Abstract
Phosphatidylinositol (PI)-transfer proteins (PITPs) have been long recognized as proteins that modulate phosphoinositide levels in membranes through their intrinsic PI/PC-exchange activity. Recent studies from flies and mammals suggest that certain PITPs bind phosphatidic acid (PA) and possess PI/PA-exchange activity. Phosphoinositides and PA play critical roles in cell signaling and membrane trafficking, and numerous biochemical, genetic and functional studies have shown that PITPs regulate cellular lipid metabolism, various signaling pathways and intracellular membrane transport events. In this mini-review, we discuss the function of mammalian PITPs at the Golgi and ER-Golgi membrane contact sites (MCS) and highlight DAG (Diacylglycerol) as a central hub of PITPs functions. We describe PITPs-associated phospho-signaling network at the ER-Golgi interface, and share our perspective on future studies related to PITPs at MCSs.
Introduction
Phosphoinositides are important constituents of cellular membranes, represent a small fraction (5–7%) of total cellular phospholipids, of which the PI pool is the largest, comprising ∼90% of total phosphoinositides. All phosphoinositides are derivatives of PI, which is synthesized at the ER from PA (Lev, 2012; Blunsom and Cockcroft, 2020b). Among the seven polyphosphoinositides (PPIns), PI(4,5)P2 and PI4P are relatively abundant and mainly found at the plasma membrane (PM) and Golgi apparatus, respectively. Other PPIns, including PI3P, PI5P, PI(3,4)P2, PI(3,5)P2, PI(3,4,5)P3 are enriched in different cellular organelles and considered as organelle-specific markers (Balla, 2013). The distinct distribution of phosphoinositides among organelle membranes is maintained despite the dynamic membrane trafficking events and metabolic processes (De Matteis and D'Angelo, 2007), and is crucial for preserving the structural and functional identity of different membrane compartments, including the Golgi apparatus.
The Golgi complex is the central organelle of the secretory pathway involved in processing, sorting and transport of lipids and cellular proteins to appropriate subcellular destinations. In addition, the Golgi functions as a membranous scaffold for diverse lipid-binding, lipid-modifying and signaling proteins, and thus, an active platform for different lipid metabolic processes and signaling events (Wilson et al., 2011). Among the different Golgi lipids, PI4P and DAG are considered as key regulators of Golgi-associated transport and signaling events (Lev, 2006; De Matteis et al., 2013).
The involvement of PITPs in Golgi-mediated transport was first evident in studies on Sec14p, the major PITP in Saccharomyces cerevisiae, and the discovery that Sec14p is essential for protein trafficking from the yeast Golgi complex (Bankaitis et al., 1989; Bankaitis et al., 1990). Subsequent studies on Sec14p, as well as on the mammalian PITPs, uncovered their important regulatory roles in lipid metabolism, membrane trafficking and phosphoinositide signaling (Cockcroft, 1999; Huijbregts et al., 2000; Li et al., 2000; Cockcroft, 2001; Wirtz, 2006; Grabon et al., 2019). The discovery that the large multi-domain PITPs, Nir2 and Nir3 bind to VAPs and function at ER-Golgi MCSs (Amarilio et al., 2005; Peretti et al., 2008) further highlighted the impact of this class of lipid-transfer proteins (LTPs) on the Golgi structure and function, and its physical and functional link of the Golgi to the ER.
In this mini-review we describe the functions of mammalian PITPs at the Golgi and ER-Golgi MCSs, and discuss how their lipid transfer activity is implicated in different lipid-metabolic pathways at the ER-Golgi interface to coordinate membrane trafficking and cellular signaling pathways.
PITP Family
There are 5 genes in the mammalian genome that encode for proteins containing a PITP domain. The family of PITP proteins comprise of Class I and Class II defined by their sequences. Class I PITPs (α and β) are small ∼35 kDa proteins comprising of a single PITP domain whilst Class II PITPs comprise of three proteins. Two are large proteins (∼160 kDa) with multiple domains variously called PITPNM1 or Nir2 and PITPNM2 or Nir3. The third-Class II protein, PITPNC1 (38 kDa), comprises of a PITP domain followed by an unstructured extension of 80 amino acids (Figure 1). Splice variants of PITPβ (sp1 and sp2) and of PITPNC1 (sp1 and sp2) have been identified that differ in their C-terminus (Figure 1). The Class I PITPs (PITPα and β) bind a single lipid, either PI or phosphatidylcholine (PC) and facilitate its transfer between separate membrane compartments. Class II PITPs also bind one molecule of PI but can also bind PA or PC as its counter ligand. A distinguishing feature of Class II PITPs is their robust PA transfer activity exhibited by the isolated PITP domain that is not shared with Class I PITPs (Cockcroft et al., 2016).

The Five Mammalian PITPs. A: Class I PITPα/β of small molecular weight proteins with a single PI-transfer domain (PITD). PITPNC1 and its two alternative spliced isoforms belong to class II. Domain architecture of the three Nir proteins: Nir1, Nir2, Nir3, is shown in the lower panel. Only Nir2 and Nir3 have an N-terminal PITD. All the Nir proteins have a FFAT motif, DDHD domain and a C-terminal LNS2 (Lipin/Nde1/Smp2) domain. Glycine rich (Gly-Rich) region is found in the middle of Nir3 protein. B: Crystal structure of PITPα. Overall fold of PITPα-PI. Functional regions are lipid-binding core (blue), regulatory loop (green), C-terminal region (red), and lipid exchange loop (orange). The 8 stranded β-sheets labelled 1-8 and helices A and F form the lipid binding core. The PKC phosphorylation site, Ser166 is part of this regulatory loop. The C-terminal region contains the helix G followed by an additional 11 amino acid which limit access to the lipid binding pocket (Tilley et al., 2004).
