Abstract
Many temporarily functioning proteins are generated during the replacement of nucleoproteins in the nuclei of late spermatids and seem to be degraded in the nucleus. This study was designed to clarify the involvement of the ubiquitin-proteasome degradation system in the nucleus of rat developing spermatids. Thus, we studied the nuclear distribution of polyubiquitinated proteins (pUP) and proteasome in spermiogenic cells and sperm using postembedding immunoelectron microscopy. We divided the nuclear area of late spermatids into two regions: (1) a dense area composed of condensed chromatin and (2) a nuclear pocket in the neck region. The latter was located in the caudal nuclear region and was surrounded by redundant nuclear envelope. We demonstrated the presence of pUP in the dense area and nuclear pocket, proteasome in the nuclear pocket, and clear spots in the dense area of rat spermatids. Using quantitative analysis of immunogold labeling, we found that fluctuation of pUP and proteasome levels in late spermatogenesis was mostly synchronized with disappearance of histones and transitional proteins reported previously. In the nuclei of human sperm, pUP was detected in the dense area, whereas proteasome was in the nuclear vacuoles and clear spots. These results strongly suggest that pUP occur in the dense nuclear area of developing spermatids and that the ubiquitin-proteasome system is more actively operational in the nuclear pocket than dense area. Thus, the nuclear pocket might be the degradation site for temporarily functioning proteins generating during condensation of chromatin in late spermatids.
Keywords
Two cellular protein degradation systems have been described: the lysosomal system and ubiquitin-proteasome system. It is likely that the latter is involved in the degradation of nucleoproteins because degradation of nuclear proteins might occur within the nuclei of spermatids. In the ubiquitin-proteasome system, the target proteins are conjugated to ubiquitin, which is a highly conserved 76-amino acid polypeptide. Ubiquitin is covalently attached to the target protein by an isopeptide bond between the C-terminal glycine of ubiquitin and the ε-amino group of a lysine in substrate protein (Chau et al. 1989; Ciechanover et al. 2000). The ubiquitination process involves activities of three enzymes: activating enzyme (E1), conjugating enzyme (E2), and protein ligase (E3) (Haas and Rose 1982; Hershko et al. 1983; Ciechanover 1994; Hershko and Ciechanover 1998). A successive ubiquitin molecule is usually added to the lysine residues of the previous ubiquitin to produce a polyubiquitin chain. Polyubiquitinated proteins (pUP) are recognized and degraded by the 26S proteasome (Hough et al. 1987; Ganoth et al. 1988; Tanaka 1998). The 26S proteasome is a giant multimeric protease present innuclear and cytosolic compartments and consists of a 20S core complex and 19S regulatory complex. In the present study we used antibodies against pUP and proteasome subunits to search for degradation sites in the nuclei of spermatids.
The nucleus of rat spermatids contains a non-condensed nuclear area in the caudal region. This area appears in step 9 and disappears in step 19, characteristically surrounded by a redundant nuclear envelope and is called “nuclear pocket” (Lalli and Clermont 1981; Clermont et al. 1993). This nuclear area has been reported in spermatids and sperm of many mammals (Franklin 1968; Fawcett and Phillips 1969; Zamboni and Stefanini 1970). However, the physiological function of the nuclear pocket is currently not clear. In the present study we divided the nuclear area into two regions: the dense area composed of condensed chromatin and the nuclear pocket to compare the localization of ubiquitin-proteasome system-related proteins in these two nuclear areas. Although the nuclear pocket is hardly observed in the rat epididymal sperm, it is found in human ejaculated sperm (Westbrook et al. 2001). Furthermore, the human sperm nucleus contains small clear spots and large electron-less dense areas known as nuclear vacuoles (Zamboni 1987; Francavilla et al. 2001). Westbrook et al. (2001) suggested that the vacuoles are in continuity with the nuclear pocket.
The aim of the present study was to map the nucleoprotein degradation sites within the nuclei of spermatids and sperm and, hence, determine the involvement of the ubiquitin-proteasome degradation system in the nucleus of rat developing spermatids. Toward this goal, we localized two proteins, pUP and proteasome, involved with the ubiquitin-proteasome degradation system, in the nuclei of rat spermatids and sperm. Moreover, considering the continuity of the nuclear vacuoles and clear spots to the nuclear pocket, we also localized these proteins in nuclei of human ejaculated sperm. Results showed the presence of pUP in the dense area and nuclear pocket and proteasome in the nuclear pocket and clear spots of rat spermatids. In the nuclei of human sperm, pUP was detected in the dense area, whereas proteasome was in the nuclear vacuoles and clear spots. These results strongly suggested that the clear spots and nuclear pocket of rat spermatids, as well as the nuclear vacuoles of human sperm, are degradation sites for nucleoproteins in the nucleus.
Materials and Methods
Animal
Twenty four 9-week-old male Wistar albino rats weighing 200–250 g were fed on appropriate standard diets and water ad libitum, until used. Five-week-old ICR mice were used. Human ejaculated sperm was obtained from two volunteers. All experiments were performed in accordance with the rules and regulations of the Committee for Research at the University of Yamanashi.
Antibodies
Mouse monoclonal antibody (MAb, clone GC3α) against gold-fish ovary proteasomes was described previously (Tokumoto et al. 2000). Mouse MAb (IgM) to polyubiquitinated proteins (clone FK1) was from Affinity Research Products (Exeter, UK). Rabbit anti-mouse IgM and horseradish peroxidase (HRP)-labeled rabbit anti-mouse IgG were from Dako (Tokyo, Japan). Alexa Fluor 568-labeled goat anti-rabbit IgG and Alexa Fluor 568-labeled goat anti-mouse IgG were from Molecular Probes (Eugene, OR). Mouse serum was prepared from the blood of ICR mouse.
Preparation of Heads of Rat Epididymal Sperm
Rat caudal epididymides were dissected out and teased in ice-cold 50 mM Tris-HCl-buffered saline (TBS) containing protease inhibitors (1 mM PMSF, 2 μg/ml leupeptin, 10 μg/ml pepstatin, 5 μg/ml antipain, and 10 μg/ml aprotinin) (all from Sigma-Aldrich; St Louis, MO) and 10 mM iodoacetamide (Sigma-Aldrich), an inhibitor of protein deubiquitination (Matsui et al. 1982; Baarends et al. 1999). Sperm suspension was filtered through 100-μm nylon mesh and centrifuged at 3600 × g at 4C. The pellet was suspended in a small volume of TBS. Sperm heads were isolated by the method of Calvin (1976). Briefly, the sperm suspension was sonicated to cleave heads from the tails. The resulting suspension was mixed with 2.2 M sucrose to yield a final sucrose concentration of 1.8 M, loadedon discontinuous sucrose gradient (2.2 M and 2.05 M), and centrifuged at 91,000 × g for 60 min with a Beckman Optima XL 80K ultracentrifuge (Beckman-Coulter; Fullerton, CA) equipped with a swinging bucket rotor (SW41 Ti) at 4C. Resulting pellets containing almost pure heads were suspended in a small volume of TBS and stored at −70C until use.
Isolation of Nuclei From Homogenate of Rat Testis
The method described by Tata (1974) was used. Rat testes were chopped in 0.32 M sucrose-3 mM MgCl2 and homogenized in a Potter-Elvehjem homogenizer with a Teflon pestle. Homogenates were filtered through two layers of nylon bolting cloth (100-μm nylon mesh), diluted with 0.6 volumes of 0.32 M sucrose-3 mM MgCl2 and 0.22 volumes of distilled water, and centrifuged at 700 × g for 10 min. Pellets were suspended in 2.4 M sucrose-1 mM MgCl2 and centrifuged at 50,000 × g for 1 hr with a Beckman Optima XL 80K ultracentrifuge equipped with a swinging bucket rotor (SW41 Ti) at 4C. Nuclei were collected into pellets, suspended in a small amount of 0.25 Msucrose-1 mM MgCl2, and stored at −70C until use.
