Abstract
Although the diagnosis of canine leukemia and lymphoma in advanced stages is usually uncomplicated, some presentations of the disease can be a diagnostic challenge. In certain situations, lymphoma and leukemia can be difficult to distinguish from a benign reactive proliferation of lymphocytes. Because clonality is the hallmark of malignancy, we have developed an assay that uses the polymerase chain reaction to amplify the variable regions of immunoglobulin genes and T-cell receptor genes to detect the presence of a clonal lymphocyte population. The assay detected clonally rearranged antigen receptor genes in 91% of the 77 dogs with lymphoid malignancy. Of the 24 dogs tested, that were either healthy or had clearly defined conditions not related to lymphoid malignancy, a clonally rearranged antigen receptor gene was found in one (a dog with Ehrlichia canis infection). Gene rearrangement was appropriate for the immunophenotype (immunoglobulin gene rearrangement in B-cell leukemias and T-cell receptor gene rearrangement in T-cell leukemias). Dilution analysis showed that the clonal rearrangement could be detected when 0.1–10% of the DNA was derived from neoplastic cells, depending on the source tissue. Potential applications of this assay include the diagnosis of lymphoma or leukemia in biopsy samples, cavity fluids, fine needle aspirates, bone marrow and peripheral blood; the determination of lineage (B or T cell); staging of lymphoma; and detection of residual disease after chemotherapy.
Detection of canine lymphocytic malignancies relies on the cytologic assessment of circulating lymphocytes or lymphoid tissue or on the histologic examination of the lymphoid tissue. The diagnosis is often straightforward but there are situations that present a diagnostic challenge. These include the early stages of lymphoma that may be difficult to distinguish from lymphoid hyperplasia; cavity fluids that contain large numbers of small, mature-appearing lymphocytes; cases with chronic or mild lymphocytosis; cases diagnosed by biopsy where the biopsy does not fully represent the lesion; and the finding of a small number of atypical lymphocytes in a fine needle aspirate or cavity fluid. In addition, mass aspirates can be limited by inadequate cell numbers or by aspirating a site that does not represent the lesion. Therefore, more sensitive and objective assays are desirable for detecting lymphoid malignancies in dogs.
Because all lymphomas, lymphocytic leukemias, and myelomas are clonal expansions of lymphocytes, each particular neoplasm contains DNA regions that are unique in both length and sequence. The CDR3 region of both immunoglobulin and T-cell receptor (TCR) genes encodes the antigen-binding region of the respective receptor and contains the majority of this unique sequence. In B cells, CDR3 is produced through the recombination of V, D, and J genes. In T cells, CDR3 of the TCRγ gene is produced through recombination of V and J genes. During the process of recombination, between 5 and 20 random nucleotides are added between each gene segment, creating genes with a broad diversity in sequence and length. 15,19
In human diagnostic pathology, a method commonly used for the identification of clonal expansions of lymphocytes uses primers specific for conserved regions of V and J genes to amplify CDR3, followed by size separation of the polymerase chain reaction (PCR) products. 16 A dominant band indicates the presence of an expanded lymphocyte population bearing a single sized receptor—a clonal population. This method is most commonly used to distinguish reactive from neoplastic populations of lymphocytes and to detect minimum residual disease. 14,22
The use of this technique has been described in dogs 23 for the purpose of examining T-cell receptor rearrangements. In this report we provide methods for amplifying canine T-cell receptor gamma and immunoglobulin heavy chain CDR3 regions. We tested a panel of samples from 101 patients with and without lymphoid malignancies. The assay detected a clonal rearrangement of antigen receptor genes in the majority of dogs with lymphoid malignancy. Furthermore, clonal rearrangements were not detected in other neoplasms or in normal tissues. One dog with Ehrlichia canis infection had a clonally rearranged T-cell receptor gene. The assay may provide a useful aid for the diagnosis of lymphoid neoplasia and can be an adjunct to immunophenotyping. Other potential clinical applications may include staging lymphomas and detecting residual disease after treatment.
Materials and Methods
Selection of cases
Samples were analyzed from dogs with the following criteria: definitive diagnosis of lymphoma, myeloma, or lymphocytic leukemia using histologic or cytologic criteria in the tissue examined or definitive diagnosis of another condition with no evidence, clinical or physical, of lymphoid malignancy. Samples were obtained from several sources. First, immunophenotyped leukemias were specifically sought from stored frozen tissue (University of California at Davis and Texas A&M University). Second, cases of lymphoma and leukemia from the Colorado State University (CSU) Veterinary Teaching Hospital were obtained as they presented to the clinic. Nonlymphoid tumors and inflammatory conditions for which there was a definitive diagnosis were selected from samples also obtained at CSU.
Samples were included only if neoplasia was detected histologically or cytologically in the tissue tested by PCR. For example if an animal had lymphoma involving a lymph node but there was no evidence of disease in the bone marrow by cytology, the bone marrow sample was not used for this study even if a rearrangement was found in the marrow. In a few cases, animals presented with diseases that could be considered antecedent to the development of lymphoma or a paraneoplastic syndrome (for example immune-mediated hemolytic anemia). These cases were only included if the animal was followed for at least 1 year without the development of lymphoma. Cases of suspected or confirmed myeloid or myelo-monocytic leukemia were not included for several reasons. First, often it can be difficult to determine the lineage of leukemic cells by cytology, and there is little consensus about the appropriate cytochemical markers to use for differentiating lymphoid from undifferentiated myeloid origin leukemias. Therefore, it can be difficult to rule out the possibility of lymphoid origin in leukemias with rearrangements. Second, in human patients, up to 40% of myeloid origin tumors can have antigen receptor rearrangements, 10 making it possible that this assay will not be useful for differentiating lymphoid from myeloid leukemias.