The Classical PITPs: α and β
PITPα and PITPβ are small soluble proteins of just over 270 amino acids residues that share 77% identity and 94% similarity. Both PITPs are widely expressed in mammalian cells and are abundant proteins. Structures of PITPα and PITPβ bound with PC and PITPα loaded with PI are available; all the lipid-loaded structures are near identical. Both PITPα and PITPβ comprise of a single PITP domain (Pfam: IP trans PF02121) and share a structural fold that can be divided into four sub regions (Yoder et al., 2001; Schouten et al., 2002; Tilley et al., 2004; Vordtriede et al., 2005). One sub region is responsible for lipid binding in its hydrophobic cavity thus allowing the transfer of PI and PC through the aqueous milieu to different membrane compartments (colored blue in Figure 1B). This transfer of lipids by PITPs do not require input of energy and is independent of ATP. The hydrophobic cavity present at the N-terminal region comprises eight strands of anti-parallel beta sheets (labelled 1-8 in Figure 1B) and two alpha helices, (labelled A and F in Figure 1B); it forms a concave structure where a single molecule of PI or PC can be embedded. The hydrophobic cavity can accommodate the two acyl chains in separate channels, the phosphate moiety and the headgroup, inositol or choline. An intervening region (colored green in Figure 1B) is the regulatory region that may participate in contacting other proteins and houses the serine residue 166 that can be phosphorylated by protein kinase C (in PITPα, see later). The third sub domain (colored gold in Figure 1B) is the lipid exchange loop. The PITP domain is compact and is referred to as the ‘closed’ form. A single structure of the lipid-free form (apo-structure) is also available and here the hydrophobic cavity is accessible as the lipid exchange loop has shifted from the main structure. This structure is referred to as the ‘open’ form. In addition, the C-terminal G-helix that terminates with 15 amino acids (colored red in Figure 1B), has also swung away allowing access to the hydrophobic cavity. The C-terminal 15 amino acids forms the lid and is known to exhibit the highest sequence variation in the different PITPs.
The binding of the lipid headgroup, inositol or choline, is located in a similar position in the hydrophobic cavity. However, the inositol ring makes a specific set of hydrogen bonds that are not shared with the choline headgroup. In contrast, residues that contact the phosphate moiety are identical for both PI and PC. Residues Q22, T97, T141 and K195 (rat numbering used in PITPα throughout) are responsible for making hydrogen bonds to the phosphate and are conserved in the majority of the PITP sequences present in the databases. Four residues that contact the inositol ring (T59, E86, K61, and N90) are also similarly conserved and mutation of any-one of the single residues results in loss of PI binding and transfer in either Class I or Class II PITPs (Tilley et al., 2004; Carvou et al., 2010; Yadav et al., 2015). Residues T59 and E86 also interact with PC, but differently to inositol. Thus, the inositol headgroup makes more interactions compared to the choline headgroup explaining why PITPα demonstrates a 16-fold higher affinity for PI in contrast to PC.
PITPβ Cycles Between the Golgi and ER
PITPα is mainly cytosolic whilst PITPβ localizes to the Golgi. Early studies to generate Pitpnb (PITPβ encoding gene) null mice or null embryonic stem cells resulted in failure, and it was concluded that PITPβ was essential (Alb et al., 2002). A recent attempt was successful in generating Pitpnb-null mice; mice appear normal and are fertile (Xie et al., 2018). PITPβ is expressed in all cell types and is normally present at much lower amounts compared to PITPα. However, liver and lung are an exception where PITPβ dominates (Cosker et al., 2008). PITPβ is present as two splice variants; the difference lies in the C-terminal 15 amino acids and this is due to the use of alternative exons. Thus, the C-terminal 15 amino acids of the splice variant 1 (PITPβ-sp1) are replaced by 16 amino acids in splice variant 2 (PITPβ-sp2). Most cells express both splice variants in equal amounts with the exception of HL60 cells which only expresses PITPβ-sp1. Both splice variants localize to the Golgi (Morgan et al., 2006). PITPβ-sp1 has a serine residue in the C-terminus (S262) that is constitutively phosphorylated by protein kinase C (van Tiel et al., 2002; Morgan et al., 2006). Early studies had suggested that localization of PITPβ to the Golgi was determined by this phosphorylation (van Tiel et al., 2004). However subsequent studies have shown that phosphorylation does not impact on localization; whether phosphorylation has a regulatory role is not clear however (Morgan et al., 2006). Golgi localization of PITPβ is determined by the C-terminus and two tryptophan residues; fusing the C-terminal 70 amino acids which encompasses these residues to GFP is sufficient to target PITPβ to the Golgi. It is notable that Golgi localization is independent of the lipid bound to the PITPβ molecule (Phillips et al., 2006).
In addition to the Golgi localization, PITPβ also localizes to the ER (Shadan et al., 2008). This was demonstrated by capturing the PITPβ in the open configuration. The ‘open’ form of PITP is transiently present at the membrane during lipid exchange and can be maintained in the open form by attacking a cysteine residue present in the lipid binding pocket with N-methylmaleimide (NEM) which gets covalently attached (Shadan et al., 2008). This modification results in a PITP that can no longer regain its ‘closed’ confirmation and the PITPβ molecule remains attached to the membrane. Thus, PITPβ is likely cycling between the ER and the Golgi. The ‘closed’ form of PITPβ is a compact structure and is soluble, whilst the ‘open’ form is membrane-attached due to the exposed hydrophobic residues. In NEM-treated cells, the ‘open’ form does ultimately detach from the membrane and forms an end to end dimer due to the exposed hydrophobic residues. It is found that within two minutes of NEM treatment, all the PITPβ molecules are present as a dimer indicating that in cells, PITPβ is constantly cycling between the soluble ‘closed’ lipid bound conformation and the membrane-associated ‘open’ conformation.
PITPβ Function at the ER-Golgi Interface
Early studies identified a role for PITPs in phospholipase C (PLC) signalling, exocytosis and vesicle formation. Using permeabilized cells to reconstitute these events, it was observed that both PITPα and PITPβ were equally functional (Cockcroft 2012). Based on these early studies, it was suggested that PITP provided the substrate, PI, for the local synthesis of phosphorylated forms of PI, including PI4P, PI3P, PI(4,5)P2 and PI(3,4,5)P3. PITPs by virtue of their lipid transfer activity could transfer PI from the ER where it is synthesised to the organelles where the phosphorylated forms were required. Furthermore, it was speculated that PITP could present PI to the lipid kinases within a signalling complex. Based on the PITP structure, it is clear that PI cannot be phosphorylated when bound to the PITP domain; the inositol ring is not accessible to the lipid kinases. Thus, the current suggestion is that classical PITPs transfer PI from regions of synthesis i.e. the ER to target membranes where PI is phosphorylated by resident lipid kinases. Thus, at the ER-Golgi MCS, the conversion of PI to PI4P by the resident PI 4-kinase IIIβ would provide for unidirectional flow of PI from the ER to the Golgi.