Isolation of 26S Proteasomes From Testes and Liver of Rat
Proteasomes were partially purified from testes and liver of rats by the method of Tanaka et al. (2006). Briefly, rat testes were homogenized in 25 mM Tris-HCl buffer (pH 7.5) containing 1 mM dithiothreitol (DTT) (Merck; Darmstadt, Germany), 2 mM ATP (Sigma; Tokyo, Japan), and 0.25 M sucrose. Homogenates were centrifuged at 70,000 × g for 1 hr with a Beckman Optima XL ultracentrifuge equipped with an angle rotor (Type 70.1 Ti). The resulting supernatants were recentrifuged at 70,000 × g for 5 hr. Precipitates were dissolved in 400 μl of 25 mM Tris-HCl (pH 7.5) containing 1 mM DDT, 20% glycerol, and 0.5 mM ATP and centrifuged at 20,000 × g for 30 min to remove insoluble material. The resulting supernatants (partially purified proteasomes) were stored at −70C.
Western Blot Analysis
pUP and proteasome subunits in rat testicular nuclei and sperm heads were analyzed by Western blotting. Protein concentration was determined using the bicinchoninic acid method (Pierce Biotechnology; Rockford, IL) with BSA as the protein standard. Protein concentration in the samples was adjusted to 10 mg/ml. Samples were mixed with an equal volume of sample buffer for SDS-PAGE. SDS-PAGE was performed as described by Laemmli (1979), using 40 μg of protein (12.5% gel). Proteins were transferred to a PVDF membrane. The membrane was soaked in PBS containing 5% (w/v) skim milk and incubated overnight with the primary antibody, followed by 1-hr incubation with the secondary antibody conjugated with HRP. Signals were visualized by DAB reaction.
Immunofluorescence and Immunoperoxidase Staining of Semithin Sections
Rat testis was fixed in a fixative consisting of 4% paraformaldehyde, 1% glutaraldehyde, and 0.15 M Hepes-KOH (pH 7.4) for 1 hr at 4C and cut into small tissue blocks. After washing in PBS, tissue blocks were dehydrated and embedded in Epon. Semithin sections (250-nm thick) were cut with a histodiamond knife (Diatome; Biel, Switzerland) and mounted on clean glass slides. After removal of epoxy resin (Litwin et al. 1984), sections were treated with trypsin (0.5 mg/ml) for 1–3 min and incubated with primary antibodies overnight. After washing in PBS, sections were incubated with HRP-labeled secondary antibodies corresponding to the primary antibodies for 30 min. HRP activity was stained with DAB reaction. For immunohistological control, sections were incubated with mouse serum instead of the primary antibody, followed by incubation with HRP-labeled secondary antibodies. For immunofluorescence staining of pUP, semithin sections were stained in three steps: first with anti-pUP (mouse IgM), second with rabbit anti-mouse IgM, and third with Alexa Fluor 568-labeled goat anti-rabbit IgG. For immunohistochemical controls, mouse serum was used instead of the primary antibody. Sections were examined with a Nikon Eclipse E600 fluorescence microscope (Nikon; Tokyo, Japan).
Immunofluorescence Staining of Sperm Smear Preparations
Rat epididymal sperm and human ejaculated sperm were smeared on clean coverglasses, air dried, and stored at −20C until used. Some smear preparations were fixed in a solution containing three parts methanol and one part acetic acid for 10 min at room temperature and treated with 30 mM DTT for 1 hr at room temperature. Other smear preparations were fixed in 0.1% glutaraldehyde and 0.1 M Hepes-KOH (pH 7.4) for 30 min at room temperature. After washing with 0.1 M Tris-HCl (pH 7.4), the preparations were treated with 0.05% SDS for 30 min (Robinson and Vandré 1997). Afterwards, smear preparations were immunostained and examined as described above.
Postembedding Immunoelectron Microscopy
Rat testes were sliced in ice-cold fixative consisting of 4% paraformaldehyde, 0.25% glutaraldehyde, and 0.15 M Hepes-KOH buffer (pH 7.4); fixed for 1 hr at 4C; and cut into small tissue blocks. Tissue blocks were dehydrated in graded ethanol series and embedded in LR White resin at −20C. Human ejaculated sperm from two volunteers was prepared in the same manner. Thin sections were cut with a diamond knife and mounted on nickel grids. Sections were then incubated overnight with the primary antibodies at 4C. Reacted antibodies were visualized by protein A-gold (15 nm in diameter). For immunocytochemical control experiments, sections were incubated with mouse serum instead of the primary antibody and then with rabbit anti-mouse IgM or IgG, followed by incubation with the protein A-gold probe. All sections were contrasted with 2% uranyl acetate for 5–10 min and lead citrate for 30 sec. Sections were examined with a Hitachi H7000 electron microscope (Hitachi; Tokyo, Japan) at an accelerating voltage of 80 kV.
(
Quantitative Analysis of Gold Labeling
For quantitative analysis, 10 electron micrographs for each selected step of spermatogenic cells were taken and enlarged to ×30,000. Each step was identified by the morphological criteria described by Russell et al. (1990). The nuclear area of late spermatids was divided into two: the dense area and nuclear pocket located in the neck region and surrounded by redundant nuclear envelope (Lalli and Clermont 1981). The nuclear areas of spermatogonia and pachytene spermatocytes, dense area, and nuclear pocket of late spermatids were estimated by a digitizer equipped with a computer using mor-phometry software (SigmaScan; Jandel Scientific, San Rafael, CA). Gold particles in each compartment were counted and the labeling density (gold particle number/1 μm2) was calculated. Background labeling was determined to measure the area without tissue and counting of gold particles in this area. Labeling density values of each compartment reported here represent values after subtraction of the background labeling density.
