Abstract
The ability to preserve the fragile ultrastructural organization of bacterial biofilms using cryo-preparation methods for electron microscopy has enabled us to probe sections through non-typeable Haemophilus influenzae (NTHi) biofilms and determine the localization of NTHi-specific lipooligosaccharide (LOS) and proteins within these structures. Some of the proteins we examined are currently being considered as candidates for vaccine development, so it is important that their distribution and accessibility within the biofilms formed by NTHi be determined. We have localized LOS to the extracellular matrix (ECM) of the biofilm and the P6 outer membrane protein to the membrane of what appear to be viable bacteria within the biofilm. The Hap and HWM1/HMW2 adhesive proteins were associated with bacteria within the biofilm and were present in the biofilm ECM. The IgA1 protease is a secreted protein that was also associated with NTHi in the biofilm and was in the ECM, but was more concentrated in the top region of the biofilm, suggesting a role in protecting biofilm bacteria from antibody attack.
Keywords
N
Attachment to both inert and living surfaces is a common property exhibited by many bacteria and is considered a first step in biofilm formation. Biofilms have been broadly defined as multicellular assemblages of microorganisms and extracellular products attached to an abiotic or biotic surface (Davey and O'Toole 2000). The extracellular products form a matrix, typically rich in sugar moieties (Neu et al. 2001), that probably confers protection from phagocytosis and antibiotic treatment (Donlan and Costerton 2002). Within the biofilm, the bacteria have different growth rates. While some are rapidly growing, others are in a dormant or sessile state, a condition that may be linked with reduced ribosome (rRNA) content (Poulsen et al. 1993).
Otitis media as a biofilm infection was first proposed to explain the observation that 40-60% of patients with chronic otitis media have sterile middle ear fluids (Rayner et al. 1998), suggesting the absence of culturable bacteria (Giebink 1989). The presence of bacterial endotoxin (Dingman et al. 1998), bacterial DNA (Post et al. 1995), and bacterial mRNA (Rayner et al. 1998) in these sterile fluids further supported the hypothesis that otitis media is a biofilm infection (Dingman et al. 1998; Costerton 1999). Direct evidence of biofilm formation on middle ear mucosa has since been presented in experimental systems (Post 2001). Bacterial biofilms in otitis media may help to explain the phenomenon of persistent infections that occur in many patients with otitis media (Pichichero 2000; Faden 2001).
Although biofilms have become an accepted concept in infection, the mechanisms by which they are formed and maintained are still being elucidated. As a contribution to the study of biofilms, we have used immunoelectron microscopy to examine the location of bacterial adhesins in NTHi biofilms. Specifically, we have examined the location and distribution of the HMW1/HMW2 proteins, the Hap protein, and IgA1 protease. The HMW1/HMW2 proteins (Barenkamp and Bodor 1990; Barenkamp and Leininger 1992) interact with eukaryotic cell receptors (St Geme et al. 1993). The Hap protein (St Geme et al. 1994) and the closely related IgA1 protease (Kilian et al. 1979,2002) are autotransporters that have both been reported to mediate microcolony formation (Plaut et al. 1992; Fink et al. 2003). The Hap protein has also been shown to mediate interactions with human epithelial cells (St Geme et al. 1994; Fink et al. 2003).
The proteins were selected to explore their distribution within a biofilm and because many of them are currently being considered as candidates for vaccine development. We are not aware of earlier reports in which specific antibodies were applied to sections through biofilms that have been cryo-fixed and freeze substituted.
Materials and Methods
Bacterial Strains, Growth, and Storage
NTHi strain 9274 was originally isolated from the middle ear fluid of a patient with acute otitis media (Gu et al. 1995) and was propagated on chocolate agar at 37C under 5% CO2 and 95% humidity. Colonies from the agar plates were transferred to 3.7% brain heart infusion medium (BHI broth; Difco Laboratories, Detroit, MI) containing nicotinamide adenine dinucleotide (NAD, 2 μg/ml) and hemin (10 μg/ml) (both from Sigma Chemical Co., St. Louis, MO). These bacterial suspensions were incubated overnight at 37C under 5% CO2 and 95% humidity, and the bacteria were then centrifuged at 15,000 × g for 5 min. Pellets were resuspended in BHI broth containing 25% glycerol, and the suspension was stored in small aliquots under liquid nitrogen.
Crystal Violet Assay for Biofilm Formation
The method used to evaluate biofilm formation by NTHi strain 9274 was essentially as previously described (Watnick and Kolter 1999; Murphy and Kirkham 2002). Aliquots of a 1:200 dilution of an overnight bacterial culture were placed into the wells of a 96-well plate and incubated overnight at 37C in 5% CO2 and 95% relative humidity. Exactly 24 hr later, 20 μl of a 1% solution of crystal violet solution was added to the wells for 15 min, after which the bacteria were washed extensively. The wells were air-dried, the dye in the wells was solubilized with DMSO, and the optical density 595 (OD595) of each well was read. Sterile BHI was substituted for overnight bacterial cultures in control experiments.
Preparation of NTHi Biofilms on Solid Substrates
Suspensions of NTHi (clone 9274) were removed from storage under liquid nitrogen, plated onto chocolate agar plates, and grown overnight at 37C in 5% CO2 and 95% relative humidity. From these plates, five individual colonies were removed and resuspended in BHI broth supplemented with hemin and NAD (Poje and Redford 2003) and incubated overnight at 37C, 5% CO2, and 95% relative humidity. After 20 hr, relative numbers of bacteria in suspension were estimated by reading the OD at 600 nm (Poje and Redford 2003).
For electron microscopy, biofilms were prepared on glass coverslips or Millipore filters. Biofilms on glass coverslips were prepared by immersing sterile coverslips in a 1:200 dilution of NTHi suspension in BHI medium and incubating overnight (37C, 5% CO2, and 95% relative humidity). In some instances, the culture dishes were placed on a slight incline so that the coverslips were only partially covered with NTHi suspension.
Colony biofilms on filter substrates were prepared using a modification of previously published methods (Anderl et al. 2000; Zahller and Stewart 2002; Thurnheer et al. 2003). Sterile Millipore filters (Millipore Corp., Billerica, MA, Catalog # GSWP 025 00) were placed onto the surface of chocolate agar plates (Hardy Diagnostics, Catalog # E-14) and inoculated with a concentration of NTHi in BHI broth that produced a density of approximately 0.3 bacteria/μm2. The filters with bacteria were incubated overnight at 37C in 5% CO2 and 95% relative humidity.
Rapid Freezing and Freeze Substitution
Biofilms on glass coverslips and on Millipore filters after 24 hr were prepared for examination by electron microscopy using rapid freezing followed by freeze substitution (Webster et al. 2004). For scanning electron microscopy (SEM), the coverslips or filters were rapidly immersed in liquid propane and transferred to dry methanol at −80C for dehydration by freeze substitution. The frozen specimens were gradually warmed to 4C, critical point dried, and given a 6-nm coating of platinum before being examined by field emission SEM (using an XL 30 SFEG SEM; FEI Company, Hillsboro, OR).
Biofilms on filter substrates were cut into 0.8-mm discs using a specially manufactured cutter (Morphew and McIntosh 2003) and placed biofilm-face down into aluminum planchettes (Ted Pella Inc.; Redwood, CA) containing a thin film of hexadecene. A second aluminum planchette was placed over each filter, and the specimen was then immediately frozen at 2100 bar in an HPF machine (HPM 010; Bal-Tec, Lichtenstein) using established methods (Sawaguchi et al. 2002,2003).
