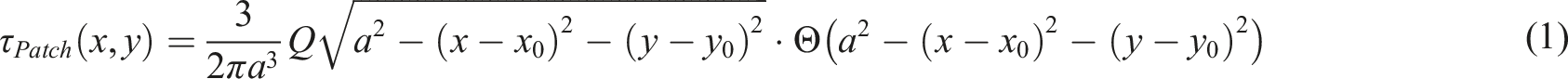Abstract
Mechanoadaptation, a process by which living cells adjust to the mechanical properties of the environment, is fundamental to their function and behavior. Generating mechanical forces is the main way a cell can acquire information about the physical nature of the immediate micro-environment. Crucially, mechanical force during this interaction is the sole stimulus upon which the cell performs mechanoadaptation. However, quantifying this mechanical interaction is notoriously challenging due to the interdependent relationship between mechanical force, mechanical substrate stiffness, and the resulting spatial displacements. A critical issue, often overlooked in common measurement techniques like traction force microscopy, is that a change in substrate stiffness inherently alters measurement sensitivity. This perspective argues that this oversight can lead to misinterpretation of cellular function and behavior, creating an ’illusion of adaptation.’ We present a conceptual framework to highlight the importance of accounting for the measurement sensitivity and advocate for a more accurate quantification of the biological processes governing mechanoadaptation to avoid pitfalls and advance the field of mechanobiology.
Keywords
Introduction
The ability of living cells to adapt to the mechanics of the immediate tissue micro-environment is fundamental to many biological processes in health and disease.1–8 This phenomenon, termed mechanoadaptation, describes how cells tune their body’s own mechanical state to fulfill function and behavior in response to a dynamically changing mechanical environment.2,9–11 The means of mechanical forces is the main way a cell can acquire information about physical changes of the envionment, mechanical properties, and mechanical force generation. Mechanical force is the stimulus upon which the cell tunes their own mechanical force generation, mechanical properties, dynamics, function, and behaviour to achieve mechanoadaptation. As we will illustrate in this perspective, the interplay between cellular behavior and measurement limitations can create an ’illusion of mechanoadaptation,’ leading to significant misinterpretation of experimental results.
Over the past two decades, the field of mechanobiology has made remarkable strides in revealing the profound impact of mechanical forces on cell function.12–18 However, this progress has also exposed a critical weakness: the quantitative assessment of mechanoadaptation remains not only a formidable technical challenge but a source of significant ambiguity.19–25 We argue that a fundamental oversight in how these adaptive responses are measured — specifically, the failure to account for variable technical measurement sensitivity — can lead to misinterpretation of cellular function and behavior, creating an ’illusion of adaptation’ that pervades the literature. This perspective introduces a conceptual framework to illustrate this issue and advocates for a renewed focus on the direct and accurate quantification of the physical parameters that report on cellular mechanoadaptation.
At the core of this fundamental oversight lies the intricate interdependence of the primary physical quantities: cellular-exerted mechanical force, environment’s mechanical stiffness, and the resulting cellular spatial displacement.18,26–30 This interdependence is a direct consequence of the constitutive laws of materials, where any measurement of mechanical force is inferred from a resulting spatial displacement, the magnitude of which is governed by the material’s mechanical stiffness. Accurately measuring relevant parameters often requires assumptions about the others, creating a measurement problem that is not unique to any single technique but is a general feature of many mechanobiological assays.30–33
While the powerful technology of Traction Force Microscopy (TFM) to quantify cellular mechanical forces by measuring substrate displacements, has been chosen as a primary example in this perspective,34–41 the principles discussed are broadly applicable to a range of measurement modalities, from atomic force microscopy to microrheology.42–45 In TFM, a cell’s response to varying mechanical substrate stiffness is a common experimental paradigm. But, the fundamental reality is that a change in mechanical stiffness inherently alters the measurement system’s sensitivity. Specifically, for a constant cellular mechanical force, the resulting substrate displacement will be smaller on a stiffer substrate and larger on a softer one. This inverse relationship between stiffness and spatial displacement for a given mechanical force can lead to profound ambiguity: is a cell failing to adapt its mechanical force, or is our measurement apparatus simply less sensitive under stiffer mechanical conditions where spatial displacements are minimal? Conversely, on mechanically very soft substrates, large spatial displacements might exceed the tracking algorithm’s limits, leading to measurement failure. This creates a scenario where an apparent change in cellular behavior could be an artifact of the measurement system.
