Abstract
BACKGROUND:
Hepatocellular carcinoma (HCC) early diagnosis remains a challenge to date. Alpha-feto protein, though less sensitive remains widely used for both diagnosis and prognosis. Recently however, a number of molecular biomarkers have been suggested as alternatives to Alpha feto protein, especially for early diagnosis.
OBJECTIVE:
To determine the role of the long non-coding RNA, LIPCAR in the pathogenesis and early diagnosis of hepatocellular carcinoma.
METHODS:
Quantitative real-time PCR, and Fluorescence in situ hybridization assays were conducted to determine LIPCAR expression in HCC vs normal blood samples, and HCC cell lines vs normal liver cell lines. Transfection was done to upregulate LIPCAR in one HCC cell line, and used to study cell proliferation, migration, apoptosis and epithelial-mesenchymal transformation. Animal experiment was finally done to determine its effect on metastasis.
RESULTS:
LIPCAR was significantly upregulated in HCC blood samples and HCC cell lines compared to their respective normal ones. Its overexpression promoted hepatocellular carcinoma cell proliferation, and migration, while inhibiting apoptosis. Its overexpression also promoted epithelial-mesenchymal transformation in hepatocellular carcinoma cells, and metastasis in vivo.
CONCLUSION:
The study demonstrated that the lncRNA, LIPCAR is significantly upregulated in hepatocellular carcinoma patients and that its upregulation promotes HCC proliferation, migration, and metastases.
Introduction
Hepatocellular carcinoma (HCC) accounts for over 90% of all primary liver cancers and has an estimated annual mortality of 782,000 [1]. It is ranked 6th globally with East Asia and sub-Saharan Africa contributing over 50% of all incidences [2, 3]. HCC 5-year survival rate in the Chinese population is just over 12% – largely attributed to its asymptomatic onset and difficulties in early diagnosis – resulting in predominantly late-stage diagnoses and subsequent poor prognoses. Its 5-year post-operative recurrence rate is estimated at between 70%–80% [4, 5, 6, 7, 8]. Prominent risk factors for HCC include Hepatitis B and Hepatitis C virus (HCV) infection, excessive alcohol consumption, liver cirrhosis, and associated metabolic disease [9, 10].
Despite progress in diagnosis and therapy, HCC prognosis remains poor, with morbidity and mortality still unacceptably high. At present, Reliable prognosis is predicted based on invasive tumor staging and histological differentiation by microscopy. These techniques are time consuming, costly and skills intensive [11]. Efforts to innovate less invasive but reliable blood biomarkers continue to be derailed by the poor understanding of the molecular mechanisms of the disease’s development and progress. As a result, serum alpha-fetoprotein (AFP), though less sensitive and specific, remains the major biomarker for prognostic monitoring in HCC to date [12].
Recently, experiments combining two or more tumor markers such as AFP with cell free molecular markers like micro RNAs have significantly improved detection sensitivity in experimental conditions, making tumor biomarkers a hot topic in HCC research [13, 14, 15]. LncRNAs are non-protein-coding transcripts with lengths greater than 200 nucleotides that have emerged as key regulators of gene expression, and appear to play vital roles in cancers development and control [16]. Studies have shown that over expression of one particular long non-coding RNA, LIPCAR has diagnostic potentials in diseases such as myocardial infraction (STEMI) and other cardiovascular diseases [14, 15]. We hypothesized that LIPCAR could play a key role in HCC pathogenesis and progress, and so designed this study and investigated its expression level in HCC patients, correlated its expression levels to the clinical characteristics of the HCC patients, and analyzed its potential utility in HCC diagnosis and prognosis.
Materials and methods
Sample collection
We collected whole Blood samples from a total of 166 patients and 64 healthy control subjects who were attending Zhongnan Hospital of Wuhan University between October 2019 and December 2021. Ethical clearance to conduct the study was sought from Zhongnan Hospital of Wuhan University Research Ethics Committee (No. 2018063). Only patients with confirmed diagnosis of Hepatocellular Carcinoma and not yet started on preoperative chemotherapy or radiotherapy were included as cases. Control subjects were healthy patients who had come for routine physical examination at the hospital. The patients were classified into three sub-groups; HCC patients (70 in total, 47 males and 23 females, mean age 59.4
RNA extraction and reverse transcription
Total RNA was extracted from patients’ plasma, HCC cell lines and a normal liver cell line using TRIzol reagent (Invitrogen, CA, USA), and blood/liquid Total RNA Rapid Extraction kit (spin- column) (Biotech, Beijing, China) according to the manufacturers’ instructions. The reverse transcription kit ReverTra Ace
Real-time PCR analysis
Quantitative real-time PCR was performed to determine the expression level of LIPCAR using SYBR
Fluorescence in situ hybridization
To determine differential gene expression. Fluorescence in situ hybridization (FISH) was conducted on the HCC cell lines and the normal liver cell line. Briefly, the cells were fixed with 10% buffered neutral formalin, washed in PBS and dehydrated in increasing concentration of ethanol (50%, 80% and 95%). Cells were then hybridized by incubation at 550C for 15 minutes in a hybridization buffer containing a 5’ biotin-labelled probe (CTTCCAGTCGAGGATGTTTACA) against hsa-lncRNA LIPCAR, (Gema Biotech Ltd). They were then removed from incubation and rinsed in probe-free hybridization buffer preheated at 550c. Cells were then counterstained for 10 minutes in the dark using DAPI, mounted onto a coverslip and examined by fluorescence microscopy.
