Abstract
Papillomas and fibropapillomas may occur in the skin and in different organs in animals. Ten different genotypes of bovine papillomavirus (BPV) have been identified. BPV-1 through BPV-10 are all strictly species-specific, but BPV-1/2 may also infect other species such as equids, inducing fibroblastic tumors. BPV-1 and BPV-2 are associated with fibropapillomas in cattle; these tumors are formed by excessive proliferation of virus-infected dermal fibroblasts and epidermal keratinocytes. Nine water buffalo (Bubalus bubalis) were examined for the presence of multiple cutaneous and perivulvar tumors. Cutaneous and perivulvar fibropapillomatosis were confirmed histologically. Negative-stain transmission electron microscopic examination revealed papillomavirus-like particles in the fibropapillomas, and papillomaviral DNA was also detected by the polymerase chain reaction. The amplified long control region (LCR) DNA sequence was identical to that of BPV-1. The BPV-1 E5 oncoprotein was strongly expressed in the tumor cells thus confirming a causal role of the virus. This article represents the first report of cutaneous, perivulvar, and vulvar fibropapilloma associated with BPV-1 infection in the water buffalo and describes another example of cross-species infection by BPV-1.
Keywords
Introduction
Cutaneous fibropapillomatosis is characterized by the presence of multiple benign exophitic proliferations of the epidermis and of the underlying derma, which is commonly associated with infection with a papillomavirus. It has been described in cattle and other ungulates such as red deer, roe deer, elk, and less frequently in horses, dog, cat, sheep, goat, pig, and bison. 3,15,21 Genital papillomavirus infection has also been described in cows and pigs. 14 Ten different genotypes of bovine papillomavirus (BPV) have been identified. BPV-1–10 are all strictly species specific, but BPV-1/2 also infects equids, where it causes fibroblastic tumors called sarcoids. 22 Furthermore, BPV-induced lesions in cattle can progress to cancer under the impact of environmental cofactors. 11
The different genotypes have been classified into 3 genera: 20 Xipapillomaviruses including the pure epitheliotropic BPV-3, BPV-4, and BPV-6; Delta-papillomaviruses including BPV-1 and BPV-2, which are associated with fibropapillomas; and Epsilon-papillomavirus comprising BPV-5, whose genome seems to share similarities with the preceding 2 BPV groups. Two additional novel BPV were characterized: one was designated BPV-7 and classified as a member of a new papillomavirus (PV) genus, and the other was designated BPV-8 and classified as a member of the Epsilon-papillomavirus genus. 5
Recently, Hamata et al. 16 identified 2 new BPV types belonging to the genus Xipapillomavirus, designated BPV-9 and BPV-10.
E5 is the major BPV transforming oncoprotein; 13 it is a short, hydrophobic, type-II transmembrane protein expressed in the deep layers of the infected epithelia 25 and is largely localized to the membranes of the endoplasmic reticulum (ER) and Golgi apparatus (GA) of the host cells. 25 BPV E5 is expressed in both basal and suprabasal transformed epithelial cells with a typical juxtanuclear pattern due to its localization in the GA. 2,8 BPV E5 induces cell transformation of cultured rodent fibroblast and keratinocytes by binding to and activating the platelet-derived growth factor β (PDGFβ) receptor. Furthermore E5 induces receptor activation by forming a stable complex with it, inducing its dimerization and phosphorylation. This mechanism has been demonstrated first in transformed cells, 10 and then in naturally occurring epithelial and vascular bladder cancer in cattle. 7
This report describes the histopathologic and ultrastructural aspects of spontaneous papillomatosis in water buffalo from Southern Italy and shows the presence of BPV-1 DNA and the expression of E5 in the fibropapillomas.
Materials and Methods
Animals
Nine water buffalo from different herds in Campania (South of Italy), 4 to 10 years old, were examined for the presence of fibropapillomas. Four buffalo showed multiple, cutaneous tumors localized in the back and in the gluteal region; the other 5 showed multiple, firm, raised, gray-to-black, verrucous tumors affecting the vulvar and perivulvar region.
Histopathology
Samples of 9 tumors, one from each affected buffalo, removed under local anesthesia were fixed in 10% buffered formalin and embedded in paraffin by routine methods. Sections (4 µm thick) were stained with HE and Van Gieson to assess the morphologic characteristics of the tumors.