All samples were read by board certified anatomic or clinical pathologists who were not aware of the PCR results. Most were tested by PCR after the diagnosis of neoplasia.
Immunophenotyping
Samples from University of California at Davis were analyzed by flow cytometry as described in Moore et al. 12 Samples from Texas A&M University were phenotyped according to Modiano et al. 11 Leukemia samples from CSU were typed as follows: 200 μl of whole blood was lysed in 1 ml of lysis buffer (0.15 M NH4Cl, 1 M KHCO3, 0.1 mM Na2EDTA, 1 N HCl to a pH of 7.2–7.4) for 5 minutes at room temperature. The cells were spun, lysed a second time, then resuspended in 200 μl of phosphate buffered saline (PBS)-2% FBS. Twenty-five microliters of the cell suspension was added to each of the six wells of a 96-well plate. Directly conjugated antibodies were added to each well in the following combinations: 1) CD5-F (FITC) and CD8-phycoerythrin (PE), 2) CD45-PE and CD4-F, 3) CD21-PE and CD3-F, 4) CD14-PE and CD4-F, 5) CD4-F and a PE-conjugated isotype control, and 6) CD8-PE and an FITC-conjugated isotype control. Samples were incubated for 15 minutes at room temperature and then washed twice. They were then suspended in PBS-1% paraformaldehyde until analysis. The antibody sources were as follows: anticanine CD4, CD8α, CD45, and CD5 from Serotec (clones YKIX302.9, YCATE55.9, YKIX716.13, and YKIX322.3, respectively), anti-human CD14 and CD21 from PharMingen (clones TUK4 and B-ly4). Lymphocyte populations were gated on the basis of the forward scatter versus log side scatter plots. Leukemias were either CD21+ or CD8+, and in all cases, the lymphocyte population contained > 80% of cells expressing the indicated marker. Cases of lymphocytosis that were not independently diagnosed by cytologic criteria were not included.
Identification of primers for amplification of immunoglobulin genes
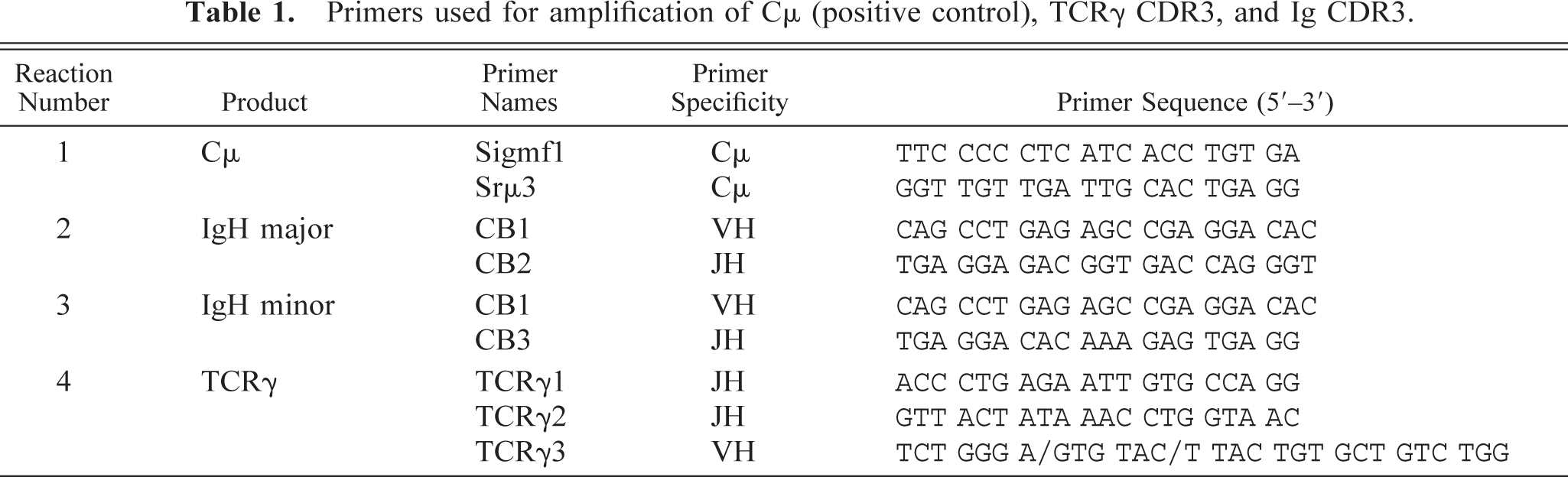
Ig primers were identified in canine VH cDNA clones (GenBank accession numbers AF067449–AF067453). The cDNA clones were generated using 5′ rapid amplification of cDNA ends 24 from RNA derived from the spleen of a healthy dog. 3′ anchor primers used for this method were designed according to the exon 1 sequence of canine Cμ (A. Avery and R. Burnett, unpublished, sequence generated using heterologous primers to amplify cDNA from the peripheral blood of a healthy dog). cDNA was inserted into the T/A cloning vector pGEM-T (Promega). Five clones were sequenced, with all these clones having V region framework 3 sequences containing a conserved region that could be amplified with a common primer (CB1, Table 1). In these five samples, two clearly distinct J region sequences were identified. Primers for each gene were selected (CB2 and CB3, Table 1). Therefore, for routine typing, a single primer V region primer was used along with two J region primers.