Cell culture and plasmid construction and transfection
Human HCC cell lines HepG2, Huh-7, Hep-3B, and normal hepatic cell line L02, were purchased from the China Center for Type Culture Collection (CCTCC, Wuhan, China) for cell culture. HCCLM3 and MHCC97L were maintained at our laboratory. All cells were cultured in Dulbecco’s modified Eagle’s medium DMEM (Gibco, USA) containing 10% fetal bovine serum (Gibco, USA) 50 U/ml penicillin, and 50 U/ml streptomycin in a humidified incubator at 37
Forward: 5’-GATTCTAGAGCTAGCGAATTCAGGCCCATTTGAGTATTTTGTTTT’-3’ And Reverse 5’GATCCTTGCGGCCGCGGATCCATGGCACATGCAGCGCAA-3’.
Finally, more copies of the reconstructed plasmids were produced and purified using the Mini Plasmid Preparation Kit (Axygen, China) according to the manufacturer’s instructions. Hep G2 Cells were cultured under normal condition in DMEM (Gibco, USA) with 10% fetal bovine serum (Gibco, USA) in a humidified incubator at 37
Cell proliferation assay
Twenty-four hours after transfection, the cells were seeded into 96-well plates (2000 cells/well) and cultured for 0 h, 24 h, 48 h, and 96 h. Then, 10
Cell apoptosis
To determine whether lncLIPCAR overexpression inhibited apoptosis, Tunel analysis was performed. In brief, after transfection, the cells were fixed with 4% paraformaldehyde and permeabilized with 0.1% Triton X-100 in 0.1% sodium citrate, then incubated with terminal deoxynucleotidyl transferase (TdT) mediated dUTP nick end labeling (TUNEL) reaction (Roche, Basel, Switzerland). After that, the cells were stained with 4’, 6-di-amidino-2-phenylindole (DAPI) (Sigma-Aldrich, St. Louis, MO, USA) and visualized. The number of TUNEL positive nuclei was then expressed as a percentage of total nuclei.
Transwell assay
To verify cell migration, Transwell assay was conducted. Following transfection and 24 h incubation in which cells were initially starved of fetal bovine serum (FBS), they were then placed into the upper chambers (20,000 cells/chamber) of Transwell assay plates (Corning, USA), and 200
Western blot assay