Electron Microscopy
For negative staining, 3 tumor samples, 2 from 2 animals with cutaneous fibropapillomas and 1 from an animal with vulvar fibropapillomas, were homogenized and then centrifuged onto glow-discharge–treated carbon films on 400-mesh copper grids using a Beckman Airfuge operated at 25 ft-lb/in 2 (∼100,000 × g) for 10 minutes. The grids were washed with distilled water, negatively stained with ammonium molybdate, and examined with a Zeiss EM 902 electron microscope operated at 80 kV.
Polymerase Chain Reaction Amplification of Viral DNA
Paraffin-embedded slices (10 µm thick) cut from 1 cutaneous and 1 vulvar fibropapilloma were used for assessing the presence of BPV-1 DNA in the tumors. DNA was extracted from paraffin sections using QIAamp DNA Mini Kit (Qiagen, Crawley, UK). In brief, a small section was placed in 1200 µl xylene and vortexed vigorously. After centrifugation for 5 minutes at room temperature, the supernatant was removed taking care not to disturb the pellet. Twelve milliliters of 100% ethanol was added to the pellet to remove residual xylene and mixed gently by vortexing, then centrifuged for 5 minutes at room temperature. The ethanol was removed, and after another ethanol wash, the tissue pellet was incubated at 37°C for 10 to 15 minutes to remove residual ethanol. The pellet was resuspended in 180 µl of Kit Buffer ATL. The DNA extraction was then carried out following the kit manufacturer's protocol. The purified DNA was resuspended in 100 µl of Kit Buffer AE.
BPV-1 DNA was amplified by polymerase chain reaction (PCR) using primers LCR-F 5′ ccg tac aca tcc tgt cca gca t 3′ (corresponding to BPV-1 nucleotides [nts] 7211–7232) and LCR-R 5′ gat ggt gtg att att gtt aac 3′ (corresponding to BPV-1 nts 21–1), which amplify a 790 base pair (bp) product from the viral long control region (LCR), as described by Nasir et al. 23 In brief, reactions were performed in the presence of Platinum Taq DNA polymerase (Invitrogen, Glasgow, UK), according to the manufacturer's instructions. Cycling conditions were as follows: 1 cycle of 94°C for 2 minutes, followed by 40 cycles of 94°C for 1 minute, 57°C for 1 minute, 72°C for 2 minutes, and a final extension of 72°C for 10 minutes.
The PCR products were purified using Invitrogen ChargeSwitch PCR clean-up kit and aliquots were run on a 1% agarose gel to assess their size.
Ten nanograms of the cleaned PCR products were used for nucleotide sequencing with LCR-F and LCR-R as primers, each at 5 pM. All sequences were performed on the ABI3100 sequencer (Applied Biosystems, Foster City, CA) using Big Dye Terminator Kit Version 3.1 (Applied Biosystems). The sequences were analyzed by BLAST search. Two rounds of PCR were required to amplify the DNA from the vulvar fibropapilloma. In all the PCR experiments, negative controls without a DNA template were also included.
Immunofluorescence and Confocal Microscopy
Nine tumor samples, one from each affected buffalo, were used for immunofluorescence studies. In brief, paraffin sections were deparaffinized and blocked for endogenous peroxidase in 0.3% H2O2 in methanol for 20 minutes. Antigen enhancement was performed by microwave heating (twice for 5 minutes each at 750 W). The primary antibody was applied overnight at room temperature in a humidified chamber. The polyclonal sheep anti-E5 antibody 7 was used at 1 : 1,000 dilution.
Slides were washed 3 times with PBS and then incubated with fluorescein isothiocyanate (FITC)-conjugated anti-sheep antibody (Jackson ImmunoResearch Laboratories, Inc., West Grove, PA) at 1 : 200 dilution for 2 hours at room temperature. Slides were then rinsed in PBS and incubated for 30 minutes at room temperature with propidium iodide 1 µg/ml (Sigma, Milan, Italy), then washed again and mounted with an aqueous mounting medium (Sigma). For confocal laser scanning microscopic investigation an LSM 510 (Zeiss, Jena, Germany) was used. Different frames were scanned separately, with appropriate installation of the optical path for excitation and emission of each scan according to the manufacturer's instructions.
Controls for staining specificity were omission of the primary antibody, its replacement with nonimmune sera or irrelevant antibody.
Results
Gross Pathology
The cutaneous and the vulvar lesions were characterized by multiple gray-white papillary growths of tough consistency. Some of the tumors were ulcerated with areas of necrosis, and the surface showed sessile vegetative outgrowths (Figs. 1, 2).