Primers used for amplification of Cμ (positive control), TCRγ CDR3, and Ig CDR3.
Identification of primers for amplification of TCRγ genes
Studies in the literature suggested that in humans TCRγ rearrangements may be easier to detect, and may also be a more sensitive measure of clonality, than TCRβ rearrangements, regardless of which gene is expressed by the tumor. 8,20,21 Therefore, we chose to examine the TCRγ locus as a means of identifying clonal T-cell populations. TCRγ cDNA was obtained from the thymus of a healthy dog using a heterologous Vγ (TCRVγ) and a canine Cγ primer (A. Avery and R. Burnett, unpublished). The Vγ primers were designed on the basis of a comparison of bovine and rabbit Vγ sequences. This comparison revealed high conservation at the 3′ end of Vγ (TCRγ3). This primer was used in the generation of V-J cDNA clones and in subsequent routine typing. Five V-J clones were sequenced (GenBank accession numbers AF79118–079123) and again, two different J sequences were identified. Thus, as with the Ig genes, one V region primer and two J region primers were used in routine typing (Table 1).
PCR conditions used for routine typing
DNA samples are prepared with the QIAamp DNA mini kit from Qiagen. For each sample four PCR reactions were run; 1) Cμ (positive control), 2) IgH major, 3) IgH minor, and 4) TCRγ. Ig major and minor are designated as such to represent the use of the two different J region primers. The specific primer sequences and combinations are given in Table 1. The “Ig major” primers amplified the majority of immunoglobulin gene rearrangements, and the “Ig minor” primers amplified a smaller proportion of these rearrangements. We found that these primers could not be combined in the same reaction. However, the three TCR primers could be mixed in the same reaction. For amplification, HotStarTaq Master Mix Kit was used. Approximately 100–500 ng of DNA was amplified with 200–400 nM of each primer in a 50-μl reaction volume. The Cμ positive control primers were taken from the constant region gene of IgM. The primer pairs used are given in Table 1.
A Perkin-Elmer thermocycler 9600 was used for all experiments. Samples were initially heated to 95 C for 15 minutes (conditions specific for the activation of HotStar Taq, Qiagen). The cycling profile was as follows: 94 C for 8 seconds, 60 C for 10 seconds, and 72 C for 15 seconds for 35 cycles. No final extension time was used. PCR products were analyzed on 10% native polyacrylamide gel electrophoresis, (1.5 mm thick) with 1× Tris borate EDTA buffer. Bromophenol blue was run until the dye front had moved approximately 14 cm. The Cμ product is approximately 130 base pairs (bp), the Ig products center around 120 bp and the TCR products center around 90 bp. A result was considered positive if one or more dominant and discreet clonal bands were present at irregular intervals. Polyclonal amplification resulted in discreet bands that were evenly spaced when separated by size. Twenty-four samples were selected randomly and repeated for purposes of quality control. In all cases, the repeat gave exactly the same result as the original sample.
We have found that changing of any variables in this protocol must be accompanied by rigorous testing of a large panel of samples. Small fluctuations in primer design, source of Taq polymerase, and amplification times and temperatures can affect the result significantly. All the results presented in this article were carried out with the reagents and conditions described here.
Results
Detection of gene rearrangements using a panel of lymphoid neoplasms
DNA was extracted from tissues taken from dogs with a variety of conditions, as described in “Materials and Methods” section, and amplified with primers specific for Ig and TCR genes. Cμ primers were used to amplify each sample as a positive control for DNA, and a negative control (no DNA added) was also carried out with each group of samples.
Figure 1A shows the amplification of DNA from 15 samples that were chosen to represent the array of samples we tested and the results we obtained. All samples had sufficient DNA for analysis, as shown by the consistent amplification of a product with the Cμ primers (panel D). The spectrum of results with the three different primer sets can be seen in panels A through C. The types of patterns seen included a single detectable rearrangement of an Ig gene (lanes 1–4 and 6), a biclonal rearrangement of Ig genes (lane 5), or single (lanes 7, 9, 12), biclonal (lane 8), or oligoclonal (lanes 13 and 14) rearrangement of TCRγ genes. One sample was found to have rearrangements in both Ig and TCRγ genes (lane 15), and in a second sample (lane 5) there was a suggestion of a biclonal T-cell receptor rearrangement (panel C) as well as of the immunoglobulin gene rearrangements. Negative results can appear either as no visible bands (lanes 10 and 11) or as a ladder-like smear (lane 9, panel A—this dog had a TCR rearrangement on a background of normal B cells that gave rise to the smear with Ig primers).
The source of material for each sample is listed in the legend for Fig. 1. Samples of all types, including biopsies, peripheral blood, bone marrow, cavity fluids, and aspirates gave sufficient material for analysis. We found that in general, aspirating the site, such as a lymph node, at least three times into physiologic saline and rinsing the syringe each time in the saline would give sufficient material for analysis. But it is also possible to obtain enough material for analysis by washing cells from a stained or unstained cytologic preparation. None of the samples analyzed as a part of this study were derived from formalin-fixed tissues.