Total protein was extracted by a cocktail of radio immunoprecipitation assay (RIPA) buffer and PMSF (Sangon Biotech Co., Ltd.), and the concentration quantified by the bicinchoninic acid (BCA) method. Appropriate amounts of proteins were then loaded and separated by 10% SDS-PAGE gel according to a standard protocol, then transferred to PVDF membranes (Merck KGaA). Background was then blocked in 5% non-fat milk for 2 h at room temperature and rinsed with PBS. Subsequently, they were incubated in primary antibodies E-cadherin (Cat: 13-5700; dilution: 1: 4000, Invitrogen, CA, USA), N-cadherin (Cat: 33-3900, dilution: 1: 1000, Invitrogen, CA, USA), Vimentin (Cat: ab137321, dilution: 1: 2000, Abcam, Beijing China), and Claudin (Cat: ab52234, dilution: 1: 1000, Abcam, Beijing, China) overnight at 4
Wound healing assay
Cells were plated in 6-well plates. A 200-
Tumor xenograft growth and metastasis model
To investigate tumor growth in vivo, Six-week-old BALB/c nude mice were purchased from Beijing Vival River Laboratory Animal Technology Co, Ltd (Beijing, China). The mice were kept and raised at Hubei Food Drugs Security Evaluation Center and acclimatized for 2 weeks before use. They were then divided into two groups at random (
Histological examination and immunohistochemistry
The mice liver and spleen tissues were stained with hematoxylin and eosin (H&E) and Masson’s trichrome stain. After histological examination, the expression on EMT markers (E-cadherin, N-cadherin) and proliferation marker (Ki-67) were performed by immunohistochemistry. In brief, paraffin embedded tissues form the mice were first deparaffinized and rehydrated, then incubated for 30 min at room temperature in 5% normal blocking serum. They were then stained with EMT markers antibodies (E-cadherin, N-cadherin) and proliferation marker (Ki-67) overnight at 4
Statistical analyses
Statistical analyses were performed using SPSS version 21.0 (SPSS Inc., USA) and GraphPad Prism 8.0 (GraphPad Software, USA). Normally distributed data were expressed as mean
Results
LIPCAR expression is upregulated in HCC plasma and cell lines
The expression levels of Lnc RNA LIPCAR were examined using qRT-PCR in the plasma of 70 HCC patients, 96 hepatitis B & cirrhosis patients, and 64 healthy controls as well as in HCC cell lines and a normal liver cell line. The results showed that the plasma levels of LIPCAR in HCC patients were significantly higher compared to healthy subjects,
