Abstract
Cryptosporidiosis is a well-known gastrointestinal disease of snakes and lizards. In the current study, 672 samples (feces and/or gastric contents or regurgitated food items) of various snakes and lizards were examined for the presence of cryptosporidia by polymerase chain reaction (PCR) assay targeting a part of the 18S ribosomal RNA gene. A consecutive sequencing reaction was used to identify the cryptosporidian species present in PCR-positive samples. Cryptosporidium varanii (saurophilum) was detected in 17 out of 106 (16%) samples from corn snakes (Pantherophis guttatus) and in 32 out of 462 (7%) samples from leopard geckos (Eublepharis macularius). Cryptosporidium serpentis was found in 8 out of 462 (2%) leopard gecko samples, but in no other reptile. The Cryptosporidium sp. “lizard genotype” was present in 1 leopard gecko sample, and 1 sample from a corn snake showed a single nucleotide mismatch to this genotype. Pseudoparasitic cryptosporidian species were identified in 5 out of 174 (3%) ophidian samples, but not in lizards. Other sequences did not show complete similarity to previously published Cryptosporidium sequences. The results stress the importance for diagnostic methods to be specific for Cryptosporidium species especially in snakes and show a relatively high prevalence of C. varanii in leopard geckos and corn snakes.
Introduction
Cryptosporidiosis is an important disease in reptiles, especially snakes and lizards. To date, 2 cryptosporidian species (Cryptosporidium serpentis and Cryptosporidium varanii (saurophilum)) are acknowledged as valid species in reptiles and have been associated with disease of the gastrointestinal tract especially in snakes and lizards. 4,5,18 Several poorly defined genotypes exist, mostly without a direct link to clinical signs. 6
Gastrointestinal passage of Cryptosporidium oocysts has been shown experimentally, 9 and, especially in snakes, the discrimination of reptilian pathogenic species from those that are ingested with the prey (pseudoparasites) and excreted as intact oocysts is important for correct diagnosis. 14,16 Several different methods for the detection of cryptosporidia have been established. Many of these methods diagnose cryptosporidia at the genus level and do not allow a distinction of different species. 8,10,16 Other methods used for the differentiation of certain Cryptosporidium species usually comprise important human pathogenic, or possibly zoonotic, pseudoparasitic species and genotypes, but not all reptilian pathogenic types. 11,13,21
Polymerase chain reaction (PCR) assays for the detection and subsequent sequencing of Cryptosporidium species have been described, usually based on a relatively long amplicon. 1,21 The primers of the present study have originally been designed for identification of cryptosporidian species from formalin-fixed, paraffin-embedded tissue samples. During such treatment, the DNA strands break into fragments, and studies have shown that amplicons of more than 300 base pairs (bp) lead to a significantly decreased sensitivity of the PCR assay. 7,19 Thus, the chosen amplicon, which had a length of approximately 122 bp that targeted a part of the 18S ribosomal (r)RNA gene, shows a sufficient number of nucleotide differences to allow identification of cryptosporidian species through sequencing. Due to the high demand of specific identification of cryptosporidian species especially in reptilian samples, the assay was then modified for live animal diagnostics.
The name C. varanii has been proposed as a substitute for C. saurophilum, since the partial 18S rRNA and actin loci of the original C. varanii isolates have been reported to be 100% identical to the corresponding sequence of C. saurophilum. 15 However, the sequences of C. varanii isolates are not available in GenBank databases. Thus, in the current report, the term C. varanii (saurophilum) is used, since the identification of the cryptosporidian species was based on GenBank sequences.
