Abstract
Gastric cancer remains the third leading cause of cancer-related mortality worldwide, and proliferation of gastric cancer represents the major reason for its poor prognosis. Recent evidence indicates that long non-coding RNAs play crucial roles in development and progression of gastric cancer. Long non-coding RNA differentiation antagonizing non-protein coding RNA is upregulated in hepatic cell carcinoma, but the role of lncRNA differentiation antagonizing non-protein coding RNA in gastric cancer has not been explored. In this article, we found that differentiation antagonizing non-protein coding RNA is also upregulated in gastric cancer. Experiments revealed that silencing differentiation antagonizing non-protein coding RNA significantly inhibited gastric cancer cell proliferation in vitro and in vivo. Overexpression of differentiation antagonizing non-protein coding RNA notably increases gastric cancer cell proliferation. From RNA-seq and gene ontology annotations, we found that differentiation antagonizing non-protein coding RNA influences the gene expression programs in cell metabolic and cycle process. Taken together, our findings suggest that the long non-coding RNA differentiation antagonizing non-protein coding RNA promotes the proliferation of gastric cancer and is a potential prognostic biomarker and therapeutic target in gastric cancer.
Keywords
Introduction
Gastric cancer (GC) is the fifth most common cancers and imposes a global cancer burden. 1 Although the mortality rate of GC has declined in recent years, 2 many patients are diagnosed at an advanced stage with lymphatic or distant metastasis in the absence of specific symptoms, especially those with early-stage GC. There are limited therapeutic approaches for advanced GC, and the 5-year survival rate of stage III GC is only approximately 15%.3–5 Therefore, it is of great important to discover new molecular mechanisms and therapeutic targets that control the severity of GC and present predictive value for prognosis.
With the development of RNA sequencing and high-throughput transcriptome, more and more studies have confirmed that most of the mammalian genome is non-coding RNAs, usually including long non-coding RNAs, which are greater than 200 nt in length and most of them without protein-coding capability. 6 Accumulating studies have investigated that dysregulation of long non-coding RNAs play critical roles in various human cancers, including tumor development, progression, and metastasis, which functions similar as oncogenes or tumor suppressors.7–14 A few lncRNAs may be applied as potential biomarkers for early diagnosis, early treatment, and prognosis. However, to the best of our knowledge, the molecular functions and clinical value of most lncRNAs are not fully discussed.
Long non-coding RNA differentiation antagonizing non-protein coding RNA (lncRNA DANCR or ANCR) is located on human chromosome 4, with the closest adjacent annotated genes USP46 and ERVMER34-1. DANCR was first reported to be involved in the suppression of progenitor differentiation. 15 Yuan et al. 16 have identified that DANCR is overexpressed in stem-like hepatocellular carcinoma (HCC) cells and increases stemness features of HCC cells to promote tumorigenesis. Zhu and Xu 17 found that upregulated lncRNA ANCR expression could inhibit Runx2 expression and osteoblast differentiation by targeting enhancer of zeste homolog 2 (EZH2). Jia et al. 18 indicated that lncRNA DANCR could promote motility of prostate cancer cells and decrease expression of tissue inhibitor of metalloproteinases 2 and 3 (TIMP2/3) synergistically with EZH2, and DANCR expression is repressed by androgen–AR signaling pathway. However, the role of lncRNA DANCR in GC still remains elusive.
In this study, we performed gain- and loss-of-function studies to determine the biological roles of DANCR. We found that DANCR functions as an oncogenic molecule in GC. We also integrated the RNA-seq data and gene ontology annotations to reveal its underlying molecular mechanism in GC. These findings indicate that the DANCR is an important regulator in the development and progression of GC and provides a candidate target for GC treatment.
Materials and methods
Patient samples
A total of 118 GC samples collected from the patients in Nanjing Drum Tower Hospital from 2006 to 2015 were used in this study. None of the patients had received preoperative chemotherapy. Clinicopathological features including age, gender, tumor site, tumor size, differentiation grade, tumor–node–metastasis (TNM) stage (American Joint Committee on Cancer classification (AJCC)), lymphatic invasion, and nervous invasion were shown in Table 1. Average follow-up time was 61.3 months (median = 65 months; range = 15–98 months). This study was approved by The Clinical Research Ethics Committee of Nanjing Drum Tower Hospital. Written informed consent was obtained from all participants.
Relationship between DANCR expression and clinicopathological factors of patients with gastric cancer.

DANCR: differentiation antagonizing non-protein coding RNA; TNM: tumor–node–metastasis.
Cell culture
SGC7901, MGC803, and MKN-45 were cultured in RPMI 1640 supplemented with 10% fetal bovine serum (FBS) and 1% penicillin–streptomycin at 37°C and 5% CO2.