In the following, to address some of these open questions, we discuss the importance of a rigorous understanding of the measurement system’s technical dynamic range, such as that of a living cell, and its limitations for the accurate interpretation of cellular mechanoadaptation. To aid the classification of the different forms of mechanical force adaptation, we introduce a conceptual framework to describe mechanoadaptive phenotypes. To disentangle the complex parameter interdependencies, computer simulations are employed to demonstrate how measurement sensitivity varies with measurement system parameters. We demonstrate that failing to account for these dynamics risks significant misinterpretation of cellular function and/or behavior, and a renewed focus on the direct and accurate quantification of the physical processes that govern mechanoadaptation is advocated to propel the field forward.
Mechanoadaptation: A measurement challenge
To illustrate the challenges in quantifying mechanoadaptation, we present a simple thought experiment. The technology of TFM is utilized as a specific example to probe the current shortcomings inherent to many measurement technologies. This model is intentionally simplified and does not purport to reflect the intricate cell biological details of our example of cell migration. Rather, its purpose is to provide a clear discussion of the frequently overlooked limitations of measurement and the potential for misinterpretation. Note, we refer to simply force and stiffness instead of mechanical force and mechanical stiffness for brevity in the following sections.
We introduce a conceptual framework for categorizing cellular adaptive behaviors. Three primary mechanoadaptive phenotypes are defined: positive (+ve), neutral (non), and negative (−ve) mechanoadaptation, based on how cellular forces respond to changes in environmental stiffness (Figure 1(a), top left). In positive mechanoadaptation (Figure 1(a), blue lines), a cell increases its force generation as the substrate stiffness increases. In contrast, a non-mechanoadaptive cell (Figure 1(a), grey lines) exerts a constant force regardless of stiffness. In negative mechanoadaptation (Figure 1(a), magenta line), the cell decreases its force output on stiffer substrates. Each of these behaviors results in a distinct substrate displacement pattern (Figure 1(a), bottom left). The Mechanobiologist’s dilemma. (a) Conceptual framework for mechanoadaptation. Top left: Force-stiffness relationships for positive (+ve, blue), neutral (non, grey), and negative (-ve, magenta) mechanoadaptation. Bottom left: Corresponding displacement-stiffness relationships. Right: Schematic of a TFM experiment showing a cell on soft and stiff substrates. (b) TFM simulation workflow. A ground truth (GT) traction field is used to generate a displacement field, which is then converted into synthetic bead images. Displacements are recovered and compared to the GT. (c) Representative TFM simulation results. Left: GT traction field. Middle: Recovered displacement field at point B (good reconstruction). Right: Reconstructed traction field. (d) Displacement fields at points A, B, and C, showing tracking errors at high displacement (C) and low signal-to-noise at low displacement (A). (e) Force-stiffness phase diagrams for SVD and accuracy metrics. The trajectories of the three mechanoadaptive phenotypes are overlaid, showing that measurement sensitivity is not uniform across the parameter space.
Consider a TFM experiment where a cell is placed on a soft (
To investigate how the measurement system influences the ability to determine these cellular behaviors, computer simulations were carried out across a range of physical parameters commonly encountered in TFM experiments. The simulation workflow is summarised in Figure 1(b) and detailed in Box 1. A ground truth (GT) traction field of two circular adhesions and the corresponding substrate displacements for different forces and substrate stiffnesses was simulated. This displacement field was then used to generate synthetic microscope bead images, from which displacements were recovered using a single particle tracking approach.
Motivated by forces typically produced by living cells, computer simulations were conducted at a range of forces (0.001–1 nN) and substrate stiffnesses (0.2–4 kPa). To assess the quality of the reconstruction, three complementary metrics were used: Singular Value Decomposition (SVD) 37 to measure the overall quality of the displacement field recovery, a ’precision’ metric to quantify the probability of recovering the correct displacement amplitude, and an ’accuracy’ metric to quantify the probability of resolving the correct number of force foci.
