Abstract
A retrospective study on reptile tissues presenting with granulomatous inflammation was performed to detect the possible presence of mycobacteria and chlamydiae in these lesions. Ninety cases including 48 snakes, 27 chelonians, and 15 lizards were selected. Mycobacteria were detected by Ziehl-Neelsen (ZN) staining and a broad-range polymerase chain reaction (PCR) followed by DNA sequencing. To detect chlamydiae, immunohistochemistry with monoclonal antibodies against chlamydial lipopolysaccharide (LPS) and a Chlamydiales order-specific PCR and sequencing were applied. Acid-fast bacilli were found in 14 cases (15.6%) by ZN staining and in 23 cases (25.6%) by PCR. Sequence analysis revealed the presence of Mycobacteria other than Mycobacterium tuberculosis complex (MOTT). Chlamydial LPS antigen was observed within granulomas from five samples (5.6%), whereas the PCR screen revealed 58 positive cases (64.4%). Of these, 9 cases (10%) showed 98–99% similarity to Chlamydophila (Cp.) pneumoniae and 49 cases (54.4%) displayed a high similarity (88–97%) to the newly described “Chlamydia-like” microorganisms Parachlamydia acanthamoebae and Simkania negevensis. Results from this study confirm, on the one hand, that MOTT are probably the most important infectious etiology for granulomatous inflammation in reptiles. On the other hand, they indicate that chlamydia infects reptiles and that Cp. pneumoniae should be considered an etiological agent of granulomatous lesions of reptiles. Because both MOTT and Cp. pneumoniae are human pathogens, the potential of zoonotic transmission from reptiles to humans has to be considered. In contrast, the significance of Chlamydia-like isolates remains completely open, and further studies are needed to evaluate their role.
Keywords
Infectious organisms, including bacterial, fungal, parasitic, and algal agents, are major cause of disease and mortality in reptiles. 18 Reptiles react to many of these infections by developing granulomatous inflammations that may be classified as heterophilic or histiocytic, depending on the etiology and the host's cellular response. 19,43 The heterophilic granuloma, usually induced by extracellular pathogens, is characterized by an accumulation of heterophils, whose degranulation and central necrosis appear to act as foreign bodies and stimulate a strong macrophage response. The histiocytic granuloma, caused by intracellular bacteria, appears to have a different pathogenesis. Early lesions are formed from organized collections of large foamy macrophages, with the centrally located macrophages often undergoing necrosis. 19,43 In reptiles surviving the acute infection stage, both types develop into chronic granulomas, in which epitheloid cells, lymphocytes, plasma cells, and eventually multinucleated giant cells surround the central lesion. Older granulomas are additionally walled-off within a fibrous connective tissue. 43
Mycobacteria are classical etiologic agents of granulomatous reactions in human and animal hosts. 5 The genus Mycobacterium (M.) comprises about 85 species, all sharing the characteristic morphologic features of gram-positive, aerobic, acid-alcohol fast bacteria. 22 The pathogenicity of different mycobacteria varies significantly, and for practical purposes they have been differentiated into two groups: 1) the M. tuberculosis complex (MTC), which includes M. tuberculosis, M. bovis, M. africanum, M. microti and M. canetti, and 2) Mycobacteria other than the M. tuberculosis complex (MOTT), comprising M. avium intracellulare, M. avium ssp. paratuberculosis, M. leprae, M. lepraemurium and atypical mycobacteria. 46
Spontaneous mycobacterial infections have been reported frequently in a wide variety of reptiles, including snakes, turtles, lizards, and crocodiles. 10,11,18,54 It is generally believed that mycobacteria are contracted through defects in the integument or by ingestion. After necropsy, grayish-white nodules may be observed in many organs and in the subcutis; histopathologic examination shows typical granulomatous inflammations with multinucleated giant cells, common features of these lesions. Unlike mammalian tubercles, calcification has not been observed. Mycobacterial diagnosis was performed by the acid-fast staining technique and, where possible, the species was identified by bacteriologic examination and included M. avium, M. chelonae, M. fortuitum, M. intracellulare, M. marinum, M. phlei, M. smegmatis, and M. ulcerans. 3,4,16,44,45,48 Polymerase chain reaction (PCR) has been used more recently to confirm diagnosis of mycobacteriosis in young freshwater crocodiles and in a royal python. 3,24
A second obligate intracellular microorganism, chlamydia, sporadically infects reptiles, sometimes inducing histiocytic granulomas. 31,32
Chlamydiae have a unique biphasic developmental cycle consisting of the infectious elementary body (EB) and the noninfectious reticulate body (RB). 52
Until 1999, four species were recognized within the genus, namely Chlamydia psittaci, Chlamydia pecorum, Chlamydia trachomatis, and Chlamydia pneumoniae. 52 They have recently been reclassified to allow for a more accurate identification of the bacteria infecting animals. The family Chlamydiaceae was divided into two genera, Chlamydia (C.) and Chlamydophila (Cp.), with a total of nine species. Furthermore, three new families, the Parachlamydiaceae, Simkaniaceae, and Waddliaceae, comprehending the lately discovered “Chlamydia-like” isolates were proposed. 15,27,50
Chlamydiae are widespread in warm-blooded hosts including humans, birds, cats, dogs, sheep, cattle, pigs, and koalas, causing a variety of diseases. 53 Only sporadic cases of chlamydiosis have been recorded in reptiles (C. psittaci at the time), including chameleons, turtles, tortoises, crocodiles, snakes, and iguanas. Necropsy findings of chlamydial infections in reptiles include lesions in the heart, liver, lung, spleen, and small intestine. Light microscopy examination revealed severe inflammatory changes in a variety of tissues, often characterized by formation of well-defined granulomas. Colonies of intracytoplasmic organisms morphologically compatible with chlamydia were present in sections stained either by the modified Gimenez's or by the modified Macchiavello's method. Under electron microscopic examination, inclusions within the cytoplasm of macrophages composed of a population of pleomorphic organisms compatible with the various developmental stages of Chlamydia were observed. In some cases, Chlamydia could be isolated by inoculating specific pathogen-free chicken eggs with the reptilian samples. 25,30,32,33,55 Recently, T. Bodetti and colleagues 8 collected reptile tissues where the presence of chlamydiae was analyzed by immunohistochemistry (IHC), direct immunofluorescence, and by PCR and DNA sequencing. Several chlamydial species including Chlamydophila (Cp.) pneumoniae, Cp. abortus, Cp. felis and Neochlamydia were found to be present in these hosts.
