Figure 2(c)). There was considerable heterogeneity among studies (overall I² for random-effects model = 93%). The Fagan plot suggested relatively high diagnostic accuracy of SNHG3 for cancers (Figure 2(b)). On the basis of funnel plot asymmetry test by Deeks, no substantial publication bias was discovered from sROC analysis (Figure 2(d); P = 0.54). The aforementioned results indicated that lncRNA SNHG3 was a potential diagnostic biomarker in human cancers.
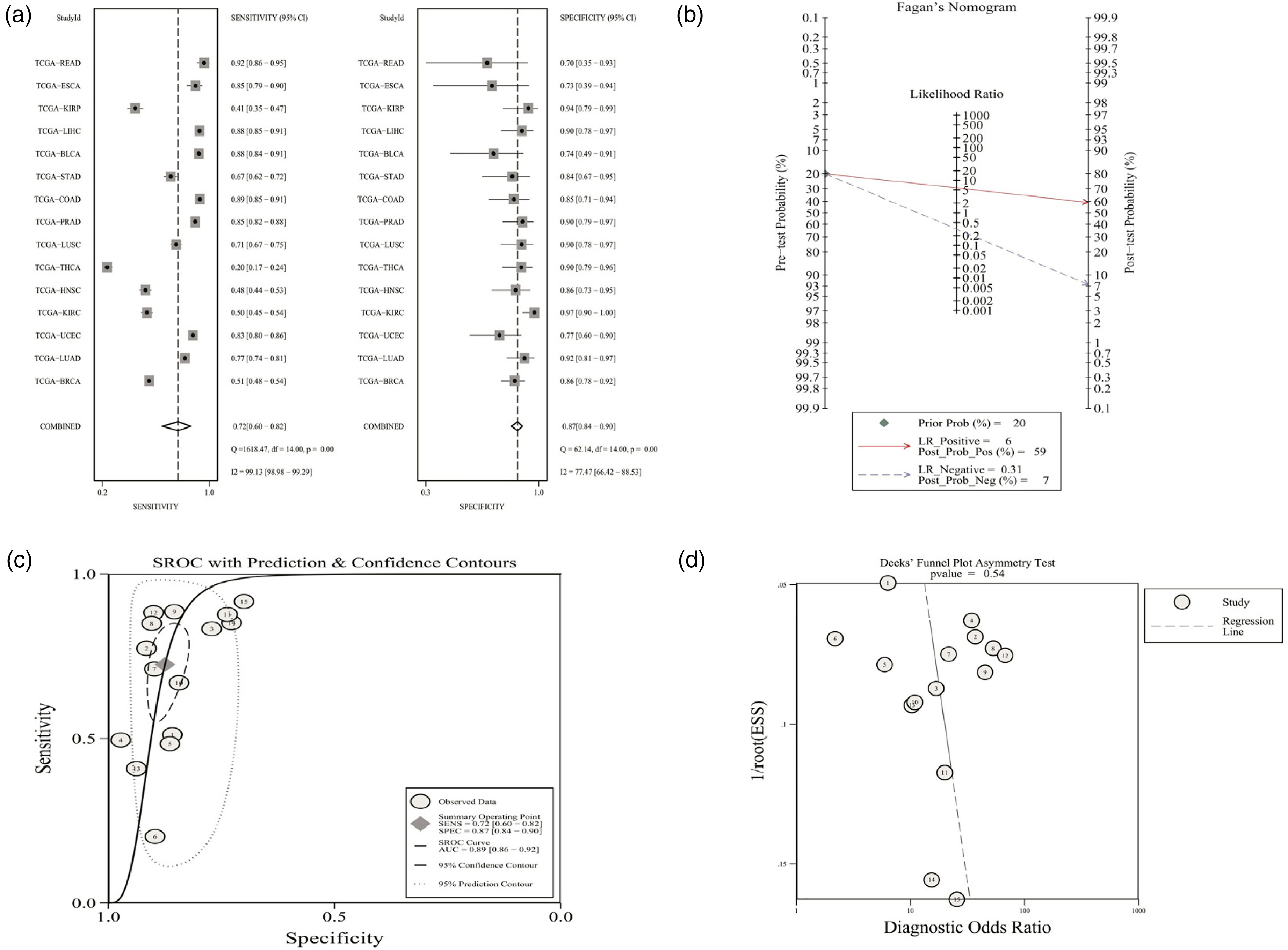
(a) Sensitivity and specificity of SNHG3 expression for the diagnosis of human cancers. (b) Fagan diagram evaluating the overall diagnostic value of SNHG3. (c) Summary receiver operating characteristic curve. (d) The Deeks’ funnel plot asymmetry test.