Frozen filters were placed into polypropylene specimen capsules (Leica Inc.; Deerfield, IL) and either stored under liquid nitrogen for further processing, or immediately transferred to cold, dry ethanol maintained at a temperature of −80C in an AFS freeze substitution machine (Leica). Specimens were left overnight at −80C and were gradually warmed to −60C over a 6-hr period. During the warming period, the specimens were exposed to two changes of fresh, dry ethanol.
At a temperature of −60C, the specimens were infiltrated with a 1:1 mixture of ethanol and Lowicryl HM20 resin (Electron Microscopy Sciences; Ft. Washington, PA) for 2 hr and then in a fresh 1:1 ethanol/resin mixture overnight. The next day, the specimens were warmed to −50C, transferred to fresh Lowicryl HM20 resin, and polymerized under UV light, following protocols provided with the resin kit.
Immunocytochemistry
Thin sections (60-80 nm) through the Lowicryl-embedded biofilms were prepared using an Ultracut S ultramicrotome (Leica) with a diamond knife (Diatome USA; Ft. Washington, PA) and collected on metal specimen grids.
The sections were labeled with antibodies using sequential labeling protocols (Griffiths 1993) by floating the grids, section side down on small drops of reagent. Polyclonal antibodies to H. influenzae (Difco Laboratories) and to the NTHi IgA1 protease (Plaut et al. 1992) were applied to sections and visualized using 10 nm protein A-gold (PAG; University of Utrecht, The Netherlands). Monoclonal antibody 3B9 against P6 outer membrane protein (OMP) (Murphy et al. 1985) and monoclonal 6E4 against lipooligosaccharide (LOS) (Campagnari et al. 1990) were applied to sections and visualized by applying anti-mouse IgG bridging antibodies produced in rabbits, and then 10 nm PAG. The Hap protein was detected using guinea pig antiserum GP74 (Cutter et al. 2002), and the HMW1/HMW2 was detected using guinea pig antisera raised against purified HMW1, but which recognizes both HMW1 and HMW2 (Buscher et al. 2004). The anti-Hap and HMW1/HMW2 were applied to sections and visualized by applying rabbit anti-guinea pig IgG antibodies and 10 nm PAG.
All polyclonal secondary antibodies were diluted in 10% goat serum containing a suspension of heat-killed, homogenized NTHi bacteria and centrifuged before use. This treatment reduced the nonspecific binding observed with some polyclonal secondary antibody preparations.
Electron Microscopy
All sections were examined by transmission electron microscopy (TEM), using a CM120 BioTwin operating at 80 kV (FEI Company). Images were collected on negative film and digitized using a high-resolution digital scanner. Quantification of immunogold labeling over NTHi biofilms was performed on images obtained using systematic sampling protocols (Lucocq 1993).
Western Blotting
Biofilms of NTHi were grown on Millipore filters as described above. The bacteria growing on the filters were solubilized either to obtain an enrichment of extracellular matrix (ECM), or to obtain a whole-biofilm homogenate. ECM proteins were obtained by solubilizing the filters in 1 × PBS using a probe sonicator. The filters were exposed to four rounds of cooling on ice, sonication for 3 min, and cooling for 10 min. Whole bacteria were removed by centrifugation, and the supernatants were subjected to an SDS-PAGE separation using a NuPAGE system and Novex bis-Tris gels (Invitrogen; Carlsbad, CA).
Whole homogenates of biofilm were obtained by solubilizing and sonicating the biofilms in lysis buffer (50 mM Tris-HCL, 150 mM NaCl, 1% Triton X-100, pH 7.4). The sonication conditions applied were the same as those used for obtaining biofilm ECM proteins (4 × 3 min sonication and 10 min cooling). The supernatants were separated from filter fragments by centrifugation. This solubilization protocol was also used to obtain whole-cell lysates from pellets of planktonic NTHi that had been grown in suspension.
SDS-PAGE-separated proteins were blotted onto nitrocellulose membranes and blocked by incubation with Tris-buffered saline Tween-20 (50 mM Tris-HCL, 150 mM NaCl, 0.1% Tween 20, pH 7.5) containing 2% fish skin gelatin. The buffer used to dilute secondary and tertiary antibodies also contained suspensions of bacterial proteins from heat-killed, lysed NTHi to remove non-relevant bacteria-specific contaminating antibodies. The blots were washed and then incubated sequentially with specific antibodies to NTHi proteins, secondary antibodies produced in rabbits, and HRP-conjugated tertiary antibodies produced in goats. The antibodies were visualized using a chemiluminescent horseradish peroxidase substrate kit (Vector Laboratories; Burlingame, CA).
Results
Formation of NTHi Biofilms on Solid Substrates: Crystal Violet Assay
To establish that NTHi strain 9274 has the ability to form biofilms, we first performed a crystal violet assay on overnight bacterial cultures grown in a 96-well plate. The crystal violet stained a significant amount of material that had been deposited on the sides of the wells by the growing bacteria (Figure 1A), compared with crystal violet staining of wells that contained only culture medium (Figure 1B). Measurement of the OD595 of solubilized crystal violet stain enabled this observation to be expressed in relative units (Figure 1C).
SEM of Biofilms on a Glass Substrate and Filters
To examine the biofilms formed by the bacteria, we grew overnight cultures of NTHi strain 9274 in plastic dishes containing glass coverslips. The material deposited on the coverslips stained with crystal violet (data not shown), and in the SEM was shown to be a thick layer of amorphous material containing embedded bacteria (Figure 2A). When coverslips were incubated half submerged in the bacterial suspension, a thick band of bacteria and associated amorphous material formed at the air-liquid interface (Figures 2A and 2B). The material that was attached to the glass substrate was a thick layer of bacteria covered by a layer of amorphous material, presumably extracellular material generated by the NTHi (Figure 2C). Large clumps of bacteria that appeared to be held together by this extracellular material were observed on the glass that had been submerged (Figure 2C) as well as on the glass surface exposed to air (data not shown).