The Golgi localization of PITPβ would suggest a role in maintaining PI4P levels at this organelle. Several phenotypes are observed when PITPβ is depleted by RNAi from HeLa cells (Carvou et al., 2010). In the knockdown cells, the Golgi apparatus forms a compacted structure and the nucleus is deformed. Coat protein complex I (COP-I)-coated vesicles mediates retrograde traffic from the Golgi to the ER and traffic of ERGIC-53 between the ERGIC (ER-Golgi intermediate compartment) and the ER. Both traffic is disrupted such that the KDEL receptor, required for retrieval of ER-resident proteins, is arrested at the Golgi and ERGIC-53 is arrested at the ERGIC. Rescue of the retrograde trafficking defect is facilitated by wild type PITPβ only; mutants deficient in PI or PC binding or lacking the membrane-interacting residues WW203/204 are inactive in rescue. These observations support the idea that both PI and PC binding contribute to the trafficking function at the ER-Golgi interface, very likely by maintaining a specific pool of PI4P, required for the assembly of COPI-coated vesicles at the Golgi apparatus.
Studies in cell-lines clearly indicate that PITPβ maintains PI4P levels at the Golgi, and a recent study has identified that PITPβ supports the development of the embryonic mammalian neocortex by maintaining PI4P levels at the Golgi. Here, PITPβ and PITPα act in a redundant fashion. Neural stem cells are bipolar cells that extend the width of the developing neocortex with an apical process contacting the ventricular surface. PITPβ or PITPα maintain a pool of PI4P for recruitment of GOLPH3 and CERT to Golgi membranes. GOLPH3 promotes MYO18A- and F-actin-directed loading of the Golgi network to apical processes of neural stem cells (Xie et al., 2018). Interestingly, PITPNC1 has also been suggested to maintain PI4P levels at the Golgi for recruitment of GOLPH3 and in this case, for vesicular release from tumor cells as discussed below (Halberg et al., 2016; Kuna and Field, 2019).
The family Picornaviridae is a group of nonenveloped positive-strand RNA viruses which includes rhinoviruses and Aichi virus. PITPβ has been identified as a host factor required for rhinovirus and Aichi virus replication. For replication, the virus builds a membrane-associated replication complex which is Golgi-derived that is tightly associated with the ER. Transport of cholesterol to these membranes requires OSBP, PITPβ, PI4KIIIβ and Sac1 all of which are also localized at this MCS, and are required for both PI4P homeostasis and viral replication (Roulin et al., 2014; Ishikawa-Sasaki et al., 2018). Whilst in mammalian cells, a cytosolic protein is used to transport PI to establish a PI4P gradient for cholesterol transport to the Golgi, in Toxoplasma Gondii, a multi-domain protein incorporating a PITP domain with an oxysterol binding domain is amalgamated into a single protein (Cockcroft and Raghu, 2018).
PITPNC1 Is Golgi-Localized and Is Required for Vesicle Formation
PITPNC1 is a Class II PITP and was the last of the PITPs to be studied. Like PITPα and PITPβ, PITPNC1 comprises of a single PITP domain with the addition of a C-terminal extension of 80 amino acids that is unstructured. Two serine residues present in the C-terminal extension, S274 and S299, are phosphorylated to form binding sites for 14-3-3 proteins. Binding to 14-3-3 increases the lifetime of the proteins (Cockcroft and Garner, 2012). Even so PITPNC1 is short lived compared to Class I PITPs. A splice variant with a shorter C-terminus that lacks the 14-3-3 binding sites has also been identified. PITPNC1 also interacts with the adapter protein, ATRAP (angiotensin II type I receptor-associated protein) and this binding site is located in the regulatory domain distal to the C-terminus, and thus, both splice variants are able to bind ATRAP. PITPNC1 also binds to Rab1B but the binding site has not been identified (Halberg et al., 2016). Whether both splice variants bind to Rab1 is not known.
Like PITPβ, PITPNC1 localizes at the TGN (trans-Golgi network). Rab1B and PI4P were proposed to recruit PITPNC1 to the TGN (Halberg et al., 2016). PITPNC1 together with PI4KIIIβ is highly expressed in tumor cells and promotes secretion of pro-tumorigenic effector proteins including IGFBP2 to maintain cancer cell survival and influence pro-metastatic process in the tumor micro-environment (Png et al., 2011; Halberg et al., 2016; Peretti et al., 2019; Tan et al., 2020). The aberrant expression of PITPNC1 is regulated by the loss of microRNA-126 in cancer cells (Png et al., 2011). PI4P synthesised by PI4KIIIβ at the Golgi appears to facilitate vesicular release by recruiting GOLPH3 (Kuna and Field, 2019). In cancer cells, PITPNC1, PI4KIIIβ and GOLPH3 are upregulated and likely work in the same pathway.
The physiological function of PITPNC1 has been explored in a zebrafish model (Ashlin et al., 2018). In zebrafish, instead of two splice variants, two separate genes code for PITPNC1. One protein is exclusively expressed in the brain and is the homologue of the long form of PITPNC1. Zebrafish PITPNC1 also binds 14-3-3 and is involved in secretion of IGFBP2, a negative regulator of IGF signalling. Thus, deletion of PITPNC1 gene results in enhanced IGF signalling in neurons resulting in enhanced neuronal activity (Ashlin et al., 2018).