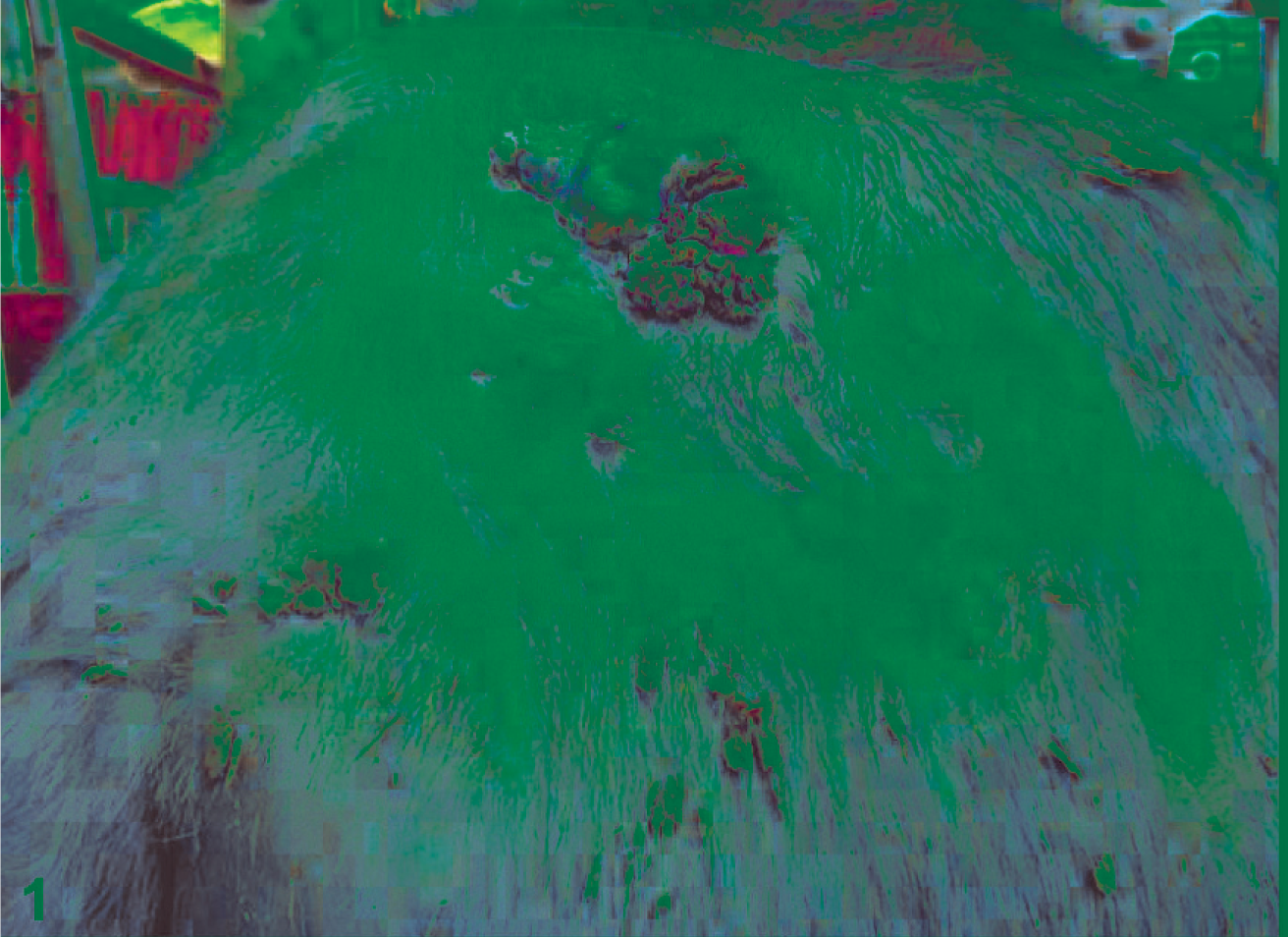

Skin; water buffalo. Cutaneous fibropapillomas of the skin in the gluteal region.
