Abstract
Bovine viral diarrhea virus (BVDV) is an important pathogen that primarily infects ruminants, leading to several clinical problems including abortion. BVDV-specific antibodies were reported in a wide range of hosts within domestic and wildlife animal populations, and serological studies also indicated BVDV infection in buffaloes. The purpose of this study was to analyze the presence of BVDV in 2 water buffalo (Bubalus bubalis) herds with a history of abortion. Virus isolation from aborted fetuses and from maternal buffy coat and the molecular characterization of the isolates confirmed the presence of BVDV in these animals. The sequence analysis based on the 5′ UTR and Npro coding regions of the Pestivirus genome revealed that the isolates belong to subgenotype 1b of BVDV. The findings of this study also suggest a possible role of BVDV in causing congenital infection in water buffalo. Its presence in fetal tissues as well as in maternal blood raises questions about the possible development of clinical disease or its influence in abortions in water buffalo.
Keywords
An indigenous breed of water buffalo, the Italian Mediterranean buffalo (Bubalus bubalis), is mainly reared in the southwestern regions of the Italian peninsula and is particularly appreciated for the production of mozzarella cheese. The effect of infectious agents on the reproduction of water buffalo has been investigated in several studies. Abortion in water buffalo has been associated with many bacteria such as Brucella spp., Coxiella burnetii, Arcanobacterium pyogenes, Bacillus licheniformis, Chlamydophila spp., and Leptospira spp. 8,9 However, little is known on the abortifacient role of viruses such as Bovine viral diarrhea virus (BVDV) in water buffalo. Bovine viral diarrhea virus (family Flaviviridae, genus Pestivirus) is an important viral pathogen of cattle that causes fatal diarrhea syndrome, respiratory problems, and reproductive failure. Infection during pregnancy can result in embryonic resorption, abortion, stillbirth calves, teratogenesis, or birth of persistently infected (PI) calves, which are the principal reservoir of the virus in nature and could later develop mucosal disease. Most of the reported BVDV infections refer to the Bovidae family, and only a few of them are related to buffalo. The studies related to these infections are mostly serological. 1,11 However, the isolation of BVDV from a pool of buffalo fetal serum, as well as the detection of BVDV nucleic acid from buffalo organs, has been recently reported. 2,5 In this study, the isolation and genetic characterization of BVDV from fetuses and maternal blood in buffalo dairy farms with a history of abortion are described.
From May 2007 to June 2007, a total of 9 cases of abortion occurred in pregnant heifers between the 4th and 7th month of gestation in 2 water buffalo herds located in Salerno's province of the Italian Campania region. The herds were officially free from brucellosis and consisted of 931 buffalo (farm A) and 99 buffalo (farm B), respectively. Only water buffalo and no cattle were present on the farms, and there was no evidence of any contact between the 2 farms. The following clinical samples were collected for laboratory investigations: 1 fetus with the respective maternal blood from farm A; 2 fetuses with the respective maternal blood from farm B. Microbiological tests were carried out on fetal tissue samples. The results were negative for the detection of Brucella spp., Listeria spp., Campylobacter spp., Mycoplasma spp., B. licheniformis, A. pyogenes, Chlamydophila spp., C. burnetii, Bubaline herpesvirus-1, Bovine herpesvirus 1, Neospora caninum, and Toxoplasma gondii. Fetal spleen and maternal blood samples (buffy coat) were also screened for BVDV antigen with the use of a commercial enzyme-linked immunosorbent assay (ELISA). a ELISA-positive samples were further examined for virus isolation and molecular assays as described below. Virus isolation was attempted on Madin-Darby bovine kidney (MDBK) cells free from contamination with any endogenous Pestivirus infection, and the cells were examined for the presence of virus by fluorescent antibody test (FAT). Total RNA was extracted from FAT-positive cultured cells with TRIzol reagent b according to the manufacturer's recommendation. Synthesis of complementary DNA (cDNA) was carried out with the use of random primers. c The genomic region encoding the highly conserved 5′ UTR of the Pestivirus genome was amplified with the use of primers 324 and 326 flanking a 288-base pair (bp) DNA fragment. 14 The genomic region encoding autoprotease Npro was amplified with the use of a nested polymerase chain reaction (nPCR). In the first PCR, the outer primers OI 100 and OL 1400R 2,3 were used, and reverse transcription polymerase chain reaction (RT-PCR) was done with the SuperScript III One-Step RT-PCR b according to the manufacturer's recommendation. In the second PCR, the first round PCR product was amplified with primers BD1/BD3 flanking a 428-bp fragment. 13 In cases of positive results, a 428-bp fragment was cut from the gel and purified with the QIAquick gel extraction kit. d Nucleotide sequences were determined by sequencing cycle with the Big Dye Terminator v1.1 Cycle Sequencing kit e and an ABI PRISM sequencing device. e Nucleotide sequences were aligned by the ClustalX version 1.83 analysis program. 12 Phylogenetic trees were calculated with the MEGA version 3.1 program on the basis of the neighbor-joining algorithm method. 10 The robustness of the phylogenetic analysis and significance of branch order were determined by the bootstrapping method and was carried out on 10,000 replicates. 7 Sequence data from this report were deposited in European Molecular Biology Laboratory nucleotide sequence data library.