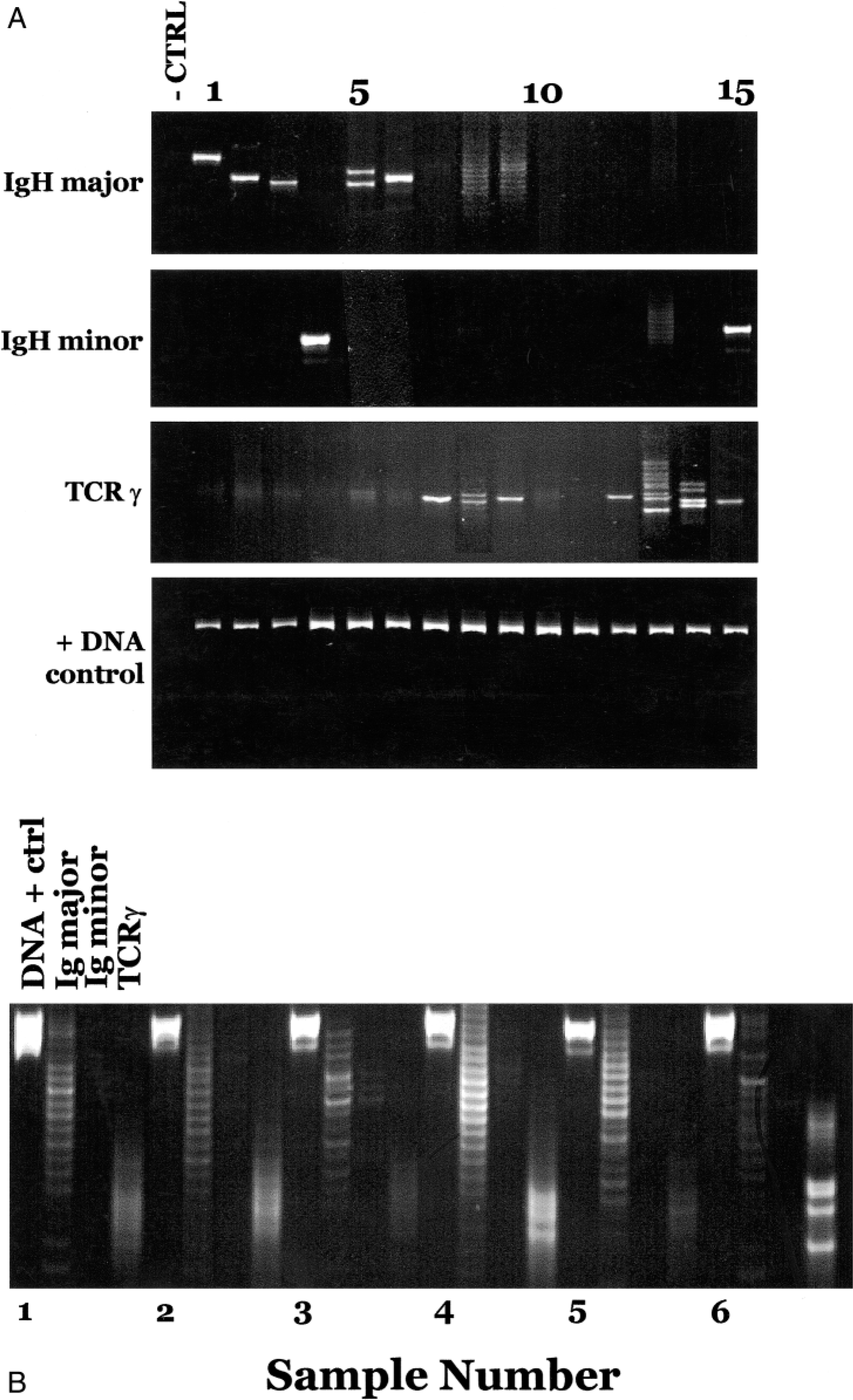
Examples of positive and negative PCR results obtained with a variety of sample types. Fig. 1A. Analysis of samples using primers for Ig major (CB1 and CB2), Ig minor (CB1 and CB3), TCRγ, and the positive DNA control, Cμ primers. Lanes are numbered starting with the first sample; hence, the negative control is not counted. Lanes 8 and 9 in the Ig major gel, and lanes 13 and 14 in the TCRγ gel were photographically enhanced because they were dim controls for DNA. All the samples that were amplified with a single primer pair were run on the same gel. The DNA was derived from the following sources (lane numbers—sample source—diagnosis); 1—bone marrow (bm)—lymphoblastic leukemia, 2—biopsy—lymphoma, 3—peritoneal fluid—lymphoma, 4—blood—lymphocytic leukemia, 5—biopsy—lymphoma, 6—aspirate—lymphoma, 7—blood—lymphocytic leukemia, 8—bm—lymphoma, 9—pleural fluid—lymphoma, 10—cerebrospinal fluid—lymphoma, 11—bm—lymphoma (same dog as 10), 12—bm—lymphoma, 13—blood—lymphoma, 14—blood—lymphoma, 15—blood—lymphocytic leukemia. The first lane represents the PCR reaction carried out without DNA, but including the primers used for the other samples in that panel, and amplified during the same experiment. Fig. 1B. Example of negative results from five different samples (1–5) analyzed in the same gel with a positive result (6). For each sample, the first lane (a) is the positive control for DNA, (b) is amplification with Ig major primers, (c) is amplification with Ig minor primers, and (d) represents amplification with TCRγ primers. The samples are from the following sources; 1–3 are blood, lymph node, and bone marrow from a dog with histologically confirmed lymphoma that could not be detected using this assay; 4 is bone marrow from a dog with pure red cell aplasia, 5 is lymph node biopsy from a dog with immune mediated arthritis, 6 is peripheral blood from a dog with T-cell lymphoma.