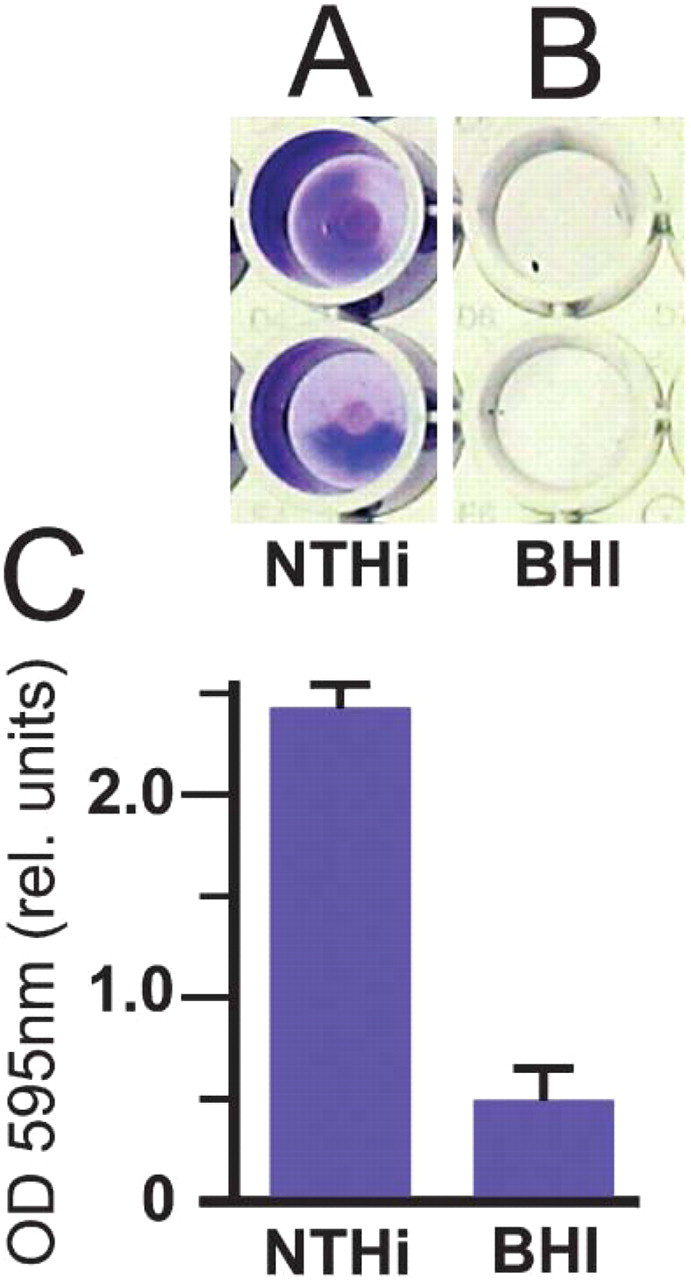
Crystal violet assay. Crystal violet was used to stain biofilm materials deposited by NTHi strain 9274. Biofilm material in the wells that contained non-typeable Haemophilus influenzae (NTHi) is stained with crystal violet (Lane
To examine NTHi biofilms in the TEM, it was necessary to prepare them on substrates that could be thin sectioned. For this reason, we generated biofilms on filters that could be frozen using a high-pressure pressure freezer and sectioned. Our first attempts, which consisted of growing NTHi biofilms on Anopore insert filters with the sterile side immersed in culture medium, produced structures with typical biofilm characteristics. Large clumps of bacteria were present on the filter surface, and in the SEM we observed what appeared to be channels running through thick monolayers of bacteria (data not shown).
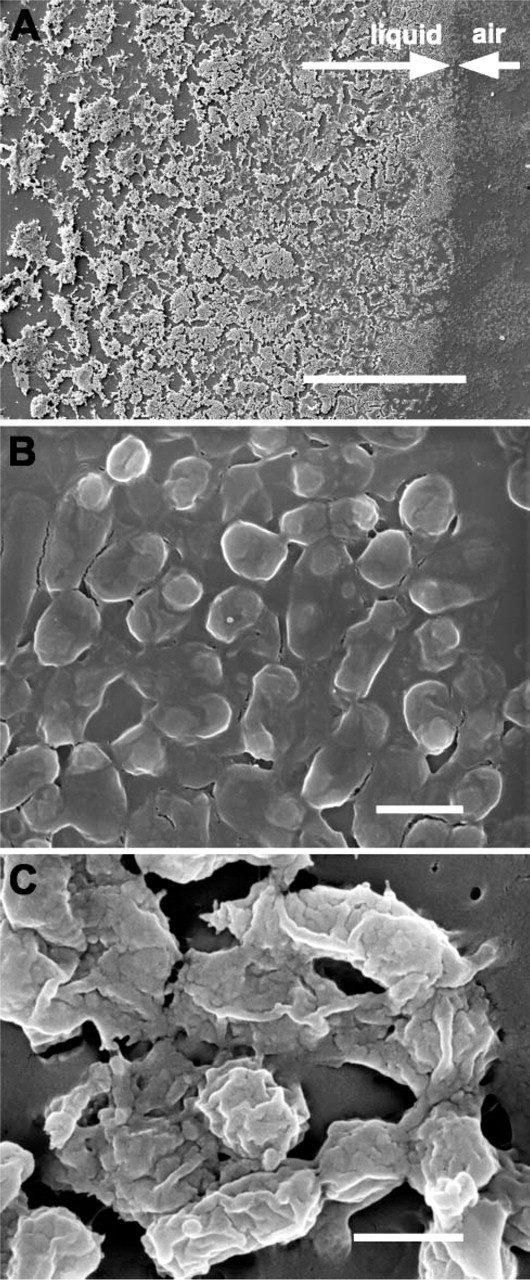
Scanning electron microscopy (SEM) images of NTHi biofilm growing on partially immersed glass substrates. (
The Anopore filters were too brittle and hard for use in the high-pressure freezer and for sectioning. For this reason, we turned to growing the NTHi biofilms on Millipore filters. Incubating filters overnight that had been inoculated with 0.3 bacteria per square micron produced an almost uniformly thick layer of bacteria on the filters (Figure 3). The biofilms were restricted to the areas of the filters that had been covered by the initial innoculum of NTHi, and spreading of bacteria beyond the edges of the biofilm was not observed. However, at the edges of the biofilm zone, the mat of bacteria tapered down to a layer that was only one bacterium thick (Figure 3A). The region immediately outside the biofilm had a layer of extracellular material covering the support filter (Figure 3A) that contained small numbers of embedded bacteria (Figure 3A). The top of the formed biofilm was uniformly flat and was covered with a layer of extracellular material over the bacteria and the spaces between them (Figure 3B). Mechanical damage to the bacterial biofilm, especially at its edge, revealed the close association of the extracellular material with the bacteria (Figure 3C), and the large amount of this extracellular material that was being produced by the bacteria (Figure 3C).
These Millipore filters could be easily manipulated for high-pressure freezing, resin-embedding, and sectioning. All subsequent sections through biofilms were of Lowicryl-embedded biofilms that had been grown on Millipore filters and prepared using low-temperature methods.
Immunocytochemistry on Lowicryl-embedded Biofilm Sections
When we examined the biofilm sections by TEM, we were able to easily identify four distinct compartments. The NTHi in the biofilm occupied 68% of the biofilm (Table 1) and consisted of the cell cytoplasm and the outer cell membrane. An electron-opaque, amorphous material that we classified as ECM surrounding the bacteria occupied 25% of the biofilm, and the remaining 7% of the biofilm was occupied by what we designated as extracellular space (ES), defined as the areas within the biofilm that did not contain stainable or electronopaque material (Table 1).
