Abstract
Long non-coding RNAs (lncRNAs) are aberrantly expressed in various cancers. Fer-1-like protein 4 (FER1L4), one of lncRNAs, plays a role as tumor suppressor in various human cancers and can be regulated by microRNA. However, the role and function of FER1L4 in human hepatocellular carcinoma (HCC) remains unknown. The aim of the present study was to annotate the role of FER1L4 and its clinical value in HCC. In the present study, we found that FER1L4 was lowly expressed in HCC tissue specimens as well as in malignant HCC cell lines, while the situation is opposite in miR-106a-5p. We found that down-regulated FER1L4 increased the expression of miR-106a-5p significantly and there was a reciprocal repression between FER1L4 and miR-106a-5p. Moreover, we identified FER1L4 as a target of miR-106a-5p by using dual-luciferase reporter assay. Knockdown of FER1L4 promoted the malignancy of HCC cells, including proliferation, migration, and invasion, and inhibited cell apoptosis. We also found that FER1L4 functions as a tumor suppressor in vivo. Together, these results suggest that FER1L4 could exert a tumor suppressive impact on HCC, which at least, in part, through suppressing miR-106a-5p expression. FER1L4, as well as miR-106a-5p, can predict the clinical prognosis of HCC alone or combined, which may be a novel therapeutic target for treating HCC.
Introduction
Human hepatocellular carcinoma (HCC) is one of the most commonly diagnosed deadly malignancies and ranks third in cancer-related death worldwide [1]. Although great advances such as liver transplantation, surgical resection, and adjuvant therapy had been achieved in HCC in the past few decades, the overall 5-year survival rate of HCC patients still remains poor [2, 3, 4]. It has been elucidated that various abnormal biological processes participating in the development of HCC, such as gene modifications [5], epigenetic alterations [6], and abnormality of protein translation [7]. Thus, for the purpose to improve the treatment and prognosis of HCC, it is of great need to identify a new potential therapeutic target underlying HCC pathogenesis and progression.
Recent publications have disclosed that long non-coding RNAs (lncRNAs), which are RNA molecules with more than 200 nucleotides (nt) and cannot be translated into proteins [8, 9]. Increasing evidence suggests that lncRNAs play important roles in tumor development and might be treated as diagnosis marker of cancers [10, 11]. LncRNAs have been verified to influence expression pattern of gene at many levels, including transcription or post-transcription processing, chromosome modification and protein regulation [12, 13]. A vast of studies have demonstrated that various lncRNAs, such as HULC, HOTAIR, ATB, HOTTIP, Dreh, URHC, UFC1, can participate in numerous biological events involved in hepatocarcinogenesis, including cell proliferation, apoptosis, and metastasis [7, 14, 15, 16]. However, up until very recently, it still remains elusive concerning the roles and function mechanisms of lncRNA in tumorigenesis [17]. It is of great value to look for novel lncRNAs as new targets for the treatment of cancer.
FER1L4 (Fer-1-like protein 4) is a newly identified tumor-related lncRNA which expressed much lower in gastric cancer patients comparing to healthy people [18, 19]. Further studies showed that lncRNA FER1L4 acts as a tumor suppressor gene by suppressing the proliferation, migration, and invasion of tumor cells [20, 21]. Tian et al. reported that lncRNA FER1L4 and miR-106a-5p functioned as competing endogenous RNA (ceRNA) that miR-106a-5p could regulate the expression of FER1L4 by targeting PTEN in human gastric cancer [20]. However, it is largely poor understood that whether FER1L4 also exhibited abnormal expression in HCC, as well as its underlying molecular mechanism.
In this study, we first detected the expression level of FER1L4 in HCC tissues and adjacent normal tissues as well as cell lines. Moreover, we investigated the role of FER1L4 in the growth, apoptosis, invasion, migration of HCC in vitro. Finally, we confirmed the role of FER1L4 in vivo.