The discussion of the computer simulations are presented as force-stiffness phase diagrams for each quality metric (Figure 1(e)). The phase diagrams reveal that the quality of displacement recovery is highly dependent on the specific experimental parameters. Three representative points are highlighted: at point C (high force, low stiffness), large displacements lead to tracking errors; at point B, there is good reconstruction; and at point A (low force, high stiffness), small displacements approach the localisation uncertainty of the measurement system (Figure 1(d)). Crucially, when the trajectories of the defined mechanoadaptive phenotypes are plotted onto these diagrams, it is seen that they are subject to non-uniform confidence levels. A non-adapting cell (grey line) shows large measurement uncertainty at low stiffness, while a negatively adapting cell (magenta line) is challenging to measure at both low and high stiffness extremes. In contrast, a positively adapting cell that maintains constant displacement (blue line) can be measured with consistent sensitivity. This demonstrates that the underlying cellular behavior directly influences the quality of the TFM experiment, and that without understanding the measurement system’s dynamic range, one could easily misinterpret the results. Together, the computer simulations reveal that TFM measurement quality is highly dependent on the specific force and stiffness conditions, meaning that different cellular adaptation behaviors are measured with varying degrees of reliability. To generate both the ground truth traction and displacement fields, previous work by Blumberg and Schwarz
46
is taken advantage of. Specifically, a dipole of two circular adhesion sites with opposing traction vectors (Figure 1(b)) is simulated. The explicit form of the so-called hertzian-like patch is described in equation (1), where x, y are the spatial coordinates, x0 and y0 the center coordinates of the patch, Q the force vector applied on the substrate, a the radius of the circular adhesion site, and If multiple adhesion sites are present, as is the case, their solution is a linear combination of the individual patches. To determine the emerging displacement field, equation (2) is used, in which u represents the displacement field, x, y, z the spatial coordinates, S the surface, and G the Green Function for the elastic halfspace (Bussinesq solution). For the simulation, a dipole of two adhesion sites of radius 1 Next, points are randomly sampled across the entire field of view representing the positions of fluorescent beads, with a spatial density of 1 To incorporate the optical characteristics of the imaging modality, the experimental point spread function (PSF) and background noise characteristic of both a confocal microscope and total internal reflection fluorescence structured illumination (TIRF-SIM) microscope were measured. Utilising this experimental calibration, the distribution of points before and after displacement were converted into synthetic fluorescence microscopy images. This was achieved using a Gaussian distribution with a full width half maximum (FWHM) of 200 nm for confocal and 100 nm for TIRF-SIM, and a photon count of up to 4096 for confocal and 8000 for TIRF-SIM based on an underlying 264 by 264 pixel grid of 30 nm spacing. Secondly, normally distributed noise was added to the images, with a sigma of six photons for confocal, and a sigma of 2100 for TIRF-SIM, representing the experimentally acquired imaging noise. Lastly, Poisson noise was added to recapture detector characteristics. Determination of the experimental displacement field was done entirely in trackpy,
47
a common tool to identify, track and link circular objects in 2D. First, beads were identified using the Crocker-Grier centroid-finding algorithm,47,48 defining an object detection diameter that varied with the expected PSF size ( The reconstruction of the example displacement field was performed using the well-established Fourier Transform Traction Cytometry (FTTC) technique. In this work the approach as implemented by Blumberg and Schwarz
46
was chosen. The singular value decomposition metric measures the distance of singular values between the GT displacement and the reconstructed displacement field.37,49 This metric was used, as shown in equation (3) to assess the general quality of the reconstructed displacement field in relationship to the ground truth, where For each parameter set, generation of a random bead distribution, synthetic image generation and displacement recovery were performed 100 times. To ease comparability, the raw average SVD values were converted into the logarithmic scale and normalized according to the dynamic range of global values, resulting in values ranging from 0 to 1. Please note that when using the SVD metric for a comparative study with changing maximum values like the SVD metric requires normalisation according to the total sum of values, not the number of pixels. The precision metric was used to quantify the probability of recovering the correct displacement amplitude compared to ground truth. Explicitly, the precision metric measures the fraction of computer simulations where the upper 20% of displacement values of a TFM recovery resided within the same regime as those from ground truth.