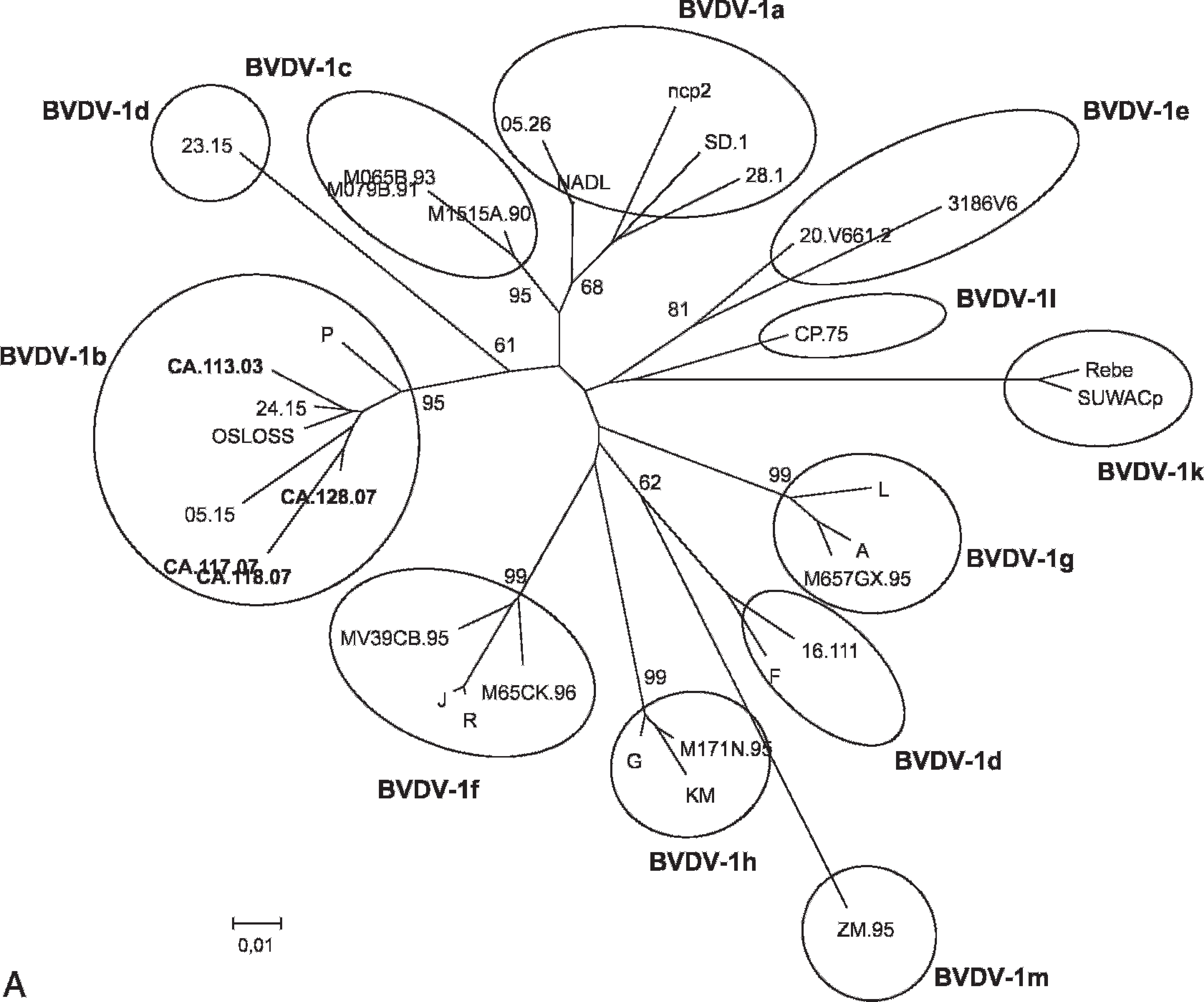
Phylogenetic trees based on analysis of 244 