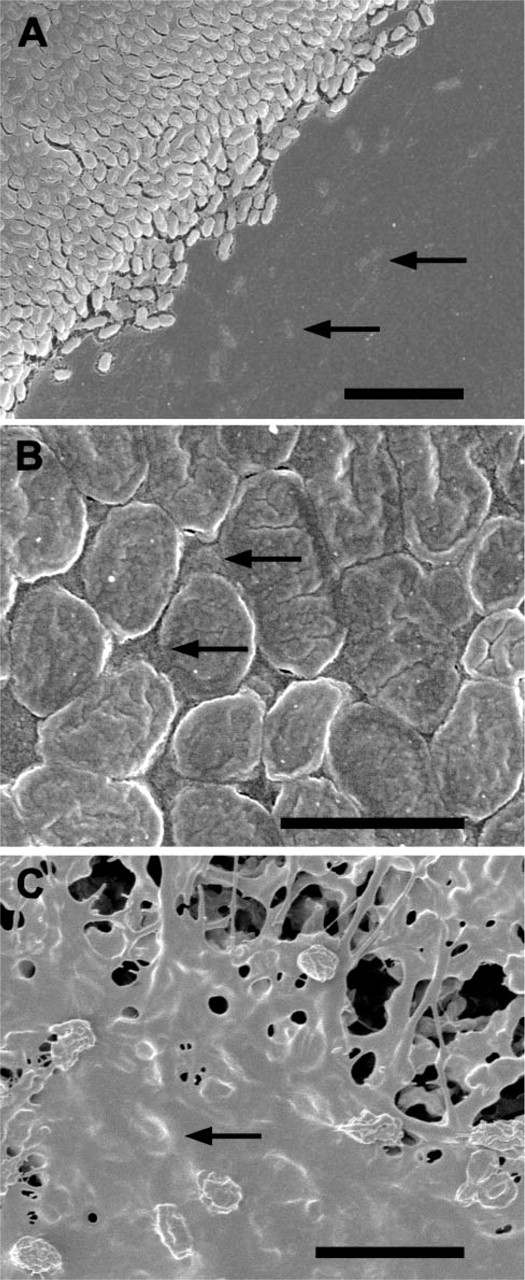
SEM images of NTHi colony biofilm growing on Millipore filters. (
Anti-NTHi, anti-P6, and anti-LOS Labeling of Biofilm Sections
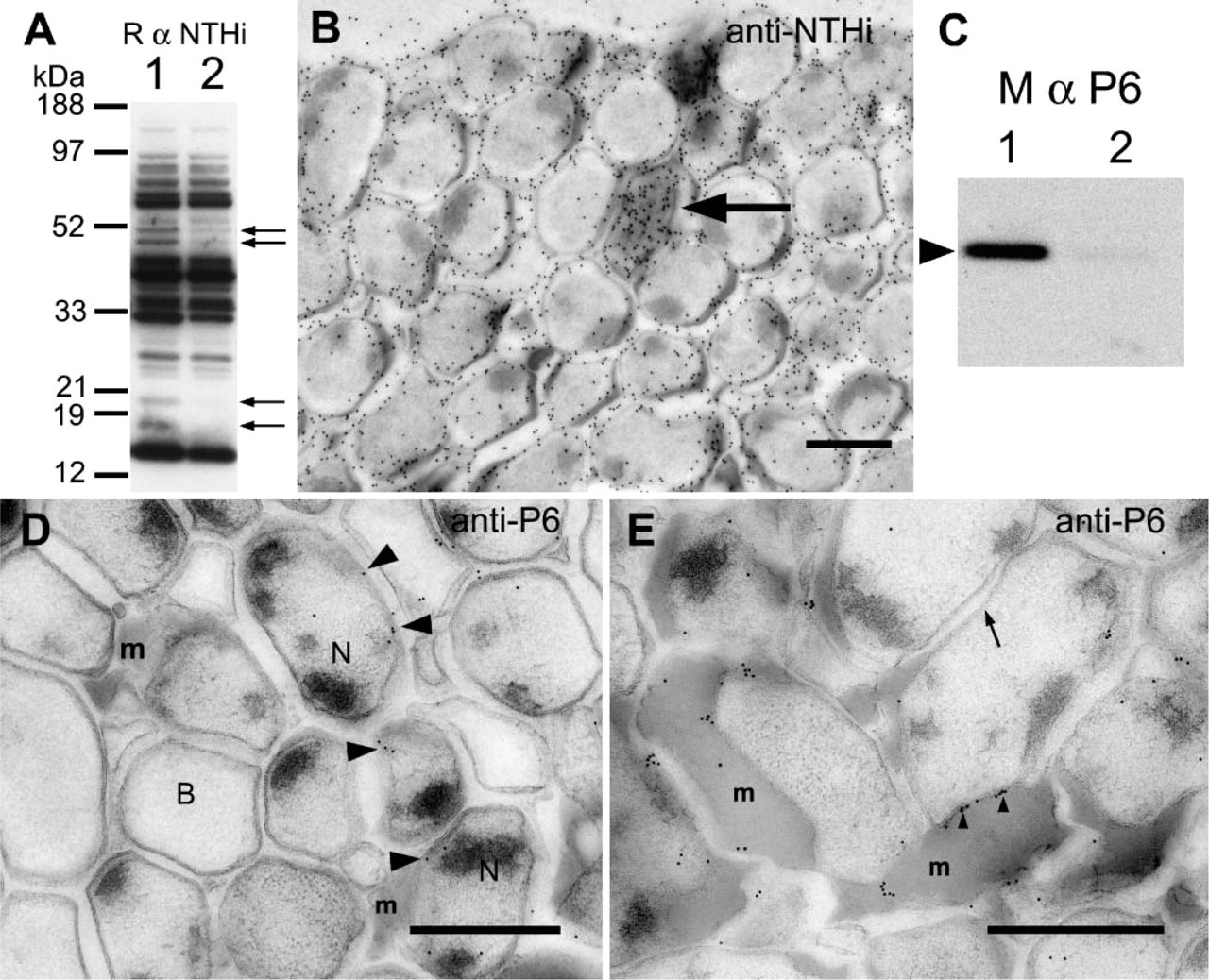
To establish that the bacteria in the biofilms were NTHi, and that the structures being formed on the support filters were biofilms, we first labeled the sections with reference antibodies. The first antibody that was applied to the sectioned biofilms was a commercial polyclonal antibody that recognizes multiple NTHi proteins. This antibody bound to multiple protein bands on Western blots of NTHi whole-cell lysates (Figure 4A) and to protein bands from ECM proteins (Figure 4A). Some protein bands that labeled well in the whole-biofilm protein lane (Figure 4A, Lane 1) were not so visible in the protein bands from the ECM (Figure 4A, Lane 2). By electron microscopy, the antibody produced substantial labeling over the biofilm sections (Figure 4B), associating with the bacteria (Figure 4B) and with the ECM (Figure 4B).
Volume densities of biofilm compartments
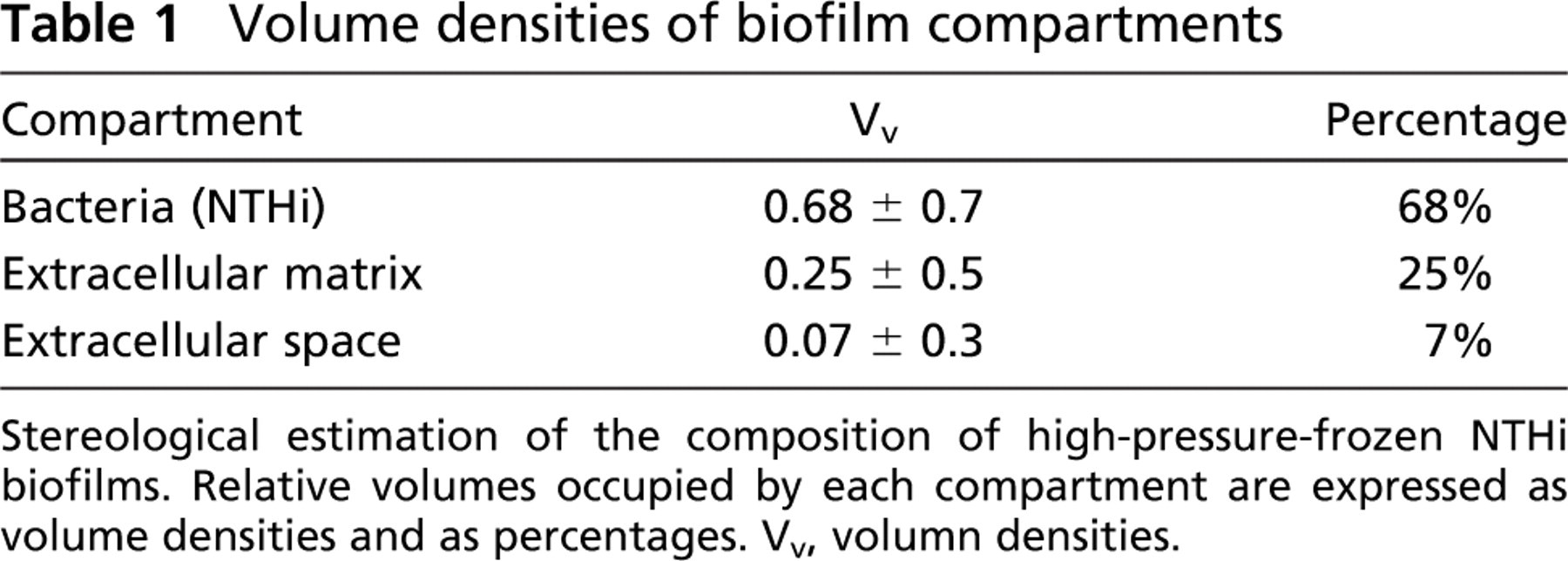
Stereological estimation of the composition of high-pressure-frozen NTHi biofilms. Relative volumes occupied by each compartment are expressed as volume densities and as percentages. Vv, volumn densities.