Materials and methods
Samples
A total of 672 samples were examined, and consisted of reptilian fecal material and/or gastric washings, and, in a few cases, of regurgitated feed items. Samples were submitted by reptile owners or veterinarians with a declaration of being single or pooled samples taken from 1 or more animals at 1 or several points of time. In the majority of cases, the genus or species of the reptile was given. Most animals were from the genera Eublepharis (all leopard geckos, Eublepharis macularius) and Pantherophis (all corn snakes, Pantherophis guttatus). In 186 out of 672 (28%) samples, information about the clinical history and/or direct (from the same enclosure) or possible indirect (from the same collection) contact with Cryptosporidium-positive reptiles was given. Sample material was usually stored for several days and sent in for examination without cooling. To determine the detection limit of the protocol defined, the numbers of isolated and heat-inactivated Cryptosporidium parvum oocysts were treated as the above samples during DNA extraction and PCR.
Extraction
Samples of DNA were extracted using a commercial kit. a Per manufacturer’s advice, an additional step of temperature shock was added before the extraction to break up the oocyst walls. During the procedure, samples were placed from a hot water bath of 95°C onto dry ice for 3 consecutive times.
Polymerase chain reaction
Primers were designed to amplify a portion of approximately 122 bp of the 18S rRNA gene. According to alignments, this portion shows variability in different Cryptosporidium species. Primer sequences were 5’-CTCGTAGTTGGATTTCTGTT-3’ (forward) and 5’-TAAGCACTCTAATTTTCTCA-3’ (reverse). Using the basic local alignment search tool (BLAST; http://www.ncbi.nlm.nih.gov/blast/Blast.cgi), each primer had a high similarity to various organisms besides cryptosporidia; however, only cryptosporidia have 100% similarity to both primers when examined as primer pair.
For the PCR reaction, 11 µl of laboratory grade water, 10 µl of master mix (2.5×), b 1 µl of forward and reverse primer each (final concentration 0.4 pmol/µl), c and 2 µl of 1:5 diluted DNA template were used. Thermocycling was performed with the following temperature program: 94°C for 2 min, 40 cycles of denaturation at 94°C for 30 sec, annealing at 55°C for 30 sec, and extension at 72°C for 1 min, followed by 72°C for 10 min. d The PCR products were evaluated by electrophoresis on a 2% agarose gel, stained with ethidium bromide, and photographed under ultraviolet light. The presence of a band of the expected length was interpreted as a positive result. Each PCR run included a negative and positive control. Laboratory grade water was used as negative control instead of DNA template, and DNA template taken from a geckonid fecal sample positive for C. varanii (saurophilum) was used as positive control.
Sequencing
Samples with a positive PCR result were sequenced in both directions. One µl of the purified e and 1:5 diluted PCR amplicon, 5 µl of a commercial sequencing master mix, f 1 µl of forward or reverse primer (final concentration 0.2 pmol/µl), c and 13 µl of laboratory grade water were used for each sequencing reaction. The thermocycling program consisted of 30 cycles of denaturation at 96°C for 30 sec, annealing at 50°C for 10 sec, and extension at 60°C for 4 min. DNA purification g after sequencing was followed by a denaturation step for 2 min at 95°C. The sequence analysis was conducted with a genetic analyzer. h,3 The obtained forward and reverse nucleotide sequences were matched and analyzed with a BLAST search and compared to GenBank sequences.
Results
Polymerase chain reaction
The detection limit of isolated C. parvum oocysts was 200 oocysts per sample, which corresponds to the DNA content of approximately 1 oocyst in the template of 1 PCR reaction. Of the 462 fecal samples from leopard geckos, 74 (16%) were positive by PCR. Similar results were achieved for other lizards (6/36 samples, 17%) and snakes excluding corn snakes (12/68 samples, 18%). The proportion of positive results was somewhat larger in corn snakes, where 27 (25%) of 106 samples yielded a positive PCR reaction (Table 1).