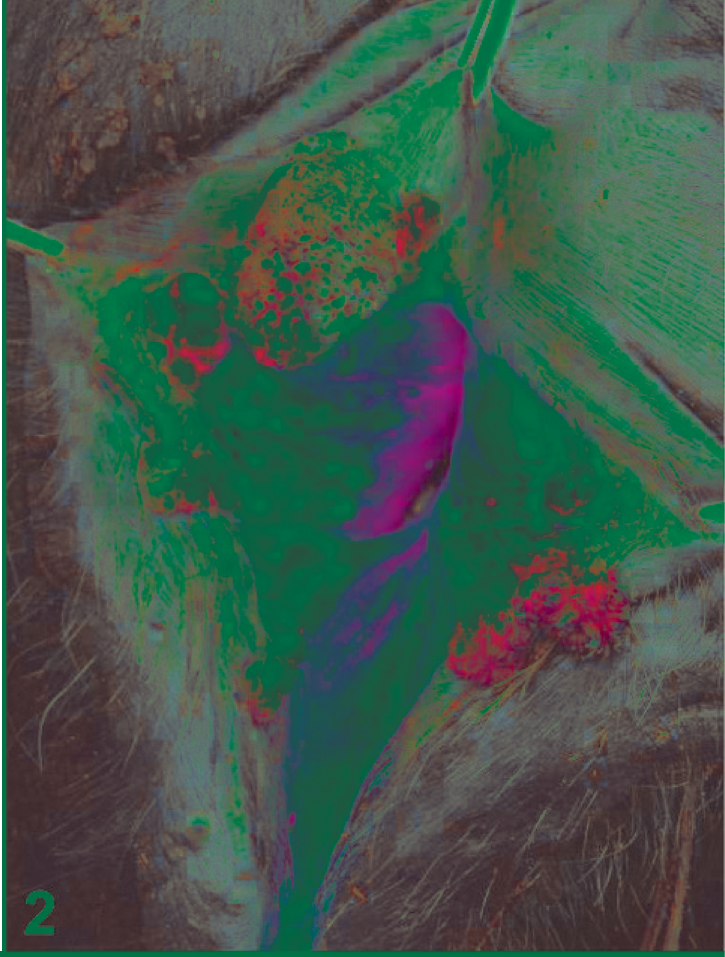
Vulva; water buffalo. Multiple cauliflower-like growths in and around the vulva.
Histopathology Examination
The tumors were diagnosed as fibropapillomas. On HE- and Van Gieson-stained sections, the tumors were seen to consist of homogeneous, smooth connective tissue covered by hyperplastic epidermis with acanthosis and orthokeratotic hyperkeratosis. The stratum spinosum was hyperplastic and many koilocytes were present, often arranged in small groups. These cells were characterized by large, mildly eosinophilic and vacuolated cytoplasm, and some cells contained small distorted nuclei with dense chromatin (Fig. 3). Nuclei in the majority of epidermocytes were vesicular, with fine chromatin and readily apparent nucleoli; some intranuclear inclusion bodies were seen (Fig. 3). Some cells displayed histologic signs of apoptosis (Fig. 3). In the basal layer, there was moderate mitotic activity and the epidermocytes in the granular layer exhibited well-developed keratohyaline granules.
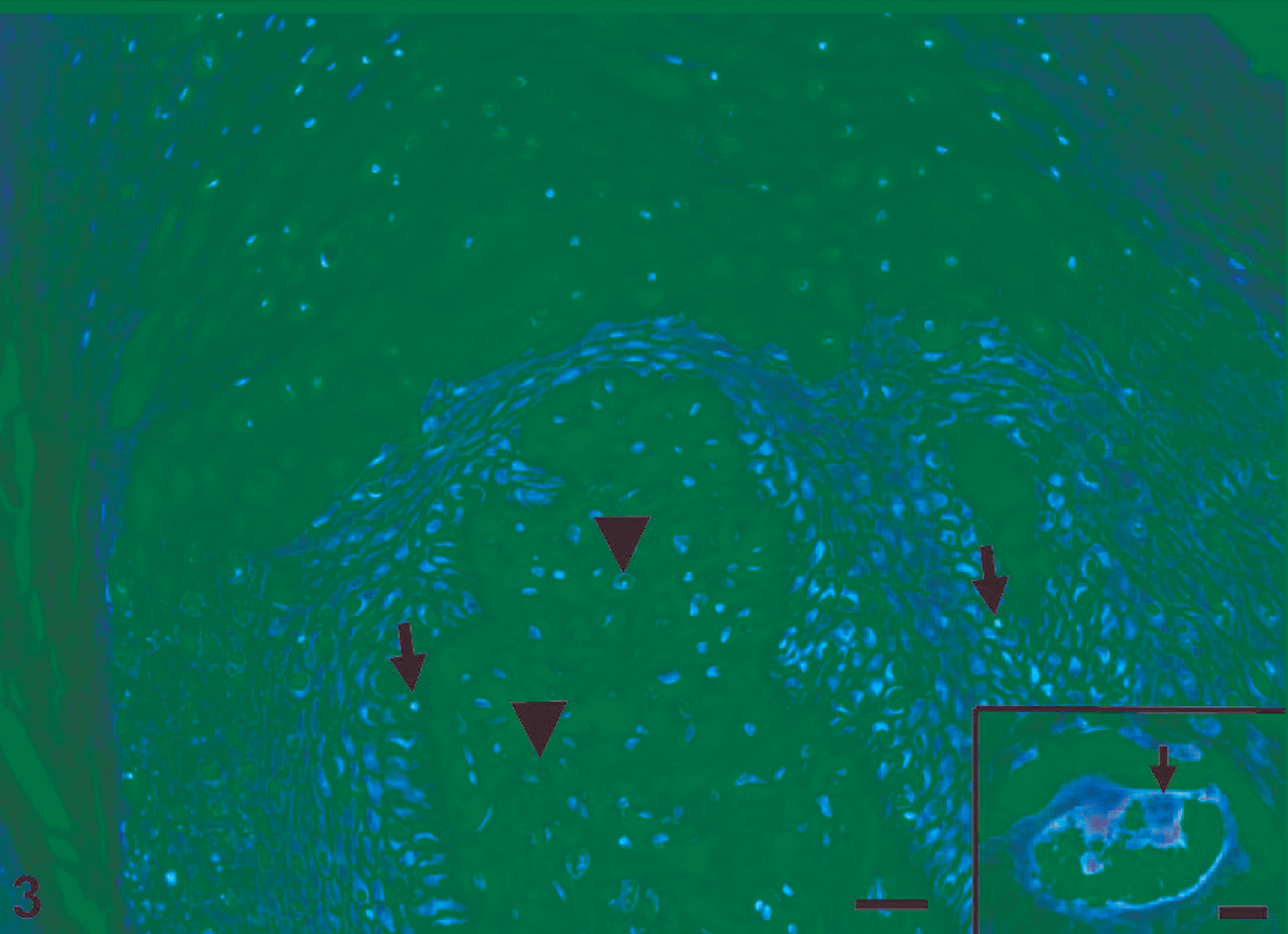