Large Multi-Domain PITPs: Nir2 and Nir3
The two mammalian large multi-domain PITPs, Nir2 and Nir3, belong to the Nir family of Nir1, Nir2 and Nir3 proteins (Lev, 2004). The Nir proteins were initially discovered as interacting proteins with the N-terminal domain of the non-receptor tyrosine kinase PYK2 using a yeast two-hybrid screen (Lev et al., 1999). Nir2, which is also known as Dres9, PITPNM1 or M-RdgB1, was independently isolated as a mammalian homolog of the Drosophila rdgB (retinal degeneration B) (Guo and Yu, 1997; Rubboli et al., 1997; Aikawa et al., 1999), and a few years later Nir3 (M-RdgB2) was isolated (Lu et al., 2001). Nir2 and Nir3 share high sequence similarity, but exhibit different tissue expression pattern and distinct cellular functions. Nir3 is more abundant in the nervous system and is highly expressed in the retina, specifically in GABAergic amacrine cells. Nir3 knockout mice developed normally (Lu et al., 2001) but failed to transduce light input from rods to bipolar retinal cells, thus displaying defect response to dim light (Walker et al., 2015). Knockout of Nir2 mice was initially thought to be embryonic lethal (Lu et al., 2001), but subsequent studies described a viable and fertile phenotype (Carlisle et al., 2013). Interestingly, Nir2, but not Nir3 could partially rescue the phenotype of rdgB mutant flies, which is characterized by light induced retinal generation, abnormal electroretinogram and a defect in light-activated PI(4,5)P2 turnover (Chang et al., 1997; Lu et al., 2001). The Nir2 and Nir3 proteins as well as the Drosophila RdgB share similar domain organization (Figure 1) including an N-terminal PI-transfer domain, followed by a FFAT (double phenylalanines (FF) in an Acidic Tract) motif, a DDHD domain and a C-terminal LNS2 (Lipin/Nde1/Smp2) domain (Selitrennik and Lev, 2016). Nir1 has similar domain composition, but lacks a PI-transfer domain. Nir2 and Nir3 are localized in the cytosol and associate with different cellular membranes (Lu et al., 1999; Litvak et al., 2002), but are not integral membrane proteins as was previously proposed for PITPNM1 and Drosophila RdgB (Vihtelic et al., 1993; Aikawa et al., 1999).
Nir2 Functions at the Golgi
Extensive studies over the past 20 years on Nir2 and Nir3 highlighted their roles as important regulators of cell signaling and membrane trafficking. While the different functions of Nir2 were previously described in several review papers (Lev, 2004, 2010; Kim et al., 2016; Selitrennik and Lev, 2016), we focus here on several critical findings related to its function at the Golgi apparatus and ER-Golgi MCSs. Early studies showed that depletion of Nir2 by siRNA (small RNA interference) affects the structural integrity of the Golgi apparatus and substantially inhibited protein transport from the TGN to the plasma membrane due to reduced DAG levels, and consequently impaired fission of transport carriers at the TGN (Litvak et al., 2005). Nir2 was proposed to negatively regulate PC production at the Golgi complex via the CDP-choline pathway. In the absence of Nir2, the consumption of DAG was increased concomitant with increased PC biosynthesis, while the level of DAG in the Golgi was decreased (Litvak et al., 2005). These strong effects on Golgi structure and secretory functions require robust and transient knockdown of Nir2 (using siRNA and not mild and sustained knockdown by shRNA), and are consistent with previous studies on Sec14p, a PITP in yeast (Skinner et al., 1995). It was previously proposed that Sec14p in its PC-bound inhibits the activity of CCT (CTP:phosphocholine cytidylyltransferase), the rate limiting enzyme of the CDP-choline pathway and consequently prevents DAG consumption in the yeast Golgi complex (Skinner et al., 1995). This inhibitory effect on PC biosynthesis at the Golgi is required for maintenance of a critical pool of DAG to preserve Golgi structure and secretory function. It is currently unclear how Nir2 negatively regulates the CDP-choline pathway, but recent studies suggest that Nir2 contains a DAG-binding like (DGBL) segment that exhibits weak similarity to the DAG binding part of C1 domains (Kim et al., 2015). Thus, it could be that this domain binds DAG and prevents DAG consumption for PC biosynthesis. Although this is a feasible possibility as the PI-transfer domain of Nir2 could partially rescue the influence of Nir2 depletion on the Golgi, it needs further investigation.
Nir2 Function at ER-Golgi MCSs
Consistent with the role of Nir2 in the Golgi, pulldown experiments and mass spectrometry analysis revealed that Nir2 interacts with the integral ER protein VAP-B (Amarilio et al., 2005). Further studies confirmed the interactions of VAP-B/A with the three Nir proteins via their conserved FFAT motif (EFFDAxE) (Loewen et al., 2003; Amarilio et al., 2005; Kim et al., 2010). As VAP-A and -B bind different LTPs at MCSs, their interactions with Nir1, which lacks the PI-transfer domain, possibly compete with other FFAT-containing proteins to impair efficient lipid transport. This could be an important mechanism to negatively regulate lipid transport at MCSs. Currently, however, the physiological functions of Nir1 remain largely unknown. It is worth mentioning that co-expression of VAP-B and Nir2 in mammalian cells induced the formation of stacked ER membrane arrays, whereas co-expression of VAP-B with Nir3 caused remodeling of the ER and bundling of thick microtubules along the altered ER membranes (Amarilio et al., 2005). Further studies suggest that the C-terminal tail of Nir3 can interact with microtubules (unpublished results). These observations imply that Nir3-VAP-A/B interaction may stabilize MCSs by linking them to cytoskeletal elements (Lev et al., 2008). These findings also highlight the differences between Nir2 and Nir3.