Figure 1B shows additional results analyzed in a format that would be appropriate for routine typing, in which each sample is subjected to four reactions that are analyzed on the same gel. One of these samples had clonally rearranged antigen receptor genes (sample 6, TCRγ lane).
Results for all 101 animals are presented in Table 2, and a breakdown of the types of lymphoid malignancies that were tested is shown in Table 3. A monoclonal, biclonal, or oligoclonal pattern was found in 91% of all lymphoid malignancies tested (n = 77). Only one sample (4%) from a dog without a definitive diagnosis of lymphoma was found to have a T-cell receptor gene rearrangement in a bone marrow sample.
Rearrangements found in samples from a panel of dogs with and without lymphoid malignancies.
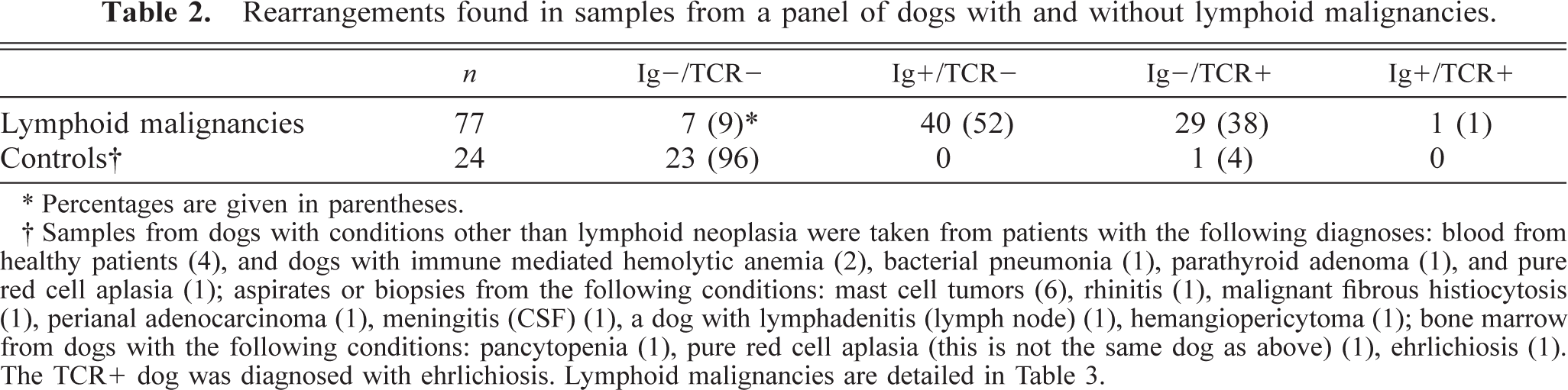

∗ Percentages are given in parentheses.
† Samples from dogs with conditions other than lymphoid neoplasia were taken from patients with the following diagnoses: blood from healthy patients (4), and dogs with immune mediated hemolytic anemia (2), bacterial pneumonia (1), parathyroid adenoma (1), and pure red cell aplasia (1); aspirates or biopsies from the following conditions: mast cell tumors (6), rhinitis (1), malignant fibrous histiocytosis (1), perianal adenocarcinoma (1), meningitis (CSF) (1), a dog with lymphadenitis (lymph node) (1), hemangiopericytoma (1); bone marrow from dogs with the following conditions: pancytopenia (1), pure red cell aplasia (this is not the same dog as above) (1), ehrlichiosis (1). The TCR+ dog was diagnosed with ehrlichiosis. Lymphoid malignancies are detailed in Table 3.
Classification of lymphoid malignancies studied.
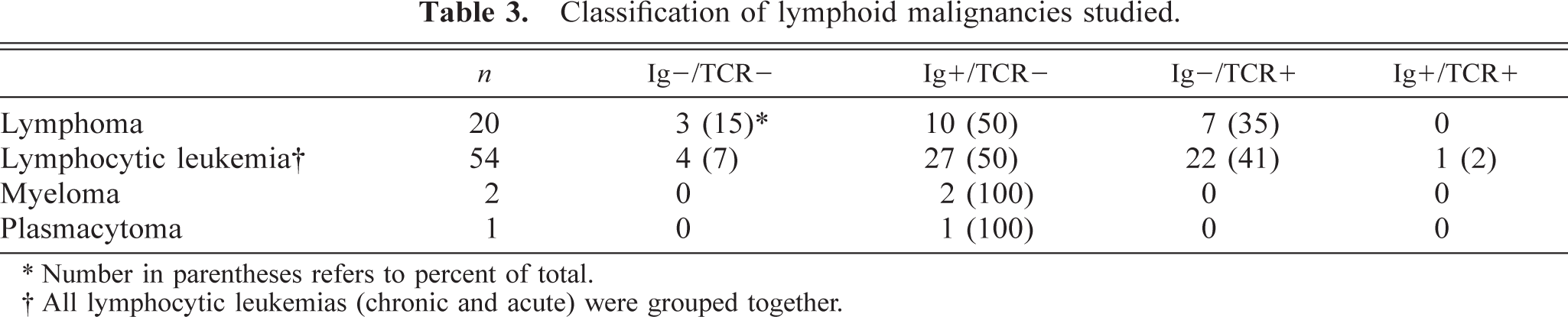

∗ Number in parentheses refers to percent of total.
† All lymphocytic leukemias (chronic and acute) were grouped together.