The aim of this study was to detect the presence of mycobacteria and chlamydiae in granulomatous inflammations of reptiles in a 10-year retrospective study.
Materials and Methods
Selection of cases
Archived formalin-fixed and paraffin-embedded tissue specimens from 90 reptiles (reptile Nos. 1–90; 48 snakes, 27 chelonians, and 15 lizards) necropsied at the Institute of Veterinary Pathology, University of Zurich between 1990 and 1999 and presenting granulomatous inflammation were investigated (Fig. 1). Of these, 67 were pets from private owners, 21 had been housed in zoologic collections, 1 was from a pet shop, and 1 was kept in a school.
Lung; snake, case No. 5. Multiple gray nodules are visible within the lung.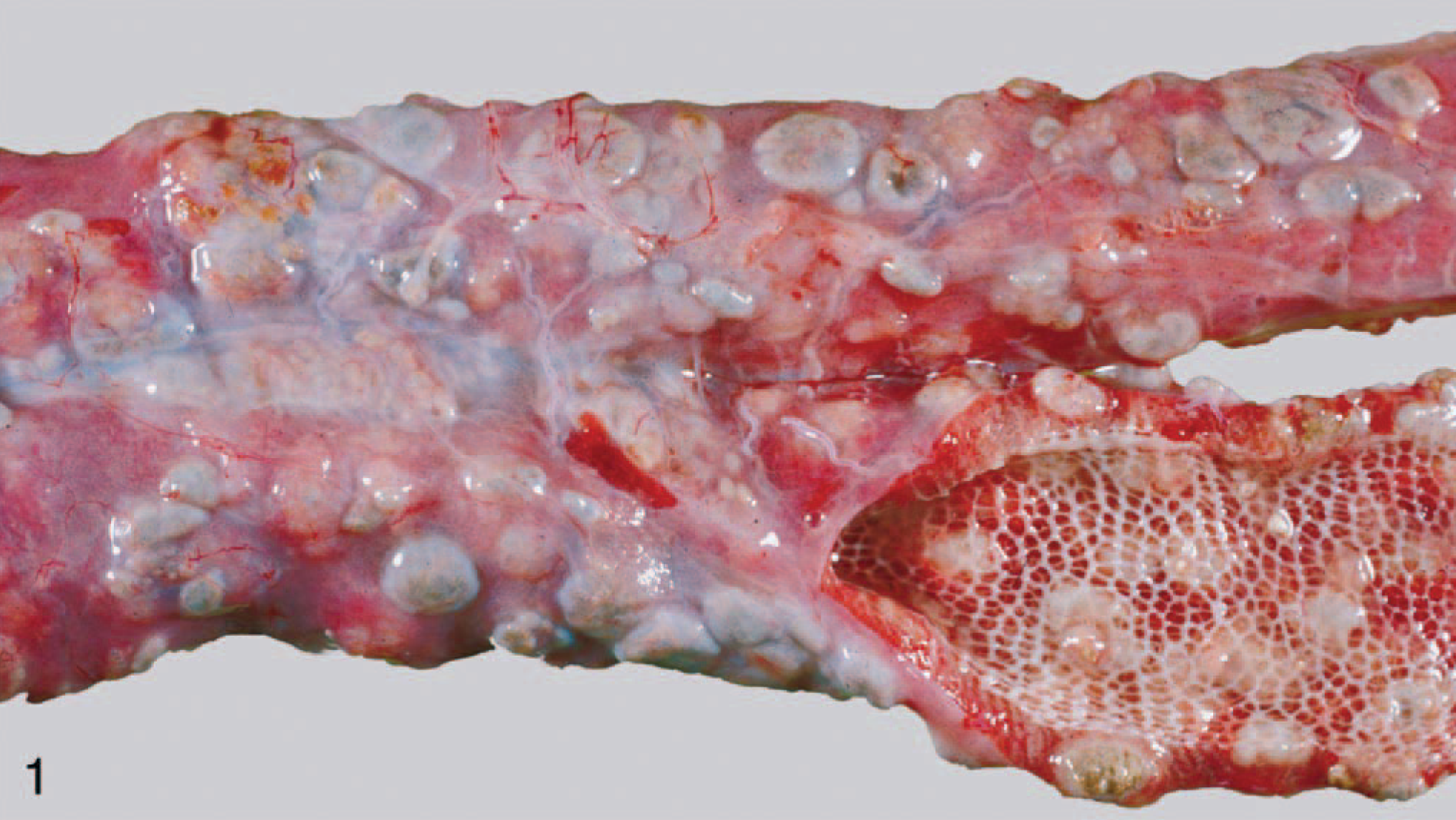
Histopathology
Histologic sections of all selected cases were stained with the hematoxylin and eosin (HE) technique and reviewed histologically to verify the presence of granulomatous inflammation and to distinguish the type of granulomatous lesion.
Screening for mycobacteria by Ziehl–Neelsen staining
Sections (2 µm) from all tissues collected were investigated for the presence of mycobacteria using the Ziehl–Neelsen (ZN) stain. Sections of intestinal tissue from a deer with Johne's disease were used as the positive control.
IHC screening for chlamydiae
Paraffin sections were investigated for the presence of chlamydial antigen using Chlamydiaceae family–specific mouse monoclonal antibodies (Abs) directed against the chlamydial lipopolysaccharide (cLPS; Clone ACI-P, Progen, Heidelberg, Germany) and the EnVision Kit (Dako ChemMate™, Glostrup, Denmark) according to the manufacturer's instructions. Briefly, paraffin sections were deparaffinated in xylene, rehydrated through graded ethanol to water, and pretreated by microwave heating (750 W for 10 minutes) for antigen retrieval. To inhibit the endogenous peroxidase activity, they were immersed in peroxidase-blocking solution for 8 minutes at room temperature (RT) and then incubated with the primary Ab at a working dilution of 1 : 200 in Ab diluent with background-reducing components for 30 minutes at RT. The sections were incubated for 30 minutes at RT with the link Ab, developed in peroxidase/3-amino-g-ethylcarbazole (AEC) substrate solution for 5 minutes at RT, counterstained with hematoxylin, and coverslipped.