Prognostic value of SNHG3 for OS in human cancers
This meta-analysis was carried out to further determine the predictive values of SNHG3 for OS in human cancers. A detailed PRISMA flowchart of the screening process is shown in Supplementary Figure 1. A total of 11 published articles15,17–26 and 33 TCGA datasets (including 11,119 patients) were collected in accordance based on the defined search strategy (Table 1). The 11 selected articles were published from 2016 to 2020 and included 795 patients diagnosed with hepatocellular carcinoma, intrahepatic cholangiocarcinoma, glioma, ovarian cancer, osteosarcoma, colorectal cancer, acute myeloid leukemia, and lung cancer. The 33 TCGA datasets were composed of 33 tumors and 10,111 patients diagnosed with different cancers.
OS characteristics of studies in the meta–analysis.
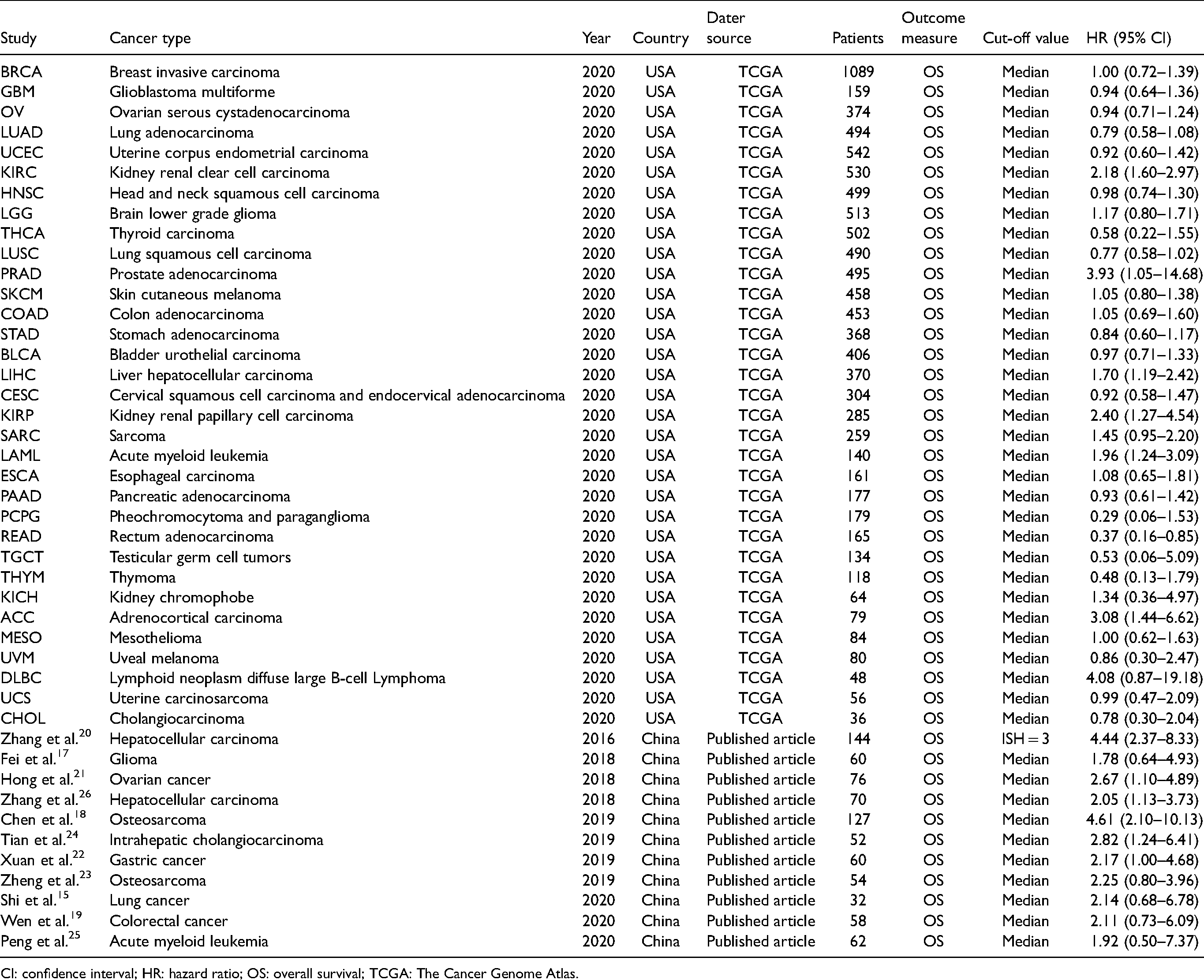
CI: confidence interval; HR: hazard ratio; OS: overall survival; TCGA: The Cancer Genome Atlas.