Materials and methods
Tissue samples
Thirty-six HCC and 36 normal liver tissue specimens were gathered from patients who accepted routine surgery in our hospital. All cases were diagnosed as HCC by histologically confirmation and not subjected to preparative radiotherapy and/or chemotherapy. These tissue specimens were snapped-frozen in liquid nitrogen and further stored at
Cell lines and culture
The HepG2, Huh7, Hep3B, and HCCM3 human HCC cell lines were purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA). LO2 human normal hepatocyte was purchased from the Cell Bank of Type Culture Collection (Chinese Academy of Sciences, Shanghai, China). All the cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and 1% streptomycin/penicillin at 37
RNA isolation and qRT-PCR
By using TRIzol reagent (Takara Bio, Inc., Shiga, Japan), we isolated total cellular RNA from tissues and cells. For FER1L4, RNA was reversely transcribed into cDNA using the PrimeScript RT-PCR Kit (TaKaRa, Dalian, China) following the manufacturer’s instructions. For miR-106a-5p, RNA was reversely transcribed using M-MLV Reverse Transcriptase (Invitrogen, Carlsbad, CA, USA). The expression level of FER1L4 and miR-106a-5p was determined by qRT-PCR using a SYBR GREEN MIX kit (Promega, Madison, WI, USA) according to the manufacturer’s protocols. Primers sequences of FER1L4: forward, 5’-CCGTGTTGAGGTGCTGTTC-3’; reverse, 5’-GGCA AGTCCACTGTCAGATG-3’. Primers sequences of miR-106a-5p: forward, 5’-AAAAGTGCTTACAGTG CAGGTAG-3’ and reverse, 5’-GAAAA GTGCTTAC AG TGCAG GT-3’. Primers sequences of GAPDH: forward, 5’-AAGGTGAAGGTCGGAGTCAA-3’; reverse, 5’-AATGAAGGGGTCATTGATGG-3’. All experiments were performed in triplicate. Relative quantification of gene expression was performed by the 2
Transfection
Three sets of FER1L4 siRNA were constructed by GenePharma Co., Ltd. (Shanghai, China). Briefly, cells were seeded in a 6-well plate. When the cell amount reached 70–80% confluency, cells were transfected with FER1L4 siRNA1, siRNA2, siRNA3 using riboFECT™ CP transfection reagents (Ribobio, Guangzhou, China) according to the manufacturer’s instructions. Cells were subjected to qRT-PCR to measure the expression of FER1L4. As FER1L4 siRNA1 possessed the most effective ability in knocking down FER1L4 expression, this vector was used for all the subsequent experiments. We also designed shRNA sequence targeted FER1L4. After annealing of the complementary shRNA oligonucleotides, the annealed oligonucleotides were ligated into pENTR vector (sh-FER1L4).
Proliferation assay
To ascertain the effect of FER1L4 in regulation of HCC cells, Huh7 and HepG2 cells were seeded into 96-well plates at a density of 4
Colony-formation assays
Huh7 and HepG2 cells were seeded into 96-well plates at a density of 4
Apoptosis assay
The transfected cells were harvested and resuspended at a density of 5–10
Wound-healing assay
Transfected cells were plated onto a 6-well plate and cultured overnight with a confluence of 80–90%. Cells were scraped using a tiptop of 200 mL tip, washed by PBS, and then replaced with serum free culture medium. The migration distances of the cells were measured at 0 h and 36 h.