A negative control of each section was performed using the Ab diluent instead of the primary Ab. Intestinal tissues from gnotobiotic piglets experimentally infected with porcine Chlamydia suis strain S45 were used as positive controls. 21
DNA extraction for PCR screening
Sections (30 µm) were cut from each paraffin block and placed in a sterile microcentrifuge tube. Paraffin was removed by extraction with 1.2 ml of xylene. After centrifugation at 13,000 × g for 5 minutes, residual xylene was removed from the tissue pellet by twice extracting with 1.2 ml absolute ethanol. The sample was centrifuged (13,000 × g, 5 minutes), and ethanol was carefully removed. DNA for PCR analysis was extracted from the tissue pellet using a commercial DNA extraction kit (DNeasy™ Tissue kit, Qiagen, Hilden, Germany).
PCR detection of mycobacterial DNA
DNA extracted from formalin-fixed and paraffin-embedded tissue samples was screened for the presence of mycobacterial DNA using a broad-range PCR assay especially developed for the detection of small DNA fragments in routine biopsies. Primers M65-Myco for (5′-GG(G/C/T)CT(G/C)AA(A/G)CG(C/T)GGCATCG-3′) and M65-Myco rev (5′-AGGCT(A/G)CCGAT(G/C)G(A/T)CTGGTC-3′) amplify a 137 bp fragment of the 65 kDa heat shock protein (hsp65) gene. 58 This gene, common to all mycobacteria, is more variable than other target genes (e.g., 16S ribosomal RNA (rRNA) gene) and is therefore adequate for the identification of genetically related species. 49 PCR reactions were performed as described by Kipar et al. 37
PCR detection of chlamydial DNA
A similar strategy was developed for the detection of chlamydial DNA. The primer pair of 23SAPF2 (5′-GAACCTGAAACCA(AG)TAGC-3′) and 23SAPR (5′-CTGGCTCATCATGCAAAAGG-3′) was designed to amplify a small 92-bp fragment of the 23S rRNA gene to account for possible DNA degradation in the formalin-fixed tissue samples while nevertheless amplifying diagnostically informative sequences for identification of members of the order Chlamydiales.
One and 10 µl of extracted DNA were added to a PCR mixture containing 1× PCR reaction buffer (2.5 mM MgCl2, 50 mM KCl and 10 mM Tris–HCl, pH 8.7; Qiagen), 200 µM of each deoxiribonucleotides (Roche, Basel, Switzerland), 1 µM of each primer (Microsynth, Balgach, Switzerland), 1× Q-Solution (Qiagen), and 1 U HotStarTaq DNA-Polymerase (Qiagen).
PCR cycling conditions consisted of an initial denaturation step at 95 C for 15 minutes, followed by 45 cycles of denaturation at 94 C for 30 seconds, primer annealing at 50 C for 30 seconds and extension at 72 C for 30 seconds, with a final extension for 5 minutes at 72 C.
Negative controls were performed using a reaction mixture with H2O substituting the template DNA. Template DNA for positive controls was obtained from formalin-fixed, paraffin-embedded tissues from gnotobiotics piglets that were infected with C. suis strain S45. 21
Sequencing
All PCR reactions were performed on a TGradient thermocycler (Biometra GmbH, Göttingen, Germany). Ten microliters of each PCR product was electrophoresed in a 3.0% TAE (40 mM Tris, 11.4% acetic acid, 1 mM ethylenediaminetetraacetic acid) agarose gel (Agarose electrophoresis grade, Gibco, Life Technologies, Carlsbad, CA) stained with ethidium bromide and observed under UV illumination. The desired fragments were excised, purified with the Qiagen's MinElute Gel Extraction Kit, and directly sequenced using the dye terminator protocol with an automated sequencer (Applied Biosystem 373A, Foster City, CA). The obtained sequences were compared with the sequences available in GenBank using the BLAST server from the National Center for Biotechnology Information. 1
Results
Histopathologic findings
In HE-stained histologic sections, heterophilic granulomas featuring masses of heterophilic granulocytes partially infiltrated with macrophages were seen in 14 cases (Fig. 2a)). Histiocytic granulomas were observed in 17 cases, with lesions being characterized by accumulation of macrophages in various stages of activation and occasional giant cells (Figs. 3a, 4a). In 58 cases, however, granulomas consisted of a necrotic center surrounded by macrophages, occasionally giant cells, lymphocytes, plasma cells, and spindle-shaped histiocytes, and were sometimes enclosed in fibrous connective tissue. Because it was impossible to discern whether macrophages or heterophils gave rise to the necrotic area, they were classified as chronic granulomas. Reptile No. 3 showed histiocytic granulomas within the lung and a chronic granulomatous pericarditis (Figs. 4a, 5a).