The pooled HR included in the current meta-analysis implied that SNHG3 upregulation was relevant to poor OS among cancer cases (pooled HR = 1.28, 95% CI:1.11–1.48, P < 0.001, random-effect model; Figure 3(b)). The Begg's funnel plot was used to evaluate publication bias, and the results indicated no substantial publication bias (P = 0.24; Figure 3(c)). According to the sensitivity analysis, the omission of individual data sets had no substantial impact on the reliability of results, confirming that they were reliable.
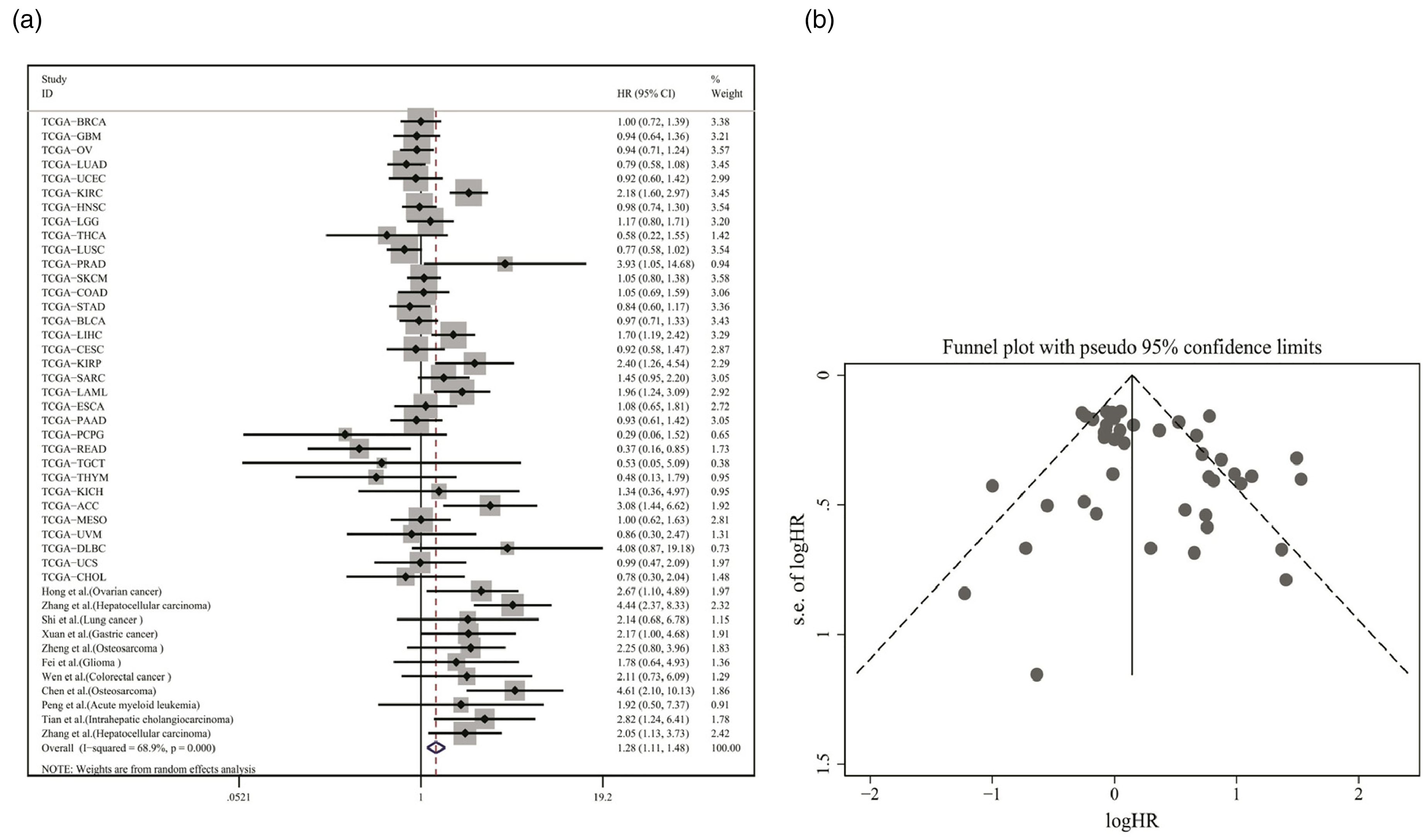
(a) Relationship between SNHG3 expression and overall survival rate. (b) Begg's publication bias plots of studies evaluating the relationship between SNHG3 expression and the survival rate of cancer patients.
Prognostic value of SNHG3 for DSS, DFI, and PFI in human cancers
We then applied the Kaplan–Meier survival analysis to assess the prognostic value of SNHG3 in human cancers from 33 TCGA datasets. As shown in Figure 4(a), several kinds of cancer types showed that SNHG3 overexpression is associated with DSS, including sarcoma (SARC), PRAD, KIRC, adrenocortical carcinoma (ACC). Furthermore, SNHG3 overexpression in