Cell lines expression was further confirmed using fluorescence in-situ hybridization (FISH). The results confirmed that LIPCAR was significantly expressed in HCC cell lines compared to the normal liver cell line, LO2,
LIPCAR expression level in plasma and cell lines (a) LIPCAR expression levels in HCC patients were significantly higher than those in hepatitis and cirrhosis, and healthy controls. One way ANOVA with LNK post-hoc test was conducted. Data were presented as mean 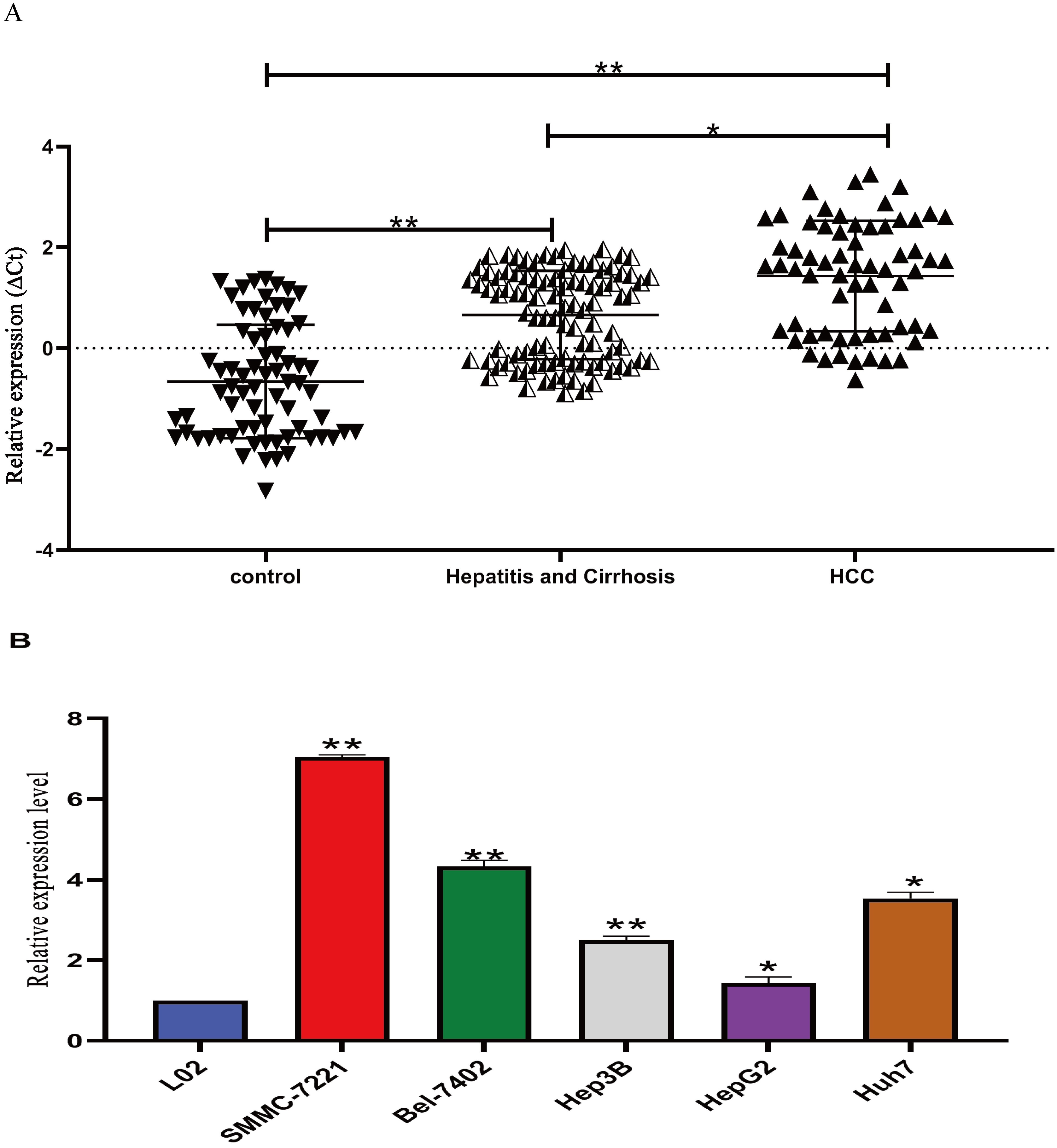
LIPCAR expression level in cell lines by Fluorescence in situ hybridization (Fish assay). LIPCAR was differentially expressed between the HCC and normal control cells. A. The fluorescence brightness of LIPCAR in HCC cell lines (Bel-7402, SMMC-7221 and hep3B) were much stronger than that of normal hepatocyte, L-02 cells, indicating upregulation in the cancer cell lines; B. The relative fluorescence intensity was significantly higher HCC cell lines (Bel-7402, SMMC-7221 and hep3B) compared to normal hepatocyte, L-02 cells. 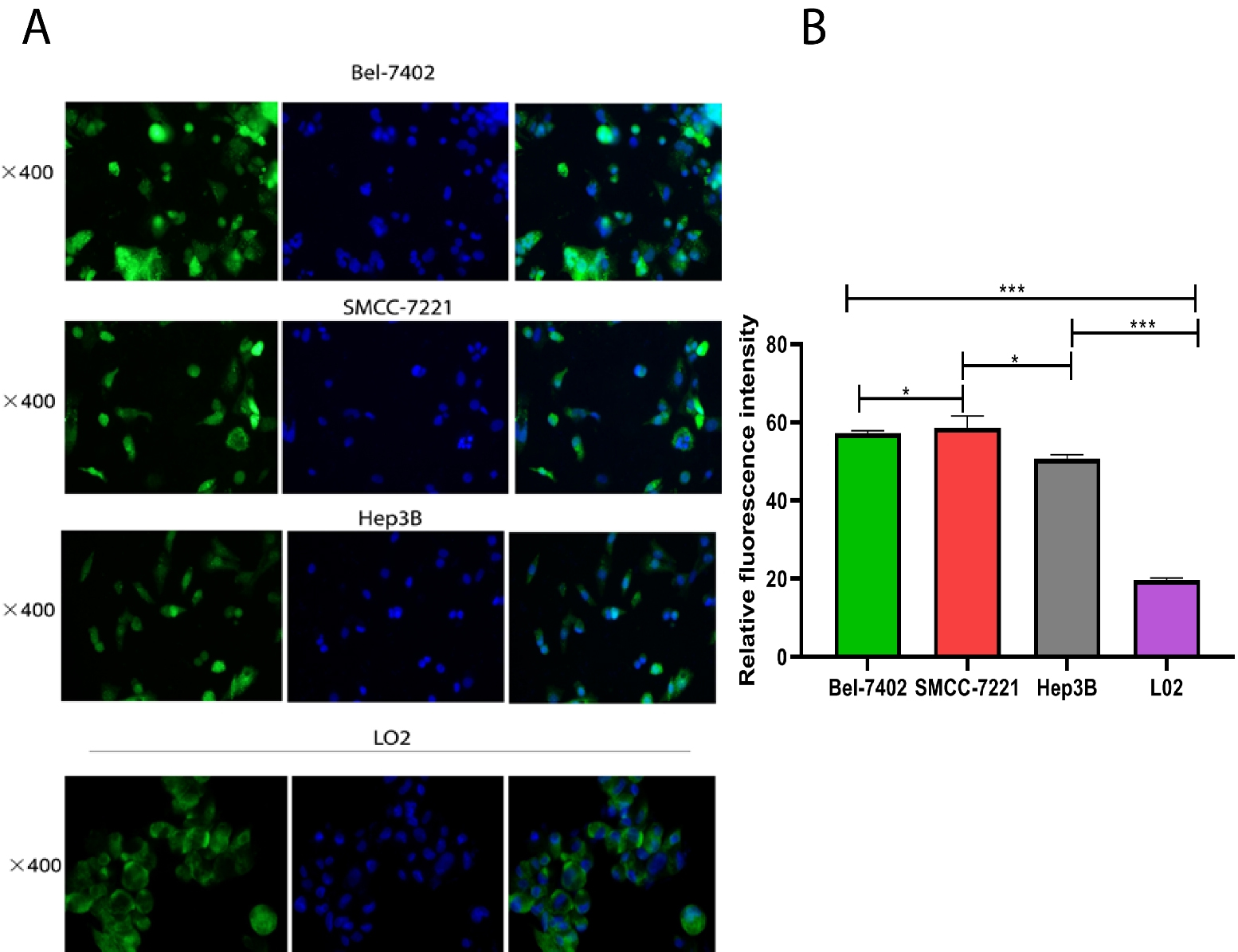
Demographic and clinical characteristics of the subjects collected included: Gender, Age, Alanine Aminotransferase (ALT), Aspartate Aminotransferase (AST),
Diagnostic value analysis
To evaluate the diagnostic value LIPCAR in plasma, ROC was performed in 4 groups: HCC vs Hepatitis B & cirrhosis, HCC (AFP
Receiver operating characteristic (ROC) curves of the Plasma LIPCAR level. A. HCC vs Hepatitis B and cirrhosis; B. Hepatitis B and Cirrhosis vs Control; C. HCC vs control and D. HCC (AFP 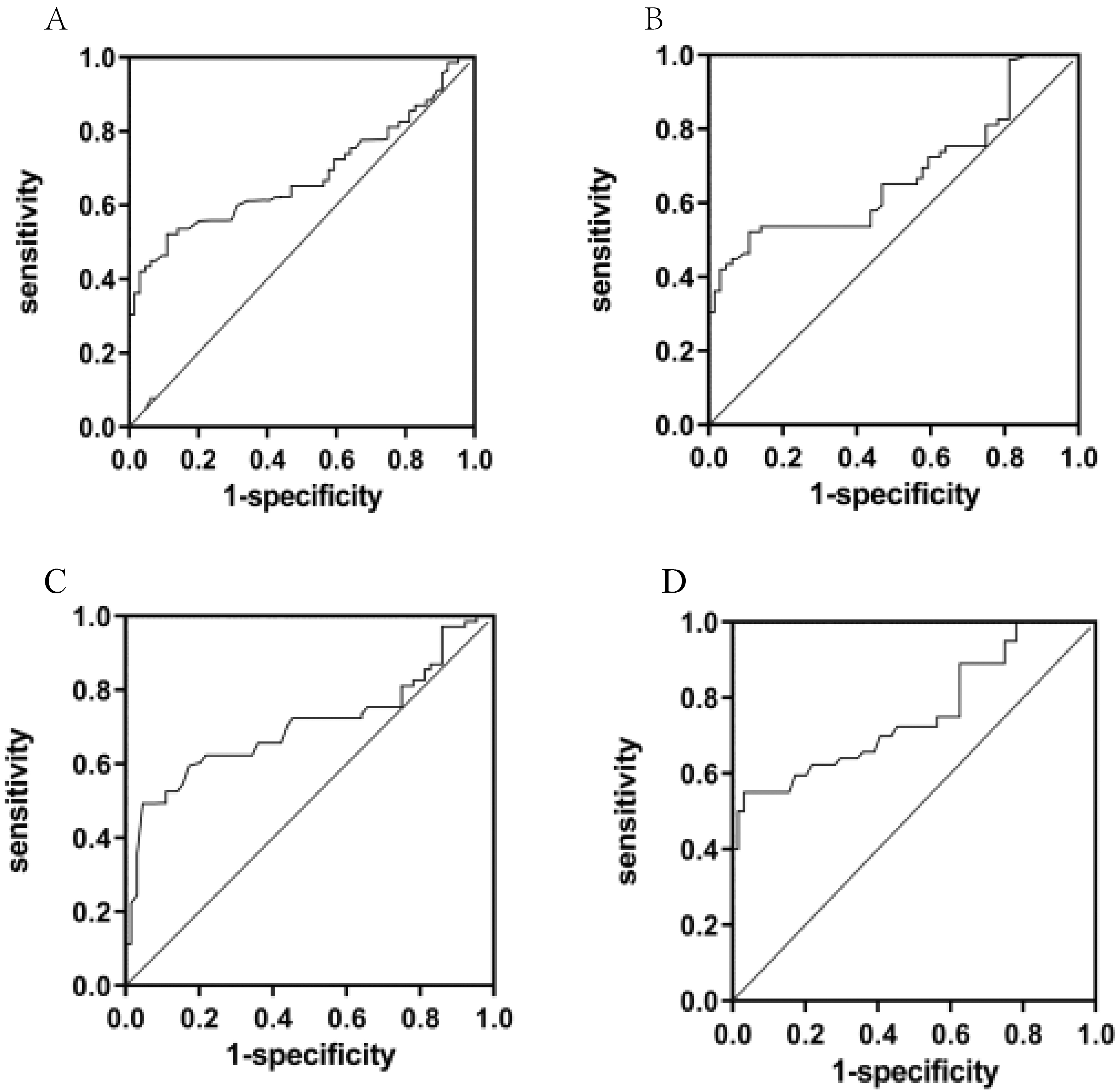
Relationship between plasma LIPCAR expression (
All data are expressed as mean
Clinically, AFP is the most frequently used biomarker for the diagnosis of HCC, but its poor sensitivity and specificity as a biomarker makes its clinical applicability very low. Therefore, when combined with plasma LIPCAR, its diagnostic efficacy for HCC detection was remarkably improved, indicating that plasma LIPCAR could serve as a diagnostic biomarker of HCC in combination with AFP (Table 2).