Small intestine; chelonian, case No. 12. Granulomatous enteritis. 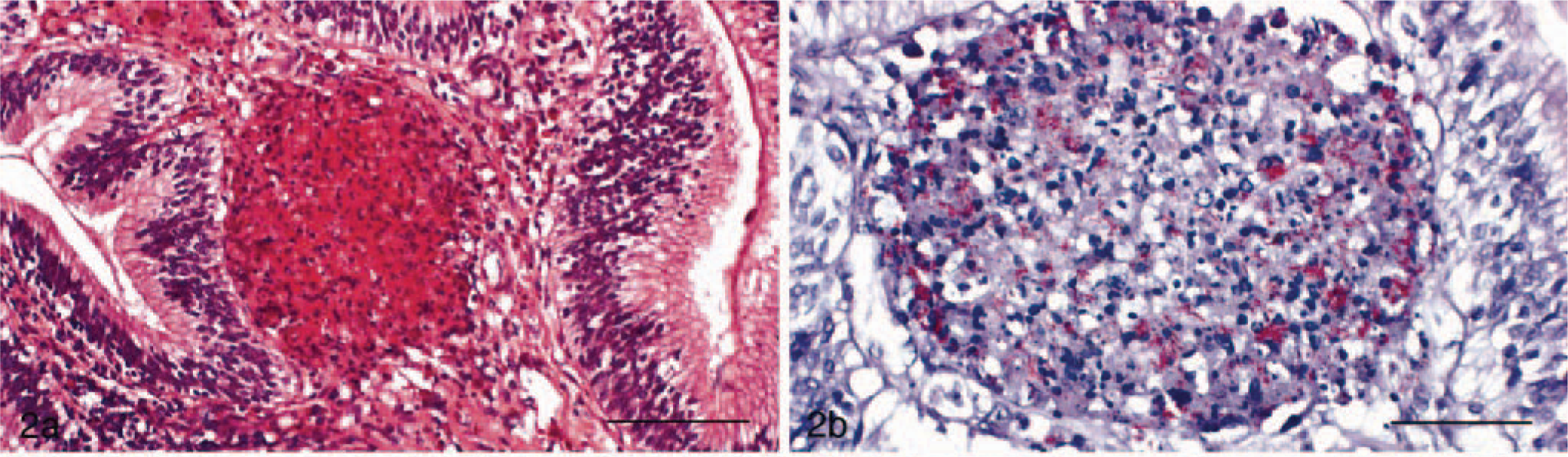
Lung; snake, case No. 27. Granulomatous pneumonia. 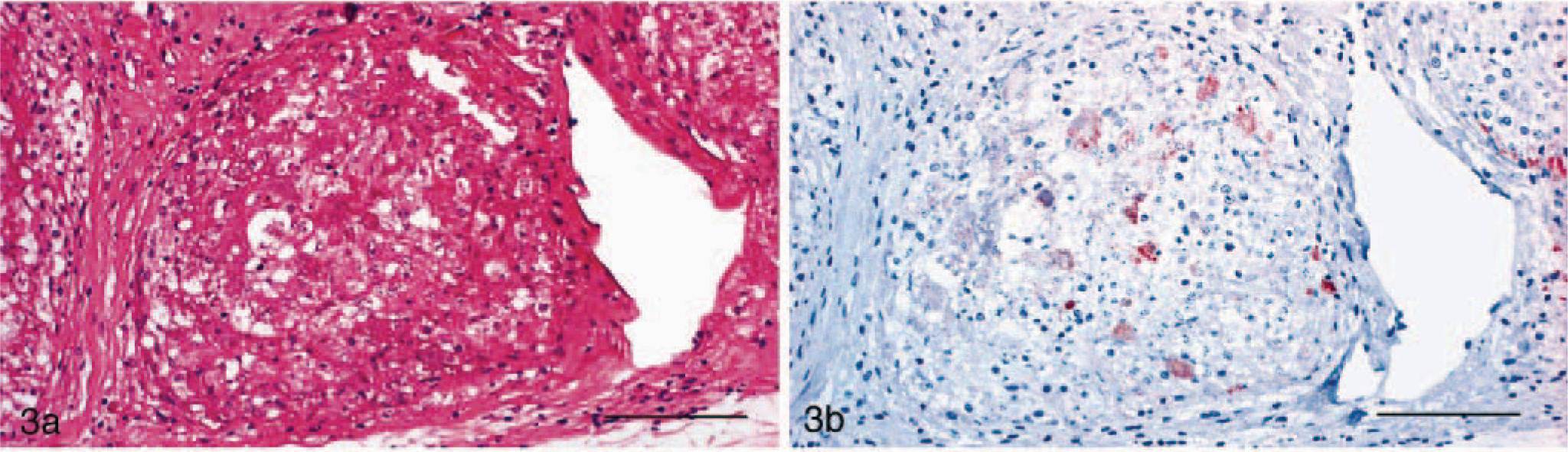
Lung; snake, case No. 3. Granulomatous pneumonia. 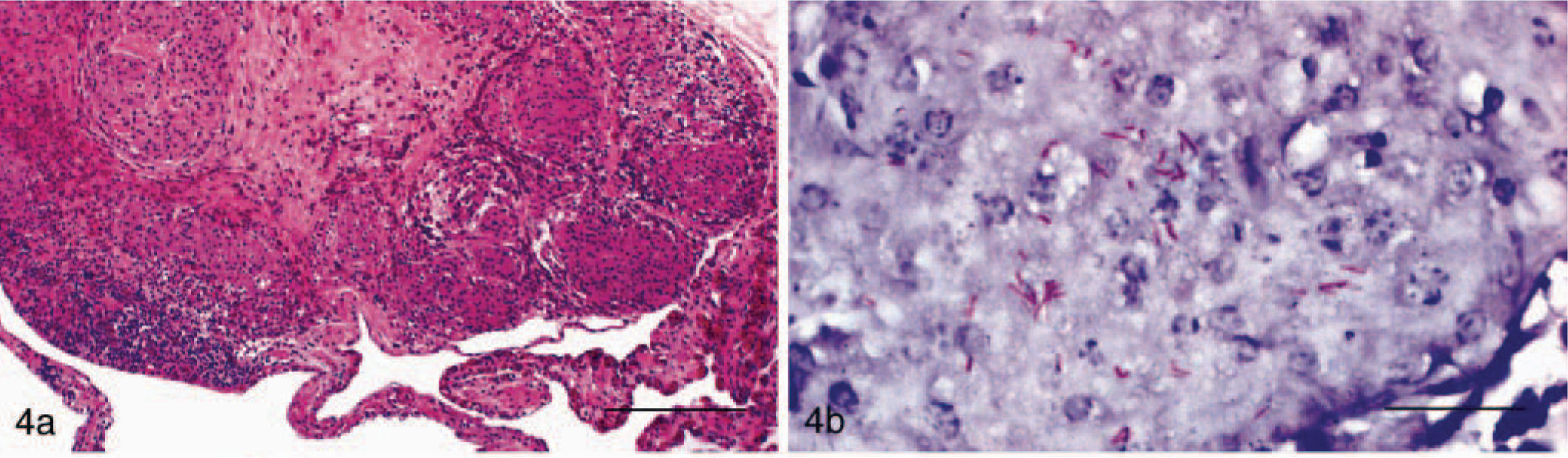
Pericardium; snake, case No. 3. Granulomatous pericarditis. 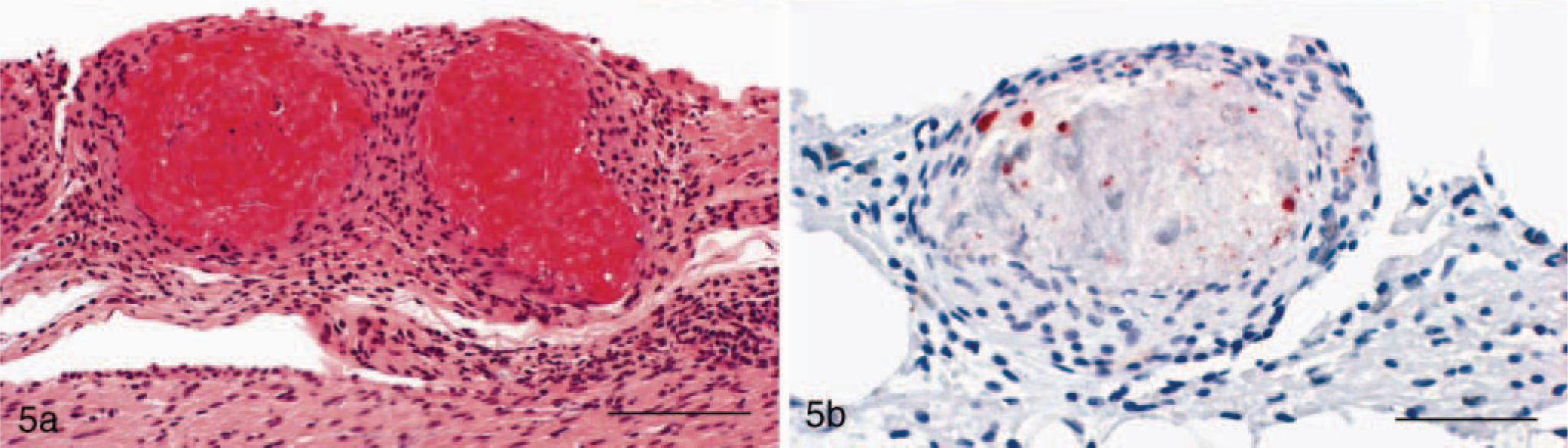
ZN and PCR detection of mycobacteria
In 14 cases (reptile Nos. 1–14), the ZN stain revealed a variable number of acid-fast organisms in the granulomatous lesions. They were present both intracellularly within the cytoplasm of macrophages and extracellularly in the necrotic centers (Figs. 2b, 4b). Mycobacteria were found in the alimentary, respiratory, and genitourinary tracts, heart, spleen, peritoneal surface, and central nervous system.
All 14 ZN-positive specimens and 12 cases without demonstrable acid-fast bacilli (reptile Nos. 15–26) yielded DNA fragments of the expected size by the broad-range mycobacteria PCR.