Subsequent studies using siRNA to deplete VAP-A and -B showed substantial effects on both the morphology of the Golgi and protein transport from the Golgi-to-PM, Golgi-to-endosome/lysosome and Golgi-to-ER (Peretti et al., 2008). In addition, VAPs depletion reduced the levels of PI4P, DAG, and sphingomyelin (SM) in the Golgi membranes. These pleiotropic effects of VAPs on the lipid composition of the Golgi and Golgi-mediated transport pathways were more profound in the presence of 25-hydroxycholesterol (25OH), the high-affinity ligand of oxysterol binding protein (OSBP) (Antonny et al., 2018). It was proposed that the effects of VAPs on the Golgi are mediated by coordinated function of three LTPs; Nir2, OSBP and CERT (ceramide-transfer protein), all contain FFAT motif and interact with VAPs. OSBP and CERT associate with the Golgi membranes through their PH domains that bind PI4P. CERT transfers ceramide from the ER to the Golgi, most efficiently at ER-Golgi MCSs (Hanada et al., 2007), and ceramide is converted into SM by SM synthase (SMS) at the TGN. SMS transfers the phosphocholine group of PC to ceramide to produce SM and DAG (Tafesse et al., 2006). It was shown that OSBP increases SM production in the presence of 25OH (Lagace et al., 1999), and is essential for sterol-dependent activation of CERT (Perry and Ridgway, 2006). Based on these observations and further functional studies using rescue experiments (Peretti et al., 2008), it was proposed that Nir2, OSBP and CERT coordinately function at the ER-Golgi MCSs and regulate the lipid composition of the Golgi through their lipid-transfer activity. Nir2, through its PI-transfer domain, transfers PI from the ER to the Golgi, which is subsequently phosphorylated by Golgi-localized PI4Ks (PI4KIIα and PI4KIIIβ) to produce PI4P (Waugh, 2019). PI4P recruits OSBP and CERT via their PH domains, OSBP possibly enhances the activity of CERT that transfers ceramide from the ER-to-Golgi, and ceramide is converted into DAG and SM at the TGN (Figure 2) (Peretti et al., 2008). It worth mentioning that not only OSBP, but additional OSBP-related proteins (ORPs) bind PI4P at the TGN and function at the ER-Golgi MCSs to deliver cholesterol (OSBP1, ORP9, ORP11) or PS (ORP10) from the ER (Venditti et al., 2020).

Lipid Transfer by PITPs at the ER-Golgi Interface. PITPα/β, PITPNC1 and Nir2 transfer lipids at the ER-Golgi interface; PI from the ER-to-Golgi and PC and/or PA from Golgi-to-ER. The metabolic pathways that regulate the production or consumption of the three transferred lipids; PI, PC and PA at the Golgi and ER are shown. The central role of DAG in these metabolic pathways are marked. DAG plays central role in the described metabolic pathways and also in vesicular fission and local signaling events. At the ER, DAG can be converted to PA by DGK (DAG kinase). PA is also produced at the ER from glycerol-3-phosphate (Lev, 2012; Blunsom and Cockcroft, 2020b) or can be transferred to the ER by certain PITPs. PA and CTP are converted into CDP-DAG (Cytidine diphosphate diacylglycerol) by ER localized CDS1/2 (CDP-DAG synthase). CDP-DAG is synthesized into PI by PIS (PI synthase). Other pathways in the Golgi includes the CDP-choline pathway that converts CDP-choline (CDP-Cho) and DAG to PC by CPT (diacylglycerol choline phosphotransferase), and production of DAG and sphingomyelin (SM) from PC and ceramide by SM synthase (SMS).
The subsequent discovery that OSBP is not only PI4P-binding protein but also a PI4P-transfer protein, which transfers PI4P from the TGN back to the ER and sterol from the ER to the TGN (Mesmin et al., 2013; Goto et al., 2016) further highlights the cooperation between Nir2, OSBP and CERT at ER-Golgi MCSs, as PI4P is hydrolyzed by the ER-localized phosphoinositide phosphatase Sac1, to produce PI, and PI can be delivered by Nir2 to replenish the depleted PI4P pool at the TGN. Likewise, the transport of ceramide by CERT from the ER-to-Golgi and the subsequent SM synthesis is coupled to production of DAG (Figures 2 and 3B), which may also be transported by CERT (Hanada et al., 2003; Kudo et al., 2008) from Golgi-to-ER. This possibility, however, needs further exploration as the efficiency of DAG transport by CERT is very low.

DAG-Dependent Phosphorylation and Lipid Transport. DAG not only affects critical lipid metabolic pathways at the ER-Golgi interface (Figure 2) but also functions as a signaling molecule that regulates the activity of multiple proteins including lipid-transfer proteins (LTPs). A: All the five PITPs have conserved phosphorylation sites of PKC (S166 major, T59 minor in PITPα), and thus, can undergo DAG-dependent phosphorylation that inhibits their PI-transfer activity. This phosphorylation can provide a mechanism to terminate DAG-associated signaling cascades, and may occur at the different cellular membranes including the PM and the Golgi. B: DAG activates PKCη which phosphorylates and activates PKD, and PKD is recruited to the Golgi by DAG. PKD modulates the activity of multiple proteins including different LTPs at the ER-Golgi interface as described in the text.
DAG at the Golgi can be converted into PA by DGKα (Xie et al., 2015), while PLD (Ktistakis et al., 1996; Chen et al., 1997) can produce PA from PC. Indeed, recent studies suggest that Nir2 as well as the Drosophila rdgB possess PI/PA-exchange activity rather than PI/PC exchange activity as was previously proposed (Cockcroft et al., 2016). Although the crystal structure of their PI-transfer domains has not been resolved yet, it could be that Nir2 transfers PI from ER-to-Golgi and PA from Golgi-to-ER in response to elevated PA levels, as was proposed for its exchange activity at the ER-PM MCs (Kim et al., 2015). At the PM PA can be produced by PLD or by PLC in response to different external stimuli. Several studies showed that stimulation of growth factor receptors (EGF, Heregulin) (Kim et al., 2013; Keinan et al., 2014) and GPCRs (histamine, angiotensin II, muscarinic cholinergic agonist) (Chang et al., 2013; Chang and Liou, 2015; Kim et al., 2015; Kirmiz et al., 2019) induce translocation of Nir2 to the PM through its C-terminal domain, and that Nir2 as well as the Drosophila rdgB function at ER-PM MCSs to replenish the PLC-hydrolyzed PI(4,5)P2 pool in response to external stimuli (Kim et al., 2013; Yadav et al., 2015). Further studies showed that the LNS2 domain of Nir2 binds PA while the DGBL binds DAG (Kim et al., 2013; Kim et al., 2015; Kim et al., 2016), and that the LNS2 and/or DGBL mediate the binding of Nir2 to the PM. However, the C-terminal region is not required for Golgi targeting of Nir2 as a truncated mutant consisting only the N-terminal PI-transfer domain was associated with the Golgi (Kim et al., 2013). Interestingly, recent studies suggest that similar to the ER-Golgi MCS, Nir2, VAPs and OSBP, also cooperate during hepatitis C virus (HCV) replication. HCV replication occurs at ER-derived cytoplasmic membranes known as HCV replication organelles (ROs). VAPs are essential for efficient HCV replication, and depletion of VAPs reduced the PI4P levels at the viral RO, and consequently the recruitment of OSBP. The integrity of RO is dependent on PI4P and OSBP, therefore, PI4P levels must be maintained despite the PI4P/Sterol exchange activity of OSBP. It was shown that Nir2, through its PI-transfer activity and interaction with VAPs, is involved in the replenishment of PI4P at the RO, thereby promoting continuous viral replication during viral infection (Wang and Tai, 2019).