nucleotides derived from the 5′ UTR (
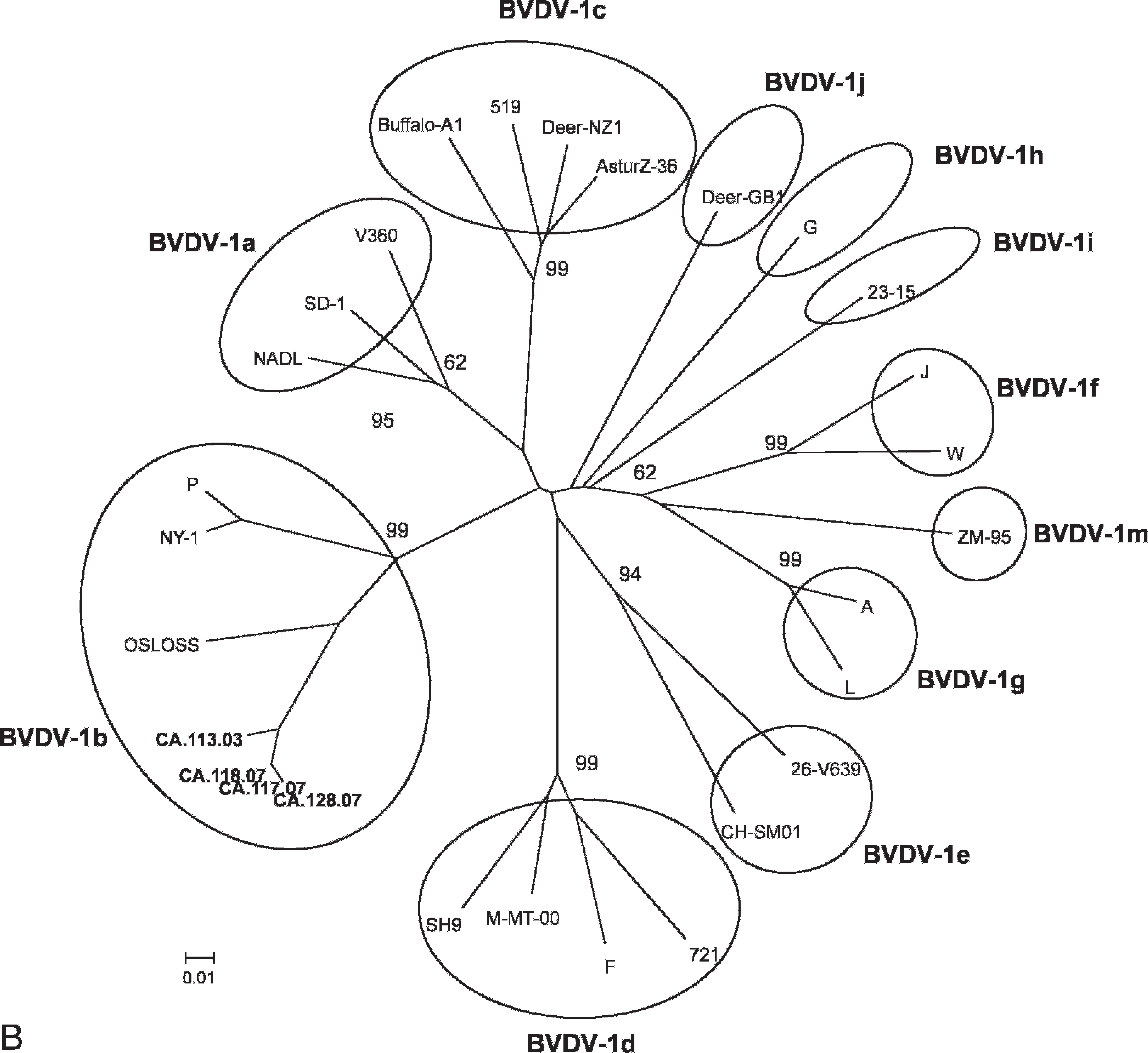
Continued.