tumor tissues was significantly related to poor DFI in SARC, PRAD, and UCEC patients (Figure 4(b)). In addition, as shown in Figure 4(c), SARC, UCEC, ACC, LIHC, pheochromocytoma and paraganglioma (PCPG), and PRAD patients with high SNHG3 expression were associated with worse PFI. High expression of SNHG3 was significantly correlated with poor DSS, DFI and PFI in human cancers (P < 0.05). Therefore, SNHG3 is a good predictor of prognosis, especially in PRAD and SARC. These results are encouraging and further prove that SNHG3 is a potential prognostic biomarker in human patients.
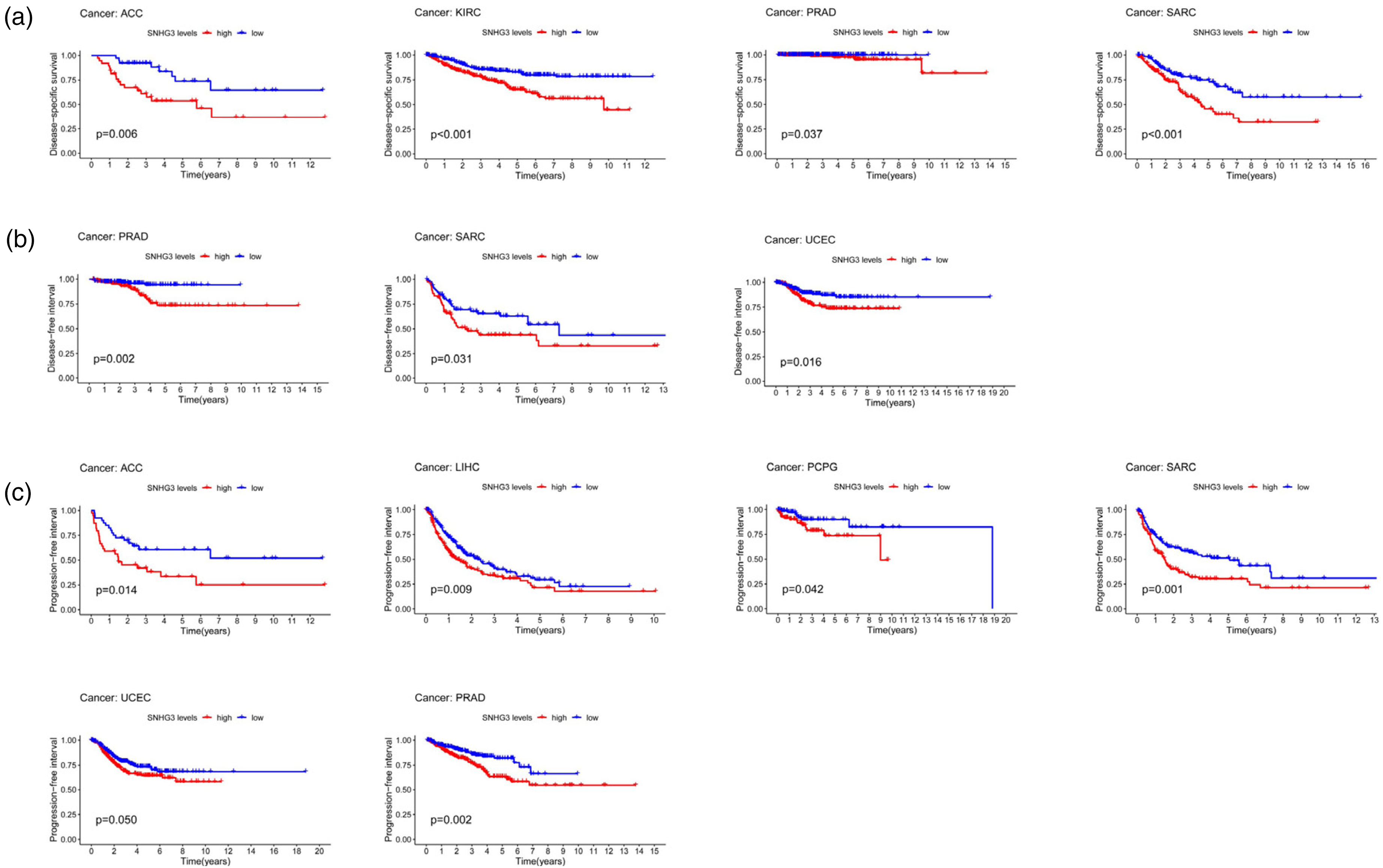
(a) Kaplan–Meier survival curves for disease-specific survival (DSS) according to SNHG3 expression. (b) Kaplan–Meier survival curves for disease-free interval (DFI) according to SNHG3 expression. (c) Kaplan–Meier survival curves for progression-free interval (PFI) according to SNHG3 expression.
Discussion
In recent years, the oncogenic property of SNHG3 was discovered from many human cancers, and some studies have showed molecular mechanisms behind correlation between high levels of SNHG3 and poor prognosis in cancer patients. Highly expressed SNHG3 promotes tumor cell proliferation and migration by activating the TGF-β pathway and the IL-6/JAK2/STAT3 pathway. 15 An overexpression of SNHG3 reportedly accelerates tumor cell growth and represses apoptosis by inhibiting KLF2 and p21. Recent studies suggest that SNHG3 was upregulated and accelerated cancer progression by sponging miR-384, miR-182-5p, and miR-139-5p.16,27,28
Also, the prognostic value of SNHG3 in human cancers has been mentioned in some of the studies. Based on previously published articles, SNHG3 was a potential cancer biomarker and a target with favorable prognostic values in the treatment of human cancers. Given the differences between these studies, the aim of this study was to comprehensively understand the prognostic and diagnostic SNHG3 values in human cancers.
With the application of the gene chip and high-throughput sequencing, freely accessible public oncology databases (e.g., TCGA and GEO) are playing an increasingly important role in human cancer research. Bioinformatics analysis, data mining, and meta-analysis are regarded as some of the most powerful technical means to study tumor genesis, which plays an important role in the development of human cancers.
