Abstract
There is an increased emphasis on visualizing neuroimaging results in more intuitive ways. Common statistical tools for dissemination of these results, such as bar charts, lack the spatial dimension that is inherent in neuroimaging data. Here we present two packages for the statistical software R that integrate this spatial component. The ggseg and ggseg3d packages visualize predefined brain segmentations as 2D polygons and 3D meshes, respectively. Both packages are integrated with other well-established R packages, which allows great flexibility. In this Tutorial, we describe the main data and functions in the ggseg and ggseg3d packages for visualization of brain atlases. The highlighted functions are able to display brain-segmentation plots in R. Further, the accompanying ggsegExtra package includes a wider collection of atlases and is intended for community-based efforts to develop additional compatible atlases for ggseg and ggseg3d. Overall, the ggseg packages facilitate parcellation-based visualizations in R, improve and facilitate the dissemination of results, and increase the efficiency of workflows.
Visualization is increasingly important for accurate explanation and interpretation of neuroimaging results, as current research is able to generate a large amount of raw data and outcomes. For MRI, neuroimaging software provides whole-brain information by dividing space into many small units (> 100.000). Nonetheless, these data are often grouped and summarized using a limited number of regions in predefined brain-parcellation atlases. Brain parcellations segment the brain into a finite set of meaningful neurobiological components, each of which reflects one or more brain features based on either structural or connectivity properties (Eickhoff, Yeo, & Genon, 2018). The use of brain atlases in neuroscience is widespread, as they facilitate interpretation and minimize the amount of data (in comparison with whole-brain data), and hence reduce statistical problems with multiple comparisons. This enables replicability and data sharing in the case of otherwise computationally expensive analyses, which are often performed in specialized software environments, such as R (R Core Team, 2019).
MRI data provide good spatial resolution, and thus an optimal representation has to respect spatial relationships across regions. Results from brain-atlas analyses are most meaningfully visualized when projected onto a representation of the brain, so it is desirable for any visual representation to take spatial relations into account. Such projections provide clear points of reference (especially when the reader is