In an attempt to further identify infecting mycobacterial strains, PCR products of hsp65 were directly sequenced. With the exception of three cases (reptile Nos. 24–26), which were ZN negative and generated sequences belonging to microorganisms other than mycobacteria, the remaining 23 specimens showed highest similarities with hsp65 sequences of MOTT, at the amino acid level (44 amino acid residues). An unambiguous classification at the species level was determined in nine cases; sequence analysis of the remaining 14 specimens displayed 93–100% similarities with corresponding sequences of several MOTT species.
ZN stain, PCR, and sequencing results are summarized in Table 1.
Positive results of the mycobacterial screen by ZN stain, PCR, and sequence analysis.∗.
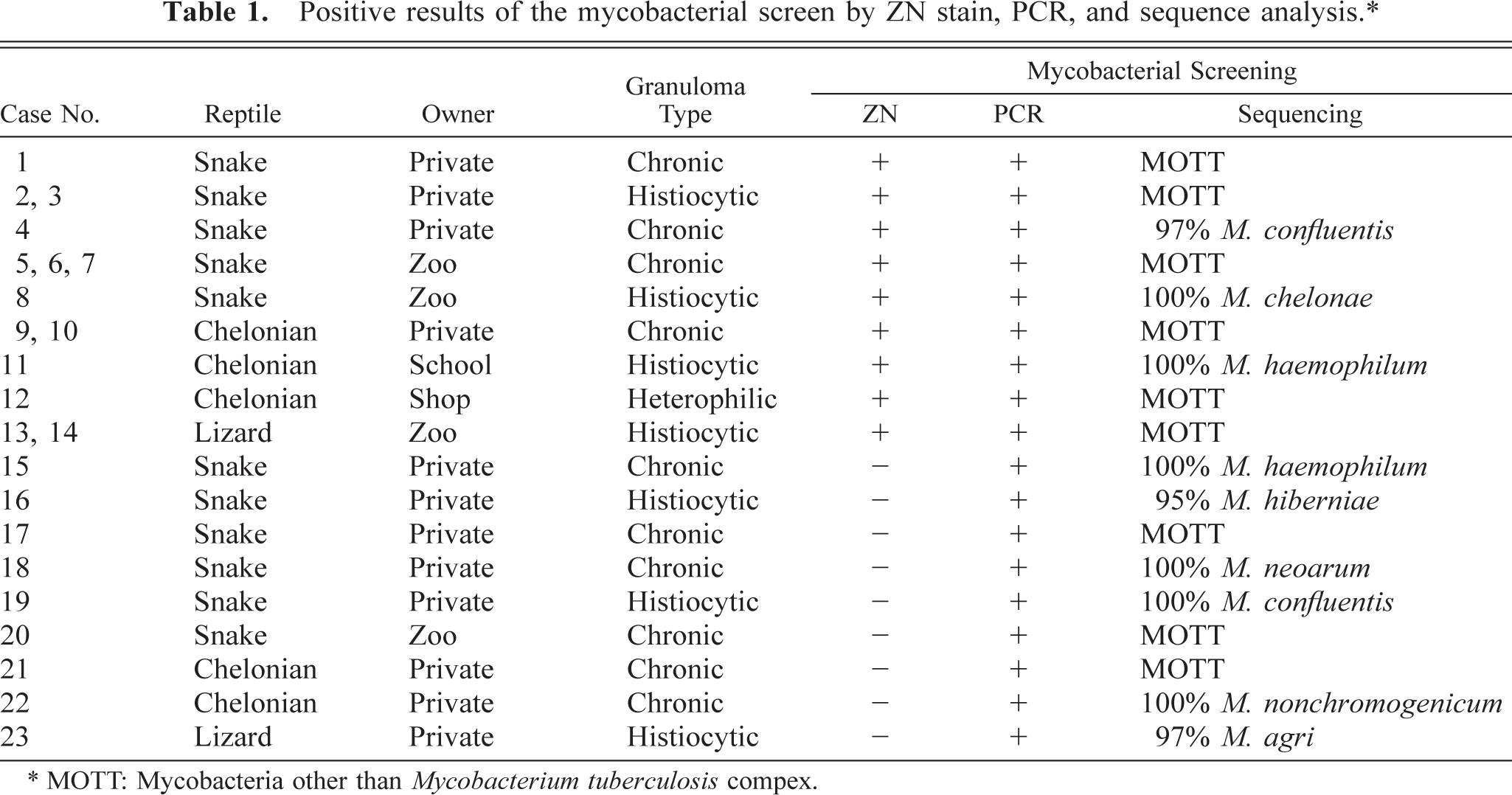
∗ MOTT: Mycobacteria other than Mycobacterium tuberculosis compex.