Results
Western Blot Analysis of Ubiquitin-Proteasome System-related Proteins in Rat Testicular Nuclei and Sperm Heads
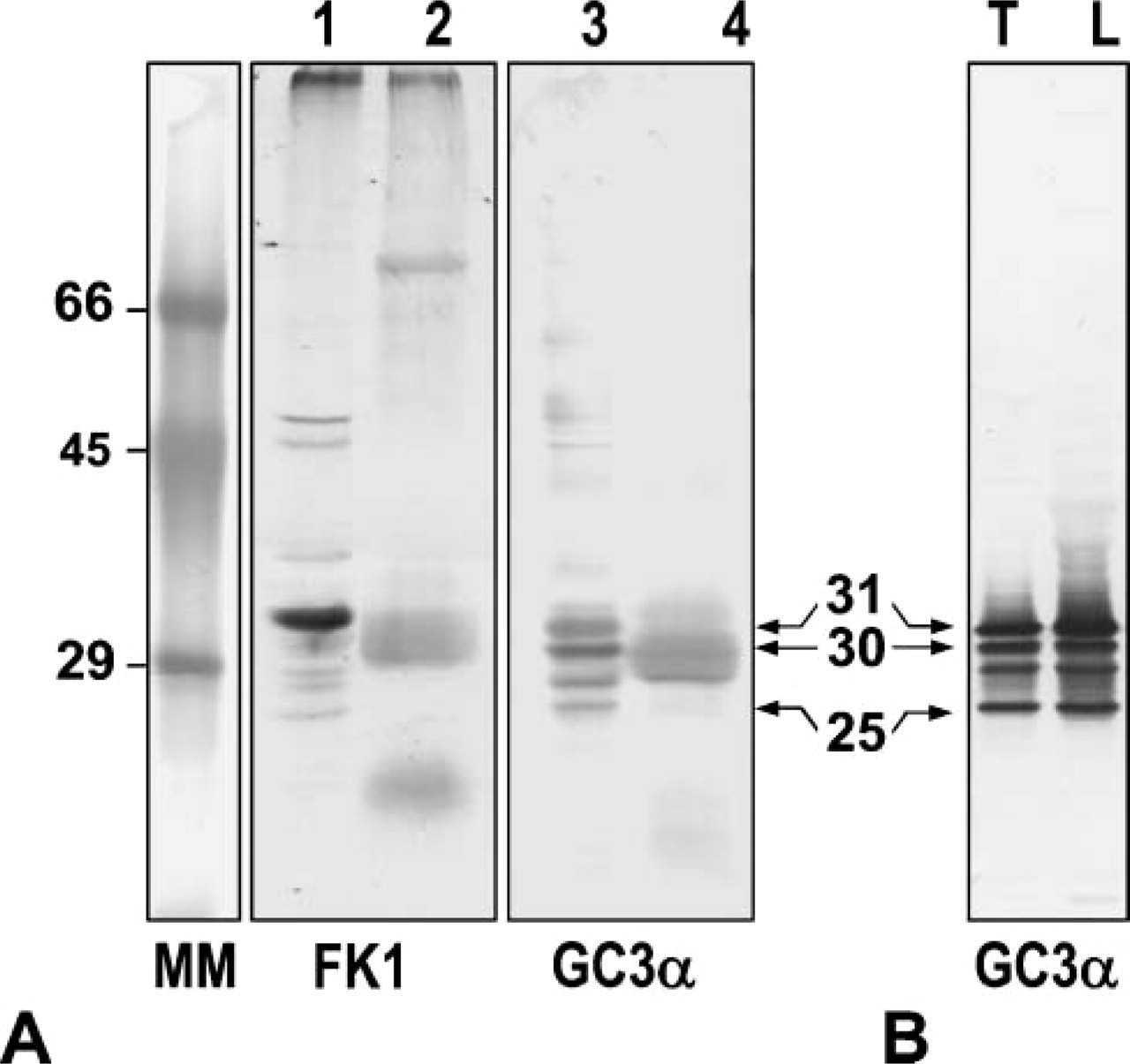
Mouse MAb (FK1) specific for polyubiquitinated proteins developed many bands of various molecular masses in the nuclear fraction of rat testis and epididymal sperm heads (Figure 1A), showing that many polyubiquinated proteins with relatively low molecular masses were contained in testis nuclear fraction and sperm heads. Mouse MAb (GC3α) to proteasome subunits detected three major bands with molecular masses of 31, 30, and 25 kDa in the nuclear fraction (Figure 1A). In sperm head extract, the 31-kDa subunit was not observed but two other subunits were recognized (Figure 1A). These three major bands were commonly observedin proteasomes partially purified from rat testes and liver (Figure 1B), showing the reactivity and specificity of the antibody (GC3α) for the subunits of 20S proteasomes.
Immunohistochemical staining of rat testis. (
Immunofluorescence and Immunoperoxidase Staining for pUP and Proteasome Subunits in Rat Testis
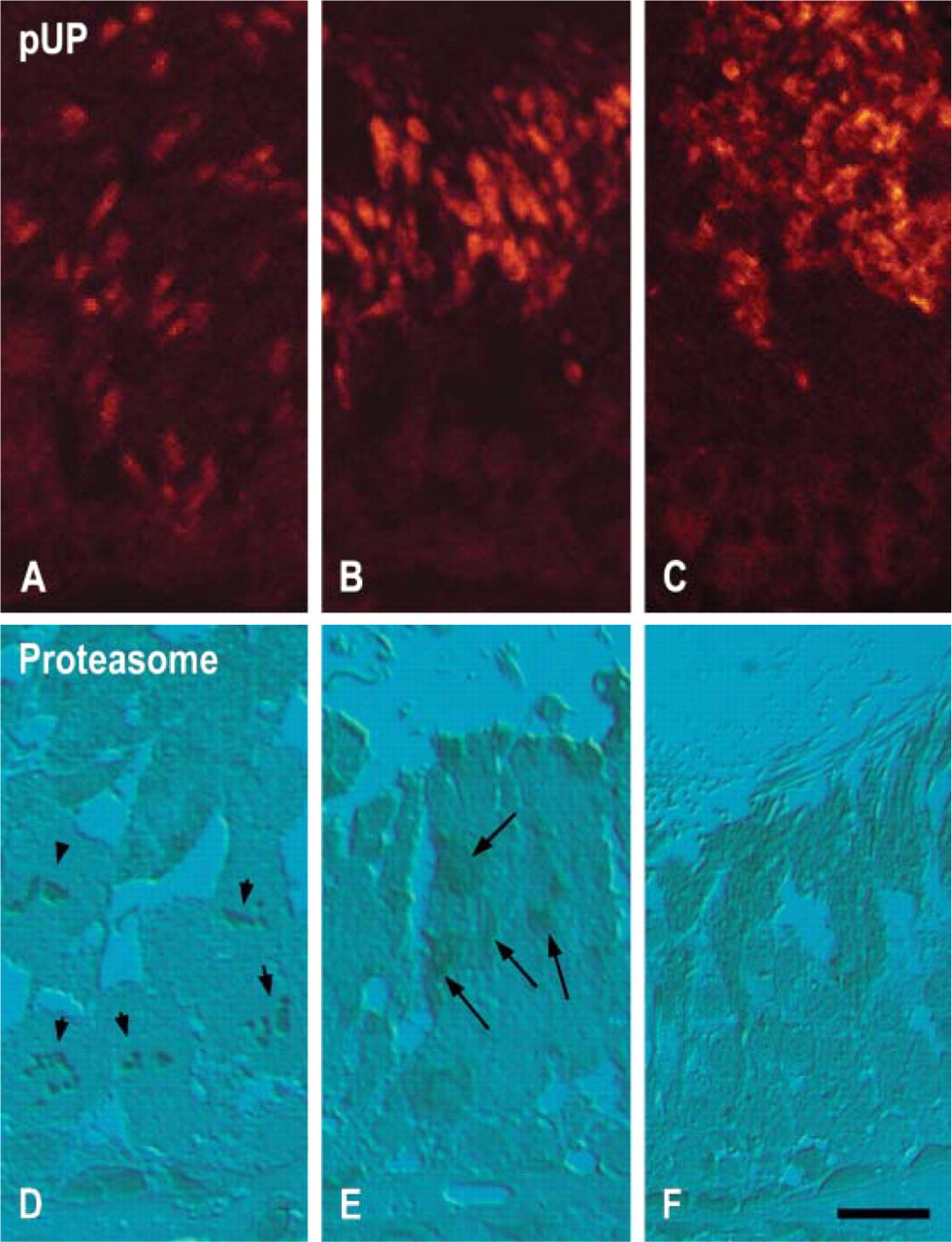
Immunofluorescence staining for pUP identified nuclei of spermatids at steps 10–19 (Figures 2A-2C). In step 10 spermatids, nuclei were weakly and partially stained for pUP (Figure 2A). In step 15 spermatids, nuclei were strongly stained for pUP (Figure 2B). In stage VII, nuclei of step 19 spermatids and residual bodies were heavily stained for pUP (Figure 2C). Proteasome was stained in the nuclei of spermatogonia, spermatocytes, and elongating spermatids (Figures 2D-2F). In addition, stained chromosomes of dividing spermatocytes were also noted (Figure 2D). Residual bodies and flagella were also stained for proteasomes (Figure 2F). No staining was noted in sections treated with only the secondary antibodies.
