Abstract
BACKGROUND:
A recent study has demonstrated that the long non-coding RNA VPS9D1-AS1 is highly expressed in colorectal cancer and predicts poor prognosis. However, roles of VPS9D1-AS1 in gastric cancer remained poorly understood.
OBJECTIVE:
The aim of this study is to decipher the expression of VPS9D1-AS1 in gastric cancer (GC) patients, so as to assess whether or not it could be used as a novel biomarker for prognosis in gastric cancer patients.
METHODS:
The expression of VPS9D1-AS1 was examined in cancer tissues and paired adjacent non-tumorous tissues from 126 gastric cancer patients using qRT-PCR. Correlations between the expression of VPS9D1-AS1 and clinicopathological parameters and patients’ survival were analyzed.
RESULTS:
VPS9D1-AS1 expression was downregulated in gastric cancer tissues than that in adjacent non-tumorous tissues (
CONCLUSIONS:
Overall, our data suggest that downregulated VPS9D1-AS1 may be used as a novel prognosis predictor of gastric cancer.
Introduction
Gastric cancer (GC) remains one of the most common malignancies and the third leading cause of cancer-related death worldwide [1, 2]. Although numerous advances in diagnostic interventions and therapeutic alternatives have been made in the past decades, gastric cancer still constitutes a major challenge because more than half of patients have stepped into a progressive stage at diagnosis [3, 4]. The 5-year overall survival rate of GC patients is estimated to be less than 30% [1, 4]. It is thus urgent to find novel molecular biomarkers for diagnosis and treatment of GC.
Recently, development in sequencing technology and bioinformatics has contributed to the findings that the vast majority of human genomes are actively transcribed into non-coding RNAs without protein coding potentials [5, 6], which are classified into long non-coding RNAs (lncRNAs) and small non-coding RNAs such as microRNAs according to their lengths [7]. Previous studies have revealed that microRNAs play important roles in cancer development [8]. Increasing evidences are emerging to show that dysregulated