Number of examined fecal or gastric samples from different lizard and snake genera, and the results of a polymerase chain reaction assay for the presence of Cryptosporidium sp., and subsequent Cryptosporidium species identification through partial sequencing of the 18S ribosomal RNA gene.*
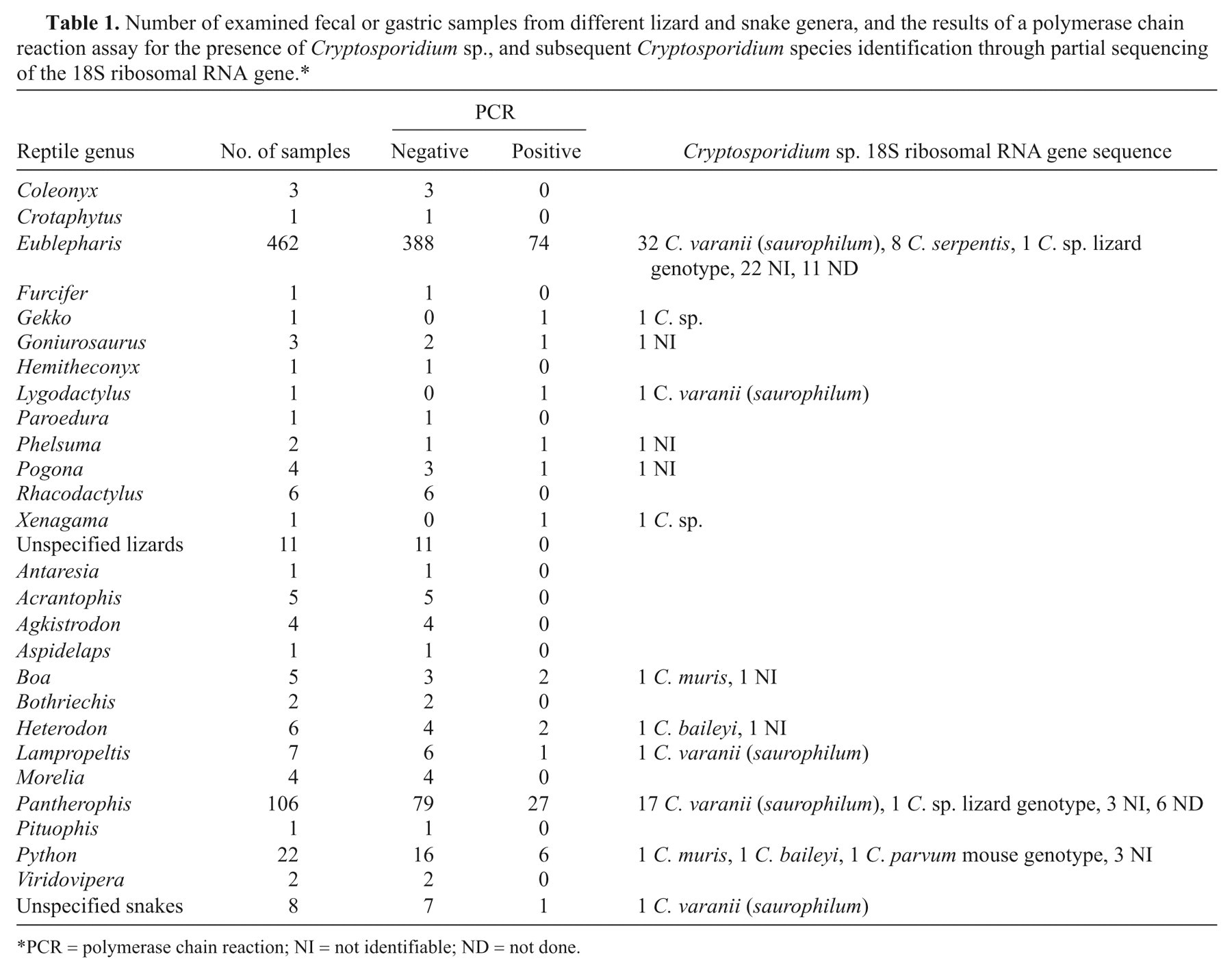
PCR = polymerase chain reaction; NI = not identifiable; ND = not done.