IHC and PCR detection of chlamydiae
Immunohistochemically, a positive staining for the cLPS antigen was observed within granulomatous tissue samples from lung, pericardium, kidney, and liver derived from five reptiles (reptile Nos. 3, 19, 27–29). Except for reptile No. 28, where in only a few cells chlamydia could be detected, strong staining of chlamydial inclusions were observed, principally localized within macrophages and in necrotic areas (Figs. 3b, 5b).
All immunohistochemically positive tissue samples were later confirmed as positive using the Chlamydiales order–specific PCR assay. Direct sequencing of the 23S rRNA PCR products of all five cases revealed sequences with 99% identity to Cp. pneumoniae. An additional 53 immunohistochemically negative specimens were found positive after PCR screening for chlamydial DNA, and their identity was determined by direct DNA sequencing. Four samples were found positive for Cp. pneumoniae (98–99% identical). Surprisingly, comparative sequence analysis of the remaining 49 PCR-positive specimens showed high similarity values (88–97%) with 23S rRNA sequences of members of the newly described Chlamydia-like families Parachlamydiaceae and Simkaniaceae. In particular, on-line BLAST searches of GenBank for similar sequences revealed that these 49 isolates were 88–97% similar with either Parachlamydia acanthamoebae or Simkania negevensis. Because a definitive identification was not possible, these isolates are referred to as Chlamydia-like. Immunohistologic, PCR, and sequencing results are summarized in Tables 2, 3.
Positive results of the chlamydial screen by cLPS IHC, PCR, and sequence analysis related to Chlamydophila pneumoniae.
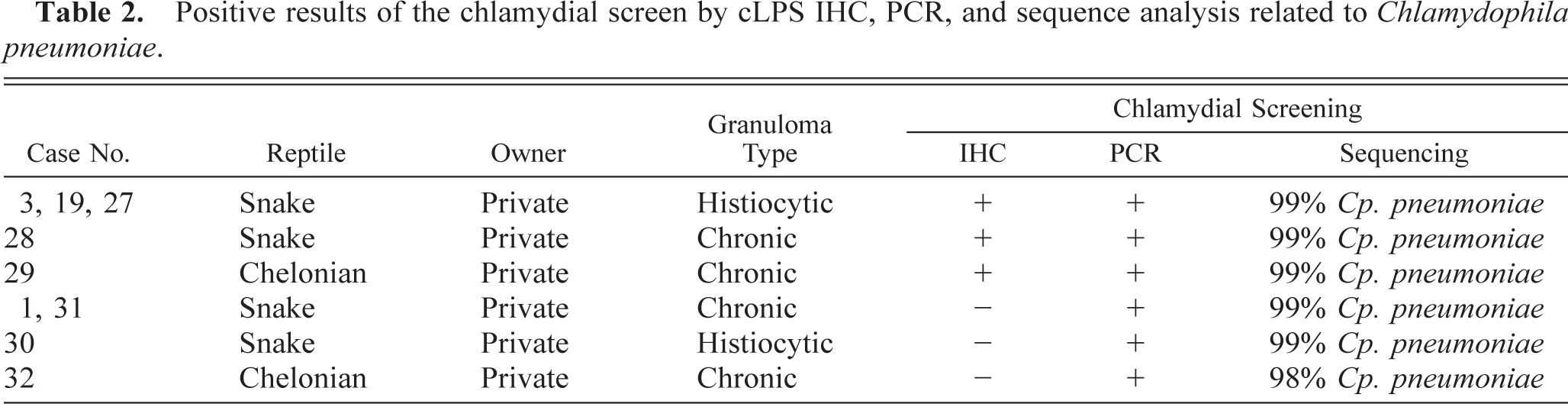
Positive results of the chlamydial screen by cLPS IHC, PCR, and sequence analysis related to “Chlamydia-like” isolates.