Oligonucleotide transfection
SiRNA was synthesized by Guangzhou RiboBio Co., Ltd. (Guangzhou, China). The siRNA sequences were as follows: si-DANCR-1: CTACAGGCACAAGCCATTG; si-DANCR-2: GCGTACTAACTTGTAGCAA. The cells were transfected using Lipofectamine RNAiMax (Invitrogen, Carlsbad, CA, USA).
Vector construction
To overexpress DANCR, the coding sequence of DANCR was amplified and subcloned into the pcDNA3.1 (+) vector (Invitrogen) according to the manufacturer’s protocol using PrimeSTAR Max DNA Polymerase Mix (TaKaRa, Dalian, China).
RNA preparation and quantitative reverse transcription polymerase chain reaction
Total RNAs were extracted from tumorous and adjacent normal tissues or cultured cells using TRIzol reagent (Life Technologies, Carlsbad, CA, USA) following the manufacturer’s protocol. Complementary DNA (cDNA) was synthesized with the PrimeScript RT Master Mix (TaKaRa) from 500 ng of RNA. The quantitative reverse transcription polymerase chain reaction (qRT-PCR) analyses were performed using SYBR Premix Ex Taq II (TaKaRa). Primers used for qRT-PCR are listed in Table 2. The qRT-PCR was performed in triplicate, and the relative expression of DANCR was calculated using the comparative cycle threshold (2−ΔCT) method with β-actin as the endogenous control to normalize the data.
Primer sequences.
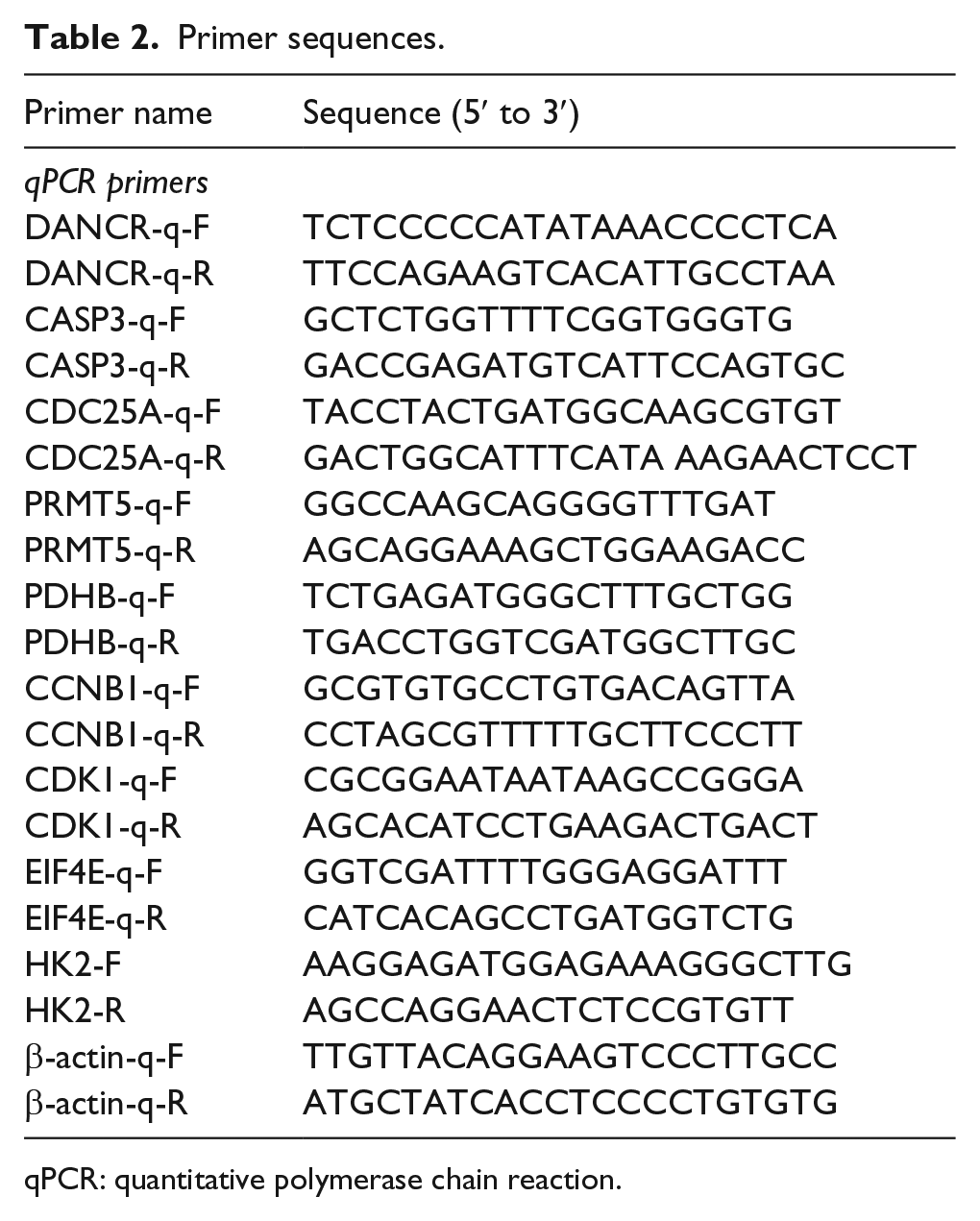
qPCR: quantitative polymerase chain reaction.