Skin; water buffalo. Histologic section of cutaneous fibropapilloma showing marked hyperkeratosis and acanthosis. Some cells with histologic signs of apoptosis (arrows) in the epidermis and large proliferating fibroblasts in the tumor stroma (arrowheads) in a skin fibropapilloma. HE. Bar = 40 µm. Inset: Large intranuclear eosinophilic inclusion in a degenerated cell (arrow). HE. Bar = 20 µm.
The connective tissue of the tumors consisted of fibrocytes of different sizes and shapes. There was mild infiltration of the connective tissue by lymphocytes and eosinophils, and presence of lymphocytes in the basal zone of the epidermis. The tumor stroma was composed of large stellate cells with extensive cytoplasm and plump nuclei with fine chromatin, immersed in a glycoproteinaceous matrix (Fig. 3).
Electron Microscopy
In all studied samples, papillomavirus-like particles were easily discriminated from tissues debris by their uniform, nonenveloped icosahedral structure of 55–60 nm in diameter and with capsids composed of symmetric capsomers (Fig. 4).
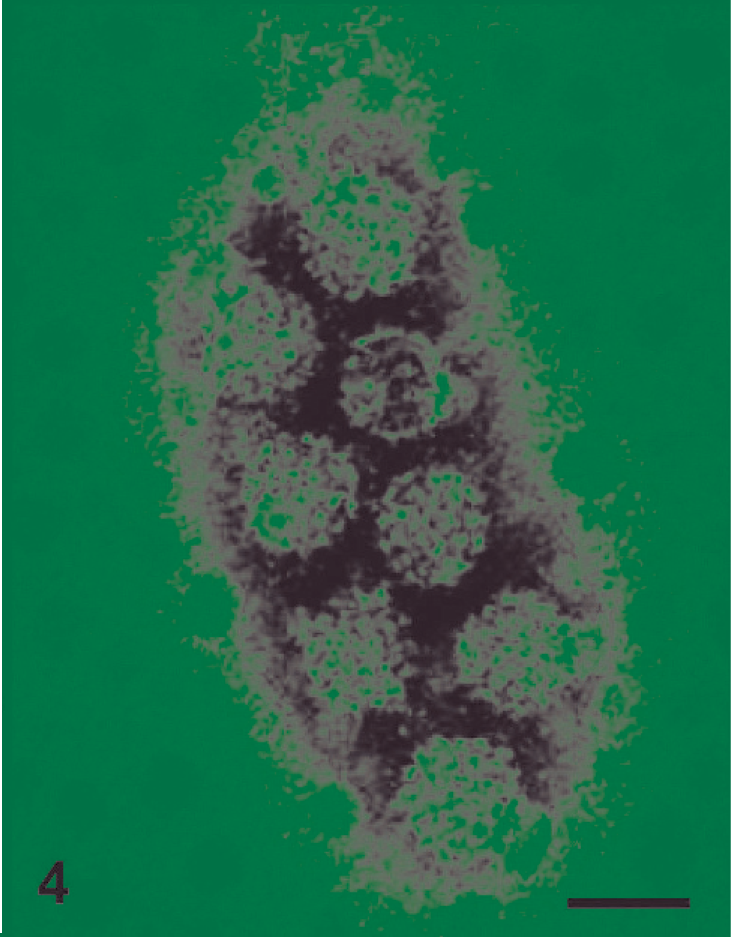
Skin; water buffalo. Negative stain transmission electron microscopy of papillomavirus virions with capsids composed of capsomers in icosahedral symmetry in a skin fibropapilloma. Ammonium molybdate stain. Bar = 100 nm.