Immunofluorescence Staining for pUP and Proteasome Subunits in Rat Epididymal Sperm and Human Sperm
Diffuse staining was observed for pUP (Figure 3A). Staining of proteasome was evident in the small dots scattered throughout the nucleus of rat sperm (Figure 3B). The nucleus of human sperm was also diffusely stained for pUP (Figure 3C). The proteasome was stained in dots of various sizes (Figure 3D). No staining was noted in sections treated only with the secondary antibodies.
Immunoelectron Microscopic Localization of pUP and Proteasome in Rat Developing Spermatids
Labeling for pUP and proteasome was detected in numerous sites of the spermiogenic cells at various labeling intensities. In this study we focused on the localization of these proteins in the nuclei of spermatids. Positive labeling was completely abolished in immunocytochemical control sections.
Round Spermatids (Steps 1–8 Spermatids) and Elongating Spermatids (Steps 9–12). The nuclei of spermatids from step 1 to step 12 stained weakly for pUP and proteasome. Labeling for pUP started to increase at steps 10–12 spermatids.
Condensing Spermatids (Steps 13–15). In step 13 spermatids, gold labeling for pUP was observed in the electron-dense area, which showed progressive chromatin condensation (Figure 4A, arrows). In addition, the labeling was associated with a rough network that appeared in the spermatids of this step. Labeling for pUP was identified in the nuclear pocket (data not shown). No gold labeling was observed in the cytochemical control sections treated with a combination of mouse serum, rabbit anti-mouse IgM, and protein A-gold probe (Figure 4B). Gold particles for proteasome were observed in the margins of electron-dense areas where the condensation of chromatin started (Figure 4C). Proteasome signals were also detected in the nuclear pocket (Figure 4C, surrounded by dotted line). In immunocytochemical control sections treated with a combination of mouse serum/rabbit anti-mouse IgG-protein A-gold probe, no gold labeling was noted (Figure 4D). In step 15 spermatids, heavy labeling for pUP was observed in the dense area as well as in the nuclear pocket (Figure 4E). In the dense area, most of the gold labeling was detected in the dense matrix but not in the