Diagnostic value of plasma LIPCAR in HCC
Abbreviations: HCC: Hepatocellular carcinoma;
To investigate the effect of LIPCAR in HCC pathogenesis and progression, we overexpressed LIPCAR in HepG2 cells by transfecting it with PCDH-LIPCAR and negative control plasmids. The transfection efficiency was observed by fluorescence microscopy (Fig. 4). The expression of LIPCAR was significantly increased in HepG2 cells after transfection with pCDH-LIPCAR compared to the negative controls after 24 h. Transfection efficiency was detected by RT-qPCR assay (
Upregulation of LIPCAR in transfected cells. (a) Representative images of hepG2 normal control cells (NC) as visualized by light microscope. (b) Representative images of hepG2 normal control (NC) as visualized by fluorescent microscopy. (c) Representative images of hepG2 transfected with overexpressing plasmid PCDH-LIPCAR as visualized by light microscope. (d) Representative images of HepG2 transfected with overexpressing plasmid PCDH-LIPCAR as visualized by fluorescent microscopy.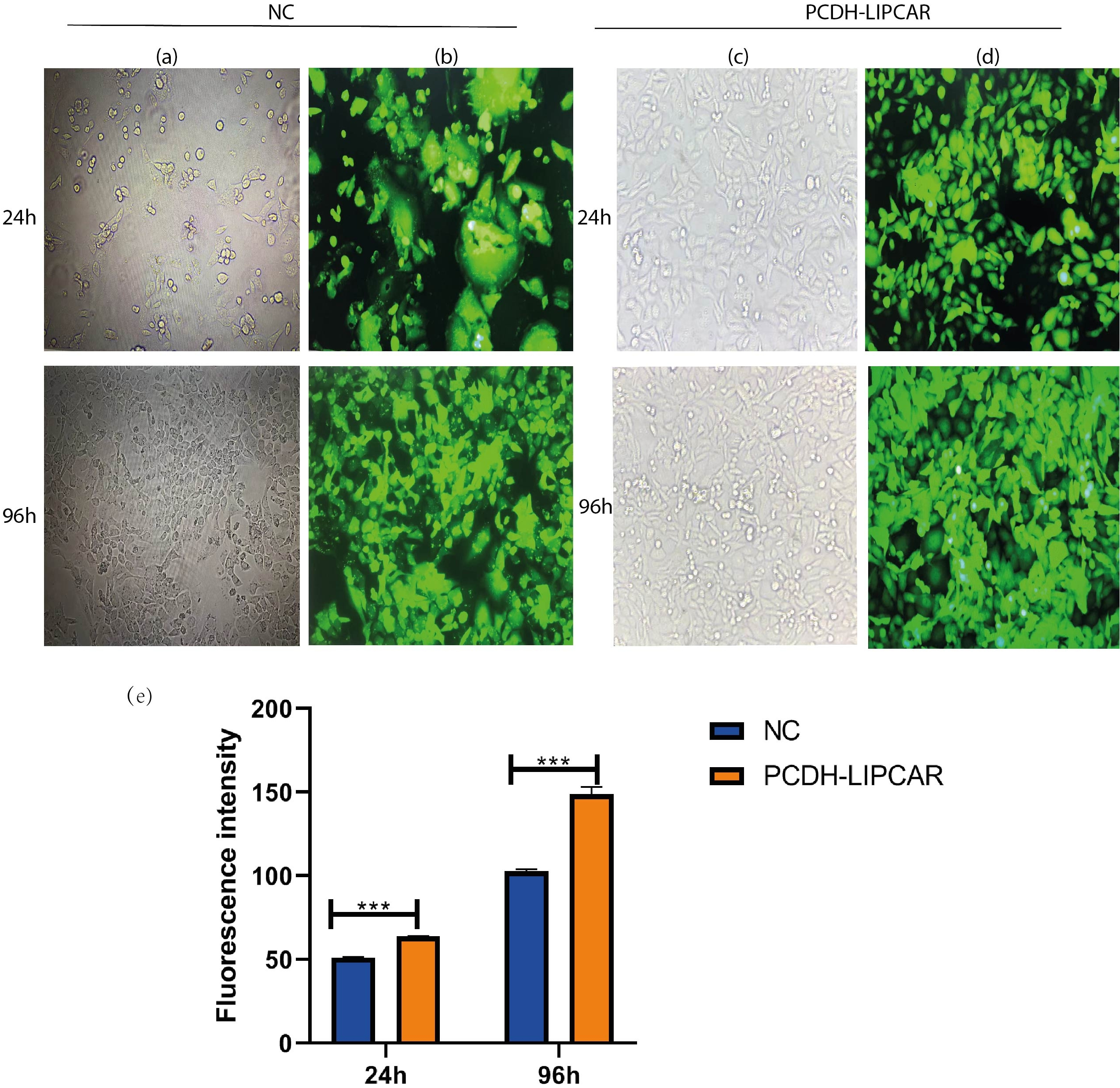
Overexpression of LIPCAR promoted cell proliferation, invasion and inhibited cell apoptosis of HCC cells in vitro. (a) Cell proliferation detected by CCK-8 assay. (b–e) Transwell and wound healing results. Overexpression of LIPCAR significantly increased the migration capacity of HepG2 cells. 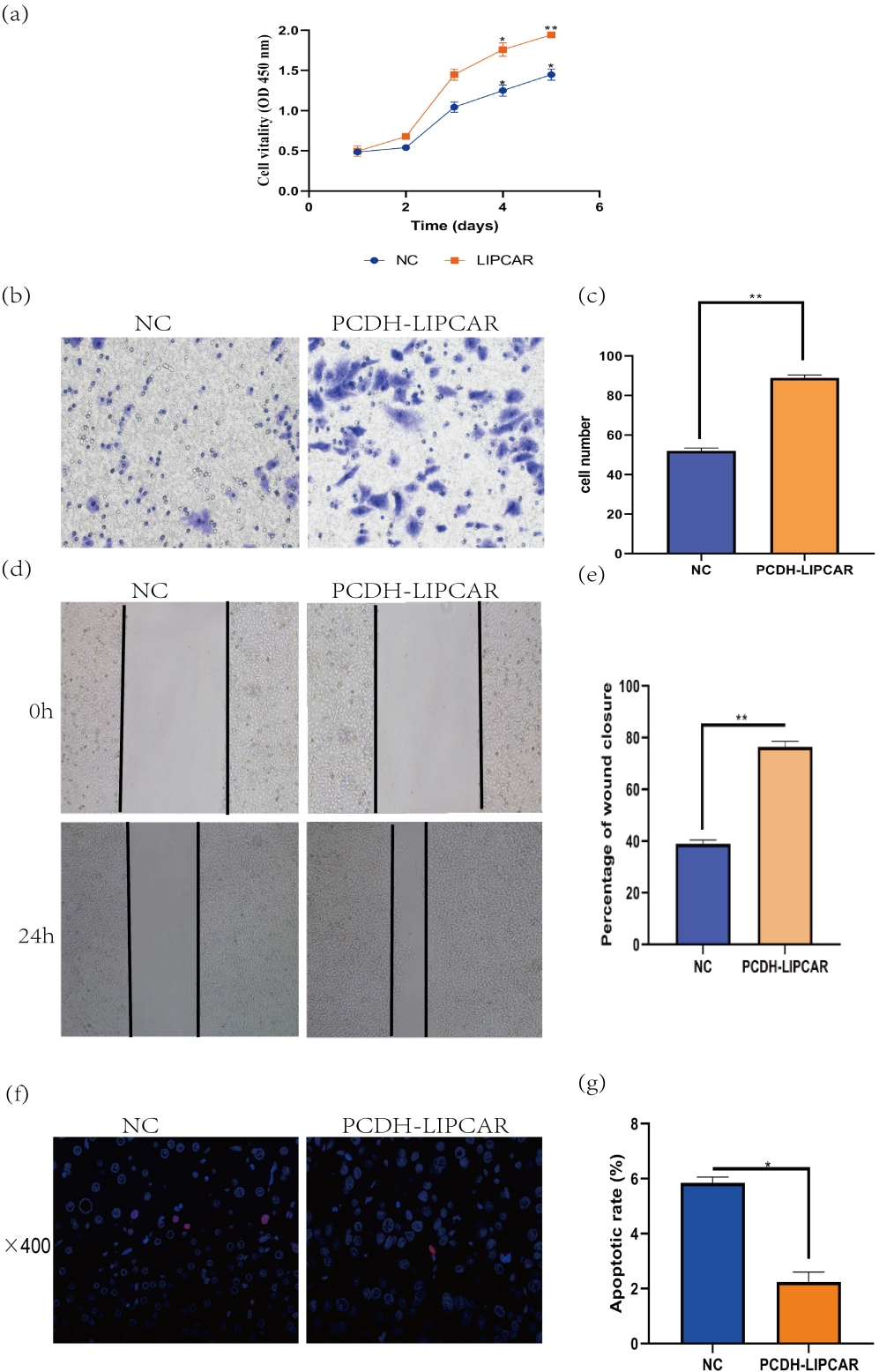
Since EMT is a crucial mechanism in tumor metastasis, we speculated that the upregulation of LIPCAR is involved in EMT. We thus determined the protein levels of several EMT markers. The results showed that the expression level of E-cadherin was reduced while levels of N-cadherin and Vimentin were increased in HepG2 cells (Fig. 6a).