We then probed the biofilm with a monoclonal antibody to the H. influenzae P6 OMP. This protein was expressed by planktonic and biofilm forms of NTHi and was restricted to the outer membrane. On Western blots, the anti-P6 antibody bound to a protein band with Mr approximately 16.6 kDa (Figure 4C), but was confined to the lane containing proteins from whole biofilms (Figure 4C, Lane 1). No similar protein was labeled in the lane containing ECM proteins (Figure 4C, Lane 2), suggesting that minimal contamination of bacterial outer membranes in the ECM preparations was occurring.
The anti-P6 labeling of sections through the NTHi biofilm (Figures 4D and 4E) was associated with bacteria that appeared to contain more cytoplasmic electrondense material (Figure 4D). Many of the NTHi in the biofilm did not label with the anti-P6 antibodies (Figure 4D), and in some regions, the anti-P6 label was associated with bacterial membranes close to the ECM (Figure 4E). In some instances, the label over the bacterial membrane was concentrated only where the membrane was in close apposition to the ECM (Figure 4E).
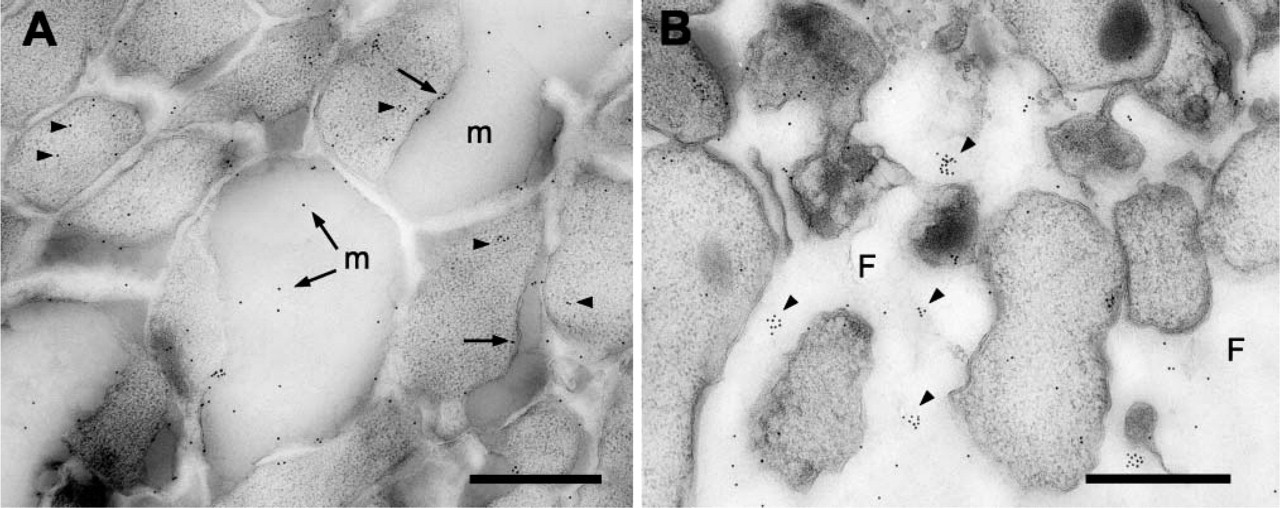
Anti-LOS labeling was also demonstrated on the biofilm sections, with the label being mostly associated with the ECM (Figure 5A). Some label was observed over the cell cytoplasm that could represent newly synthesized LOS or nonspecific background label. In the bottom region of the biofilm, the anti-LOS label was mostly associated the ECM (data not shown), and in some instances, anti-LOS label was observed in regions of the support filter (Figure 5B), close to bacteria that had migrated into the support filter (Figure 5B). From these observations it is possible to speculate that the LOS being shed by the NTHi plays a role in the initial attachment of bacteria to substrates.
Antibodies to Proteins that May Be Involved with Biofilm Formation
Of particular interest to the study of biofilms is the process of bacterial adhesion. For this reason we treated biofilm sections with antibodies to known adhesins to examine the distribution of these molecules within the biofilm.
Anti-Hap Protein Labeling
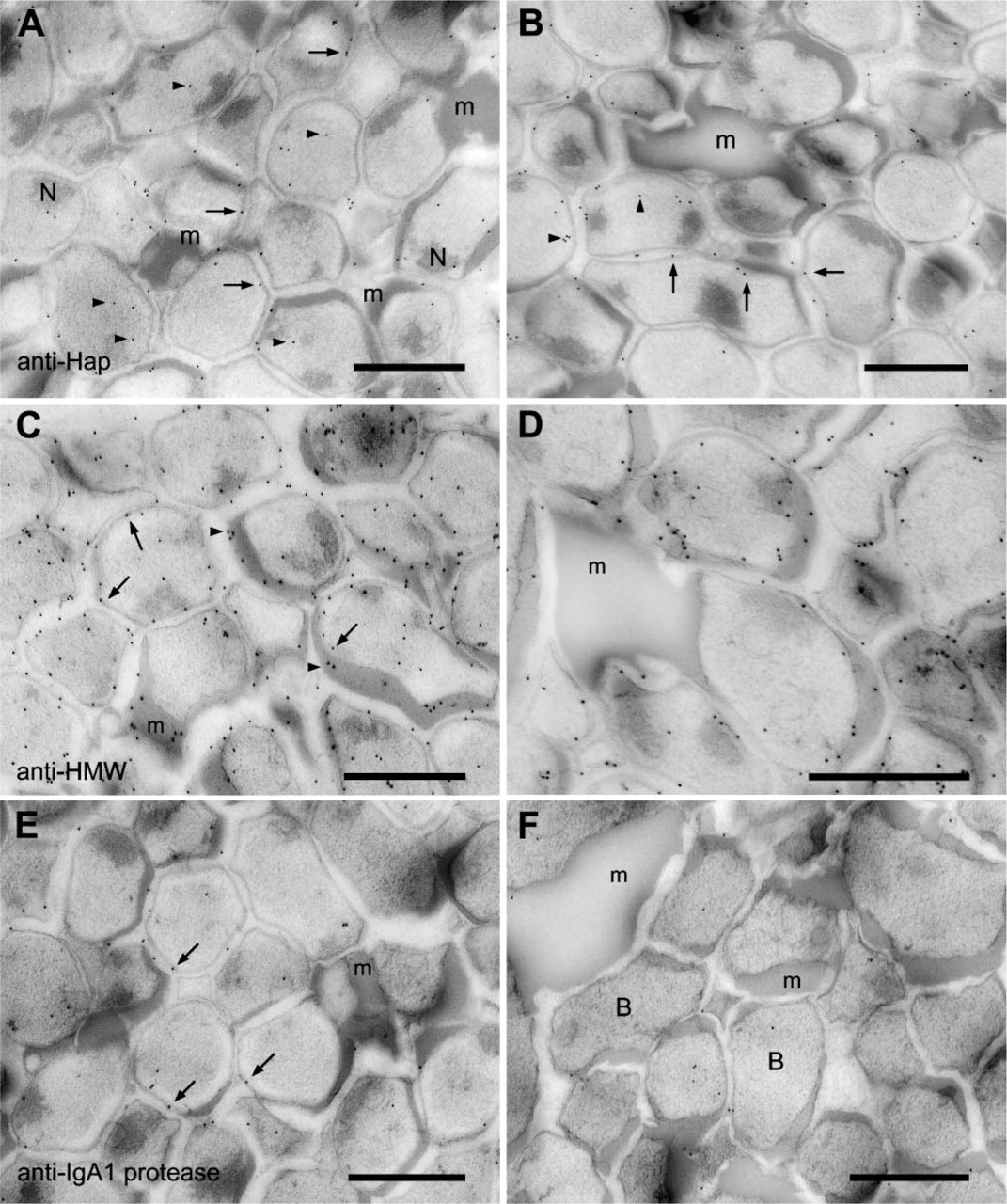
Labeling with antiserum to the Hap protein was associated with the bacteria in the biofilm (Figures 6A and 6B). The label was observed over the bacterial membrane and over the cell cytoplasm (Figures 6A and 6B). Although the ECM was labeled with the antiserum, (Figure 6B), the label was confined to material in close proximity to the bacteria (Figure 6B). The large accumulations of ECM in the biofilm did not appear to be labeled (Figures 6A and 6B). In comparison to labeling with the anti-NTHi antibody, the anti-Hap labeling was much less, suggesting a lower amount of this antigen in the biofilm.