Polymerase Chain Reaction and Nucleotide Sequencing
Amplification of the LCR by PCR is diagnostic of the presence of papillomavirus DNA. Following PCR amplification for BPV-1 LCR, the expected band of approximately 800 bp was detected in the DNA from a cutaneous fibropapilloma (Fig. 5A, left panel) but not in DNA from a vulvar fibropapilloma. However, following a second round of amplification, the 800 bp band was also detected in the vulvar sample (Fig. 5A, right panel). No band was ever amplified from controls having no DNA, showing that the reaction was specific.
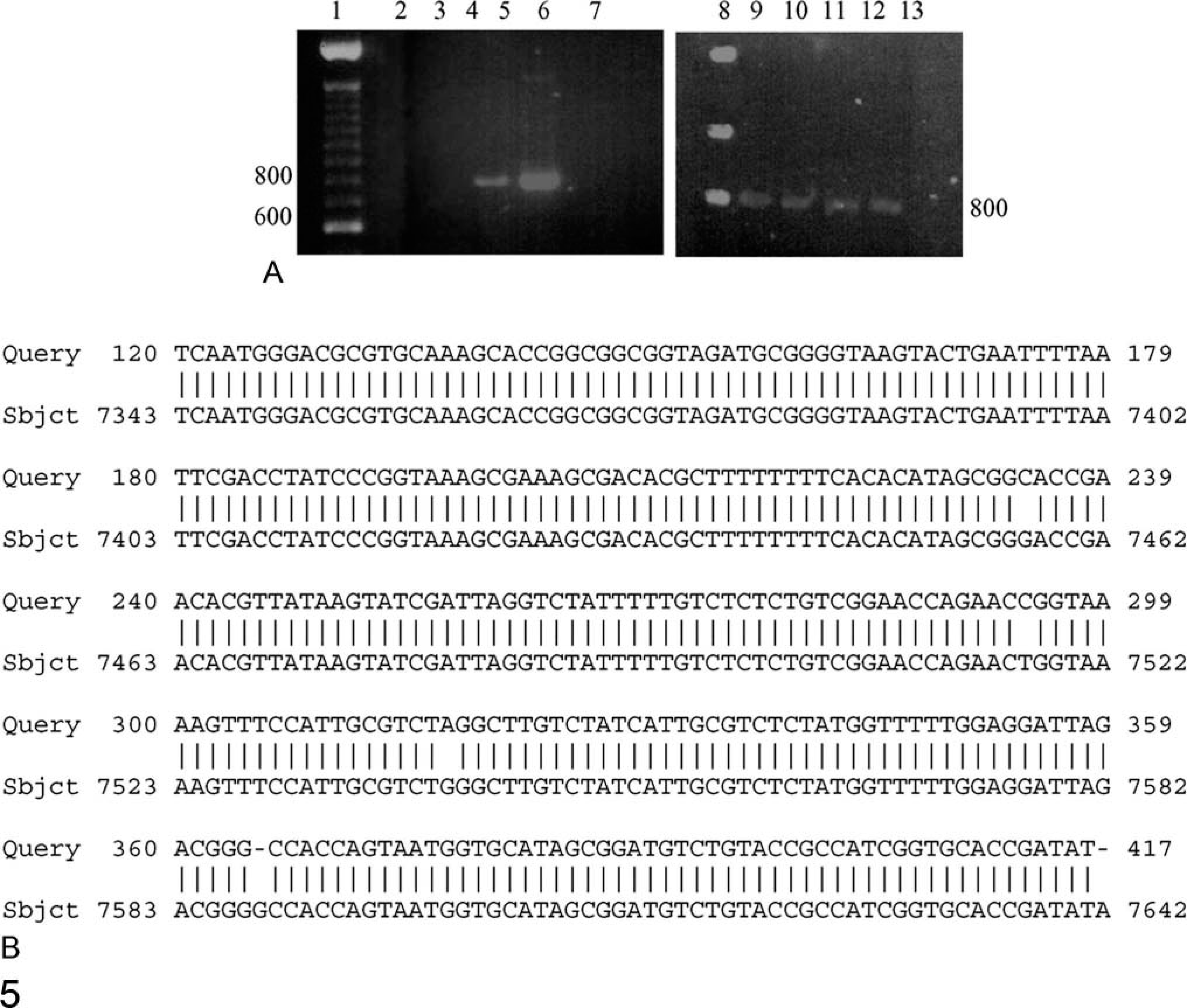
Water buffalo fibropapillomas harbour BPV-1 DNA. Fig. 5A. PCR amplification of a 790 bp fragment of the viral LCR in a cutaneous (left panel) and a vulvar (right panel) fibropapilloma. Lane 1, 100 bp ladder molecular weight marker (Gibco BRL, UK). Lanes 2, 7, and 13, negative controls (no DNA). Lane 3, negative control (koala DNA). Lane 4, vulvar fibropapilloma, after one round of amplification. Lane 5, cutaneous fibropapilloma. Lane 6, positive control (molecularly cloned BPV-1 DNA). Lane 8, mass ladder molecular weight marker (Gibco BRL, UK). Lanes 9–12, vulvar fibropapilloma after 2 rounds of DNA amplification. (The numbers next to the panels are base pairs.) Fig. 5B. Sequence homology between the amplicon from the vulvar fibropapilloma (Query) and BPV-1 LCR (Subject). The same level of homology was found for the amplicon from the cutaneous fibropapilloma. Both amplicons were sequenced in both orientations.
The PCR bands from both tumor samples were sequenced in the forward and reverse orientation in the ABI3100 sequencer using the LCR-F and LCR-R primers, respectively, and the resulting 4 nucleotide sequences were subjected to a BLAST search. The 2 sets of sequences from the cutaneous and the vulvar fibropapillomas were indistinguishable from each other and largely homologous to the LCR of BPV-1 (prototype sequence, accession number X02346) (shown for 300 nt of the vulvar sample in Fig. 5B).