clear spots (Figure 4E, arrows). In control sections, almost no gold particles were observed (Figure 4F). Labeling for proteasome was detected in both areas (Figure 4G). In the dense area, approximately half of the gold labeling was associated with clear spots (Figure 4G, arrows). Labeling for proteasome in the nuclear pocket was stronger than in the dense area. No gold signals were observed in control sections (Figure 4H).
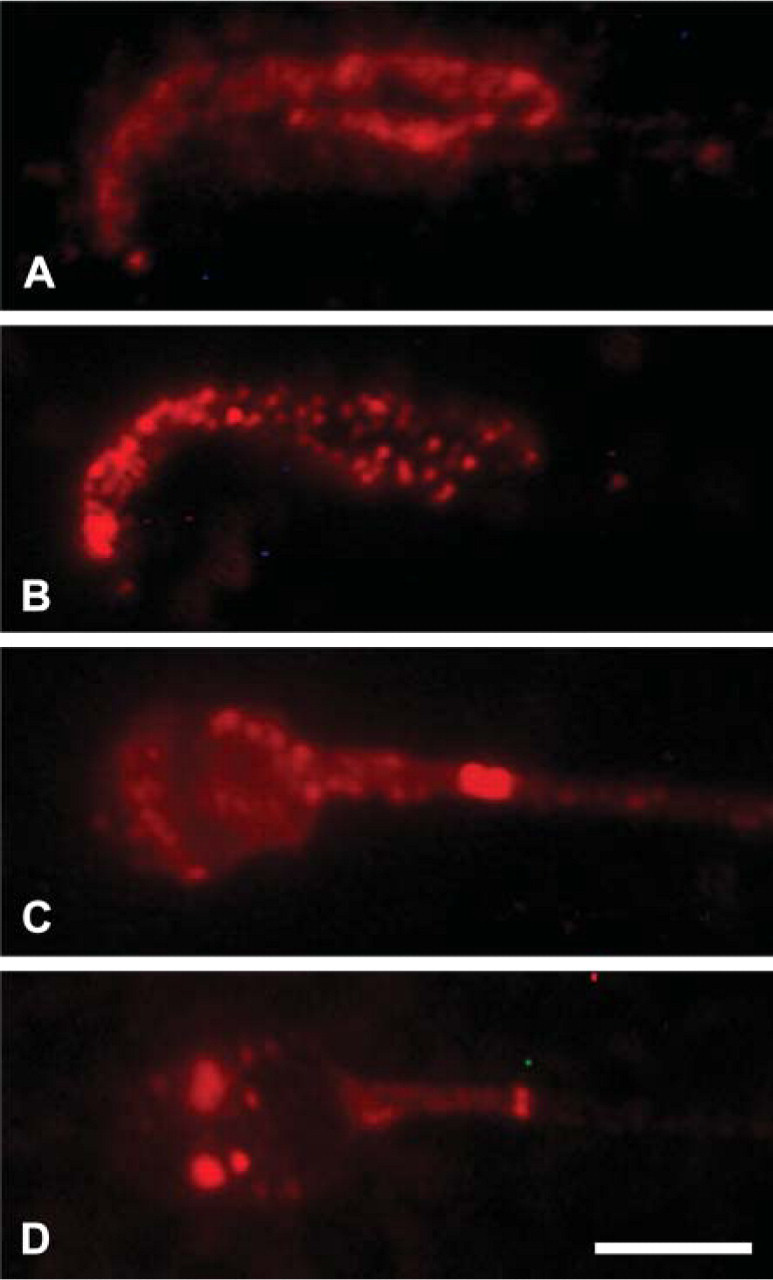
Immunofluorescence staining of rat epididymal sperm and human ejaculated sperm. (
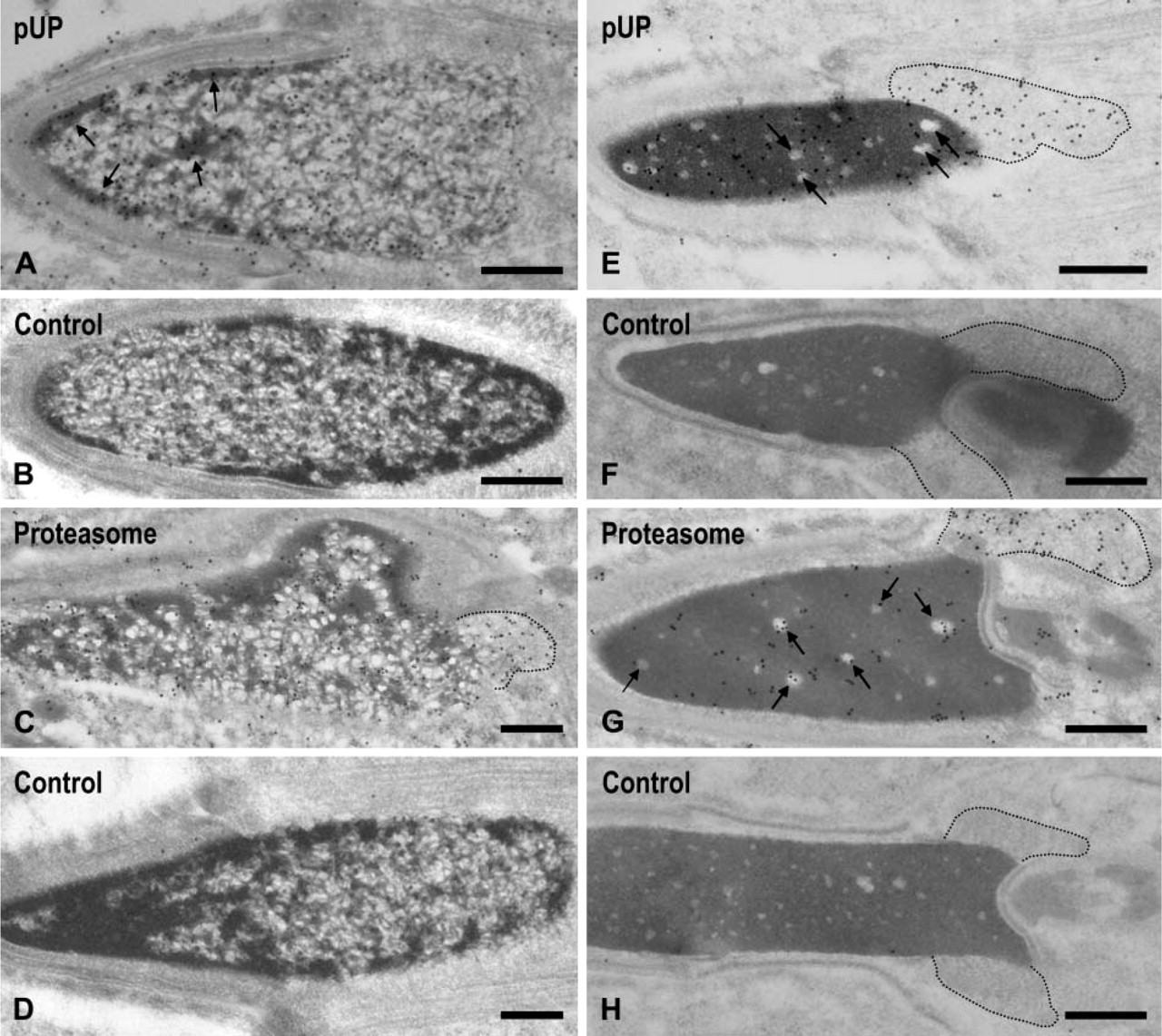
Immunoelectron microscopic localization of pUP and proteasome in the nuclei of steps 13 and 15 spermatids. (
Condensed Spermatids (Steps 16–19). Strong labeling for pUP was observed in the dense area of spermatids at step 19 (Figure 5A). Most clear spots in the dense area were devoid of gold labeling (Figure 5A, arrows). The size of the nuclear pocket decreased gradually and almost disappeared in step 19 spermatids, and labeling for pUP in this area also decreased. Moderate labeling for proteasome was observed in the dense area of step 19 spermatids, in which some gold signals were present in the clear spots (Figure 5B, arrows). Labeling for proteasome in the nuclear pocket diminished also in this step.
Immunoelectron Microscopic Localization of pUP and Proteasome in Rat and Human Sperm
Rat Epididymal Sperm. Sperm nuclei were strongly stained for pUP (Figure 6A) but weakly or not stained for proteasome (Figure 6B). Some gold particles for proteasome were associated with small clear spots (Figure 6B, arrows).
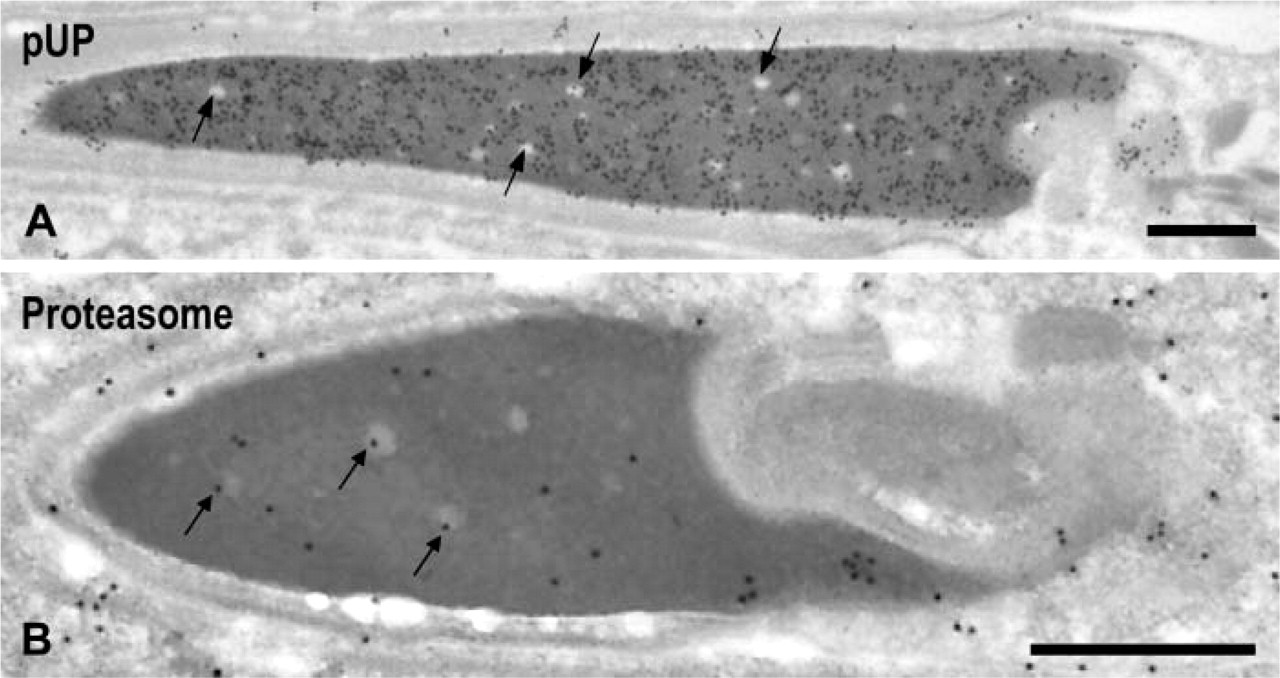
Immunoelectron microscopic localization of pUP and proteasome in step 19 spermatids. (
Human Sperm. The nucleus of human ejaculated sperm contained small clear spots that were also observed in the nuclei of rat sperm, as well as a large clear area representing the vacuoles (Westbrook et al. 2001). In the next step we studied the localization of pUP and proteasome in these areas. Heavy labeling for pUP was noted in the dense area but not in the clear spots or in the vacuoles (Figure 7A, arrows). Gold signals for proteasome were detected exclusively in the clear spots and vacuoles (Figures 7B, arrows).