DAG, a Central Hub of PITPs Function
Given the remarkable effects of yeast and mammalian PITPs on DAG levels at the Golgi (Lev, 2006; Bankaitis et al., 2012), DAG can be considered as a central hub of PITPs function, as it lies at the intersection of metabolic pathways that regulate the production of the three PITPs exchanged lipids, PI, PC and PA (Figure 2). DAG can be phosphorylated by DAGKα to PA at the Golgi. Similarly, DAG can be phosphorylated by ER-specific DAGKs (δ, η and κ (Xie et al., 2015)) to produce PA in the ER, which can be further converted into CDP-DAG by CDS (CDP-DAG synthase) and subsequently to PI by PIS (PI synthase) (Blunsom and Cockcroft, 2020a), to be transport by Nir2, PITPNC1 and/or PITPβ back to the Golgi. DAG can be consumed by the CDP-choline pathway for PC synthesis at the Golgi, and together these metabolic pathways highlight the central role of DAG in regulating the production of PI, PC, PA at the ER-Golgi interface. Importantly, DAG does not only regulate the levels of PI, PC and PA production, but also, it modulates the lipid-exchange activity through DAG-dependent phosphorylation of specific residues within the PI-transfer domain (Figure 3A) as further discussed. DAG-dependent phosphorylation of other LTPs at the ER-Golgi MCSs contributes, through feedback mechanisms and crosstalk between metabolic pathways, to modulate lipid-transfer activity. This central role of DAG may coordinate the lipid exchange of different Golgi localized PITPs including Nir2, PITPNC1 and PITPβ to ensure vectorial transport of PI, which is exclusively synthesized at the ER, either to the Golgi or the PM and also to coordinate non-vesicular and vesicular transport to and from the Golgi.
PITPs Associated Trafficking and Lipid Metabolism at ER-Golgi Interface
As described, PITPs are involved in a network of lipid metabolic pathways at the ER-Golgi MCSs that modulate the levels of key Golgi lipids, including PI4P, DAG, PA, SM and cholesterol (Figure 2) through interplay with other Golgi-ER proteins including CERT, ORPs, VAPs and Sac1. The roles of these lipids on trafficking events from and to the Golgi complex have been described in numerous studies and review papers (Lev, 2010; Bankaitis et al., 2012; Venditti et al., 2020). In brief, PI4P has pleotropic effects on Golgi-mediated transport through recruitment of different regulatory proteins. It recruits Arfaptin1 to the TGN to prevent premature fission of secretory vesicles (Gehart et al., 2012; Cruz-Garcia et al., 2013), as well as FAPP2, a glucosylceramide (GlcCer) transfer protein, which together with Arf1 regulate vesicular transport from the TGN to plasma membrane (Godi et al., 2004). It also regulates the levels of SM and sterols at the TGN, by recruiting CERT and ORPs, to establish a segregated SM-cholesterol-rich domain at the TGN for anterograde transport of specific vesicles (Deng et al., 2016). In addition, PI4P is required for COPI vesicles formation and thus Golgi-to-ER retrograde transport as well as TGN-to-endosome trafficking (Waugh, 2019).
DAG and PA also regulate different Golgi-mediated transport routes. The conical shape of DAG and PA can induce negative curvature, membrane bending and the formation of highly curved intermediates to facilitate membrane invagination and vesicular fission (Bard and Malhotra, 2006). In general, PA has a short lifetime in intracellular membranes, and is rapidly converted into DAG by PAP (Lev, 2006). PA is involved in fission of COPI vesicles as well as transport carriers from the TGN. DAG also regulates anterograde transport from the TGN and is also required for COPI vesicle formation, and thus, Golgi-to-ER retrograde transport (Roth, 2008; Bankaitis et al., 2012). In addition, DAG recruits PKD to the TGN and activates PKCη, which phosphorylates and activates PKD. Both PKCη and PKD remarkably affect vesicular transport from the TGN (Baron and Malhotra, 2002; Bard and Malhotra, 2006). DAG is produced at the Golgi through PI4P hydrolysis by certain PLCs (such as PLCε; (Zhang et al., 2013)), through dephosphorylation of PA by phosphatidic acid phosphatase (PAP), in particular PAP2b (also known as LPP3),which localizes at ER export sites and Golgi complex (Gutierrez-Martinez et al., 2013), and from PC and ceramide by SMS (Figure 2).
The major lipid metabolic pathways at the ER-Golgi MCSs that have been discussed are interconnected and regulated by different enzymes including kinases, phosphatases and phospholipases as illustrated in Figures 2 and 3B and further described by several comprehensive reviews (Lev, 2012; Holthuis and Menon, 2014).