expression of lncRNA may be associated with a variety of biological processes including cell growth, differentiation, senescence, tumorigenesis and treatment response [6, 7, 9]. For example, the lncRNA XIST, which was essential for X chromosome inactivation during female development [10], was found to be overexpressed in GC and promote malignant phenotype of GC cell lines in vitro and in vivo [11]. HOTAIR acted as a scaffold to recruit proteins required for chromatin remodeling [12] and high expression of HOTAIR predicted poor prognosis of several human cancers [13, 14]. The lncRNA VPS9D1-AS1 was recently found to be upregulated in human colorectal cancer tissues and considered to be a potential prognosis biomarker via re-mining the Affymetrix array data [15]. However, roles of VPS9D1-AS1 in GC remain poorly understood.
In this study, the expression of VPS9D1-AS1 was investigated in GC cancer tissues and corresponding adjacent non-tumorous tissues. The association between expression of VPS9D1-AS1 and clinicopathological parameters was investigated. This could highlight the possibility whether VPS9D1-AS1 could be used as a novel diagnosis biomarker or treatment target of GC.
Materials and methods
Clinical samples
A total of 126 primary GC cancer tissues and matched adjacent non-tumorous tissues were obtained from patients who underwent gastrectomy in the Sun Yan-sen University Cancer Center between April 2007 and September 2010. No patients enrolled into our study received radiotherapy or chemotherapy prior to surgery. Informed consent of all the patients was obtained for using the tissue samples for research purpose. Our study about the human specimens was approved from the Ethics Committee of the Sun Yan-sen University Cancer Center and in accordance with the principles set out in the Declaration of Helsinki 1964 as modified by subsequent revisions. The samples were immediately snap-frozen in liquid nitrogen and stored in
Cell lines and cell culture