Cell invasion assay
The 8
Immunofluorescence analysis
HepG2 cells were seeded and cultured in an Immunofluorescence plate. After transfection with siRNA, cells were washed and fixed in 4% paraformaldehyde for 20 min. After washing with PBS, cells were fixed in 4% paraformaldehyde for 20 min and blocked with 3% BSA in PBS for 1 h at room temperature. After that, cells were incubated with the primary antibodies overnight at 4
Western blot analysis
After transfection, cells were harvested, and protein was extracted. Protein concentrations were measured using a BCA protein assay kit (Beyotime, Beijing, China). Equal amounts (30
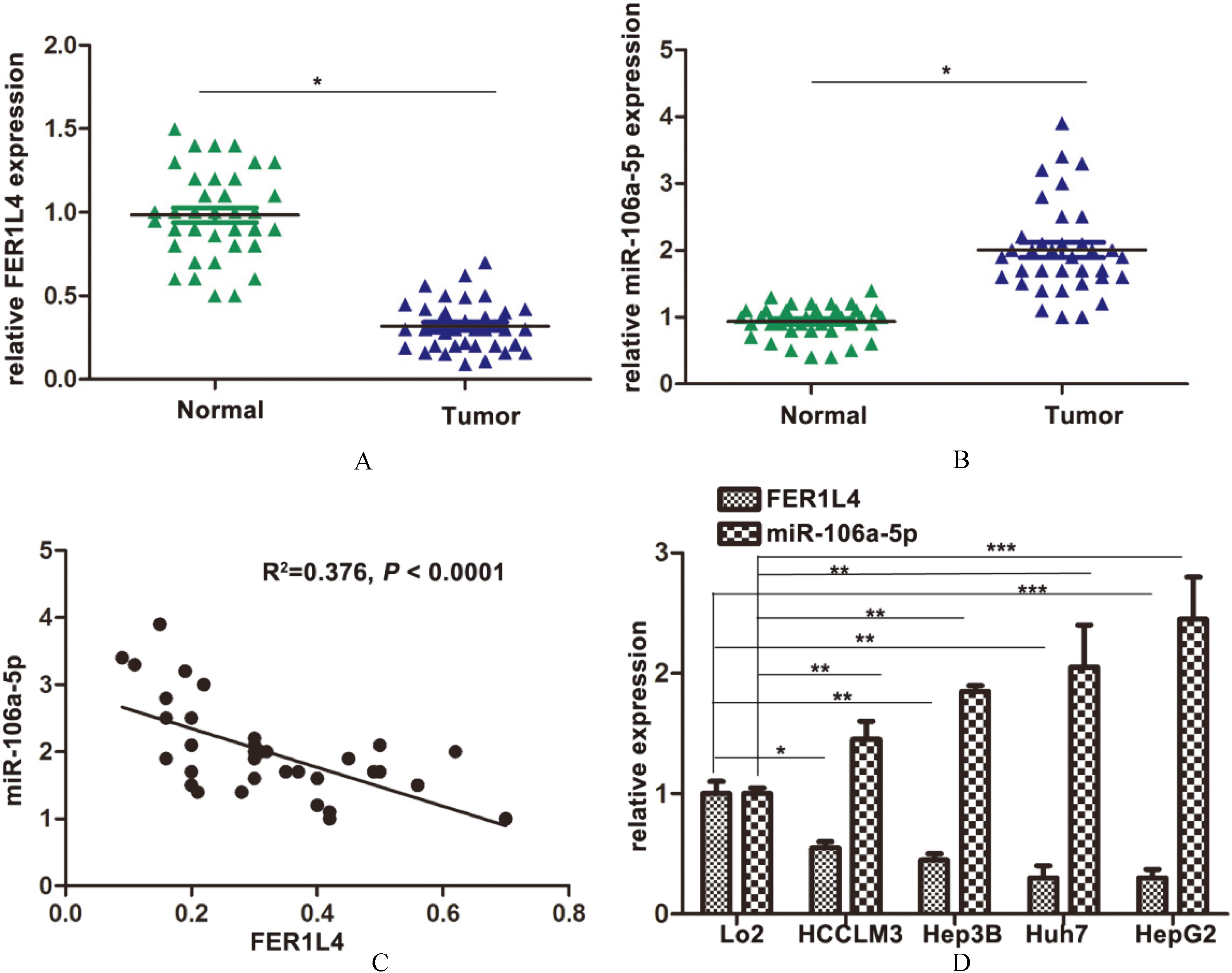
Expression of FER1L4 and miR-106a-5p in HCC tissues and cell lines. A. Relative expression of FER1L4 in HCC tissues (

Effect of FER1L4 on cell proliferation and apoptosis in HCC cells Huh7 and HepG2. A. Huh7 and HepG2 were transfected with con-siRNA or FER1L4-siRNA 1, FER1L4-siRNA 2, or FER1L4-siRNA 3 for 24 h, the expression of FER1L4 was determined by qRT-PCR. B, C. Huh7 and HepG2 were grown and transiently transfected with si-NC or si-FER1L4, then subjected to CCK-8 assay. D, E. Relative number of Huh7 and HepG2 colonies after transfected with si-NC or si-FER1L4. **

F, G. Flow cytometry apoptosis assay. Huh7 and HepG2 were grown and transiently transfected with si-NC or si-FER1L4, then subjected to flow cytometry analysis of cell apoptosis assay. **

Effects of FER1L4 on migration and invasion in Huh7 and HepG2. A, B. Detection for cell migration ability of Huh7 and HepG2 cells after knockdown of FER1L4. C, D. Transwell chamber assay was employed to examine the invasion ability of Huh7 and HepG2 cells after knockdown of FER1L4. *

Downregulation of FER1L4 suppresses tumor growth in vivo. HepG2 cells were transfected with sh-FER1L4 or control vector, then injected into left subaxillary of the mice (4/group). A. Tumor growth was measured every 3 days of injection. B. Tumors were harvested on the day 18 and weighed. *

Identification of FER1L4 as a target of miR-106a-5p. A. Expression of miR-106a-5p after FER1L4 siRNA transfection by using qRT-PCR in Huh7 and HepG2. *