Immunofluorescence: E5 Expression
Next, the tumors were analyzed for the expression of the oncoprotein E5 by immunofluorescence. This process demonstrated clearly the presence of the E5 protein in the tumor cells (Fig. 6). Typically E5 was located mostly within the cytoplasm of basal and suprabasal cells and the superficial granular cell layer. Furthermore, some of the cells showed accentuated staining in a perinuclear region, consistent with localization in the Golgi apparatus. E5 staining was also observed within mid-spinous cell layers but was much less intense than basal E5 staining.
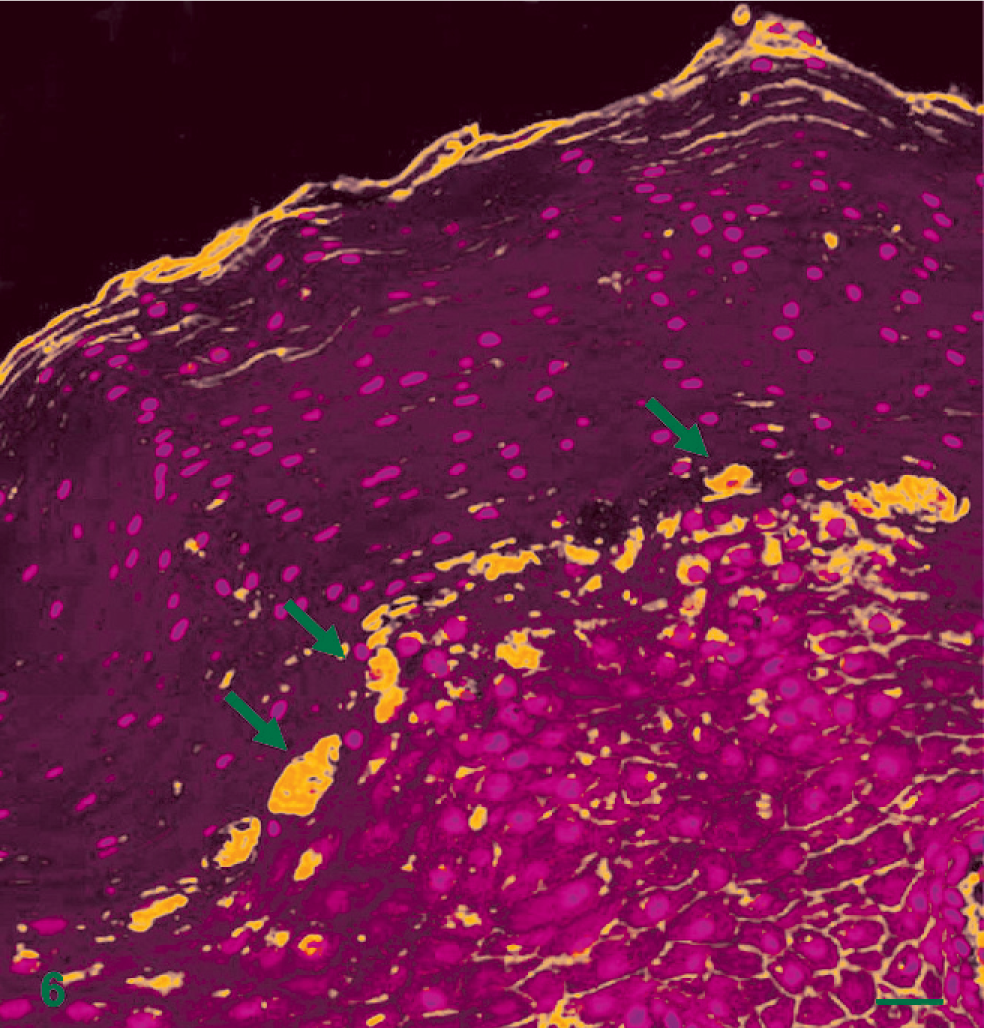
Skin; water buffalo. Strong immunoreactivity for E5 protein predominantly in the epithelial cells of the fibropapilloma. Nuclei counterstained with propidium iodide. Immunofluorescence and confocal microscopy. Bar = 200 µm.
Discussion
It has been established that both BPV-1 and BPV-2 are the etiologic agents of cutaneous and genital fibropapillomas in cattle. 9,17–19 However, it has been demonstrated that both BPV-1 and BPV-2 can jump species and infect bison and equids. 21,24
This study documents the presence of cutaneous and vulvar fibropapillomas also in water buffalo. Infection of buffalo by BPV-1 is based on several pieces of evidence: first, the histologic appearance of buffalo fibropapillomas is reminiscent of that of cattle fibropapillomas; second, papillomavirus-like particles can be detected in the fibropapillomas; third, BPV-1 DNA is present in the tumors; and fourth, the viral oncoprotein E5 is abundantly expressed. This last observation proves that BPV-1 is an active agent of fibropapillomatosis also in the water buffalo.