PITPs and Phospho-Signaling Landscape of ER-Golgi MCSs
DAG is not only a central hub for PITPs-associated lipid metabolic functions, but it is also a key signaling molecule that activate Golgi-localized protein kinases, in particular PKCη and PKD (PKCµ) among other PKCs, such as PKCδ and PKCθ (Mayinger, 2011). Hence, it is conceivable that Golgi DAG levels are coupled to dynamic changes in phosphoproteome at the ER-Golgi interface. CERT, for example, is phosphorylated by PKD at S132 within its serine-repeat motif (SRM) adjacent to the PH domain, which triggers sequential multisite phosphorylation by Casein Kinase 1γ2 (CK1γ2), and consequently suppression of CERT activity. Dephosphorylation of SRM sites by PP2Cε (protein phosphatase 2 Cε) concomitant with phosphorylation of S315, which flanks the FFAT motif, induce full activation of CERT in response to cellular requirements of SM (Kumagai and Hanada, 2019).
Interestingly, PP2Cε is an integral ER protein which interacts with VAP and dephosphorylates CERT in VAP-dependent manner, possibly to ensure its dual binding to ER and Golgi at ER-Golgi MCSs and efficient ceramide transport (Yamaji et al., 2008). OSBP is also phosphorylated by PKD at S240, and this phosphorylation reduced its TGN targeting in response to 25OH and cholesterol depletion, and impaired Golgi localization of CERT as well as proteins transport to the PM (Nhek et al., 2010), implying a feedback regulation of DAG via PKD-mediated phosphorylation of both CERT and OSBP (Figure 3B). Importantly, similar to CERT, also OSBP is phosphorylated on multiple sites; three (S381, S384, and S387) are located near the PH domain and are sequentially phosphorylated by CK1, while the other sites (S192, S195, S200) flank the FFAT motif. These phosphorylation sites can influence sterol binding and transfer as well as VAPs binding (Goto et al., 2012). Recent studies suggest that OSBP interacts with PKD1 and inhibits its autophosphorylation at S916 and consequently its activity, further demonstrating a negative feedback regulation (Goto et al., 2018).
On the other hand, PKD phosphorylates PI4KIIIβ (S294) and stimulates its lipid kinase activity to enhance protein transport to the plasma membrane (Hausser et al., 2005). Elevated PI4P levels, which triggers the targeting of CERT and OSBP to the Golgi, concurrently control SM and cholesterol levels through their PKD-dependent phosphorylation of OSBP and CERT. PI4P in the TGN also triggers the recruitment of PITPNC1 (Halberg et al., 2016), and its phosphorylation at the C-terminus (S274 and S299) induces 14-3-3 binding and consequently protein stabilization (Garner et al., 2011). 14-3-3 also binds and stabilizes phospho-PI4KIIIβ. Importantly, in addition to the DAG-dependent phosphorylation of PITPNC1 by PKC at its C-terminal tail (S274 and S299), PITPNC1 and other mammalian PITPs contain several conserved PKC phosphorylation sites (S166 major, T59 minor) within their PITP domains (Morgan et al., 2004) that markedly affect their lipid-transfer activity (Figure 3A), further demonstrating the link between DAG-dependent phosphorylation and PITPs activity, and highlight a possible DAG-dependent feedback mechanism that regulates PI/PC- or PI/PA-transfer activity at the ER-Golgi and/or ER-PM MCSs.
Currently, it is unclear whether any of these phosphorylation events occur by Golgi localized PKCs, but previous studies suggest that stimulation of different receptor tyrosine kinases (RTKs) can induce phosphorylation of PITPs by PKCs, and that RTKs can also activate Golgi-associated Ras signaling through Src, PLCγ and RasGRP1 (Bivona et al., 2003). Accordingly, activation of PLCγ by Src triggers an increase in Golgi DAG and consequently the translocation of the GEF RasGRP1, which binds DAG via its C1 domain, to activate Ras and downstream MAPK at the Golgi complex. This mechanism can coordinate signaling between ER-PM and ER-Golgi MCSs. Intriguingly, Src also phosphorylates and activates DGKα, induces its translocation to the TGN and consequently the phosphorylation of DAG to PA (Xie et al., 2015), which can be transported by Nir2/3 or PITPNC1 to the ER.
Additional phosphorylation sites that were identified through different proteomic screens might play important roles in regulating signaling, trafficking and lipid metabolism at the ER-Golgi MCSs. The most relevant are multiple Ser/Thr phosphorylation sites in VAP-B (some are conserved in VAP-A; PhosphoSitePlus) at the serine rich segment within the linker region between the MSP domain and the coiled-coil domain (Figure 4). At least one site fits the PKD target motif (L/V/I)X(R/K)XX(S/T) (Franz-Wachtel et al., 2012), while the others might be phosphorylated by CKs or other kinases, and based on their proximity could be involved in a primed phosphorylation event. Currently, the role of these phosphorylations are not known, but may influence VAPs oligomerization and/or interaction with other proteins, which will be important to explore in the future. In this context, it will be very intriguing to apply a proteomic and phospho-proteomic approaches and map the interactome of PITPs and possibly other critical LTPs at the ER-Golgi MCSs (Figure 4).