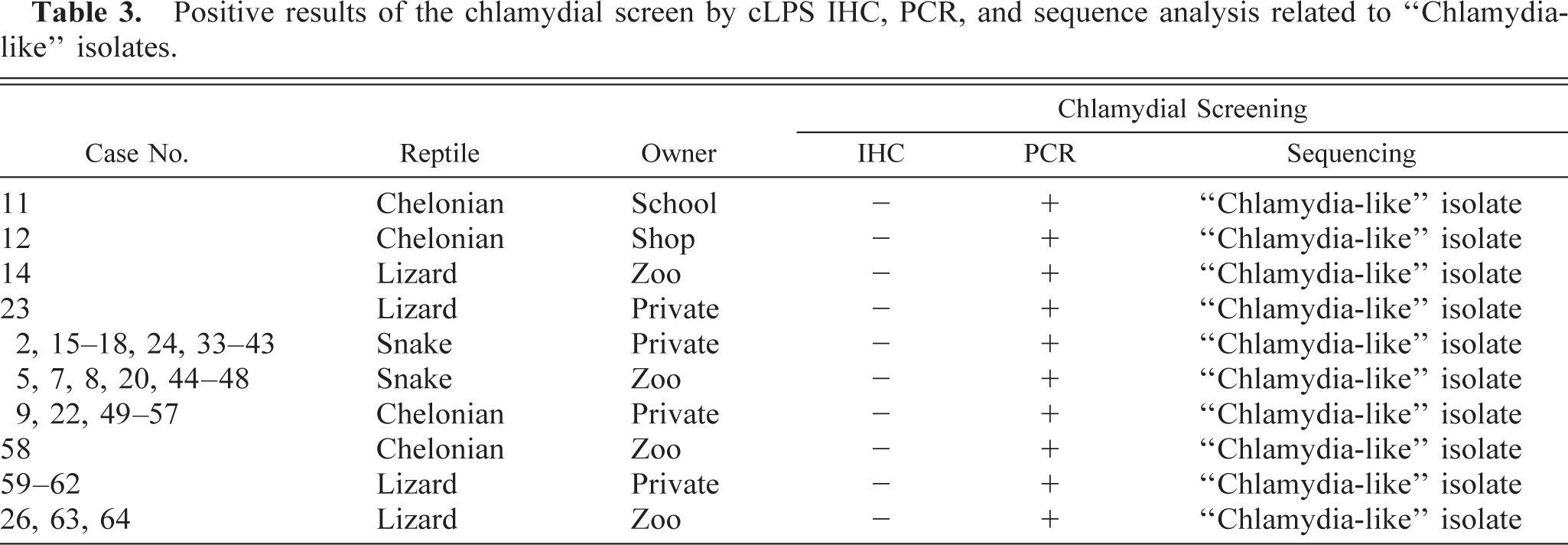
Mixed and dual infections
In two cases (reptile Nos. 1 and 19), both Cp. pneumoniae and MOTT were detected either by staining methods and PCR or by PCR alone. Reptile No. 3, however, presented a dual infection: the granulomatous pneumonia was caused by MOTT, whereas Cp. pneumoniae induced the pericardial granulomatous lesions (Table 4, Figs. 4, 5).
Dual and mixed infections.∗.
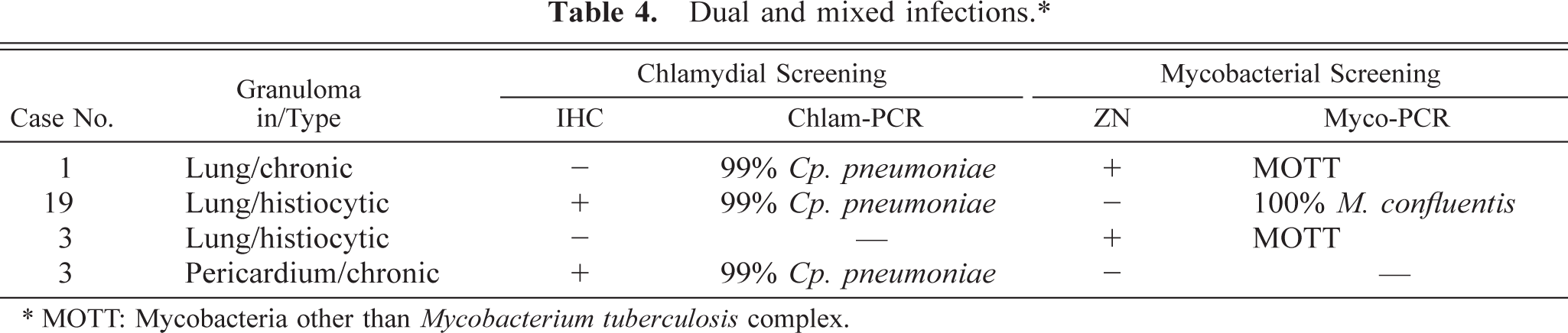

∗ MOTT: Mycobacteria other than Mycobacterium tuberculosis complex.
In a total of 27 cases (reptile Nos. 25, 65–90) neither mycobacteria nor chlamydiae could be detected.
Discussion
In this study, we investigated the presence of mycobacteria and chlamydiae in granulomatous inflammation of reptiles. For the detection of these two intracellular microorganisms a similar methodology was developed and applied: first a staining procedure (ZN and IHC, respectively) followed by broad-range PCR assays and gene sequencing. This proved to be successful.
The histologic classification of granulomatous inflammation was rather unproductive because in only 31 cases a clear distinction between heterophilic and histiocytic granulomas was possible and represented only the first step for etiologic diagnosis. Moreover, the detection of acid-fast bacilli within heterophilic granulomas (reptile No. 12) demonstrates that grouping these etiologic agents (i.e., extracellular and intracellular bacteria with heterophilic and histiocytic granulomas, respectively) by lesion type is not always correct. In contrast, chronic granulomas involve the wide spectrum of etiologies. In both cases, the necessity of specific investigations is clearly required.
The microscopic identification of acid-fast bacilli and chlamydiae by ZN and IHC, respectively, was less sensitive than PCR, because it requires a large number of organisms in the sample. 5 In addition, a nonhomogeneous distribution of microorganisms within the granulomatous inflammation must be taken into account. 12,41 Regarding the chlamydial IHC, a third reason could be a possible reduction in steady-state levels of outer membrane constituents, including LPS in persistent chlamydial infections. 40 In contrast, the nondetection of the 49 Chlamydia-like cases was an expected result because Chlamydia-like organisms appear not to be recognized by monoclonal Abs specific for the LPS of the Chlamydiaceae. 15
Although formalin-fixed and paraffin-embedded tissues represent an invaluable resource for research, a major problem is the physical and chemical alteration of DNA during tissue processing, which limits the length of PCR products that can be amplified. 7 Therefore, for a study aiming to maximally detect the presence of microorganisms, we suggest the use or the development of PCR assays that amplify relatively short DNA fragments.