unfamiliar with the atlas), improve readability, guide interpretation, and convey the spatial patterns of the data. Adopting the grammar of graphics implemented in ggplot2 (Wickham, 2016), one can plot neuroimaging data directly in R with several tools, such as ggBrain (Fisher, 2019) and ggneuro (Muschelli, 2017; see Muschelli et al., 2019, for curated neuroimaging packages for R). Yet these tools display whole-brain image files and are not well suited for representing brain-atlas data.
In this Tutorial, we introduce two packages for visualizing brain-atlas data in R. The ggseg and ggseg3d packages—plus the complementary ggsegExtra package—include precompiled data sets for different brain atlases that allow for 2D and 3D visualization. The 2D functionality in ggseg is based on polygons and ggplot2-based grammar of graphics (Wickham, 2016), whereas the 3D functionality in ggseg3d is based on tri-surface mesh plots and plotly (Sievert, 2019).
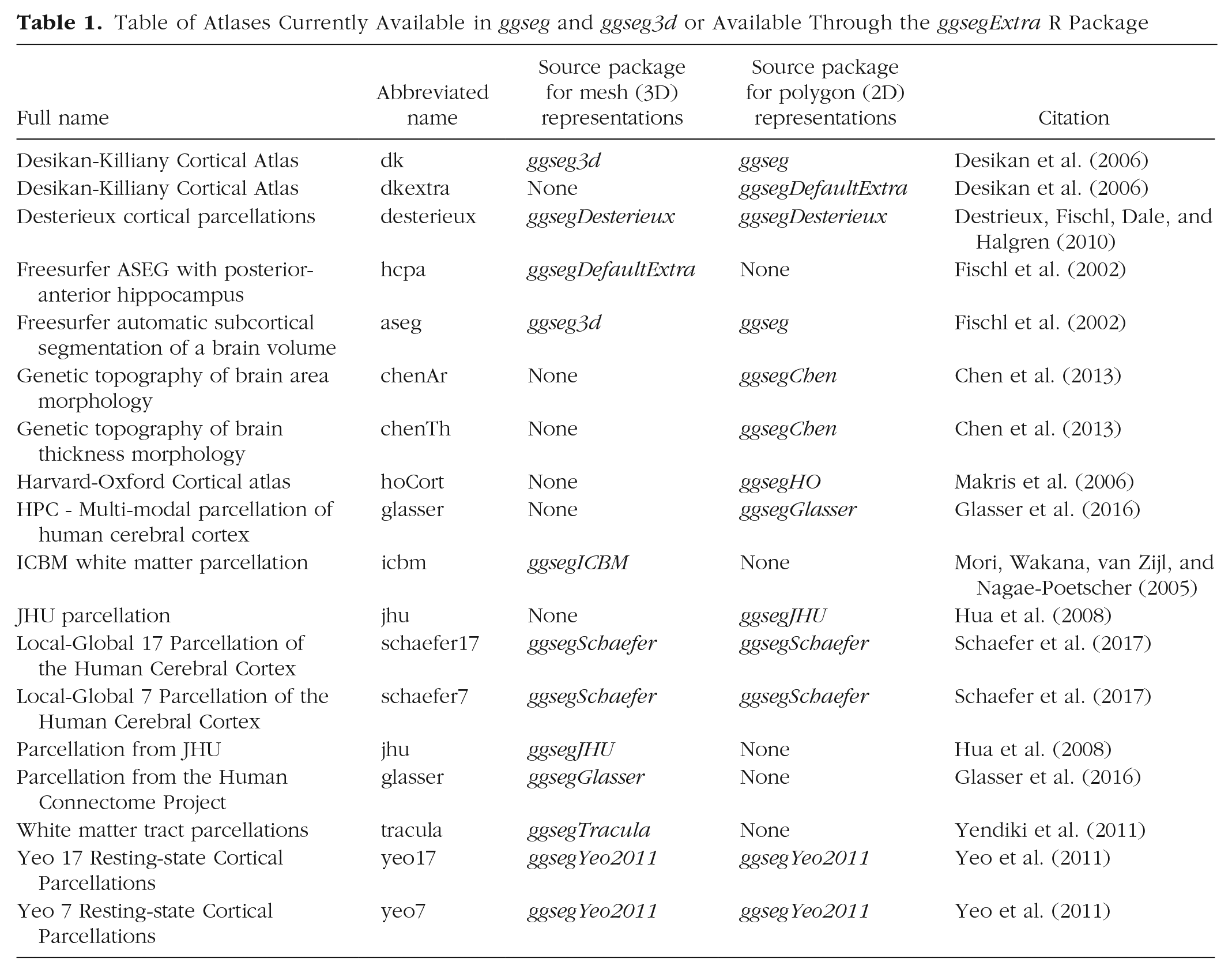
In addition to the compiled data sets, the ggseg and ggseg3d packages provide tailored functions that allow brain-data integration and plotting, as well as other minor features, such as custom color palettes. The data featured in the packages are derived from two well-known parcellations: the Desikan-Killiany cortical atlas (dk; Desikan et al., 2006), which covers the cortical surface of the brain, and the Automatic Segmentation of Subcortical Structures (aseg; Fischl et al., 2002), which covers the subcortical structures. Both atlases are implemented in a variety of neuroimaging software, such as FreeSurfer (Dale, Fischl, & Sereno, 1999; Fischl & Dale, 2000; Fischl, Sereno, & Dale, 1999), and are commonly used to study developmental changes, disease biomarkers, genomic data, and cognition (Amlien, Sneve, Vidal-Piñeiro, Walhovd, & Fjell, 2019; Pizzagalli et al., 2009; Walhovd et al., 2005). The ggsegExtra package contains a list of online repositories of precompiled atlases (currently 11 additional polygon atlases and 10 additional mesh atlases), and it is frequently updated. A summary of all available atlases compatible with the ggseg packages as of June 2020 can be found in Table 1.
Table of Atlases Currently Available in ggseg and ggseg3d or Available Through the ggsegExtra R Package
This Tutorial introduces the ggseg, ggseg3d, and ggsegExtra packages and familiarizes the reader with the main functions and the general use of the packages. We focus on the two main functions:
Disclosures
All materials for this Tutorial, including the R Markdown document used to write it, may be found online at https://github.com/Athanasiamo/ggsegpaper.