Anti-HMW Protein Labeling
The localization of the anti-HMW1/HMW2 protein antiserum labeling was also distributed through the whole of the biofilm (Figures 6C and 6D), and the extensive labeling seen over the biofilm sections suggested the presence of a large amount of antigen. The label, although concentrated around the surface of the bacteria, was also observed in small amounts over the cell cytoplasm (Figures 6C and 6D) and over the ECM (Figures 6C and 6D). The large accumulations of ECM in the biofilm did not label with the antiserum (Figure 6D). However, the anti HMW1/HMW2 antiserum did bind to ECM that was in close association with the bacteria (Figure 6D).
Anti-IgA1 Protease Labeling
Anti-IgA1 protease labeling was present over all regions of the biofilm, with the majority of the labeling appearing to be associated with the cell membrane (Figures 6E and 6F). What was immediately obvious from initial observations was that the top region of the biofilm had more gold particles over the bacteria than did the bottom region. In the top region of the biofilm, labeling was obvious and mostly associated with the cell membrane (Figure 6E), although small amounts of label were observed associated with the cell cytoplasm (Figure 6E) and with the ECM (Figure 6E)
The density of the gold label over the bottom region of the biofilm using anti-IgA1 protease was less than that observed for the top region. Many of the biofilm bacteria appeared to have no label associated with them, even across large areas of the sections examined (data not shown). However, within the labeled regions, the gold labeling pattern, although at a lower density when compared with the top region, was similar to that observed over the top of the biofilm. The antibody labeled the cell membrane (Figure 6F), the cell cytoplasm (Figure 6F), and the ECM (Figure 6F). Again, although labeling of ECM was observed, the large aggregations of ECM that had accumulated between the bacteria did not label with the IgA1 protease antibody (Figure 6F).
Labeling of NTHi biofilms with anti-NTHi and anti-P6 antibodies. (
Quantification of Gold Labeling
To present an unbiased view of the immunolabeling distribution within the different compartments of the biofilm, we estimated the labeling density of gold particles for each labeling experiment and correlated these with the stereological parameters of compartment area and membrane length, generating estimates of gold particles per unit area of NTHi, ECM, and ES (Table 2). The image areas we sampled were approximately 10-μm2 areas, allowing for an easy estimation of the total gold labeling density over the biofilm sections for each experiment. The labeling density for each antibody was expressed as gold/1 μm2 of the complete biofilm, and these values (Table 2) allowed us to compare the labeling density produced by the different antibodies over the biofilm, thus providing a rough estimate of the amount of each antigen present. To further compare antigen amounts, we also estimated the gold labeling density per unit length of NTHi membrane (Table 2). These values give an indication of the relative amounts of antigen present over the membrane surface for each antigen.
Anti-lipooligosaccharide (LOS) labeling of NTHi biofilms. (
The Hap protein-labeling pattern obtained by quantifying the immunogold signal (Table 2) confirmed the initial observations we made of the labeled sections. The quantified result showed labeling of the bacteria and the ECM. Interestingly, in our initial observations, the large areas of unlabeled ECM, and anti-Hap protein labeling only over the ECM that was in close proximity to the bacteria, gave the false impression that the Hap protein was associated more with the NTHi. However, the quantitative results showed that Hap protein label was slightly higher over the ECM (19.3 gold/1 μm 2) than over the NTHi profiles (14.6 gold/1 μm 2). The labeling density of the anti-Hap protein antibody was much lower than that observed for the anti-NTHi antibody.
The anti-HMW1/HMW2 protein antisera produced the most labeling over the biofilm. This was demonstrated by the high labeling density on the total biofilm as well as by the labeling densities over the NTHi, the NTHi membranes, and the ECM (Table 2).
The HMW proteins appeared to be evenly distributed between the NTHi in the biofilm and the ECM (Table 2). By comparing the gold labeling over the whole-NTHi profiles (membrane and cytoplasm) with the labeling density observed only over the cytoplasmic regions of the NTHi, we were able to determine that proportionally fewer gold particles were associated with the NTHi cytoplasm (16.5 gold/1 μm 2) than with the NTHi membrane (by subtracting the gold associated with the cytoplasm from the gold associated with the whole-NTHi compartment; i.e., 50.2 gold - 16.5 gold = 33.7 gold), suggesting a high efficiency of secretion, with newly synthesized proteins in the cytoplasm being rapidly transported to the cell membrane. A high labeling density over the extracellular space was present.