The disadvantages of such a strategy for the mycobacteria were the difficulty in clearly distinguishing between members of the MOTT and the amplification of DNA belonging to organisms other than mycobacteria. For the chlamydiae, the limited length of the amplified 23S rRNA gene fragment did not allow comparative studies and phylogenetic analysis with corresponding Cp. pneumoniae sequences detected in human, horse, koala, amphibian, and reptile hosts.
Within these limits, we could demonstrate the presence of MOTT in a total of 23 out of 90 reptiles (25.56%). Therefore, although considered an uncommon and sporadic disease of reptiles by some authors, they represent probably the most important infectious etiology for granulomatous inflammation. 10,24,44
All 90 reptiles investigated involved either zoologic collections or private owners. These captive reptiles are frequently mismanaged and maintained in suboptimal environmental conditions, circumstances that may repress the immune system and favor the invasion of opportunistic pathogens. MOTT normally live in soil and aquatic habitats, and most of them are saprophytes. However, some species are opportunistic pathogens and can infect human and animal hosts with underlying immunodeficiencies. 29 Contaminated soil, water, and food are the most likely sources of infection for immunosuppressed reptiles, and infected reptiles themselves could represent an additional reservoir for humans.
The results of the chlamydial screen can be subdivided into two parts: the detection of Chlamydia-related 23S rRNA sequences in 49 reptile samples and the 9 cases of Cp. pneumoniae infection. The high prevalence of Chlamydia-like organisms found in reptile samples most likely reflects opportunistic infection by an environmental pathogen.
Chlamydia-like organisms are obligate intracellular bacteria with developmental stages similar to RBs and EBs of the known Chlamydiaceae, but they are morphologically and antigenetically different. 34–36 The organisms are distributed widely across diverse environments. 13,26 Ossewaarde and Maijer detected more than 100 Chlamydia-like sequences in human, animal, and environmental specimens, including distilled water, phosphate-buffered saline, DNA isolation, and PCR controls (results presented at the International Chlamydia Conference at Antalya, Turkey in June 2002). Although the first described Chlamydia-like organisms, S. negevensis and P. acanthamoebae, were considered environmental contaminants, serologic studies and the molecular detection of Chlamydia-like rRNA gene sequences in clinical samples suggest their possible association with respiratory tract diseases and even arteriosclerosis. 2,14,20,35,42 Whether Chlamydia-like organisms are involved in the granuloma formation of reptiles remains unknown. The use of a specific staining procedure would be of great value to determine their localization within tissue specimens. 17 Nevertheless, further studies are needed to clarify their potential role as newly emerging pathogens.
Apart from these environmental Chlamydia-like organisms, the primary aim of this study was the detection of pathogenic Chlamydia species in granulomatous inflammations of reptiles. From a veterinary pathology perspective, the nine cases of Cp. pneumoniae clearly demonstrate that chlamydial infection occurs in reptiles and its prevalence may be greater than indicated in the literature. In addition, the immunohistochemical association of cLPS antigen to specific lesions such as granulomas confirms the pathogenicity of Cp. pneumoniae for reptile hosts. This is a significant finding for our understanding of reptiles diseases and for the extension of the differential diagnosis of granulomatous inflammation to include also Cp. pneumoniae. The source of infection remains unknown. Chlamydiosis in reptiles and amphibians was frequently presumed to be caused by C. psittaci and consequently a transmission from feral birds was proposed. 25,32,33 The application of PCR and gene sequencing to detect and identify the chlamydial species infecting poikilothermic animals demonstrated the presence of Cp. pneumoniae, a result that raises interesting questions regarding their origin. 8 There could exist either an independent feral reservoir or a separate biovar of Cp. pneumoniae; it is also possible that humans could be the reservoir of infection. An additional interesting finding was our detection of both MOTT and Cp. pneumoniae within the same reptile and in two cases within the same tissue, indicating the presence of dual and mixed infections. We believe that the pericardial granuloma in reptile No. 3, induced by Cp. pneumoniae, was older than those observed in the lung, suggesting that the chlamydial infection was primary, lowering the snake's defense mechanisms and facilitating the mycobacterial infection. In contrast, in reptile Nos. 1 and 19, it was difficult do predict the progression of infection.
The presence of Cp. pneumoniae isolates in reptiles is also noteworthy from a purely microbiologic view. Originally, the host range of Cp. pneumoniae was believed to be limited to humans. 38 Our results, along with the detection of Cp. pneumoniae in horse, koalas, amphibians, and other reptiles, provide convincing evidence for a wide host range. 6,8,9,28,47,51,56
The presence of Cp. pneumoniae strains in reptiles is important from a human perspective because Cp. pneumoniae is a common cause of pneumonia and bronchitis throughout the world and has been recently associated with several chronic diseases, including coronary heart disease, Alzheimer's disease, and multiple sclerosis. 23,38,57 In our study and other studies the animal isolates of Cp. pneumoniae revealed few differences at the DNA level compared with the human biovar. 6,28,47,51,56 Although transmission between humans is believed to occur from person to person through respiratory secretions, 39 until more is known about the epidemiology, pathogenicity, and transmission of the Cp. pneumoniae strains recovered from captive and wild populations of cold-blooded animals, their zoonotic potential must be considered. Moreover, because all nine reptiles infected with Cp. pneumoniae belonged to private owners and because of the increasing ownership of reptiles as pets, caution must be exercised in the direct handling of these animals, especially by immunecompromised individuals.
Footnotes
Acknowledgements
We are grateful to the laboratory technical staff of the Institute of Veterinary Pathology, University of Zurich, for technical assistance. This work represents part of the requirements to obtain the degree of Dr. Med. Vet. from the Veterinary Faculty of the University of Zurich (G. Soldati).