Overexpression of LIPCAR promoted EMT in HCC cells: (a and b). Protein levels of E-cadherin, N-cadherin, Vimentin and Claudin detected by Western blot. Image analysis was conducted by Image J software. Independent t-test was conducted and data expressed as mean 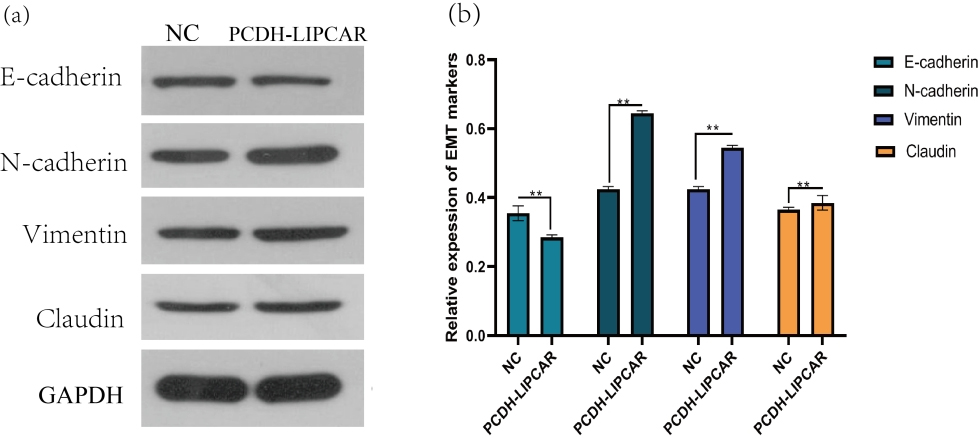
Overexpression of LIPCAR promotes HCC proliferation and metastasis in vivo. (a) Tumors isolated after 30 days. (b) Difference in tumor weight between the two groups, 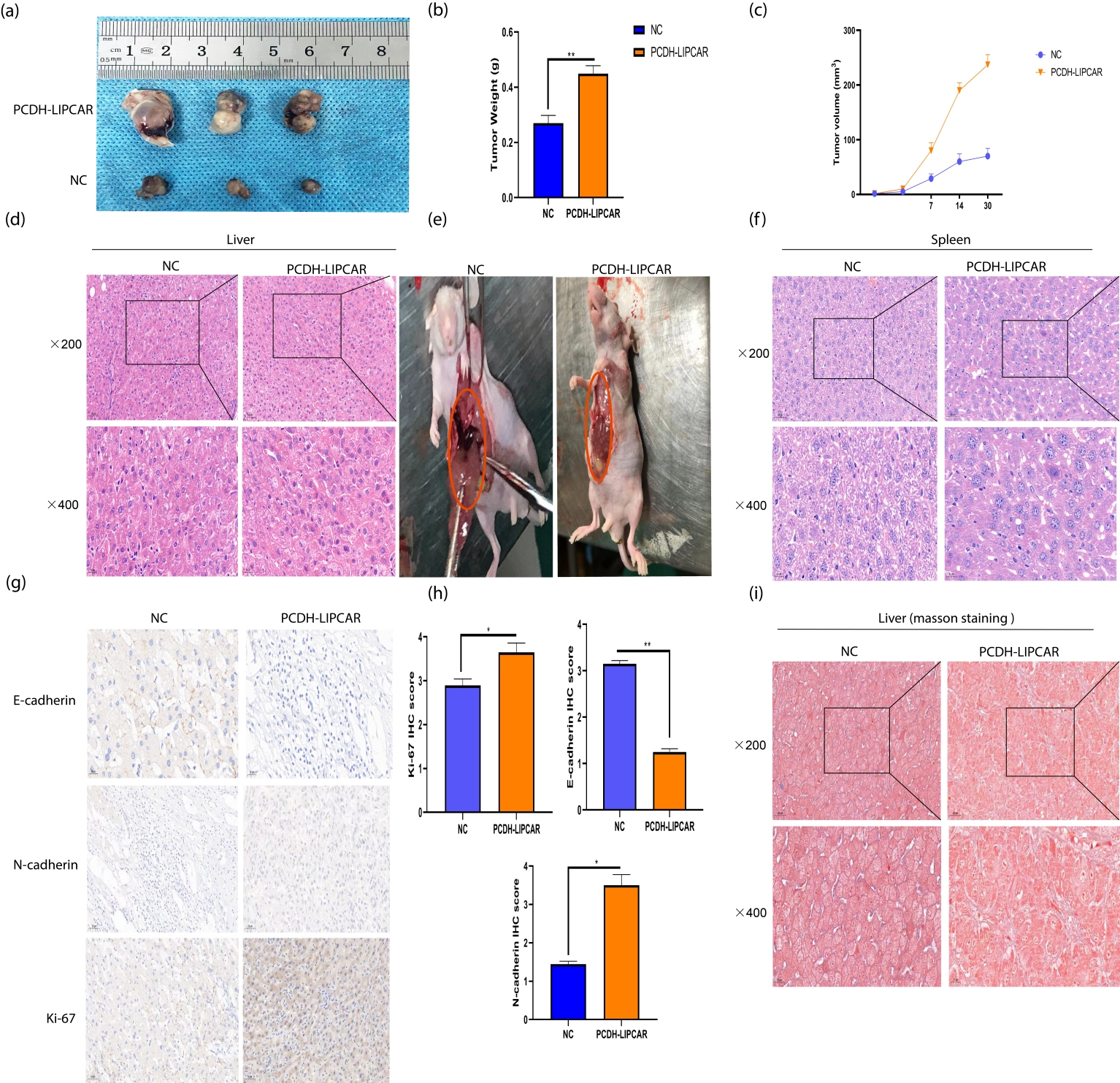
To determine the role of overexpressed LIPCAR (PCDH-LIPCAR) in HCC progression, Normal control cells, hep3B cells, and hep3B cells with LIPCAR overexpression model were injected into mice and their liver monitored. The LIPCAR overexpression model resulted into bigger HCC tumors compared to the control tumors, demonstrating that LIPCAR overexpression significantly promotes liver cancer progression in mice. Both the tumor volume and weight of the LIPCAR-overexpressing hep3B cells group significantly increased (Fig. 7a–c).