37
For a given recovery and ground truth, both images were thresholded for 80% of their respective max value. Afterwards, the mean value of the remaining pixels was compared. If this average displacement peak of the recovery was within 20% of the ground truth the recovery was deemed successful. For visualization, the ratio of successful recoveries was taken from the 100 repetitions, resulting in a probability range from 0 to 1. The accuracy metric was utilised to determine the ability of TFM to recover the correct number of displacement foci resulting from the Hertzian dipole simulation. To quantify the ability of TFM to spatially resolve nearby foci, the displacement peaks of the hertzian dipoles are detected by applying a threshold of 80% to an image and counting the number of objects consistent with the size of the original ground truth foci (Area >250 pixels = min. 0.225 The calculations of the sensitivity metrics were repeated eight times for the respective computer simulation. The respective standard deviations were computed from these repeats. In each run the synthetic TFM experiments were repeated 100 times for each force stiffness pair.Box 1: TFM computer simulations
Ground truth and experimental displacement field
Synthetic fluorescent bead image generation
Displacement field recovery
Force reconstruction
Box 2: Metrics & statistics three complementary sensitivity metrics are computed to evaluate the sensitivity of the force-stiffness phase diagrams
Singular value decomposition
Precision
Accuracy
Statistics
Summary and outlook
Measurement dilemma of mechanical force and stiffness
The computer simulations demonstrated how the ability to accurately measure cellular forces is fundamentally linked to substrate stiffness. 37 The inherent limitations of the measurement system creats a “Goldilocks zone” for any given force level, where the stiffness is just right for an accurate measurement. The changing measurement sensitivity is accopined by mechanoadaptation of the cell. This means that if you can measure it, the cell might move out of your optimal window. When you can not measure it, this is where the cell might do more interesting things.
A critical challenge arises from the dynamic interplay between measurement sensitivity and cellular mechanoadaptation. As measurement sensitivity changes, cells may undergo adaptive responses that shift their behavior outside the optimal detection range. Consequently, when cellular activity is measurable, it may no longer reflect the most informative or physiologically relevant state. Paradoxically, the most intriguing or significant cellular behaviors may occur precisely when they fall below the threshold of detection.36,37,39
The illusion of mechanoadaptation
The interplay between cellular behavior and measurement limitations can create an ’illusion of mechanoadaptation,’ leading to significant misinterpretation of experimental results. The illusion arises when experimental observations are confounded by the technical constraints of the measurement system. A primary source of this illusion is the failure to directly measure all relevant physical parameters, particularly cellular forces. For example, if one observes constant substrate displacement across substrates of varying stiffness, it might be tempting to conclude that the cell is not adapting. Nevertheless, maintaining the same displacement on a stiffer substrate requires a significant increase in force generation. Relying solely on displacement would therefore create the illusion of a non-adapting cell, masking a potent biological response. Also, the inherent variability in measurement sensitivity across different mechanical conditions can create misleading results. As the computer simulations show, the ability to accurately measure force is highly dependent on substrate stiffness. On stiff substrates, displacements may be too small to be reliably detected, leading to an apparent decrease in force that is merely a measurement artifact. Conversely, on very soft substrates, large displacements can exceed the limits of tracking algorithms, again leading to inaccurate measurements. A researcher might observe that a cell’s measured traction forces decrease on stiffer substrates and conclude that the cell is negatively mechanoadapting. In reality, this could simply be an artifact of the measurement system becoming less sensitive. Without a clear understanding of the system’s dynamic range, it is easy to mistake measurement artifacts for true biological phenomena, creating a compelling but false ’illusion of mechanoadaptation’.
Implications of misinterpreting mechanoadaptation
Accurately quantifying mechanoadaptation is not merely a technical exercise; it is fundamental to the understanding of cell biology.50–54 From immune cell activation and cancer metastasis to stem cell differentiation,12,14,55–57 the ability of cells to adapt to their mechanical environment is a key ability to maintain, or even adapt their function and henceforth purpose.21,58–61 Misinterpreting a cell’s adaptive response can lead to flawed models of these critical biological processes.23–25 For example, an experiment might show that T cells generate the same substrate displacement on soft and stiff substrates. Concluding that a T cell does not adapt its force during activation on a stiff substrate might lead to incorrect assumptions about the role of mechanics in the immune response. This conclusion is indeed wrong, but seems trivial. If the displacement is the same, the force must be higher. Therefore, a rigorous, quantitative understanding of mechanoadaptation is essential for progress in these fields. In short, misinterpreting mechanoadaptation can lead to flawed models of critical biological processes like immune response and cancer metastasis, highlighting the need for accurate quantification.