miR-106a-5p expression plasmid (GV268) as well as the wild-type and mutant DNA sequences of FER1L4 were purchased from GeneChem Co., Ltd. (Shanghai, China). The wild-type and mutant DNA sequences of FER1L4 were synthesized by cloned into GV272 Firefly luciferase plasmid (GeneChem). The miR-106a-5p MRE of wild-type or mutant FER1L4 was 5’-GCACUU-3’or 5’-UACAGG-3’, respectively. Firefly luciferase plasmid and miR-106a-5p expression plasmid were cotransfected with pRL-TK Renilla luciferase vector (Promega, Madison, WI, USA) for normalization. After transfection for 48 h, Dual-Glo Luciferase Assay System (Promega) was performed to measure luciferase activity. Experiments were done independently three times.
Tumor xenografts
Female nude (BALB/c-nu) mice (4–5 weeks old) were purchased from Vital River (Beijing, China). HepG2 cells were transfected with scrambled control 200
Statistical analysis
All data were expressed as mean
Results
FER1L4 was down-regulated and miR-106a-5p was up-regulated in patients with HCC and malignant HCC cell lines
The expression of FER1L4 and miR-106a-5p was detected in tumorous tissue and adjacent normal tissue of 36 patients using qRT-PCR. The results showed that the level of FER1L4 in tumor tissues was significantly lower than that in the normal tissues (Fig. 1(A)). On the contrary, miR-106a-5p expressed more in the tumor tissues (Fig. 1(B)). A reverse relationship between FER1L4 and miR-106a-5p was also observed in HCC tissues (
Downregulation of FER1L4 promotes cell proliferation and inhibits apoptosis of HCC cells
To investigate the biological role of FER1L4 in HCC cell lines, a series of experiments were performed for detection. Firstly, Huh7 and HepG2 cells were transfected with FER1L4 siRNA. As FER1L4 siRNA 1 possessed the most potent knockdown ability of FER1L4 in Huh7 and HepG2 cells, it was chosen for the following studies. We first determined the effect of FER1L4 on HCC cell growth. CCK8 was performed after transfection of FER1L4 by FER1L4 siRNA, compared to NC-siRNA. It was found that the proliferation of Huh7 and HepG2 cell lines increased in FER1L4 transfection group compared to the growth of si-NC group (Fig. 2(B) and (C)). Besides, the colonies number, after a 2-week culture, also showed markedly increase in FER1L4 down expressing Huh7 and HepG2 cells, compared to si-NC cells (Fig. 2(D) and (E)). To elucidate the mechanism of FER1L4-mediated cell growth inhibition in HCC cells, apoptosis analysis was performed. The results showed that, when compared with the si-NC group, FER1L4 down expression inhibited the apoptosis of HCC cells (Fig. 2(F)).