The GC cell lines and one normal gastric epithelial cell line (GES1) were obtained from the Shanghai Cell Bank of the Chinese Academy of Sciences (Shanghai, China). The gastric cancer cells were cultured in RPMI1640 (MGC803, BGC823 and MKN45) or in DMEM (SGC7901 and AGS) supplemented with 10% fetal bovine serum (FBS), 100 U/ml penicillin, and 100 mg/ml streptomycin (Invitrogen, Carlsbad, CA, USA) at 37
Quantitative real-time PCR (qRT-PCR) analysis
Total RNA from tissue samples or cells was extracted with TRIzol Reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. Reverse transcription from 1
VPS9D1-AS1 forward, 5’-CTCAGCAGTAAGTA ACAGTGGTAGA-3’ VPS9D1-AS1 reverse, 5’-CTTCAGCATCTTGGA GGTGTC-3’ GAPDH forward, 5’-GGAGCGAGATCCCTCCA AAAT-3’ GAPDH reverse, 5’-GGCTGTTGTCATACTTCT CATGG-3’
The average value from three biological replicates with three technical replicates was used to calculate the relative amount of VPS9D1-AS1. The ABI PRISM Cycler software (ABI, Foster City, CA, USA) was used to calculate the threshold cycle number (Ct) value for GAPDH and VPS9D1-AS1 during the log phase of each cycle. VPS9D1-AS1 levels were normalized to the expression level of GAPDH (
Statistical analysis
All statistical analyses were performed using SPSS 18.0 (IBM, Chicago, IL, USA) and GraphPad Prism 5.0 (GraphPad Software, La Jolla, CA). All data were presented as mean
Results
VPS9D1-AS1 is downregulated in GC cancer tissues and cell lines
Expression level of VPS9D1-AS1 in 126 paired GC cancer and adjacent non-tumorous tissues was detected by qRT-PCR assays. As shown in Fig. 1A, expression of VPS9D1-AS1 was significantly downregulated in cancer tissues compared with that in non-tumorous tissues. Underexpression of VPS9D1-AS1 (fold change (Tumor/Normal)
VPS9D1-AS1 is downregulated in gastric cancer tissues and cell lines. (A) Expression level of VPS9D1-AS1 in gastric tumor tissues are significantly lower than that in non-tumorous tissues. Results were presented as cycle threshold (Ct) of Ct (VPS9D1-AS1) relative to that of Ct (GAPDH). (B) Expression of VPS9D1-AS1 in 126 GC samples were shown as log2 (Ct (tumor)/Ct (normal)). (C) Expression of VPS9D1-AS1 in GC cancer cell lines including SGC7901, MKN45, BGC823, MGC803 and AGS and one gastric epithelial cell GES1 was examined. Values are from three biological replicates with three technical replicates and presented as mean 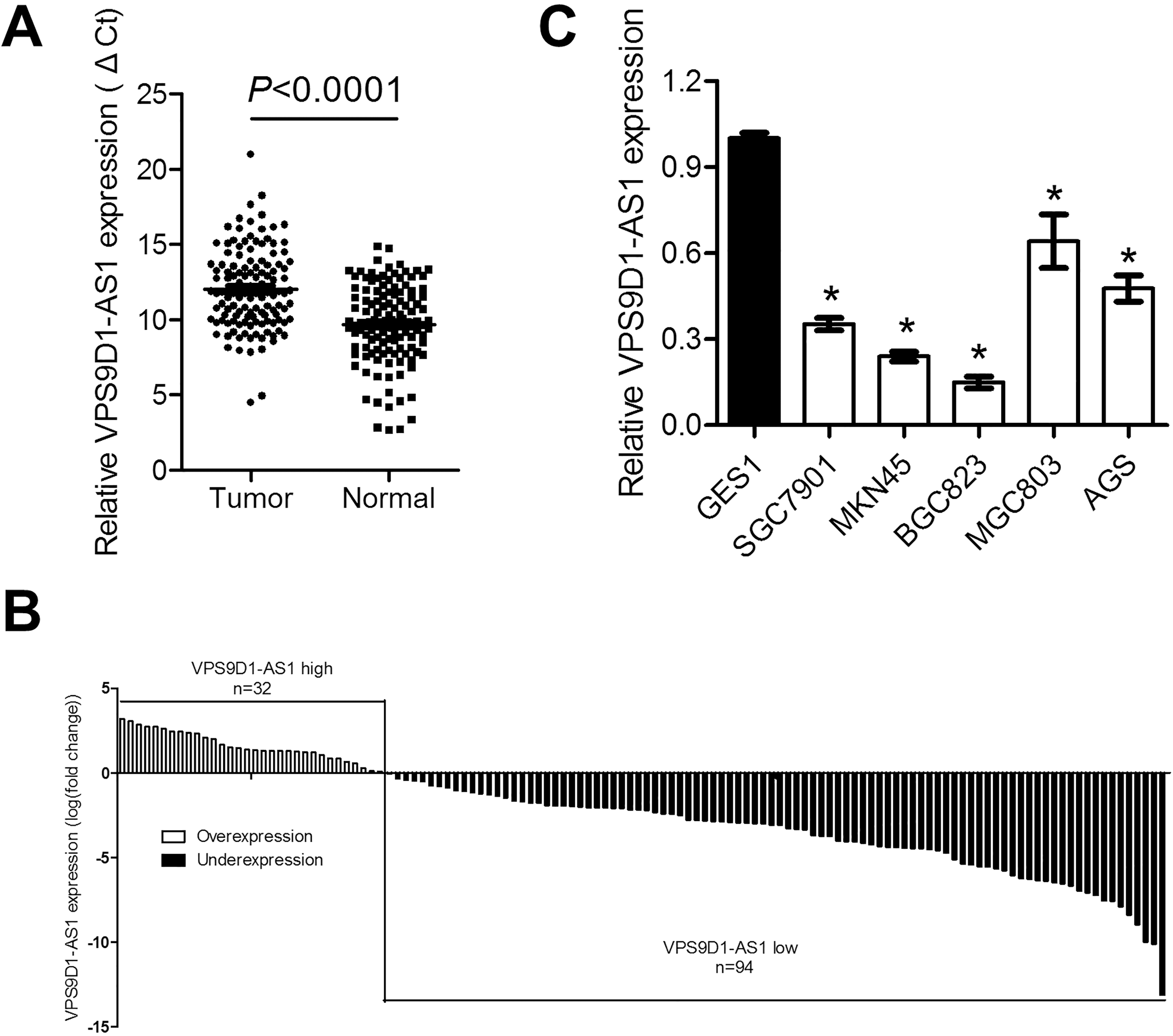