Plotting Polygon Data (ggplot2)
The main function for plotting 2D data is
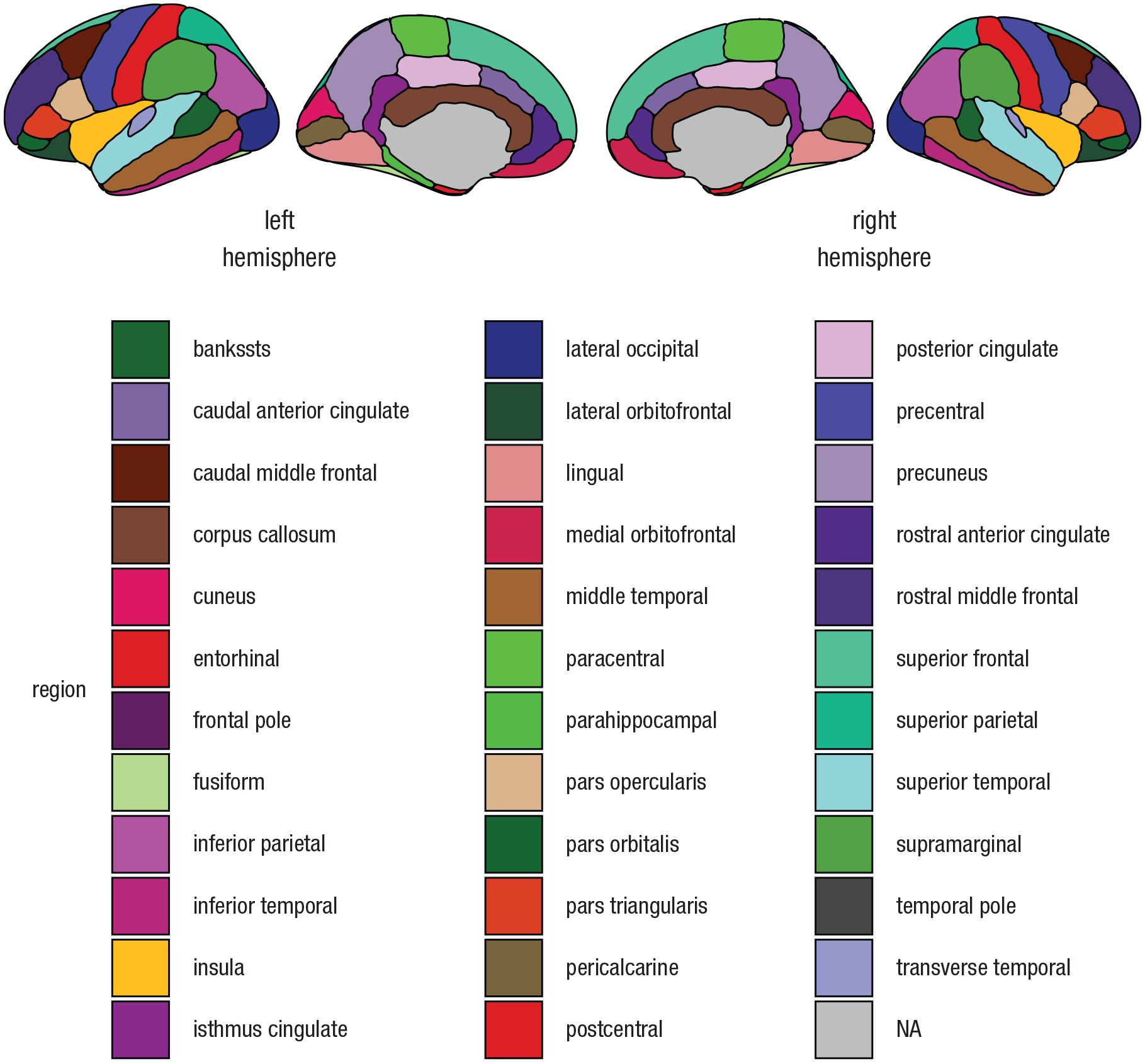
To start the Tutorial, use the following code, which loads the necessary R libraries and plots the dk atlas using ggseg (Fig. 1):


Plot of the Desikan-Killiany atlas (Desikan et al., 2006) in gray polygons. This plot was created by running the
In addition to the standard options for ggplot2 polygon geoms, the function has several options for plotting the main brain representations. These options are atlas-specific. For cortical atlases, such as the dk, one can stack the hemispheres, display only the medial or lateral side, choose either one or both hemispheres, or choose any combination of hemisphere and view (see Fig. 2 for examples). For subcortical atlases, such as the aseg atlas, the options are more limited, but