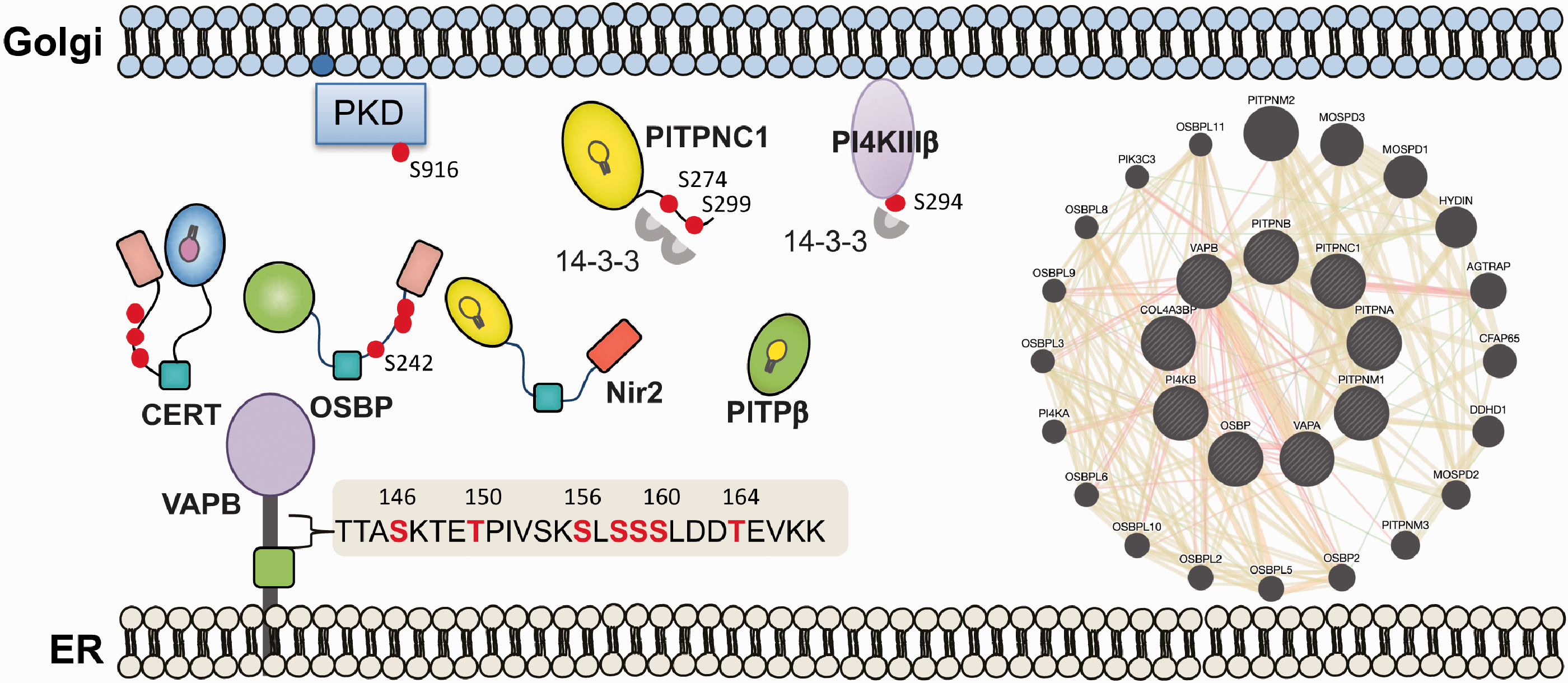
Signaling landscape of the ER-Golgi MCSs. A perspective view on the signaling landscape of the ER-Golgi MCSs. The ER-Golgi MCSs undergoes dynamic metabolic changes that are regulated by multiple lipid biosynthesis enzymes, LTPs, kinases and phosphatases, which all together establish a dynamic, highly regulated signaling landscape that is crucial for maintenance the homeostasis of the Golgi and the ER. Characterizing the dynamic properties of this signaling landscape by analyzing (1) the interactomes of key components as illustrated (performed by GeneMANIA; a schematic example for interactions of VAPS, PITPs, OSBP, CERT), and (2) the phospho-proteome as described for a few proteins in the text, could advance the current understanding of ER-Golgi MCSs function in lipid-homeostasis and lipid-trafficking.
Summary and Perspective
The available evidence supports a fundamental role for PITPs to function at the ER-Golgi interface and regulate a multitude of different lipids mainly through their ability to modulate PI4P and DAG levels. PI4P together with DAG signaling at the Golgi integrates multiple aspects of signaling and lipid metabolism with membrane trafficking between the ER and Golgi. PITPs can dynamically regulate the amount of PI4P produced at the Golgi by facilitating the transfer of PI from its biosynthesis compartment. At the Golgi, PI4P is used to power the movement of ER-synthesized cholesterol and ceramide to the Golgi. At the Golgi, ceramide conversion to sphingomyelin will increase DAG levels. Thus, the functional coordination of both phospholipid and sphingolipid metabolic pathways with phosphoinositide signaling is a center point for membrane traffic from the Golgi.
Nevertheless, many unanswered questions remain. For example, it is unclear how different PITPs, including PITPNC1, PITPβ and Nir2 coordinately function at ER-Golgi MCSs at steady-state and in response to different stimulatory or stress signals. Depletion of two or three of them could provide interesting insights. Characterizing the effect of their depletion/knockout on genome-wide transcriptome as well as their interactome by quantitative proteomic approaches such as BioID could uncover important regulatory mechanisms. Further structure-function analysis on specific domains of Nir2 and Nir3 including the DDHD domain or the hydrophobic patches, which were initially considered as transmembrane, could be very informative, especially if they mediate membrane association or interact with different lipids.
Currently, much of our knowledge is gleaned from studies in cultured cells and for further progress to be made, lipid and protein activity needs to be examined within a physiological context that takes into consideration the expression patterns of specific proteins and how the lipid regulation is coordinated. Taking lessons from the fly phototransduction system, it is clear that RdgBα in phototransduction plays a specific role in phospholipase C-mediated signal transduction where its role in maintaining PI(4,5)P2 is unchallenged (Cockcroft and Raghu, 2016). Studies with the Class I PITPs clearly indicate PITPα and PITPβ have both overlapping roles as well as specific roles in mice. In contrast, in zebrafish, pitpnc1a is exclusively expressed in the brain in a subset of neurons whilst pitpnc1b is expressed in pronephric ducts (Ashlin et al., 2018). The distinct tissue distribution of pitpnc1a and pitpnc1b supports the idea that PITPs have nonoverlapping functions in vivo. Similarly to zebrafish, in C.elegans, the single multi-domain PITP (PITP-1, homologue of Nir2/Nir3) is expressed in a subset of neurons where it participates in sensory transduction and behavioral plasticity (Iwata et al., 2011). Does Nir2 and Nir3, the mammalian homologues, play specific roles in animal organisms? In mice, Nir3 expression is highly restricted in specific set of cells in the retina (Walker et al., 2015). Analysis of model organisms of Nir2 knockouts would help in clarifying the importance of these proteins at the ER-Golgi interface.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Israel Science Foundation (ISF) grant No. 1530/17 and by the Estate of Emile Mimran. It is also supported by grants to Shamshad Cockcroft from BBSRC (FS/15/73/31672) and BHF (BB/J005606/1).