The correlation between clinicopathological parameters and VPS9D1-AS1 expression
Univariate and multivariate analyses of various potential prognostic factors in GC patients
HR: hazard ratio; CI: confidence interval;
VPS9D1-AS1 correlated with tumor size and TNM stage. (A) Relative expression level of VPS9D1-AS1 in GC cancer tissues with tumor size larger than 5 cm and smaller than 5 cm compared with tumor size smaller than 5 cm. 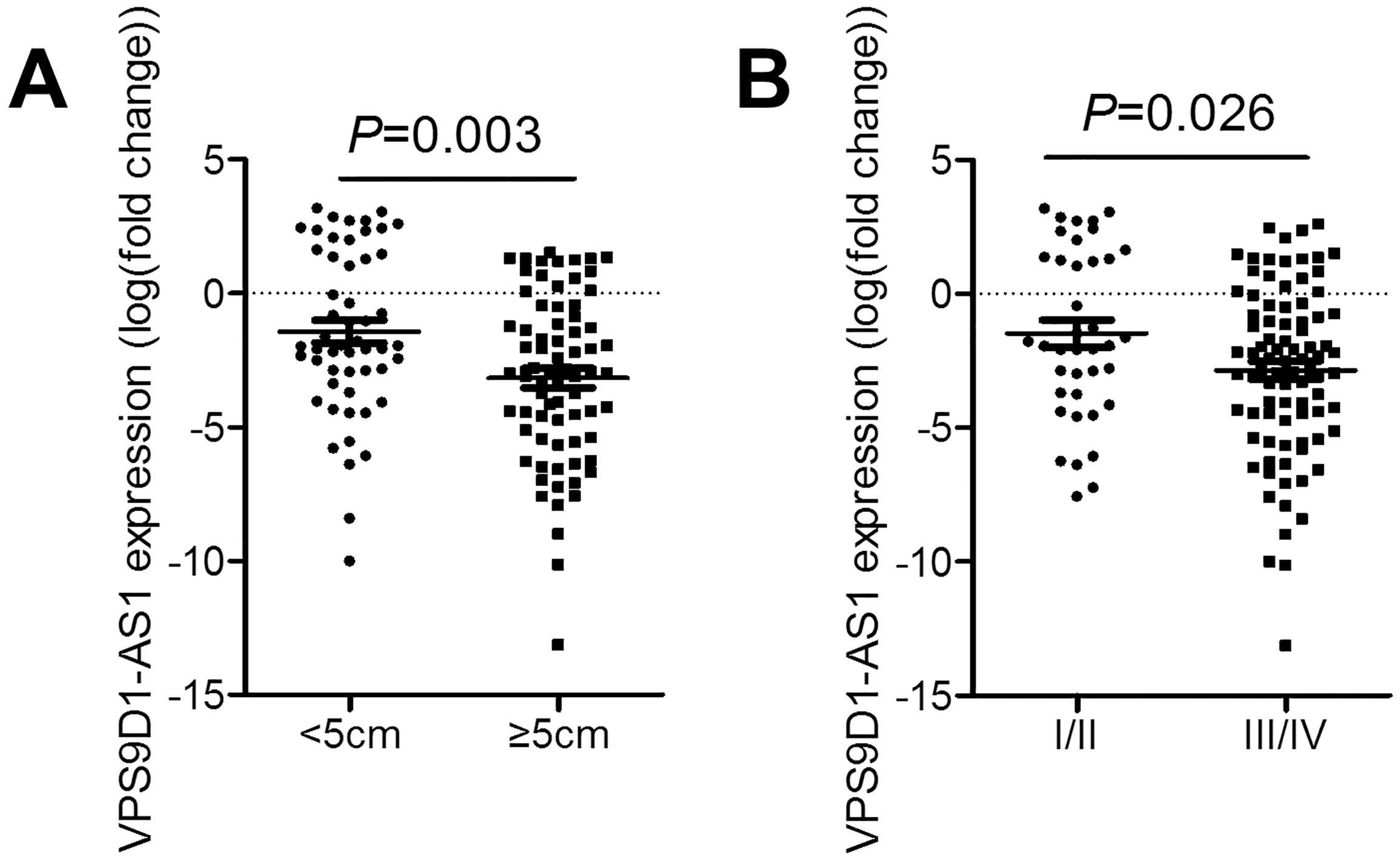
To analyze correlations between VPS9D1-AS1 expression and the clinicopathological features of 126 GC patients, our patient cohort was divided into high expression arm (expression of VPS9D1-AS1 above the cutoff,
Decreased VPS9D1-AS1 expression is an unfavorable prognostic factor in GC patients
To further explore whether VPS9D1-AS1 expression predicted prognosis of GC patients, we used Kaplan-Meier methods with log-rank tests. As shown in Fig. 3A, the median overall survival of patients with low VPS9D1-AS1 expression was significantly shorter than that of patients with high VPS9D1-AS1 expression (
Discussion
VPS9D1-AS1 predicts prognosis of GC patients. Kaplan-Meier overall survival (A) and disease-free survival (B) curves according to VPS9D1-AS1 expression level. Patients with low VPS9D1-AS1 expression (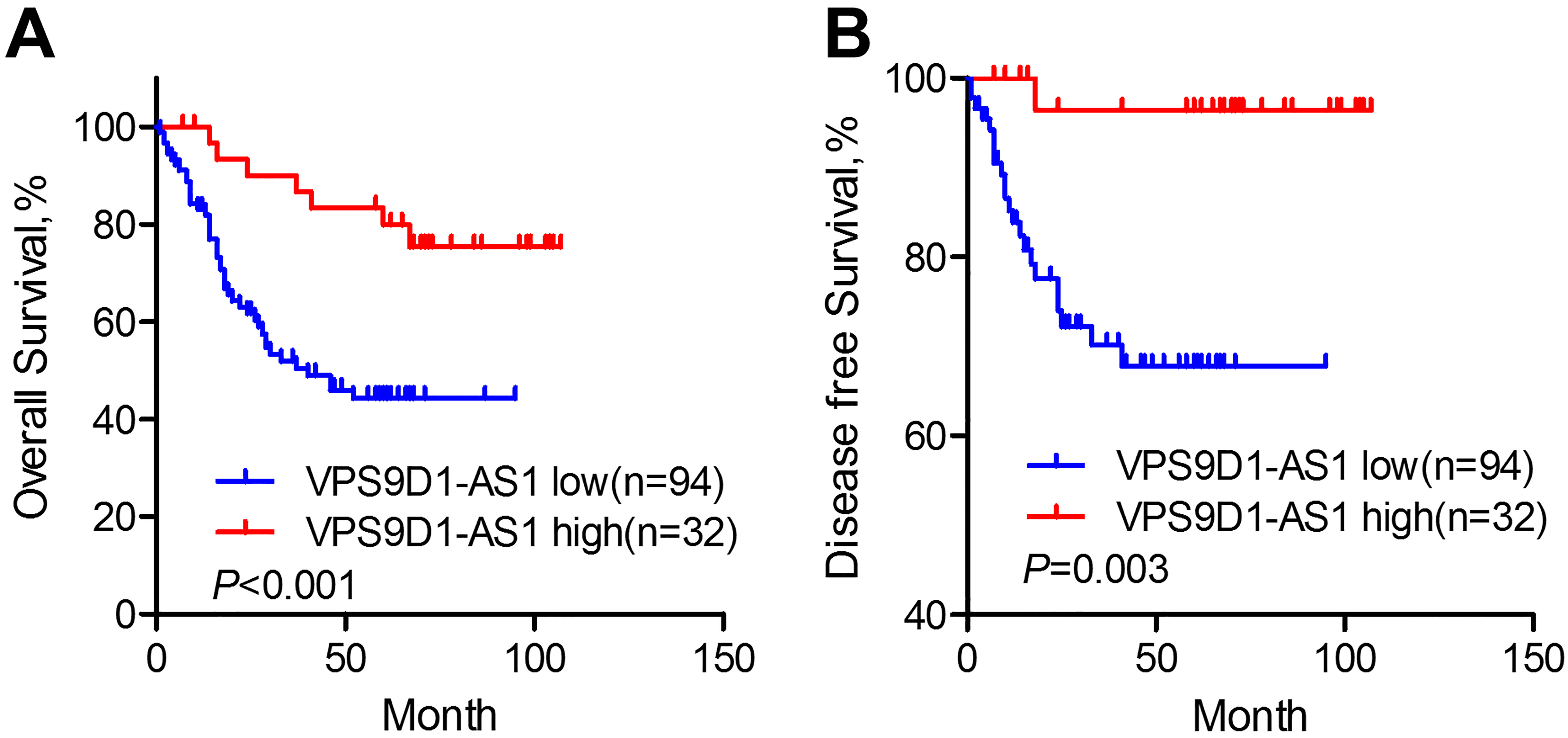
It has been recognized that in addition to protein-coding genes, non-coding genes exist primarily in human genomes [6, 9], of which lncRNA was recently identified. Generally, lncRNAs play oncogenic or suppressive roles in cancer biology, although the underlying mechanisms were diverse [9, 16, 17]. One of the well-characterized models is that lncRNAs act as endogenous competitive RNA for microRNA leading to release of downstream target genes. For instance, lncRNA PVT1 promotes gastric cancer progression via