Cell Counting Kit-8 assay
Cell proliferation was performed with Cell Counting Kit-8 (CCK-8; Dojindo Laboratories, Kumamoto, Japan). Cells (1 × 103) were seeded into 96-well plates. A volume of 10 µL of CCK-8 solution was added into each well on days 1, 2, 3, and 4. After 2 h of incubation at 37°C, the absorbance at 450 nM was measured using an automatic microplate reader (Synergy 4; BioTek, Winooski, VT, USA). The experiment was repeated three times.
Colony formation assay
The cells were plated in six-well plates (1000 cells per plate) and cultured for 10–14 days. The colonies were stained with 0.1% crystal violet for 30 min after fixation with ethanol for 15 min.
In vivo tumor formation assay
For in vivo tumor formation, 2.0 × 106 control cells or the cells that stably expressed DANCR were suspended in 200 µL of serum-free RPMI 1640 and subcutaneously injected into one flank of each mouse (nine male BALB/c-nu/nu in each group for 5 weeks). After 5 weeks, the mice were sacrificed, and the parameters were measured.
Statistical analysis
Statistical analyses were performed using SPSS 20.0 (IBM, SPSS, Chicago, IL, USA) and GraphPad Prism 6 (GraphPad Software, San Diego, CA, USA). Student’s
Results
DANCR is upregulated in GC tissues and confers a poor prognosis in GC patients
To address the significance of DANCR in the clinic, we estimated the expression level of DANCR in GC tissues. Interestingly, DANCR was significantly upregulated in the GC tissues of the 118-patient cohort (Figure 1(a)). Kaplan–Meier analysis was used to evaluate the effects of DANCR expression and clinicopathological features on OS. The result shows that patients in the low DANCR expression group (

DANCR expression was upregulated in gastric cancer tissues compared with non-cancerous tissues through qRT-PCR. (a)The expression level of DANCR in 118 paired tumor and adjacent non-tumor gastric tissues (p = 0.002, n = 118). (b)Kaplan-Meier analysis of overall survival based on DANCR expression in all 118 gastric cancer patients (p = 0.035). (c) Expression levels of DANCR in 3 gastric cancer cell lines.
Silencing of DANCR inhibits GC cell proliferation and induces G2/M arrest
To investigate the biological function of DANCR in GC cells, we examined the effects of DANCR on cell proliferation and found that in vitro experiments silencing DANCR substantially reduced the rate of cell proliferation through the CCK-8 assay (Figure 2(a)). In addition, colony formation assay in both SGC7901 and MGC803 shows that silencing DANCR was also substantially reduced (Figure 2(b)). Flow cytometry analysis showed that silencing DANCR caused a dramatic increase in G2 cell population in SGC7901 and MGC803 cells, indicating DANCR was able to arrest the cell cycle in G2/M phase (Figure 2(c)).

Silencing of DANCR inhibits gastric cancer cell proliferation and induces G2/M arrest. (a) Proliferation curves of SGC7901 and MGC803 transduced with the silencing negative control (si-NC) or with the silencing DANCR (si-DANCR-1 and si-DANCR-2). Each time point represents the mean ± SEM (n = 3 independent experiment, each performed in triplicate, *P < 0.05, **P < 0.01, ***P < 0.001). (b) Colony formation assay with SGC7901 and MGC803 transduced si-NC or DANCR siRNA. The quantification of the foci per well is shown in the bottom. Error bars represents the mean ± SEM. (c) Flow cytometry analysis determined the relative cell numbers in each cell cycle phase. Numbers inside bars represent percentages of cell in each phase (*P < 0.05).