Four pestiviruses were isolated on confluent monolayers of the cell line MDBK: 2 from the fetus and the respective maternal blood from farm A and 2 from the 2 fetuses from farm B. The viruses were sequenced in the 5′ UTR and Npro regions. Genetic typing revealed that the isolates were related to BVDV-1 species, and the topology of the trees indicates that they belong to the subgenotype BVDV-1b (Fig. 1). To determine the presence of Pestivirus antibodies, the maternal serum samples obtained from 3 dams from which the virus-positive fetuses originated were tested with the ELISA BVDV Antibody test kit f and found negative. Thirty days later, a second sampling was performed. The result of the ELISA test this time indicated that 2 dams seroconverted, whereas the virus-positive dam still resulted serologically negative to BVDV. Interestingly, this latter was simultaneously tested for BVDV antigen and still found virus-positive. Unfortunately, it was not possible to obtain a larger number of blood samples to monitor the serological and virological situation at the farm level.
In a study on buffalo calves experimentally infected with BVDV, it has been shown that the clinical signs are mild compared with those in cattle, but they are very similar in temporal relation (Hegazy AA, Shelaby MA, Bagouri, et al.: 1991, Studies on buffalo calves experimentally infected with bovine viral diarrhoea virus. Proceedings of the Third World Buffalo Congress, 13–17 May, Varna, Bulgaria, 9:105–115). However, little is known about BVDV pathogenesis. Assuming that the pathogenesis in buffalo is similar to cattle, the fetuses from which viruses were isolated probably originated from immunotolerant and PI dam (farm A) or with an acute transient form of BVDV (farm B). Cattle are in fact seronegative at this stage and develop serum antibodies later on. 4 The serological and virological investigations ruled out the possible role of a PI dam in at least 1 herd. According to our findings, because the fetal age of the immune system development in buffalos is unknown, it is difficult to predict the gestational age at which BVDV could cause persistent infection in this species. Immune system competence of bovine fetuses occurs before day 125–150 of gestation, and fetuses in the first and early second trimester are most susceptible to become PI. The gestation length for water buffalo is about 1 month longer, therefore the gestational timeframe at which persistent infection in buffalo can occur could be different from cattle. In any case, the PI status in a buffalo herd might need consideration when planning BVDV eradication or control strategies. For this reason, further research is needed to determine when persistent infection in buffalo fetuses might occur and whether it could act the same way as in cattle. Reproductive deficiency and abortions are commonly associated with BVDV infection of pregnant animals. In the examined cases in which common pathogens causing pregnancy loss were not detected, the evidence of BVDV seems to be associated with abortion, although that can only be speculated.
In conclusion, the main interesting finding was the detection of BVDV-1b in 2 buffalo herds. The subgrouping of BVDV-1b might be correlated with the geographical origin of these viruses. The majority of the Italian isolates to date characterized are related to the same subgenotype. 6 The findings of the current study also suggest a possible role of BVDV in the etiology of abortion and underline the need for the adoption of adequate prevention and control measures. Even more important, the presence of BVDV could represent a threat to buffalo health that was previously unknown in the Mediterranean buffalo.
Footnotes
a.
BVD/MD Ag Mix, Institut Pourquier, Montpellier, France.
b.
Invitrogen Corp., Carlsbad, CA.
c.
Amersham Biosciences Corp., Piscataway, NJ.
d.
Qiagen GmBH, Hilden, Germany.
e.
Applied Biosystems, Foster City, CA.
f.
Institut Pourquier, Montpellier, France.