Knockdown of FER1L4 promotes migration and invasion of HCC cells in vitro
We further performed the cell migration and invasion assay to determine the malignant characteristics of FER1L4. Wound healing assays indicated that the migratory potential of FER1L4-silenced cells was significantly increased when compared with control-siRNA treated HCC cells (Fig. 3(A) and (B)). Invasion assays demonstrated that FER1L4 siRNA transfected Huh7 and HepG2 cells had higher invasive capability (Fig. 3(C) and (D)). We then detected the expression of EMT related genes including E-Cadherin, ZO-1 in HepG2 cells by immunofluorescence. We found that knockdown of FER1L4 significantly down-regulated E-Cadherin and ZO-1 (Fig. 3(E)). To further confirm the role of FER1L4 in EMT of HCC cells, we detected the expression of EMT-related factors by Western blot. It was found that knocking down of FER1L4 considerably down-regulated E-Cadherin and ZO-1 (Fig. 3(F)). The above results indicated that FER1L4 played a functional role in mediating cell migration and invasion in HCC.
Knockdown of FER1L4 promotes tumorigenicity in vivo
To further investigate the oncogenic role of FER1L4, we injected HepG2 cells infected with sh-FER1L4 into the female nude mice. As shown in the result, tumors derived from FER1L4 transfected HepG2 cell grew little faster than those in the empty vector group (Fig. 4(A)). In addition, the average tumor weight was heavier in the sh-FER1L4 group compared with the control group at the end of the experiment (Fig. 4(B)). The dissected tumor tissues were further performed by qRT-PCR analysis of FER1L4 expression. The results presented that the levels of FER1L4 expression in tumor tissues formed from sh-FER1L4 cells were lower than those of the tumors formed from normal cells (Fig. 4(C)). Besides, we found that miR-106a-5p was upregulated in the sh-FER1L4 tumor tissues (Fig. 4(D)).
FER1L4 is a target of miR-106a-5p
As the result above showed that expression levels of FER1L4 and miR-106a-5p had a negative correlation, qRT-PCR was performed after transfection of FER1L4 and miR-106a-5p in Huh7 and HepG2 cell lines for a further study. Our studies showed that knockdown of FER1L4 increased miR-106a-5p levels (Fig. 5(A)). At the same time, miR-106a-5p inhibitor enhanced FER1L4 expression (Fig. 5(B)). To further investigate whether FER1L4 was a functional target of miR-106a-5p, dual-luciferase reporter assay was performed (Fig. 5(C)). Dual luciferase reporter assay showed that the mutant FER1L4 plasmid had a much higher luciferase activity than that of the wild-type plasmid (Fig. 5(D)). This indicated that the mutations introduced in the seed matches weaken the binding ability of miR-106a-5p to FER1L4. Taken these results, FER1L4 can directly regulate miR-106a-5p in HCC cells.
Discussion
In the current study, we found that FER1L4 was down regulated in tumor tissues and the HCC cell lines, while the situation is opposite for miR-106a-5p. Furthermore, FER1L4 depletion promoted cell viability, migration, and invasion in vitro. In addition, FER1L4 suppression led to the inhibition of HCC cell apoptosis. We find that FER1L4 is the direct target of miR-106a-5p. These results indicate that FER1L4 is a tumor suppressor lncRNA and that miR-106a-5p is an onco-miRNA in HCC.