Correlation between rearranged gene and immunologic phenotype
Some of the leukemias in this study were immunophenotyped by flow cytometry. Only those cases for which a definitive phenotype (T versus B in origin) could be established were included in this part of the study. The myelomas and plasmacytoma were also included, even though they were not immunophenotyped, because these tumors are known to be of B-cell lineage. The majority of cases were either not phenotyped or did not react with any of the panel of antibodies tested. We found concordance between the phenotype obtained by flow cytometry and the rearrangement detected by PCR in all cases but one, a T-cell leukemia with both immunoglobulin and TCR rearrangements (Table 4). This suggests that the PCR method can be useful for immunophenotyping. The exception would be in those rare cases (one in this study) where both types of rearrangements are identified.
Correlation between immunophenotype and type of rearrangement.
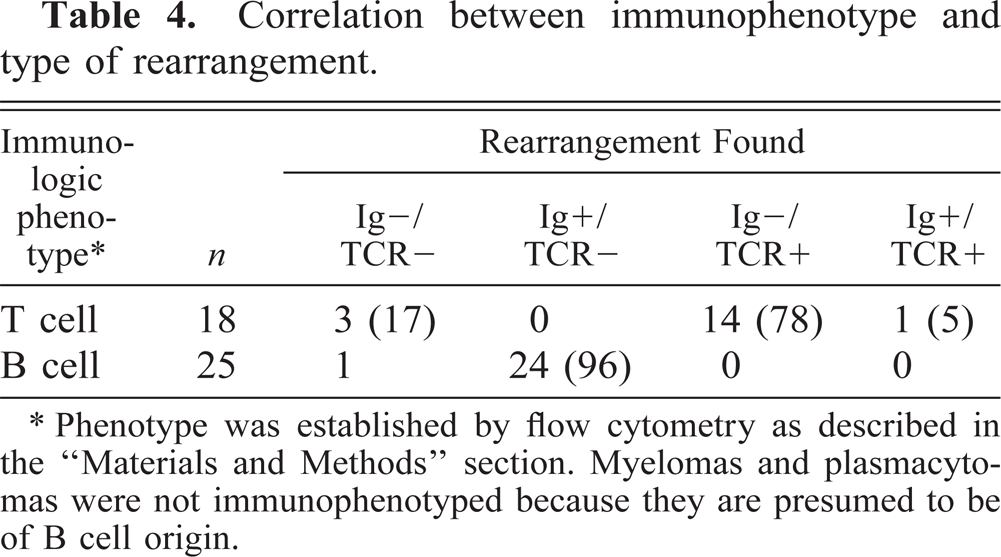

∗ Phenotype was established by flow cytometry as described in the “Materials and Methods” section. Myelomas and plasmacytomas were not immunophenotyped because they are presumed to be of B cell origin.
Limits of detection of gene rearrangements within different tissues
One important use of this assay will be to establish the presence of lymphoid malignancy in tissue biopsies and bone marrow samples. Therefore, it was important to determine the sensitivity of the assay when used in a background of normal lymphoid or nonlymphoid tissue. To measure sensitivity, DNA from a B-cell leukemia that exhibited a clean, single band in the IgH PCR was diluted serially by 10-fold from 100 to 0.1 ng (Fig. 2). A single rearrangement was still detectable when as little as 0.1 ng of DNA was used. A similar result was seen when DNA from the leukemia was diluted into a constant amount of DNA extracted from normal liver, which demonstrates that in a nonlymphoid organ this assay can detect a malignant clone that comprises as little as 0.1% of the tissue. When the same dilutions were performed in DNA from normal spleen, a clear dominant rearrangement was seen at 1 ng of DNA (representing 1% of the total DNA present), but this band was obscured by background bands at 0.1% of the total DNA. Comparable results were observed with TCRγ amplification, except when diluted into DNA from the thymus. In this case, the neoplastic band could only be seen at 10% of the input DNA. Thus, the nature of the tissue from which the sample is taken influences the sensitivity of the assay.
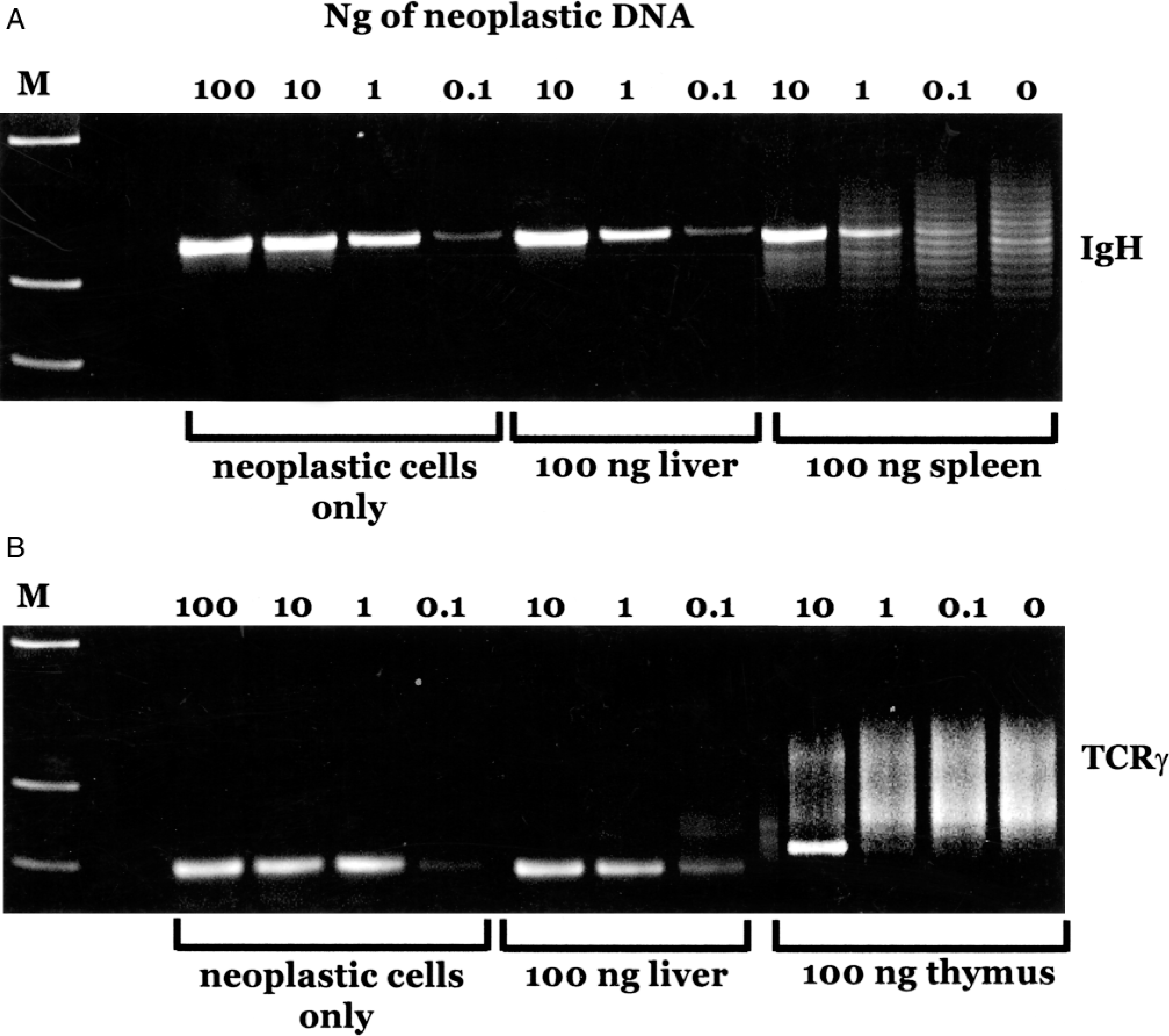
Assay sensitivity. Fig. 2A. DNA from a neoplastic sample was mixed with 100 ng DNA from the indicated tissue in the amounts shown, and PCR for the Ig major rearrangement was carried out. Fig. 2B. Similar to A, except that PCR for TCRγ was used. M represents the molecular weight markers, and the next lane was left blank.