Quantitative Analysis of Labeling Density for pUP and Proteasome in Developing Spermatogenic Cells
To verify the changes in the relative amounts of nuclear pUP and proteasome during spermatogenesis, we quantified the labeling density for these proteins in the dense area and nuclear pocket. The dense area consists of clear spots and dense matrix. We determined the density of gold particles in the dense area without dividing both regions. The nuclear pocket appeared in spermatids after step 9 and almost disappeared in 19 spermatids. Results are shown in Figure 8. Labeling density for pUP in the dense area was low during early phases of spermatogenesis but started to increase in steps 10–12 spermatids and reached the maximum at step 15. Afterwards, labeling density remained stable until epididymal sperm. Labeling density for pUP in the nuclear pocket was at a low level in steps 10–12 spermatids, increased gradually, and reached a maximum at step 15, decreasing thereafter. Labeling density for pUP in the nuclear pocket was always lower than in the dense area. Proteasome labeling density in the dense area remained at ~40 from spermatogonia to step 12 spermatids and started toincrease from steps 13 to 14, reached a peak value at step 15, and then gradually decreased. In epididymal sperm, labeling density remained stable at ~30. In the nuclear pocket, labeling density for proteasome was detected first in steps 7–9 spermatids and then started to increase, reaching a maximum value in step 15 spermatids, decreasing thereafter.
Discussion
In the present study, Western blot analysis of proteasomes showed several bands in the nuclear fraction isolated from rat testes, which had the same molecular mass as described for proteasomes (Tokumoto et al. 2000) and were commonly observed in two 20S proteasome preparations partially purified from rat testes and liver, respectively. In rat sperm heads, the 31-kDa subunit was not observed but two other subunits were detected. It was shown that the 31-kDa subunit was the phosphorylated form of the 30-kDa subunit (Tokumoto et al. 2000). Thus, subunits of sperm nuclear proteasomes might be modified for their function so that they show molecular masses slightly different from those in testis nuclei. Antibody FK1 reacting with pUP, but not with free ubiquitin (Fujimuro et al. 1994), detected many bands of various molecular mass in testicular nuclei and in sperm heads. The molecular mass of some bands observed in the testicular nuclei was similar to those in sperm head. Results indicate the specificity of the antibodies used.
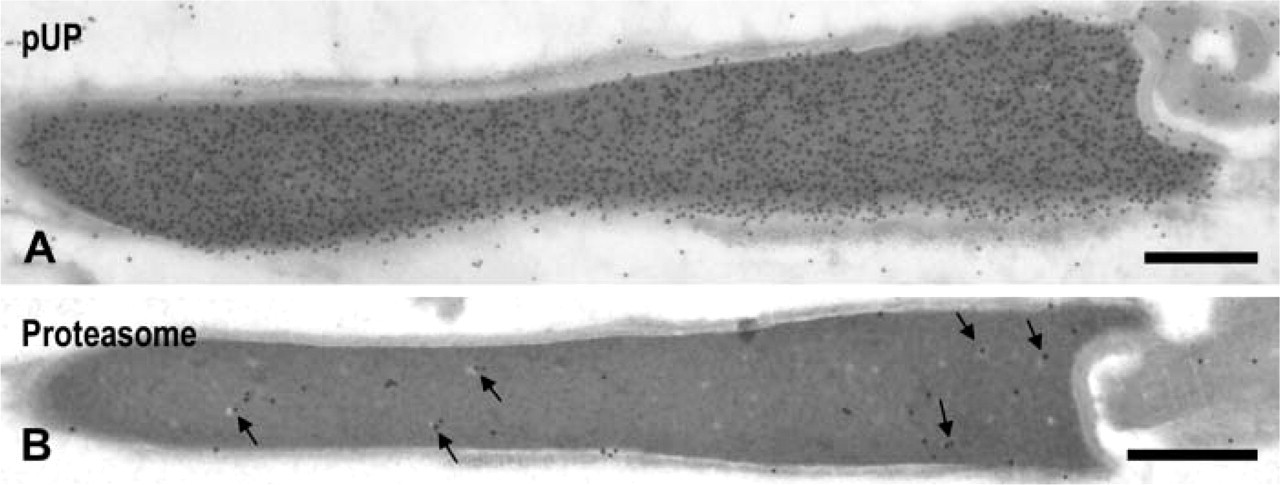
Immunoelectron microscopic localization of pUP and proteasome in the nuclei of rat epididymal sperm. (
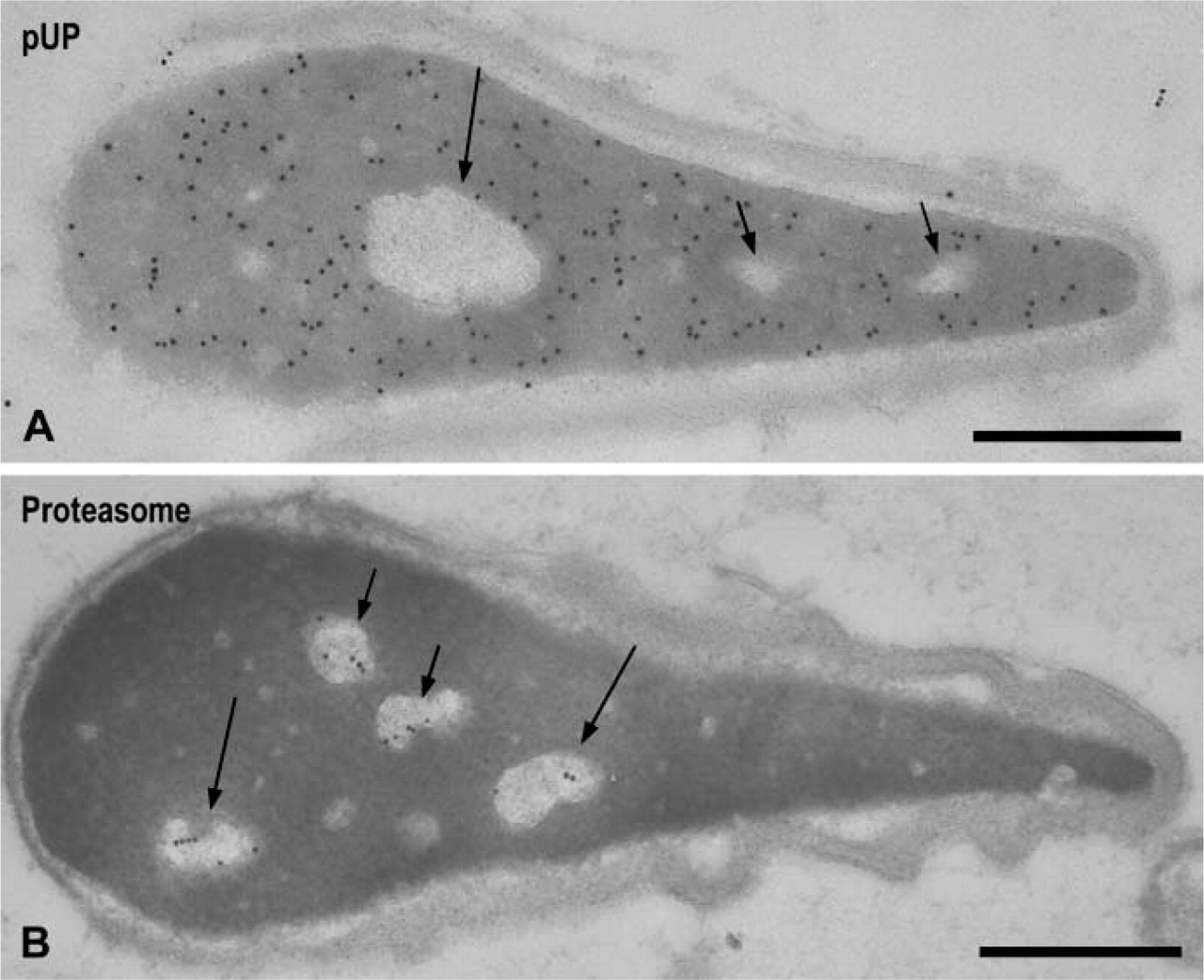
Immunoelectron microscopic localization of pUP and proteasome in the nuclei of human ejaculated sperm. (
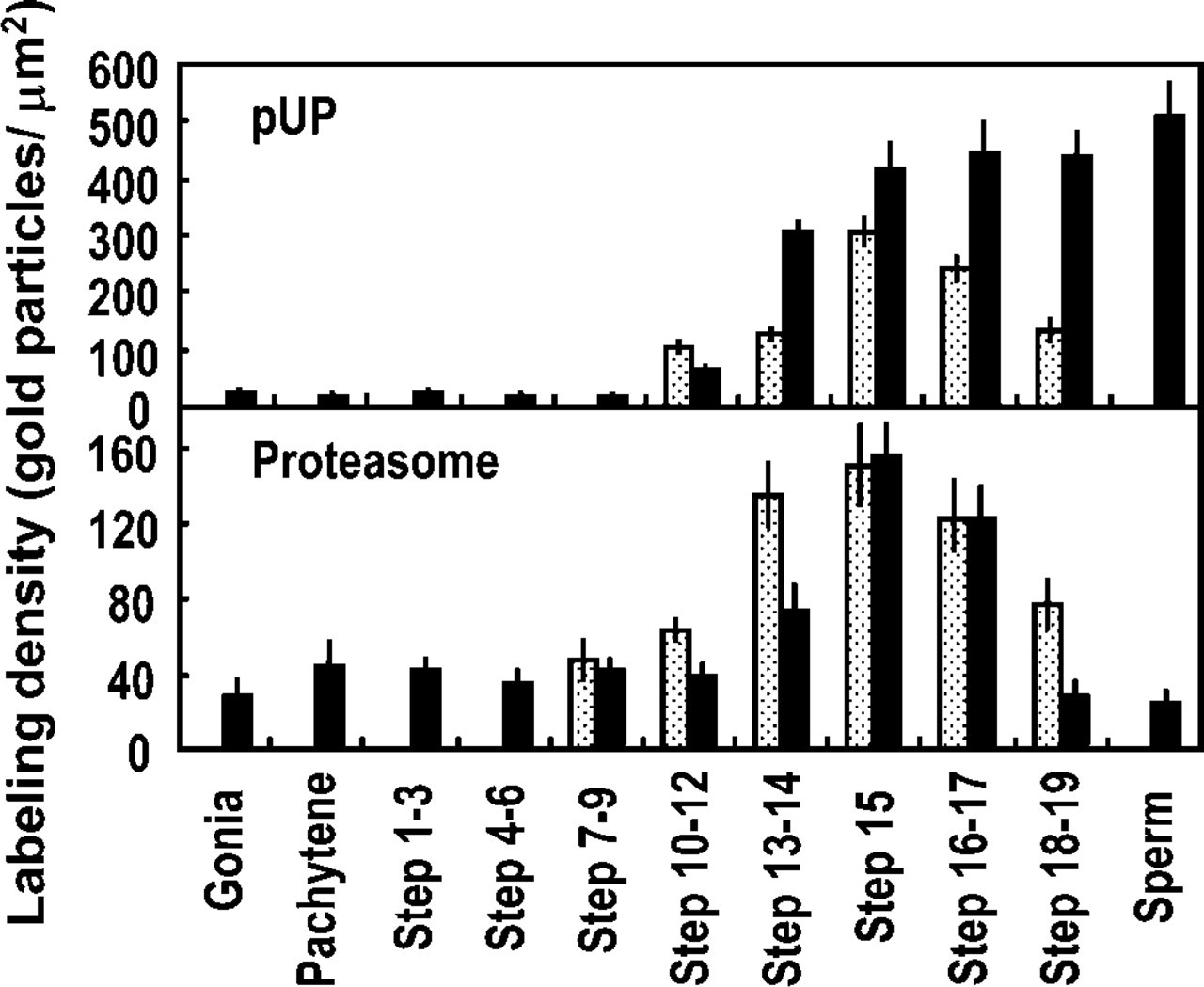
Quantitative analysis of ubiquitin-proteasome system-related proteins in developing spermatogenic cells. Solid bars, labeling density in dense area; dotted bars, labeling density in nuclear pocket. Data are expressed as mean ± SD (n = 8).
Our immunofluorescence studies showed localization of pUP in various areas of spermiogenic cells including Golgi apparatus, residual bodies, flagella, and nuclei. Localization of pUP in the Golgi region was also reported previously (Haraguchi et al. 2004). Our study identified proteasome in the nuclei of spermatocytes and late spermatids and in flagella. Considered together, our results suggest that pUP are produced at multiple sites of spermiogenic cells and are degraded by the ubiquitin-proteasome system.