Sequencing
The results from partial sequencing of the 18S rRNA gene are shown in Table 1. Unambiguous sequencing data indicating the presence of reptilian-specific Cryptosporidium species included the following: sequences identical within the region sequenced to C. varanii (saurophilum) were found in 32 out of 462 (7%) samples from leopard geckos, 17 out of 106 (16%) corn snakes, 2 out of 68 (3%) other snakes (1 Lampropeltis sp. and 1 unspecified snake), and 1 out of 36 (3%) other lizards (Lygodactylus sp.); 1 sample from Xenagama sp. resulted in a sequence with 97% similarity to C. varanii (saurophilum) sequences. Sequences identical to the correlating region of C. serpentis sequences were found in 8 out of 462 (2%) leopard gecko samples, but in no other lizard or snake. One leopard gecko sample showed identity of the sequenced region with a Cryptosporidium sp. previously detected in a leopard gecko and designated “lizard genotype”. 22 Also, 1 sample from a corn snake showed a 99% similarity to this Cryptosporidium sp. “lizard genotype” sequence.
One sequence with a 98% similarity to sequences of a Cryptosporidium species from 2 cockatiels (Nymphicus hollandicus) designated “avian genotype V” 2 was found in a Gekko sp. sample. Other sequences were completely identical within the region sequenced to designated nonreptilian or pseudoparasitic Cryptosporidium species, and these sequences were found in 5 out of 174 (3%) of ophidian samples, but not in lizards: 2 samples were positive for Cryptosporidium baileyi, 2 for Cryptosporidium muris, and 1 for C. parvum mouse genotype.
Thirty-three out of 672 (5%) samples from both lizards and snakes resulted in a chromatogram, which showed no single distinct sequence or no significant similarity to GenBank sequences. Furthermore, in 11 out of 462 (2%) leopard gecko samples and 6 out of 106 (6%) corn snake samples, sequence analysis was not done. Sequences of the different cryptosporidian species and genotypes detected in the current study are shown in Figure 1 and have been submitted to GenBank with the accession numbers HM750250–HM750259.
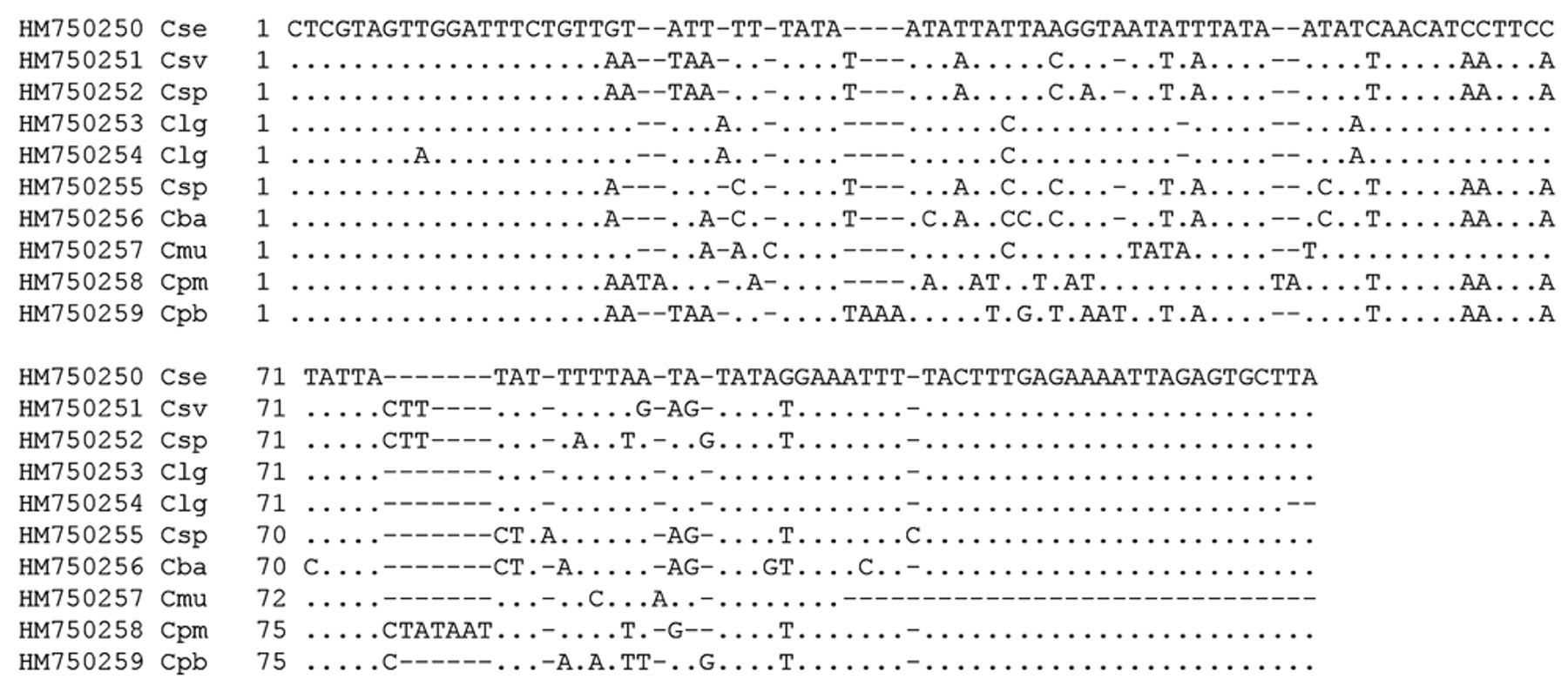