regulation of microRNA-186 [18]. LncRNAs such as HOTAIR or XIST may function as a scaffold to recruit proteins involved in chromatin remodeling or histone modification [12, 19, 20]. In addition, the lncRNA NBR2 directly binds to the metabolic master AMPK and regulates tumor cell metabolism under glucose deprivation [21]. Through interactions with other RNA, DNA or proteins, lncRNAs can regulate cell function and are proposed to be diagnostic markers or therapeutic targets of human cancer [22, 23, 24].
Over the past decades, the possible molecular alterations underlying gastric carcinogenesis have been thoroughly explored [25, 26]. However, particular pathogenesis of GC is still largely vague. Multitudinous reports have demonstrated dysregulated lncRNAs in GC. For example, the lncRNA GClnc1 was found to be overexpressed in GC and acted as a modular scaffold for WDR5 and KAT2A complexes [27]. Li et al. reported that MALAT1 promoted malignant transformation of gastric cancer by regulating vasculogenic mimicry and angiogenesis [28]. Yang et al. found higher VPS9D1-AS1 expression in the colorectal cancer tissues and CRC cell lines than in normal tissues and cell lines. Upregulated VPS9D1-AS1 indicated shortened OS time [15]. In this study, we found downregulated expression of VPS9D1-AS1 in GC tissues and cancer cell lines compared with that in adjacent non-tumorous tissues and gastric epithelial cells. Moreover, downregulation of VPS9D1-AS1 predicted poor overall and disease free survival of GC patients. These results showed that VPS9D1-AS1 may play different functions in colorectal cancer and GC. However, exact mechanisms underlying VPS9D1-AS1 in GC progression warrant further investigation.
In conclusion, we found for the first time that VPS9D1-AS1 was underexpressed in GC. Our data suggest that VPS9D1-AS1 may be used as a novel prognosis predictor of GC.
Footnotes
Acknowledgments
This work was supported by Guizhou Provincial People’s Hospital Youth Funding (No. GZSQN [2016] 20).
Conflict of interest
The authors declare that they have no conflict of interest.