Immunofluorescent staining in the present study also identified pUP and proteasome in the rat sperm head and in the human sperm head. Results are consistent with those of Western blotting. Immunofluorescent localization of proteasome in the sperm of other mammals has been reported by several groups (Wojcik et al. 2000; Sutovsky et al. 2004; Pasten et al. 2005); however, it was detected in the neck region or acrosome region but not within the nucleus. These results are inconsistent with ours. Differences in the results of our study and those of others are probably based on differences in the methods used, such as cytochemical treatments including fixation and staining procedures, as well as the antibodies used. Immunofluorescent staining results of human sperm were quite consistent with the immunoelectron microscopic results and clearly showed the distribution of proteasome in the clear spots and in the vacuoles and that of pUP in the dense nuclear matrix.
Relationship Between Nucleoprotein Replacement and Fluctuation of pUP and Proteasome Levels in Late Spermiogenesis
Ubiquitinated histones are detected in late spermatids of chicken and trout (Agell et al. 1983; Nickel et al. 1987). In rooster, levels of ubiquitinated H2A increase in late spermatids, just before replacement of histones by protamine (Agell et al. 1983). In the mouse testis, ubiquitinated histone H2A was detected in steps 8–12 spermatids (Baarends et al. 1999), whereas poly-ubiquitinated forms of histone H2A and H3 were identified in steps 9–12 spermatids of rat (Chen et al. 1998). Our immunoelectron microscopic results showed localization of pUP and proteasome in two distinct nuclear sites in spermiogenic cells: the dense area and nuclear pocket. The latter was recognized in steps 9–19 rat spermatids and surrounded by so-called redundant nuclear envelope (Lalli and Clermont 1981; Clermont et al. 1993). The appearance and increase of pUP in the dense area during late spermiogenesis shown in the present study are similar to those of nucleoproteins described previously (Kistler et al. 1996; Oko et al. 1996; Dadoune 2003). In rats, replacement of histones by transition proteins begins at steps 11–12 and is complete at steps 13–15 (Kistler et al. 1996; Oko et al. 1996). In addition, replacement of transition proteins by protamines commences at about steps 13–15 and is almost complete at steps 16–19 (Kistler et al. 1996; Dadoune 2003). On the other hand, proteasome in the dense area increases at steps 13–14, reaches peak values at step 15, and then decreases to low levels at step 19. Together the increase of pUP in late spermatids suggests polyubiquitination of histones and transition proteins. Moreover, based on the fluctuation in proteasome level, it is likely that these proteins are involved in the degradation of nucleoproteins.
In addition, our data showed high levels of pUP in rat epididymal sperm. Protamine is a major nucleoprotein found in mature sperm. It is known that protamine is replaced rapidly by oocyte-derived histones after fertilization, but little is known about the mechanisms of this replacement. The released protamine might be degraded by the ubiquitin-proteasome system. It is then suggested that the protamines might be polyubiquitinated and targeted for degradation by the ubiquitin-proteasome system present in the oocyte. Results of preliminary studies in our laboratories showed polyubiquitination of some population of protamine molecules in late spermiogenesis.