GenBank accession numbers and alignment of partial 18S ribosomal RNA gene sequences from different Cryptosporidium species obtained in the current study. HM750250 Cse: Cryptosporidium serpentis from Eublepharis macularius; HM750251 Csv: Cryptosporidium varanii (saurophilum) from Pantherophis guttatus; HM750252 Csp: Cryptosporidium sp. from Xenagama sp.; HM750253 Clg: Cryptosporidium sp. lizard genotype from E. macularius; HM750254 Clg: Cryptosporidium sp. lizard genotype from P. guttatus; HM750255 Csp: Cryptosporidium sp. from Gekko sp.; HM750256 Cba: Cryptosporidium baileyi from Python regius; HM750257 Cmu: Cryptosporidium muris from P. regius; HM750258 Cpm: Cryptosporidium parvum mouse genotype from P. regius; HM750259 Cpb: C. parvum bovine genotype–isolated oocysts. Dot (·) indicates the same nucleotide as in the reference sequence in the uppermost line; dash (–) indicates that there is no nucleotide at this place.
Correlation with additional information
Only in 186 out of 672 (28%) samples was information about the health status and/or previous contact to Cryptosporidium-positive reptiles reported. In animals showing clinical signs indicative of cryptosporidiosis and/or with reported contact with positively tested reptiles, PCR examination yielded positive as well as negative results.
Clinical signs in C. varanii (saurophilum)–infected animals included emaciation, inappetence, regurgitation, and apathy for leopard geckos, and inappetence, regurgitation, diarrhea, and maldigestion for corn snakes. The C. varanii (saurophilum)–positive Lampropeltis sp. of snakes did not show any signs of disease.
Of the C. serpentis–positive leopard geckos, 1 suffered from inappetence and 2 came from a mixed collection in which corn snakes also were kept. Clinical signs of reptiles that tested negative for cryptosporidia were identical to those above, with the addition of stunting, malodorous feces, and thickening of the gastric wall.
Discussion
The sensitivity of the presented PCR protocol for isolated Cryptosporidium oocysts is comparable to conventional PCR assays for C. parvum. 20 Sequencing detected various important Cryptosporidium species, especially those expected to be present in reptilian samples either as pathogenic parasites or as pseudoparasites. The results underline the importance of cryptosporidian species identification especially in ophidian samples to prevent making a diagnosis with potentially far-reaching consequences on the basis of the mere presence of cryptosporidial oocysts in gastrointestinal contents. Pseudoparasitic cryptosporidian species included species commonly found in mice as well as birds.