Labeling of NTHi biofilms with anti-Hap, anti-HMW, and anti-IgA protease antibodies. (
Quantification of the gold labeling density and its distribution within the biofilm
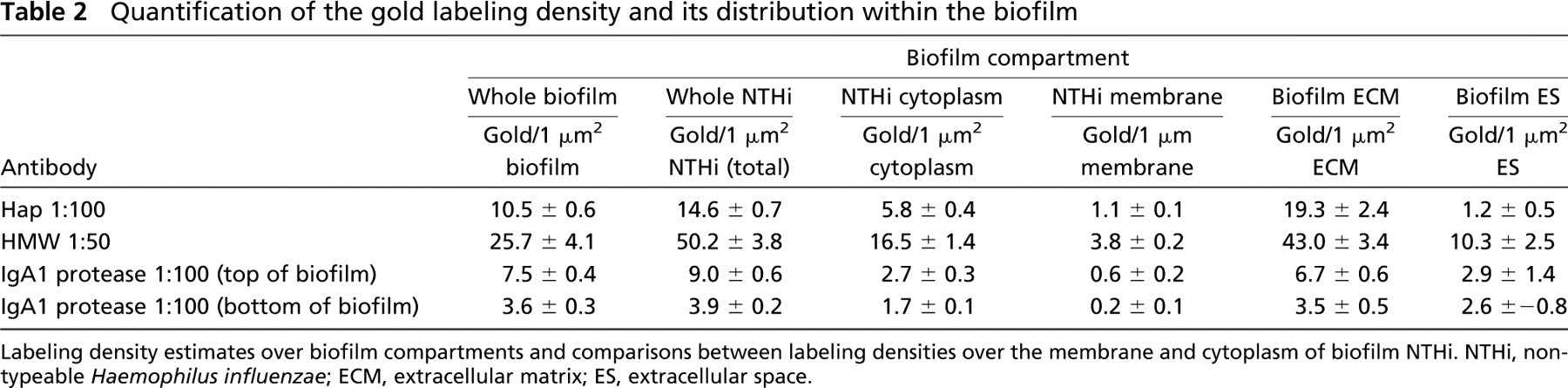
Labeling density estimates over biofilm compartments and comparisons between labeling densities over the membrane and cytoplasm of biofilm NTHi. NTHi, non-typeable Haemophilus influenzae; ECM, extracellular matrix; ES, extracellular space.
In our initial examination of labeled sections, it was apparent that the distribution of the anti-IgA1 protease labeling was uneven over the biofilm sections. In addition to a patchy distribution of label throughout the biofilm section, there also appeared to be more gold labeling in the top region of the biofilm as compared with the bottom. For our quantitative analysis, instead of following previous sampling protocols in which random images were taken from all regions through the biofilm, we sampled only the very top and the very bottom of the biofilm. Random images were taken from regions that were within 500 nm of the top or bottom edges of the biofilm (10-μm2 areas), and the labeling densities of each were compared.
The antibody labeling density of the IgA1 protease antibody over the top region of the biofilm was much higher when compared with the labeling density at the bottom. This observation was supported by our unbiased estimates of labeling density (Table 2). Although the labeling densities were lower than for the other proteins being examined, the label appeared to be specific. The total gold/1 μm2 at the top of the biofilm (7.5 gold/1 μm2) was almost double that observed at the bottom of the biofilm (3.6 gold/1 μm2), suggesting the presence of a concentration gradient through the biofilm.
There were also differences in distribution of anti-IgA1 protease antibody at the top and bottom of the biofilm. In the top region of the biofilm, more of the antibody associated with the NTHi (9.0 gold/1 μm2) than with the ECM (6.7 gold/1 μm2). At the bottom of the biofilm, the label appeared to be more evenly distributed between the NTHi and the ECM (3.9 and 3.5 gold/1 μm2, respectively). There was little difference in the extracellular space labeling at the top and bottom of the biofilm.
In this study, antibody dilutions were determined using an iterative process of labeling experiments and assessing the levels of nonspecific binding. In this way, we were able to determine the most concentrated dilution of each antibody and the visualization probe that would produce the highest signal-to-noise ratio. In some instances, nonspecific binding, as determined by the binding observed over regions of the specimen support grid and over extracellular space in the biofilm, was not eliminated even when the antibodies were significantly diluted. In these instances, we used the most concentrated antibody dilutions that did not significantly increase background, or nonspecific, labeling observed over the support film, resin, or extracellular spaces.
We examined the biofilm sections further in a non-quantitative way and found that the anti-IgA1 protease antibody labeling was confined to specific but not easily identified regions throughout the biofilm. The relative number of the labeled regions diminished when moving from the top of the biofilm to the bottom (data not shown). There appeared to be no morphological differences among the bacteria that occupied regions of the biofilm that labeled with the anti-IgA1 protease and those regions that did not (data not shown).
Discussion
Until recently, routine specimen preparation methods for electron microscopy have not been suitable for biofilms. Chemical cross-linking followed by ethanol dehydration at ambient temperature is known to disrupt bacterial morphology and to dissociate extracellular material from biofilms (Webster et al. 2004). Rapid freezing followed by freeze substitution (Gilmour et al. 1993; Webster et al. 2004; Hunter and Beveridge 2005) is one approach that can be used to preserve biofilm morphology for examination, both in the scanning and in the transmission electron microscopes. We have used these methods to preserve bio-films formed by NTHi in vitro to enable us to determine the distribution of specific proteins within the biofilms, using immunocytochemical methods.
We first established that the structures we were immunolabeling were truly biofilms and were being formed by NTHi bacteria. For TEM, although we faced technical restrictions on the size and shape of specimens acceptable for high-pressure freezing, we aimed to produce biofilms that mimicked those that may form in the middle ear. During an infection, such biofilms colonize biotic surfaces within or on top of a mucous layer that covers epithelial cells. Newly forming biofilms have the possibility of being exposed on one surface to a non-liquid environment of humid air. Watnick and Kolter (1999) have previously shown that the morphology of biofilms formed in static culture (colony biofilms) is similar to that of biofilms formed in flow cells. Colony biofilms have thus become a useful tool for antibiotic testing (Anderl et al. 2000,2003).
The crystal violet assay established that the NTHi strain used in this study has the ability to form biofilms. An SEM examination of the biofilm material formed by NTHi on solid substrates revealed layers of bacteria embedded in ECM. The morphology of the biofilms formed on the glass surfaces was similar to that of those formed on Millipore filters, and those that had been embedded and sectioned in Lowicryl.
Applying NTHi-specific antibodies to the sections confirmed that NTHi, and not contaminating bacteria, were forming the biofilms. The anti-NTHi antibodies labeled all the bacteria we examined, confirming the presence of an axenic population of NTHi. The antibody also detected NTHi proteins in Western blots, enabling the protein profiles reacting with the antibody to be compared between those in the whole biofilm with those in the ECM. Although slight differences between the two protein preparations were present, the Western blots confirmed the presence of a large number of proteins that reacted with the anti-NTHi antibodies in both the whole biofilm and in the ECM.
Anti-P6 OMP and anti-LOS antibodies both labeled the NTHi biofilm sections. Expression of P6 OMP is preserved in NTHi biofilms, and monoclonal anti-LOS 6E4 labeling is increased when compared with planktonically grown NTHi (Murphy and Kirkham 2002).
The importance of LOS in biofilm formation by NTHi has been established in previous work (Greiner et al. 2004; Swords et al. 2004). Examination of the distribution of LOS in NTHi biofilms is therefore an important step in understanding how biofilms are formed. From our labeling experiments, it appears that the bacteria secrete LOS over the substrate on which the bacteria have attached. At the base of the biofilm, the LOS is mostly extracellular and accumulates in the ECM at the base of the biofilm. LOS was also observed to penetrate into the supporting filter at the biofilm base. Throughout the biofilm, the LOS was also associated with NTHi membranes. Although LOS is important for maintaining biofilms as they develop, perhaps by maintaining biofilm architecture, the accumulation of LOS at the biofilm base may suggest that it also plays a role in the initial attachment of bacteria to substrates. LOS binding to ear tissue may be an important pathogenesis factor (Demaria 1999). The possibility of LOS-based vaccines being used in otitis media treatments (Gu et al. 2003) suggests that the localization of LOS within NTHi biofilms will be important information for understanding how such vaccines may work.