DANCR overexpression promotes GC cell growth and tumorigenicity in vitro and in vivo
When we transfected with the vector that DANCR was overexpressed by transfecting the pcDNA3.1-DANCR vector into the SGC7901 cell line and MKN-45 cell line, which harbored the lower expression level of DANCR, we found that both the proliferation and colony formation substantially increased than that observed in controls transfected with empty vector (Figure 3(a) and (b)). Also, flow cytometry analysis shows that overexpressing DANCR resulted in a substantial decrease in the number of cells of SGC7901 and MKN-45 in G2 phase (Figure 3(c)). To validate the effects of DANCR on proliferation of GC cells in vivo, MKN-45 cells, which have naturally lowest DANCR expression, compared to SGC7901 cell line and MGC803 cell line (Figure 1(c)), transfected with pcDNA3.1-DANCR were subcutaneously inoculated into the flanks of nude mice. As shown in Figure 3(d), the mice that carried the pcDNA3.1-DANCR have much bigger tumor volume than those that carried an empty vector. Above all, overexpression of DANCR promotes GC cell growth and tumorigenicity in vitro and in vivo.

DANCR overexpression promotes gastric cancer cell growth and tumorigenicity
DANCR influences gene expression programs in cell metabolic and cycle process
Our results establish that DANCR influences GC cell proliferation and tumorigenicity in vitro and in vivo. To identify what gene expression programs with DANCR ectopic expression, we examined gene expression changes upon DANCR overexpression and knockdown by high-throughput RNA-seq. 806 genes that significantly changed with both overexpression and knockdown changes reciprocally (Figure 4(a); see Supplementary Material for gene list). Consistent with the RNA-seq data, the quantitative polymerase chain reaction (qPCR) data showed significant differential expression of all genes tested (Figure 4(b)). The genes that were upregulated with DANCR overexpression and downregulated with DANCR knockdown were enriched for gene ontology annotations associated with cell metabolic and cycle process (Figure 4(c)).

DANCR influences gene expression programs in cell metabolic and cycle process. (a) Log2 fold change of all genes that significantly change (absolute value of log2 fold change > 0.5 at FDR < 0.1) with both DANCR knockdown and overexpression by RNA-seq. (b) Validation of RNA-seq data by qRT-PCR. (c) Functional Gene ontology annotations associated with the genes that significantly change with both DANCR knockdown and overexpression. Ten GO terms most associated with the change of the genes and their association p-values are shown as bar plots.
Discussion
LncRNA is commonly defined as a kind of non-coding RNA, which cannot translate into protein and with transcripts greater than 200 nt in length. They were dismissed as simply transcriptional noise in the past decades and without biological functions. Emerging studies have revealed that lncRNAs are corrected with the cancer development and progression, including metastasis, chemotherapy resistance, apoptosis, and epigenetic regulation in the area of tumor research.19,20 Therefore, there are no doubts that the area of lncRNA research offers a new perspective to investigate the cancer initiation, progression, and metastasis. Increasing evidence has revealed that long non-coding RNAs (lncRNAs) may serve as useful and potential biomarkers for the early diagnosis, early treatment, and prognosis of human cancers.
In cancer prognostic and therapeutic exploitation, emerging studies have suggested that significantly different lncRNA profiles can serve as phenotypic signatures for relevant cancers. In this study, using a cohort of GC patients, we determined that the DANCR upregulation confers poorer clinical outcome. Our data indicated that DANCR might be an attractive biomarker for risk prognostication. Collectively, the results suggest that clinical-oriented research on lncRNAs in GC should be undertaken and further research should be designed to discover more tumor-related lncRNAs as candidate prognostic biomarkers and therapy targets.
As high DANCR expression was associated with an expansion tumor phenotype in GC, we speculated that DANCR could play a significant role in tumor biology. Initially, we determined whether DANCR expression influenced tumor-like characteristics. knockdown of endogenous DANCR expression significantly suppressed GC cell proliferation and induces cell G2/M arrest, whereas ectopic expression of DANCR significantly enhanced these capacities. Moreover, increased DANCR expression significantly promoted tumor growth in nude mice.
To explore the molecular mechanism through which DANCR contributes to growth and tumorigenicity in GC, we examined gene expression changes upon DANCR overexpression and knockdown by high-throughput RNA-seq. Our data suggest that incorporating features of nitrogen compound metabolic and cell cycle process program into the DANCR expression influences growth ability. DANCR also involves in a regulatory network of stem-cell–like properties of HCC cells, 16 and metabolism at a stemness ground state is unable to fulfill the needs of differentiated progeny; conversely, metabolic metamorphosis underlies pluripotent induction during nuclear reprogramming. 21 So, DANCR may regulate the nucleic acid metabolic process to support biosynthesis of cell building blocks required for replication.22,23
In summary, according to the results of this study, DANCR is dramatically upregulated in GC tissues and the high expression of DANCR is significantly associated with worse OS in GC patients. Moreover, overexpression of DANCR promotes GC cell growth and tumorigenicity in vitro and in vivo, whereas knockdown of DANCR has the effect of suppressing GC cell growth. Thus, the expression of DANCR may be promising prognostic biomarker in GC.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