Discussion
We have developed an assay that can be used to detect clonally rearranged antigen receptor genes in tissues of dogs with lymphoid malignancies. The purpose of this assay is to identify the presence of a clonal population of lymphocytes within a heterogeneous mixture of cells to distinguish neoplastic lymphocytes from nonneoplastic ones. In the present study, we established assay conditions and carried out an analysis of 77 lymphoid neoplasms and 24 samples from animals that had conditions other than lymphoid neoplasia. Clonal rearrangements of either immunoglobulin or T-cell receptor genes or both were found in 91% of the lymphoid neoplasms tested and in 4% of conditions other than lymphoid neoplasia.
We have found that we can detect a high percentage of gene rearrangements using very few primer sequences (two J primers and a single V primer for both TCR and Ig genes). This sensitivity is greater than what is found in studies with human patients, even when a large number of primer combinations are used. 8 This finding suggests that the dog may use a limited repertoire of V region genes from both types of antigen receptors. Alternatively, there may simply be less diversity in the available V region genes in dogs, with enough conservation that a single primer can bind to the vast majority of them. A final, but unlikely explanation is that canine neoplasms express only a restricted set of antigen receptors. But there is no precedent for this idea in human patients or in experimental animals.
It is important to note that for this preliminary study we only used samples in which neoplastic cells could be unequivocally observed by cytology or histology. Therefore, the reported sensitivity should be an accurate reflection of the percentage of rearrangements detectable with these primers. The 9% of samples that gave a negative result despite the presence of confirmed lymphoid neoplasia are likely to be samples that use V or J region genes to which our primers do not bind (for example a T-cell neoplasm expressing TCRαβ genes that has deleted Vγ genes), tumors that have deleted antigen receptor genes, or NK-derived lymphomas or leukemias that do not have rearranged antigen receptor genes. When used for routine diagnostic situations, the sensitivity of the test will reflect not only the number of rearrangements that are bound by our primers but also the number of neoplastic cells in the sample. This in turn will be affected by the extent of disease, the cellularity of the sample, and how well the sample reflects the neoplasm. Therefore, it is likely that as a clinical tool, the sensitivity of the assay will be lower than 91%.