The implications of misinterpreting mechanoadaptation extend to the subcellular scale.19–22 The fields of genomics, proteomics, and other ’omics are increasingly being integrated with mechanobiology to understand how mechanical cues regulate gene expression and protein activity. If the mechanical input to a cell is incorrectly characterized—for example, if a cell is assumed to be experiencing low force when it is in fact generating high force on a stiff substrate—then the resulting correlations between mechanical state and molecular profiles will be flawed. This can lead to the misidentification of mechanosensitive genes and signaling pathways, sending researchers down fruitless paths. A correct interpretation of the cell’s mechanical state is a prerequisite for a meaningful integration of ’omics data and for building accurate models of mechanotransduction.62,63
Advanced tools to quantify mechanoadaptation
While we focus on TFM in this manuscript it is by no means the only technique capable of probing for mechanoadaptation. Other popular techniques include Förster Resonance Energy Transfer based molecular tension sensors (FRET-TS), atomic force microscopy (AFM) or the newly emerging brillouin microscopy (BM). FRET-TS are genetically encoded constructs that consist of a spring-like linker protein or DNA hairpins, linking together two fluorophores. Upon mechanical strain the fluorophores are separated and the FRET efficiency drops. Quantifying this change in FRET efficiency with fluorescence microscopy allows for the investigation of intra- or intercellular forces in situ. These novel tension sensors have already been employed to quantify cytoskeletal tension, focal adhesions and cellular junctions, two of which were even shown to work in vivo.64–68 AFM is one of the most widely used techniques in mechanobiology and has been extensively applied to quantify the material properties of cells and their components.31,69 Measuring the displacement of an elastic cantilever with an interferometer, while approaching closer and closer to the specimen system, the immediate mechanical response of the cell 70 can be quantified. However, AFM remains limited to the cell surface and 2D cell cultures. The most recent addition to the mechanobiological toolbox is Brillouin Microscopy. BM is a non-invasive optical technique that probes the mechanical properties of materials by analyzing light scattered from acoustic phonons within the sample. When a focused laser beam interacts with these thermally induced density fluctuations, the scattered light undergoes a small frequency shift known as the Brillouin shift.71,72 By scanning the sample and measuring these frequency changes with a high-resolution spectrometer, BM generates three-dimensional maps of local viscoelasticity without requiring physical contact or labeling. This capability makes it particularly valuable for studying the biomechanics of living cells and tissues under physiological conditions.
Future quantification of mechanoadaptation
In conclusion, this perspective highlights a critical, yet often overlooked, challenge in the quantification of cellular mechanoadaptation. The central tenet of the argument is that the sensitivity of measurement tools, particularly in TFM, is not constant across substrates of varying stiffness. We call for a more critical and sophisticated approach to the quantification of cellular mechanoadaptation with a focus on the adaption of cellular force generation. The principles discussed, though exemplified by TFM, are not confined to it; they represent a universal challenge in a field where the act of measurement can influence the specimen system under observation. By acknowledging the inherent technical limitations and sensitivity of measurement systems, the “illusion of mechanoadaptation” can be avoided and a move towards a more accurate and comprehensive understanding of how cells navigate their mechanical world can be made. To foster more rigorous and reproducible mechanobiological assays, we propose the following guidelines: • • • • •
The development of new technologies to quantify mechanoadaptation in vivo and in complex three-dimensional environments remains a major challenge for the field.7,8,10,54,73,74 Furthermore, it is important to recognize that cells can dynamically change their own mechanical properties, such as stiffness, adding another layer of complexity that has not been considered in this perspective. Nevertheless, a clear understanding and reporting of the measurement system’s measurement sensitivity, as outlined, is a necessary and immediate step that will foster more rigorous and reproducible mechanobiological assays. Tuning cellular mechanical force generation is fundamental to cell function, and only through adequate quantification can the field fully unravel the biological mechanisms underlying mechanoadaptation in health and disease.
Footnotes
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: M.F. and H.C.-Y. acknowledge support from the UK Research and Innovation Medical Research Council (APP23835)), Engineering and Physical Sciences Research Council (2204AZ003/CP5 and EPSRCEP/X033015/1), and Cancer Research UK (DRCPFA). M.I. was supported by Erasmus+ (Generation 2014-2020).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.