For example, Zhang et al. discovered that the lncRNA TUBA4B had good prognostic value in human cancers on the basis of data downloaded from the GEO database. 29 In addition, Yue et al. showed that the pseudogene DUXAP10 showed significant overexpression in human cancers that high DUXAP10 expression was relevant to poor outcomes in human cancers based on a meta-analysis using data extracted from the GEO and TCGA databases. 30
In this study, the data mining, sROC, and meta-analysis were carried out to explore the predictive and diagnostic value of SNHG3 in human malignancies. Initially, the expression information of SNHG3 from 11,093 cancer patients was collected from original data based on TCGA database, and results showed that SNHG3 was overexpressed in 16 out of 33 human cancers. ROC and sROC curve analyses were then performed to determine the diagnostic value of SNHG3, and it was found that SNHG3 had good diagnostic value in 14 types of human cancers, especially LIHC (AUC was 0.984), COAD (0.928), and PRAD (0.917). Moreover, in accordance with information collected from TCGA database and published articles, HR with 95% CIs of cancer patients were calculated accordingly. It was found that OS of patients with high SNHG3 expression was much shorter than that of patients with low SNHG3 expression, which was consistent with the results reported by Jiang and Le. 31 In addition, we found that SNHG3 high expression was significantly correlated with poor DSS, DFI, and PFI, specifically in PRAD and SARC. All in all, the above results showed that SNHG3 was significantly overexpressed in various human cancers of great values for diagnosis and prognosis.
Nevertheless, our study had several limitations that need to be addressed. First, substantial heterogeneity was found during the calculation of sensitivity and specificity from diversified TCGA datasets. And further analysis is required to find out the major source of heterogeneity. Second, this research extracted some HRs from the Kaplan–Meier curve, which may result in a small statistical error because they are not provided in the published papers. Third, subgroup analyses of prognostic meta-analyses are not available due to studies on specific cancers rather than all cancers.
In conclusion, SNHG3 shows significant overexpression in human cancers. Therefore, high SNHG3 expression means poor prognosis in human cancers. Also, SNHG3 might be a potential biomarker in the diagnosis of cancers. This comprehensive analysis brings a brand-new perspective to study the relationship of lncRNA SNHG3 and human cancers. However, further research needs to be conducted to verify the predictive value of SNHG3 in patients with various cancers.
Supplemental Material
sj-jpg-1-jbm-10.1177_03936155221077121 - Supplemental material for Potential diagnostic and prognostic value of the long non-coding RNA SNHG3 in human cancers: A systematic review and meta-analysis
Supplemental material, sj-jpg-1-jbm-10.1177_03936155221077121 for Potential diagnostic and prognostic value of the long non-coding RNA SNHG3 in human cancers: A systematic review and meta-analysis by Dingting Wang, Longfei Zou, Jian Luo, Conghong Zhang, Huajun Feng and Gang Qin in The International Journal of Biological Markers
Supplemental Material
sj-docx-2-jbm-10.1177_03936155221077121 - Supplemental material for Potential diagnostic and prognostic value of the long non-coding RNA SNHG3 in human cancers: A systematic review and meta-analysis
Supplemental material, sj-docx-2-jbm-10.1177_03936155221077121 for Potential diagnostic and prognostic value of the long non-coding RNA SNHG3 in human cancers: A systematic review and meta-analysis by Dingting Wang, Longfei Zou, Jian Luo, Conghong Zhang, Huajun Feng and Gang Qin in The International Journal of Biological Markers
Supplemental Material
sj-docx-3-jbm-10.1177_03936155221077121 - Supplemental material for Potential diagnostic and prognostic value of the long non-coding RNA SNHG3 in human cancers: A systematic review and meta-analysis
Supplemental material, sj-docx-3-jbm-10.1177_03936155221077121 for Potential diagnostic and prognostic value of the long non-coding RNA SNHG3 in human cancers: A systematic review and meta-analysis by Dingting Wang, Longfei Zou, Jian Luo, Conghong Zhang, Huajun Feng and Gang Qin in The International Journal of Biological Markers
Supplemental Material
sj-docx-4-jbm-10.1177_03936155221077121 - Supplemental material for Potential diagnostic and prognostic value of the long non-coding RNA SNHG3 in human cancers: A systematic review and meta-analysis
Supplemental material, sj-docx-4-jbm-10.1177_03936155221077121 for Potential diagnostic and prognostic value of the long non-coding RNA SNHG3 in human cancers: A systematic review and meta-analysis by Dingting Wang, Longfei Zou, Jian Luo, Conghong Zhang, Huajun Feng and Gang Qin in The International Journal of Biological Markers
Footnotes
Acknowledgments
Not applicable
Author contributions
Dingting Wang and Longfei Zou contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (grant nos. 81773529 to G.Q).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