Function of the Nuclear Pocket During Spermiogenesis
Our idea for the function of the nuclear pocket is summarized in Figure 9. The morphology of the nuclear pocket in mammalian spermatids has been described in detail (Franklin 1968; Fawcett and Phillips 1969; Zamboni and Stefanini 1970; Lalli and Clermont 1981), but its function is not clear. We detected pUP and proteasome in the nuclear pocket. Our quantitative analysis showed that labeling for proteasome appeared at steps 7–9 and then increased, whereas pUP labeling started at step 10 and also increased thereafter. The process of nucleoprotein replacement seems to require the degradation of a large amount of proteins such as histones and transition proteins, as discussed above. We postulate in this study that these proteins might be polyubiquitinated in the dense area as mentioned and degraded in the clear spots and in the nuclear pocket where proteasome is localized. Westbrook et al. (2001) reported the presence of SPAN-X, a sperm protein associated with the nucleus, in the nuclear vacuole and in the nuclear pocket of human sperm, suggesting continuity between these structures. They suggested that this continuity facilitates transport of materials such as mRNAs or proteins targeted for degradation into the cytoplasmic droplet for disposal. In rat sperm nucleus, no nuclear vacuoles are observed but, instead, small clear spots are present. In our study we showed the presence of proteasome in these clear spots. It is likely that the clear spots could move to the nuclear pocket or continue to the nuclear pocket as considered in human sperm (Westbrook et al. 2001). Moreover, our results showed the presence of proteasome in the clear spots as well in the vacuoles in human sperm. If pUP were transported to the nuclear pocket by clear spots or vacuoles, one would expect pUP signals in both structures. However, no such pUP signal was present in the clear spots and in the vacuoles. It is possible that pUP are degraded rapidly in the clear spots or transported rapidly to the nuclear pocket, so that they are not observed in these spots. Thus, our results suggest that the nuclear pocket functions as a site for degradation of nucleoproteins.
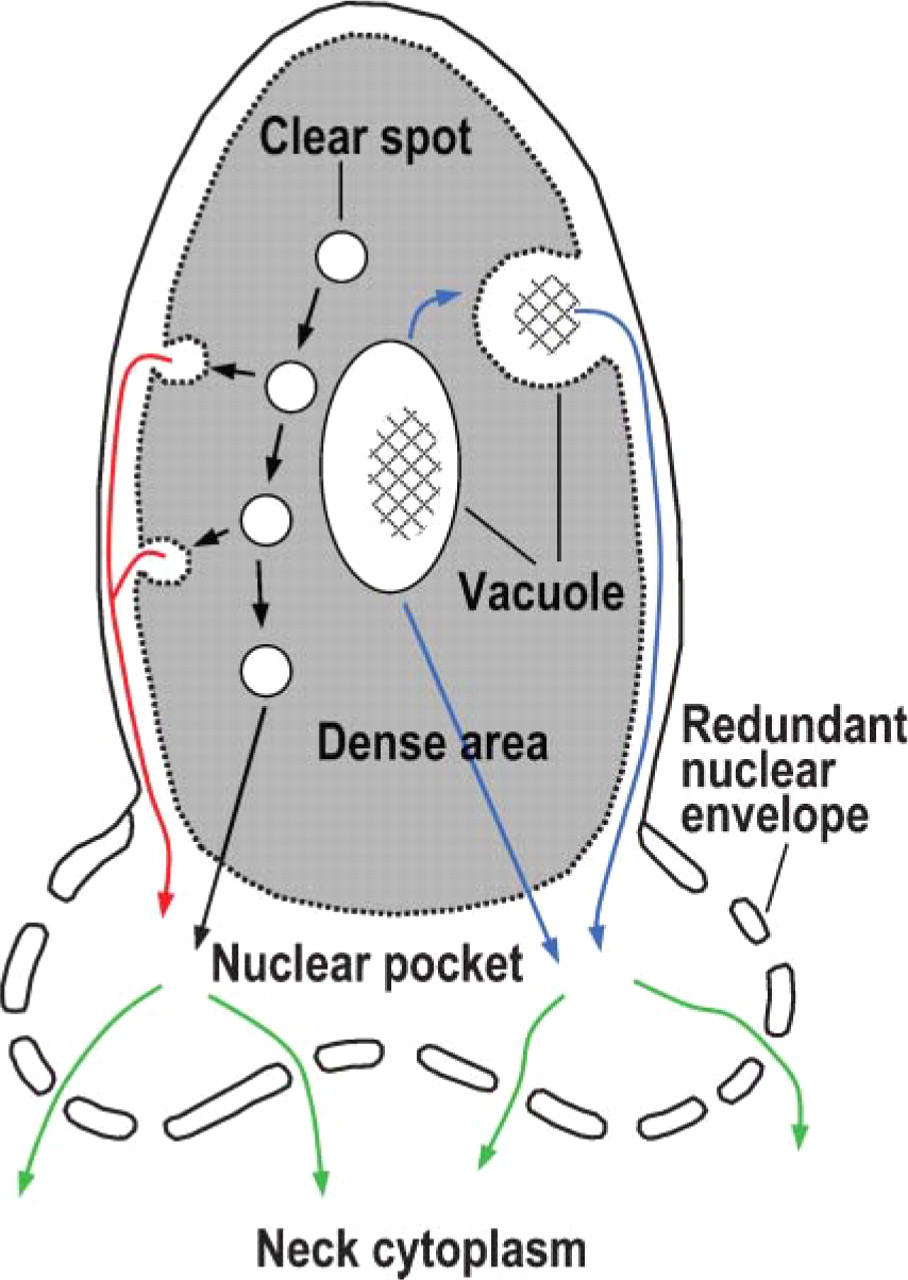
Schematic diagram of degradation sites in late spermatid. Cell membrane, acrosome, and neck cytoplasm are not drawn. Nuclear pocket is the nuclear area surrounded by redundant nuclear envelope. Proteasome was present in the clear spots, nuclear vacuoles, and nuclear pocket. Polyubiquitinated proteins are detected in the dense area and nuclear pocket but not in the clear spots and nuclear vacuoles. We propose that clear spots, nuclear vacuoles, and nuclear pocket are nuclear degradation sites in late spermatids. Clear spots could move directly to the nuclear pocket (black lines) or to the periphery of the dense area and be released into the subacrosomal space between dense area and nuclear membrane (red line), which is open to the nuclear pocket. Nuclear vacuoles could move directly to the subacrosomal space and the nuclear pocket (blue line), the sites of the most extensive degradation of unnecessary proteins generated during replacement of nucleoproteins. The resultant degradation products could be transported through nuclear pores into the neck cytoplasm (green lines) connecting to residual body.
In conclusion, in the present study we showed the presence of ubiquitinated proteins and proteasome in the clear spots, nuclear vacuoles, and nuclear pocket of spermatids and sperm. These results suggest the involvement of the ubiquitin-proteasome system in degradation of temporarily functioning proteins that appear following replacement of nucleoproteins. The sites for degradation of the proteins are the clear spots within the dense nuclear area and the nuclear pocket.
Footnotes
Acknowledgements
This work was supported in part by a Grant-in-Aid (17570158) and the Center of Excellence (21COE18999997) program from the Ministry of Education, Science, Culture and Sport (to SY).