To determine the role of LIPCAR in Liver cancer metastasis in vivo, BALB/c nude mice were injected with HepG2 cells overexpressing LIPCAR. Three weeks later, the mice were sacrificed and their liver and spleen dissected to observe metastasis (Fig. 7e). The specimens were paraffin embedded, sectioned and stained with H&E and Masson’s trichrome stain (Fig. 7d, f and j). The results demonstrated that high expression of LIPCAR significantly promoted tumor metastasis. Moreover, immunohistochemistry results showed that expression levels of the pro metastasis proteins Ki-67 and N-cadherin were increased, while E-cadherin was decreased by LIPCAR overexpression. These results suggest that LIPCAR promotes tumor growth and metastasis through activating the EMT process in HCC (Fig. 7h and i).
Discussion
We conducted this study to investigate the LncRNA, LIPCAR’s expression level in the plasma of hepatocellular carcinoma (HCC) patients, correlate its expression to the clinical characteristics of the HCC patients, and determine its potential utility in HCC diagnosis and prognosis. Our results found out that LIPCAR was significantly upregulated in the plasma of HCC patients compared to hepatitis B & cirrhosis patients and healthy controls. Similarly, its expression level was significantly upregulated in different HCC-derived cell lines including HepG2, Hep3B, HCCLM3, MHCC97L and Huh-7 compared to normal hepatic cell line, L02. Furthermore, our in vitro experiments revealed that overexpression of LIPCAR by hep3B cells promoted cell proliferation, migration, and invasion indicating that LIPCAR possibly promotes HCC progression.
Our findings are in agreement with several studies that have pointed out key connections among various lncRNAs with liver cancer development and progression. For instance, Wang et al., found out that various LncRNAs promote HCC metastasis [17], Fang et al., also discovered that some lncRNAs are differentially expressed in HCC and appear to promote its development and progression [18], while Zhou et al. [19], indicated that the long noncoding RNA, ZFAS1, was significantly upregulated in both HCC tissue and cell lines, and promoted hepatocellular carcinoma proliferation.
To date, early diagnosis of HCC remains elusive, as a result most patients are diagnosed at middle or late stage when therapy is ineffective. As a possible biomarker for early diagnosis, we compared LIPCAR to AFP – the currently used biomarker for HCC diagnosis – first alone, then in combination with AFP. The results revealed a statistically significant diagnostic value for LIPCAR to distinguish HCC patients from healthy controls with AUC as high as 70.2% [(95% CI: 60.8–79.5%)]. The sensitivity was quite high at 87.4% but specificity a little low, at 52.0%. In combination with AFP, both sensitivity and specificity showed modest increments; to 88.3% and 64.0% respectively. Studies that have compared AFP with other biomarkers to improve HCC diagnosis have consistently showed that AFP has a low sensitivity but very high specificity ranging between 88.1%–93.3%, in some case up to 98% with increasing threshold of detection [20, 21, 22]. This means that LIPCAR and AFP can be used in a panel of tests complementing each other, since LIPCAR is more sensitive but less specific while AFP is vice versa. Further evaluation is required to explore this possibility.
Cognizant of the fact that E-cadherin prevents EMT while N-cadherin, Vimentin and Claudin all promote EMT, leading to cancer metastases, we evaluated the effect of LIPACR upregulation on their expression levels in vitro and correlated it with HCC metastases in the nude mice in vivo [23, 24]. The role of EMT in HCC development has gained attention in recent years with studies demonstrating that EMT enhances HCC development, while therapies targeting EMT appear to reduce or prevent metastases [25, 26, 27, 28]. Indeed, our results demonstrated that overexpression of LIPCAR significantly downregulated E-cadherin while upregulating N-cadherin and Vimentin in vitro. In-vivo experiment showed that overexpression of LIPCAR regulates EMT to promote metastasis in HCC as seen by the numerous metastatic cancer nodules in the mice liver. This result gives a strong indication that LIPCAR could be involved in EMT in HCC and thus regulates HCC progression, in agreement with Wang et al. [29] who also demonstrated that LIPCAR promotes proliferation, migration, and change in the phenotype of vascular smooth muscle cells. The exact molecular mechanism of LIPCAR overexpression in HCC however needs to be further investigated.
Conclusion
In a nutshell, our study comprehensively demonstrated that the non-coding LncRNA LIPCAR is significantly upregulated in hepatocellular carcinoma (HCC) patients and that this upregulation promotes HCC proliferation, migration, invasion and metastases. We further demonstrated that plasma LIPCAR is diagnostically useful for HCC diagnosis with diagnostic accuracy significantly improved in combination with AFP. We however could not verify the exact molecular mechanism of its overexpression in HCC. Our study thus sets the basis for further evaluation of the LncRNA LIPAR for both diagnosis and possible therapeutic use in HCC.
Footnotes
Acknowledgments
Ms. Bena Binoga: The authors acknowledge her immense support in the grammatical review of this manuscript.
Author contributions
Conception: Christian Cedric Bongolo.
Interpretation or analysis of data: Christian Cedric Bongolo and Erick Thokerunga.
Preparation of the manuscript: Christian Cedric Bongolo and Erick Thokerunga.
Revision for important intellectual content: Nyimi Bushabu Fidele, Tapara Dramani Maman Souraka, Peter Kisembo, Simon Peter Rugera, Paul F Worley, and JianCheng Tu.
Supervision: JianCheng Tu.