As HCC is still prevailing in China, it is an urgent clinical challenge to elaborate molecular mechanisms of development of HCC, as well as to develop a new therapeutic protocol for this deadly disease [22]. In recent years, targeted cancer therapy becomes a new potential protocol. More and more studies indicated that mechanism carcinogenesis has a connection with not only protein-coding genes but non-coding genes [23]. Some long non-coding RNAs are proven to play considerable role in the development and progression of cancer [24], which may offer new opportunities to completely conquer HCC. FER1L4 is a novel long non-coding RNA which was first identified in gastric cancer, with a lower expressed pattern in gastric cancer tissues compared with matched adjacent normal tissues [19]. After that, several studies have indicated the tumor suppressor role of FER1L4 in various cancers such as colon cancer [21], endometrial carcinoma [25]. However, there are few reports regarding the relationship between FER1L4 and HCC.
In order to highlight the impact of altered expression and function of FER1L4, we showed the biological role of FER1L4 in HCC cells by applying loss-of-function approaches. Knockdown of FER1L4 promoted the HCC cell growth and led to the inhibition of cell apoptosis in vitro. Our results also indicated that FER1L4 promoted the migration and invasion, which is similar to the previous research that FER1L4 is associated with migration and invasion of cancer cells [25]. In addition, the decrease of EMT biomarkers E-cadherin and ZO-1, further illustrated that FER1L4 did involve in the regulation of cell migration and invasion.
Mounting evidence suggests that lncRNAs are indispensable in the competitive endogenous RNAs (ceRNA) regulatory networks, which act as endogenous miRNA sponges by binding to miRNAs and thus affect their function [26, 27]. That is to say, there exists a reciprocal repression in ceRNA network between lncRNA and miRNA [28, 29, 30]. It has been reported that FER1L4 is the target of miR-106a-5p, a typical onco-miRNA [20]. However, whether the relationship between FER1L4 and miR-106a-5p is similar in HCC needs further research.
In the present research, we first found that downregulation of FER1L4 was a common event in HCC tissues, and that expression of miR-106a-5p was increased in HCC tissues. Our results indicated that there was a reverse relationship between FER1L4 and miR-106a-5p in HCC clinical tissues. In addition, we also observed there exists a higher miR-106a-5p level in tumors grown from the sh-FER1L4 transfected cells. To illustrate whether FER1L4 is the target of miR-106a-5p in HCC, we knocked down FER1L4. Interestingly, we found that knockdown of FER1L4 led to an increase of miR-106a-5p. Similarly, downregulation of miR-106a-5p also increased the level of FER1L4. To assess whether FER1L4 serves as a miR-106a-5p sponge, we performed the dual-luciferase reporter assay to ascertain whether FER1L4 is the direct target of miR-106a-5p. The results showed that downregulation of FER1L4 enhanced the luciferase activities of the cells than that of the wild-type plasmid. Taken together, FER1L4 is the target of miR-106a-5p. Previous study has illustrated miR-106a-5p could regulate the expression of FER1L4 by targeting PTEN in human gastric cancer, whether Fer1L4 acts as a miRNA sponge of miR-106a-5p and thus impairs the miR-106a-5p action on PTEN still remains explored.
In summary, we have found that FER1L4 is markedly down-regulated while miR-106a-5p is upregulated in human HCC tissues and cell lines. Silencing of FER1L4 promotes cell growth, invasion and migration as well as inhibits cell apoptosis in vitro. We also found that FER1L4 acts as a tumor suppressor in vivo. FER1L4 is identified as direct target gene of miR-106a-5p. Therefore, our study highlights the significance of the microRNA-lncRNA interaction in tumorigenesis that FER1L4 suppresses malignancy of HCC mainly by inhibiting miR-106a-5p. Besides, FER1L4 may be a biomarker and therapeutic target in HCC therapy in the future.
Footnotes
Conflict of interest
None.