The localization of E5 in the transformed epithelial cells of buffalo fibropapillomas is similar to that reported for papillomas in cows during early stages of infection; namely, expressed strongly in the basal and granular layers and sporadically in the spinous cell layer. 1,5 Recently E5 has been found expressed not only in benign tumors but also in cancers of the urinary bladder of cattle, strengthening its role in neoplastic transformation in vivo. 4,6
In the buffalo, fibropapillomas are found on the back, in the gluteal region, and in the vulvar and perivulvar regions, all common sites of scraping and scratching. It is known that papillomavirus tumors tend to manifest themselves at site of trauma, 10,12 probably due to re-activation of latent virus, due to the release of inflammatory cytokines and stimulation of cell proliferation leading to tumor development. 10,12
The presence of BPV-1 in water buffalo tumors raises the question of the provenance of infection. Buffalo are closely related to cattle, both species belonging to the Bovidae family, and very often buffalo and cattle herds are kept together. These 2 facts would explain cross-species infection from cattle to buffalo. Moreover, buffalo fibropapillomas contain virus and are therefore productive for virus, presumably infectious. Thus, once established in a buffalo, infection could spread from buffalo to buffalo, without cattle intermediary.
More studies are needed to establish if BPV-1 infection takes place from cattle to buffalo, between buffalo or even from buffalo to cattle.
Footnotes
Acknowledgements
We wish to thank Loredana Cozzolino for the excellent technical assistance in electron microscopy. This study was partially funded by the Grant 2008 (Ricerca corrente) from the Istituto Zooprofilattico Sperimentale del Mezzogiorno di Portici (IZSME).