In the reptilian population examined in the current study, C. varanii (saurophilum) had a relatively high prevalence in corn snake (17/106, 16%) as well as leopard gecko (32/462, 7%) samples. Only in 2 out of 68 (3%) other snakes (1 Lampropeltis sp. and 1 unspecified snake) was C. varanii (saurophilum) detected. Results suggest an especially high susceptibility of corn snakes to C. varanii (saurophilum), which is generally considered to be present mainly in lizards. 12 The first report of C. varanii (saurophilum) in a corn snake has only recently been published. 17
Surprisingly, C. serpentis was detected in leopard gecko samples but not in snake samples, which is in contrast to previous findings. 16 Overall, cryptosporidian species with known pathogenicity for reptiles were present in 40 out of 462 (9%) leopard gecko samples, indicating a high rate of infection in the captive population of this species, which is in accordance with results from other studies. 5 The specific pathogenic potential of C. serpentis in leopard geckos has not yet been determined; however, since reptilian keepings often comprise several different reptilian species, leopard geckos carrying C. serpentis pose a potential threat to other reptiles and especially to snakes.
As reported earlier, the clinical signs reported in some Cryptosporidium-positive reptiles are consistent with chronic gastrointestinal disease of subtle clinical onset. 11,12,17 However, not all animals with gastrointestinal diseases were positive for cryptosporidia. In particular, swelling of the abdomen, gastrointestinal wall thickening, malodorous feces, and diarrhea can be caused by other microorganisms, such as flagellates or enteropathogenic bacteria. 18 Thus, reptilian cryptosporidiosis should not be diagnosed merely based on the presence of gastrointestinal clinical signs.
Thirty-three out of 672 samples with a positive PCR result did not yield an identifiable sequence. Cross-reactivity of the PCR with unspecified microorganisms present in the samples resulting in an amplicon of similar size to cryptosporidia cannot be ruled out. However, occasional technical problems during the sequencing reaction or purification steps have to be considered. The dilution of the template for the sequencing reaction resulted in a more distinct chromatogram than was achieved with undiluted template. Likewise, a dilution of DNA template was used for the PCR assay to prevent possible inhibition of the reaction by components present in the sample. However, this additional step might reduce the sensitivity of the assay.
The described combination of PCR assay and sequencing reaction proved to be useful in the diagnostics of cryptosporidiosis in live reptiles, although not every PCR-positive sample yielded a distinct sequence. Smaller laboratories are capable of performing the conventional PCR reaction themselves and the sequencing reaction can be outsourced to specialized facilities to identify the cryptosporidian species. It should be noted that similar to other methods used for cryptosporidian diagnostics, a single negative sample could not rule out an active infection, since the shedding of oocysts might temporarily be below the detection limit of the assays. Thus, a combination of different methods might lead to a more reliable result than a single approach. The cryptosporidian-free status of an animal or collection can only be ascertained by the examination of repeated samples, as shown previously. 5,10
Footnotes
Acknowledgements
The authors thank Dr. Franz Petry from the Johannes Gutenberg-University Mainz for the provision of C. parvum oocysts. The close cooperation with the Clinical Virology of the University of Veterinary Medicine Vienna is also gratefully acknowledged.
a.
QIAamp DNA Stool Mini Kit, Qiagen GmBH, Hilden, Germany.
b.
Eppendorf HotMasterMix, 5Prime; Eppendorf AG, Hamburg, Germany.
c.
Microsynth AG, Balgach, Switzerland.
d.
GeneAmp PCR system 9700, Applied Biosystems, Vienna, Austria.
e.
MinElute PCR Purification Kit, Qiagen GmBH, Hilden, Germany.
f.
ABI PRISM Big Dye Terminator Cycle Sequencing Ready Reaction kit, Applied Biosystems, Vienna, Austria.
g.
DyeEx 2.0 Spin Kit, Qiagen GmBH, Hilden, Germany.
h.
ABI PRISM 310 genetic analyzer, Applied Biosystems, Vienna, Austria.
The authors declared that they had no conflicts of interest with respect to their authorship or the publication of this article.
The authors declared that they received no financial support for their research and/or authorship of this article.