The single case in which a clonal rearrangement was found in a dog without lymphoma was in a 9-year-old male Akita. At the time that the bone marrow sample from this dog was taken, he was undergoing evaluation for pancytopenia that was an incidental finding on routine geriatric blood work. The dog was clinically normal. As part of the evaluation, he was found to have an elevated E. canis titer, and was treated with doxycycline. After treatment, his leukocyte, erythrocyte, and platelet counts returned to normal. The dog remains clinically healthy 2 years later. We have not had the opportunity to retest his bone marrow to determine if the rearrangement persists. A similar finding was made by Vernau and Moore, 23 who also identified clonal T-cell receptor rearrangements in a dog with ehrlichiosis, and recently we have identified several other E. canis–positive dogs with TCRγ clonal rearrangements (R. C. Burnett, unpublished), but these dogs have not been followed long enough to determine if they have occult malignancy. These T cells may be a population of transformed T cells that have not yet evolved to overt neoplasia, such as can be seen in human T-cell lymphotropic virus (HTLV) infection in humans. 6 Alternatively, they may represent an antigen-specific population that will recede over time. Epstein-Barr virus infection in humans has been shown to stimulate this kind of clonal and oligoclonal T-cell expansion that appears to be antigen-reactive rather than a preneoplastic state because the clones cannot be detected once the infection is controlled. 4,5
The lower limits of detection of neoplasia with the PCR assay depend on the tissue from which the sample was taken. In nonlymphoid tissue, the assay is very sensitive and allows detection of one neoplastic cell in 1,000. In lymphoid tissue, detection is less sensitive. One likely explanation is that the rearranged sequences from normal lymphocytes compete with amplification of the neoplastic DNA. Because of the level of sensitivity, we believe that this assay will allow us to diagnose lymphoid neoplasia in cases where that diagnosis cannot be made by microscopy because too few cells are present. For many of the cases used in this study, we have carried out PCR analysis on multiple tissues from the same dog. Even when no malignant cells were evident by microscopy, we were able to detect a clonal rearrangement in several tissues. These results will be published as part of a separate study. The detection limits of the assay are similar to that reported in human patients. 10 Sensitivity can be increased at least 10-fold if primers specific for the individual tumor, rather than consensus primers, are used. 22 Although not practical for routine clinical use, this method is frequently used experimentally for the detection of minimum residual disease after chemotherapy in human patients. 14,22
A number of samples contained either two TCR or two Ig rearrangements. These findings can be explained by the presence of two populations of neoplastic cells or by the presence of rearrangements on both chromosomes. More intriguing was the finding that some samples had multiple rearrangements, as shown in Fig. 1A, lanes 13 and 14. There are at least two different explanations for the presence of multiple rearrangements. The first is that the tumor began as a single clone and evolved to multiple clones through serial transformations. This series of events has been described for B-cell lymphomas in humans. 7 An alternative possibility is that multiple clones arose from a transforming process. Exposure to a carcinogen or transformation by a virus could account for the expansion of multiple clones at one time. An example of such a viral process can be found in the study of clonal expansion of T cells by HTLV-1. When a host is infected with HTLV-1, multiple T-cell clones are initially expanded. 6
Although we anticipate that the primary use of the assay described is to establish the presence of a malignancy, the finding of exclusively TCR or Ig rearrangements may help to distinguish B-cell from T-cell origin lymphomas and lymphocytic leukemias. Determination of B- or T lineage has been shown in several studies to offer valuable prognostic information; for example, lymphomas of T-cell origin are associated with poorer survival and remission times than are B-cell lymphomas, 9,18 and immunophenotype may have more prognostic significance than histologic phenotype. 1 We anticipate that the majority of lymphoid malignancies will have rearrangements in the appropriate receptor loci (immunoglobulin for B-cell or T-cell receptor for T cell). Our findings demonstrate that some tumors will have rearrangements in both loci. Similar findings have been made in humans. In one study, for example, 4% of B-cell lymphomas and 9% of T-cell lymphomas (defined by immunophenotypic criteria) had rearrangements of both IgH and TCRγ loci. 8 In these few cases, lineage assignment must rely upon immunophenotyping.
One of three acute myelogenous leukemias we tested contained an Ig gene rearrangement (data not included in the results shown here). This finding is not surprising because in one study, 40% (of 35 cases) of human AML also exhibit this type of rearrangement, as well as expressing some B-cell–specific antigens. 10 In this case series, the presence of an IgH rearrangement was associated with poorer survival. In another study, T- or B-cell receptor rearrangements were found in 2 of 16 patients with myelodysplastic syndrome that progressed to myelogenous leukemia. These findings demonstrate that this assay may not be useful in assigning lineage to a leukemia that is cytologically ambiguous with respect to origin (myeloid versus lymphoid). Other methods, such as immunophenotyping and cytochemical stains, will be important in establishing the origin of these tumors.
In using this assay, we have made the assumption that malignancy is always clonal. But the reverse is not necessarily true. That is, all clonal expansions of lymphocytes are not necessarily malignancies. An excellent example of this is the finding that the single cutaneous plasmacytoma examined in this study had a clonal immunoglobulin rearrangement. This neoplasm is generally considered benign. 2 Humans develop a number of benign lymphoproliferative disorders that can be clonal, including large granular lymphocytosis, certain AIDS-related lymphoproliferative disorders, and angioimmunoblastic lymphadenopathy. 17 In addition, a small percentage of elderly humans develop benign monoclonal gammopathy, and using a sensitive PCR technique, a clonal population of plasma cells can be found in the peripheral blood of these individuals. 3 Benign clonal T-cell expansions may also occur rarely in humans. 13 Although neither of these conditions has been described in dogs, it will be important to consider this possibility when evaluating samples that have clonal rearrangements but no other evidence of leukemia or lymphoma.
We have described a method of detecting malignant lymphocytes that can be used as an adjunct to morphology and immunophenotyping for the diagnosis of leukemia and lymphoma. Other potential applications of the technique include assignment of lineage to B- or T-cell origin, staging of disease, and detection of minimum residual disease. We are currently evaluating the usefulness of some of these applications.
Footnotes
Acknowledgements
This study was supported by AVMF grant #81–98 to ACA. We thank the oncology service at the Veterinary Teaching Hospital of Colorado State University for their generous help in obtaining samples for analysis.