The presence and distribution of the P6 OMP, a highly conserved protein in H. influenzae (Murphy et al. 1985,1986), is important because it is currently being considered as a basis for a vaccine (Murphy et al. 1986; Wu et al. 2005) and is able to stimulate macrophage and epithelial cells (Chen et al. 2004; Berenson et al. 2005). In the labeled biofilm sections, the anti-P6 antibody was only associated with the membranes of bacteria containing observable amounts of intracellular particulate electron-opaque material. Previous studies of cryo-fixed bacteria have demonstrated that electronopaque particles in the cell cytoplasm are mostly ribosomes (Hobot et al. 1985; Bohrmann and Kellenberger 2001). Thus, a large number of these particles within the bacterial cell would indicate a high level of protein synthesis. The P6 OMP is known to be associated with viable bacteria, so it is possible that the non-labeled cells in the NTHi biofilms could represent a population of either dormant or dead cells. Reduced amounts of electron-opaque material in the cell cytoplasm of non-labeled cells further suggests reduced metabolic levels. If dormant cells are present in NTHi biofilms and they do not express the P6 OMP on their surface, then vaccines targeting this molecule may not be effective against some of the bacteria present in the biofilm. They could then emerge from their dormant state ready to reestablish an infection.
Although the anti-P6 antibody label was confined to the bacterial membranes, there was an association between labeled membranes and the ECM. The anti-P6 label was often associated with parts of the bacterial membrane that were also in close apposition to the ECM. The exact meaning of this association is currently unknown. Immunization of animal models with purified P6 OMP resulted in the production of NTHi-specific bactericidal antibodies but did not reduce the incidence of NTHi-induced otitis media (DeMaria et al. 1996). The meaning of this result in the context of NTHi biofilm infections is not clear, but it may suggest that the P6 OMP may not play an important role in the establishment of NTHi biofilms.
Of particular interest to us when we first demonstrated an ability to preserve biofilm ultrastructure for sectioning and immunolabeling for electron microscopy was the localization of proteins involved in bacterial adhesion. Proteins such as the Hap protein and the HMW1/HMW2 proteins are adhesins known to play a role in bacterial attachment. We also examined the location of the NTHi IgA1 protease, because this protein has also been implicated in bacterial microcolony formation.
The Hap protein belongs to the autotransporter family of proteins and is synthesized as a large precursor protein with three functional domains. The N-terminal signal sequence directs the protein from the cytoplasm into the periplasm, a C-terminal 45-kDa domain inserts into the outer membrane to form a pore, and through this pore the 110-kDa N-terminal domain is translocated to the cell surface. Through a final step called autoproteolysis, the N-terminal domain, which includes a serine protease active site, cleaves the precursor to release the 110 kDa fragment from the cell surface (St Geme et al. 1994; Hendrixson et al. 1997; Henderson et al. 1998).
The Hap protein has also been proposed as a potential candidate vaccine (Liu et al. 2004). It is involved with binding to human epithelial cells in vitro (St Geme et al. 1994), binds to extracellular mammalian proteins (Fink et al. 2002), and promotes microcolony formation of the bacteria (Hendrixson and St Geme 1998; Fink et al. 2003). These properties suggest that the Hap protein may play an important role in the process of biofilm formation.
In the TEM, the amount of specific anti-Hap protein label we detected was low, considering the possible role this protein has in adherence. The specific signal was distributed between the bacteria and the ECM, with more Hap protein present in the ECM than over the NTHi. Inasmuch as the passenger domain of Hap is released extracellularly by autoproteolysis, it is not surprising to find that it is present in the ECM of the biofilm. However, the ability of the Hap protein to bind bacteria to each other to form microcolonies depends on the passenger domain being present on the NTHi membrane surface (Fink et al. 2001).
The HMW proteins also mediate binding of NTHi to macrophages and human epithelial cells (St Geme et al. 1993; Noel et al. 1994). These proteins are localized to the NTHi cell surface (Bakaletz and Barenkamp 1994), and their interaction with glycosylated proteins suggests that carbohydrate moieties within the biofilm ECM may have an important role in biofilm formation. The HMW proteins on the cell surface may have the ability to bind to extracellular glycosylated proteins to form complex cross-linking bridges with proteins on adjacent bacteria. The distribution of HMW proteins labeling over the NTHi and the ECM makes it possible to speculate any number of roles for these proteins within the biofilm. Examination of HMW proteindeficient mutants for their biofilm-forming abilities will be important if we are to assign any links between the HMW proteins and biofilms.
All human isolates of NTHi secrete IgA1 proteases (Plaut et al. 1992). They are autotransporters with significant sequence homology to the Hap protein (St Geme et al. 1994), and like the Hap protein, IgA1 protease has been reported to mediate microcolony formation (Plaut et al. 1992). The patchy labeling of the anti-IgA1 protease antibody over the biofilm sections suggests that some parts of the biofilm, where the IgA1 protease is present, may have properties that differ from those where there is no labeling. Previous observations have already suggested that some parts of the NTHi biofilms grown on support filters are more susceptible to removal when exposed to aqueous medium (Webster et al. 2004). As yet, little is known about the characteristics of the parts of the biofilm that are not washed away. However, it is possible that increased adhesion resulting from concentrations of adhesins may help to explain the local differences found within the biofilms.
Although we examined the distribution of the IgA1 protease within biofilms looking at the possibility that the protease had a role in bacterial adhesion, we are also aware of the proposed role of this enzyme as an inactivator of serum and secretory forms of mammalian IgA antibodies (Kilian et al. 1979; Plaut et al. 1992). With this in mind, it is reasonable to speculate that the increased levels of IgA1 protease in the top region of the NTHi biofilm may form a useful part of a defense mechanism for protecting against antibody attack.
In this study, we have demonstrated that fragile biofilm structures can be easily preserved and embedded in resin for subsequent sectioning and immunolabeling. We have examined biofilms formed in vitro by NTHi, an exclusively human pathogen that is a common cause of otitis media, sinusitis, pneumonia, and chronic bronchitis (Turk 1984). The increasing reports of biofilm formation by NTHi and the role of biofilms in disease suggest an important need to understand how bacteria are able to establish biofilms on epithelial cell surfaces. If we are to develop vaccines against bacterial proteins, we also need to know the composition of biofilms, as well as the subcellular distribution of antigens that are being used in vaccine development. Only then will we be able to fully appreciate the efficacy of potential vaccines.
Because this may be the first time that antibodies have been applied to sections through biofilms prepared using cryo-methods, we believe that we have revealed new information on the distribution of NTHi proteins on bacteria in a biofilm and the association of these proteins with ECM that has been shed by the bacteria.
The goals of our study were to determine whether it was possible to preserve NTHi biofilms so that they could be probed with antibodies at the ultrastructural level and to examine the distribution of antibodies directed to NTHi-specific proteins. In achieving these goals, we have obtained information on the distribution of NTHi proteins that are being considered as suitable candidates for vaccine development and on their possible accessibility for antibody attack. We have also obtained results that may reveal some of the early stages of NTHi biofilm formation, and these may stimulate questions about how the bacteria are able to maintain the biofilm phenotype. These methods for preserving biofilm ultrastructure for subsequent immunochemical analysis will enable us to continue examining the composition of NTHi biofilms. In addition, these methods may also be of value for preserving the ultrastructure of biofilms formed by other bacteria, revealing important structural and immunochemical information on biofilm formation in general.
Footnotes
Acknowledgements
This work was funded in part by the National Organization for Hearing Research (NOHR) and the Deafness Research Foundation (PW); National Institutes of Health Grant NIH R01 DE-015844, and the core facilities of the Center for Gastroenterology Research on Absorptive and Secretory Processes (GRASP), NIH Grant P30 DK-034928 (AGP); and NIH Grant RO1AI AJ-44322 to J.W.S.G. G.G. was supported by a scholarship from the Hope for Hearing Foundation of Los Angeles.