one can often choose among axial, sagittal, and coronal views. The following code produces the six different ways of customizing ggseg plots illustrated in Figure 2:

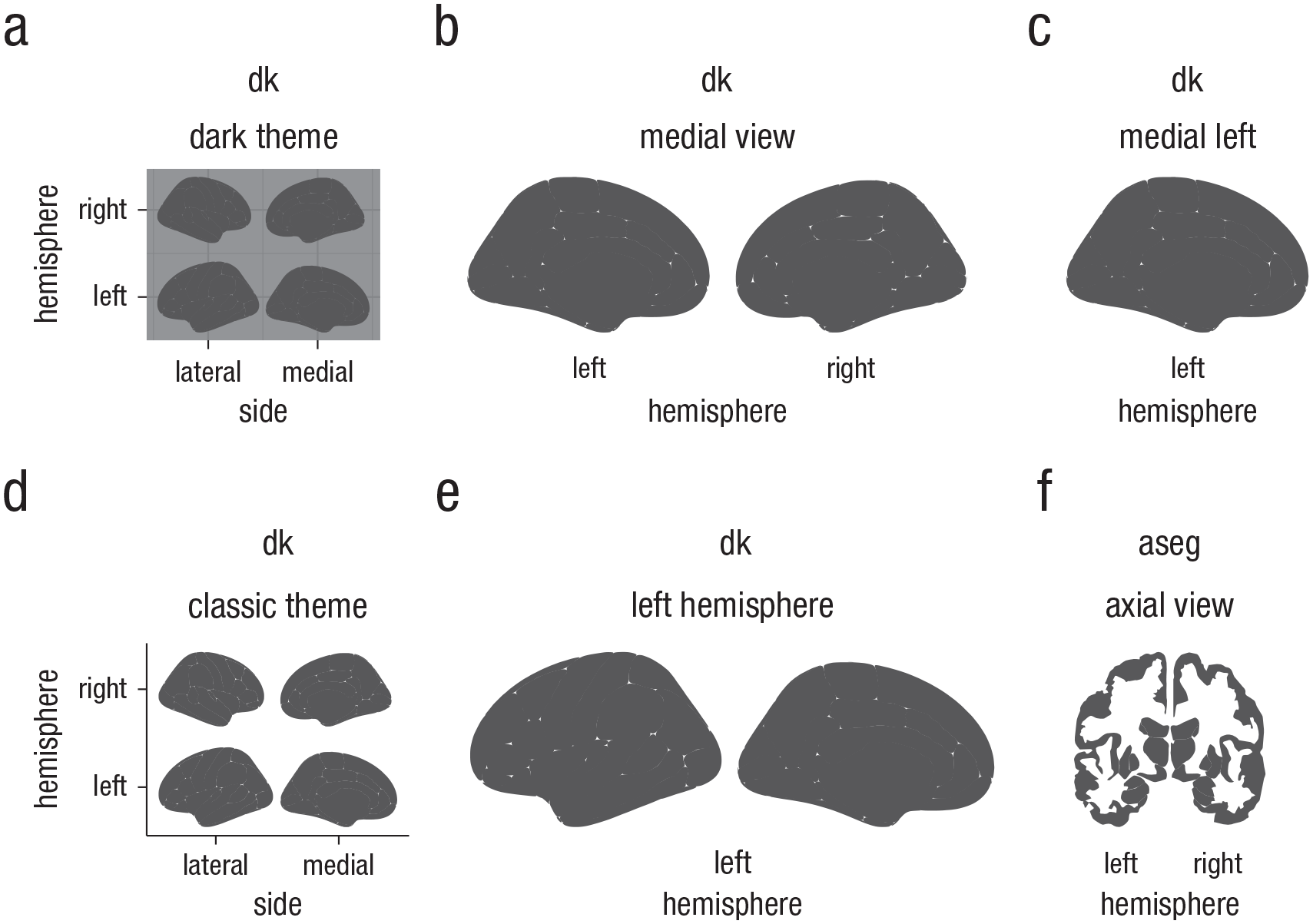
Illustration of some ggseg display options for cortical atlases. The displays here are representations of (a) the Desikan-Killiany (dk) atlas (Desikan et al., 2006), stacked, with the dark theme; (b) the dk atlas with the medial view only; (c) the dk atlas with only the left medial display; (d) the dk atlas, stacked, with the classic theme; (e) the dk atlas with the left hemisphere only; and (f) the axial view of the Automatic Segmentation of Subcortical Structures (aseg atlas; Fischl et al., 2002).
Using one’s own data with fill and colour
The

Plot created by supplying
Most users will use

In any atlas, the column
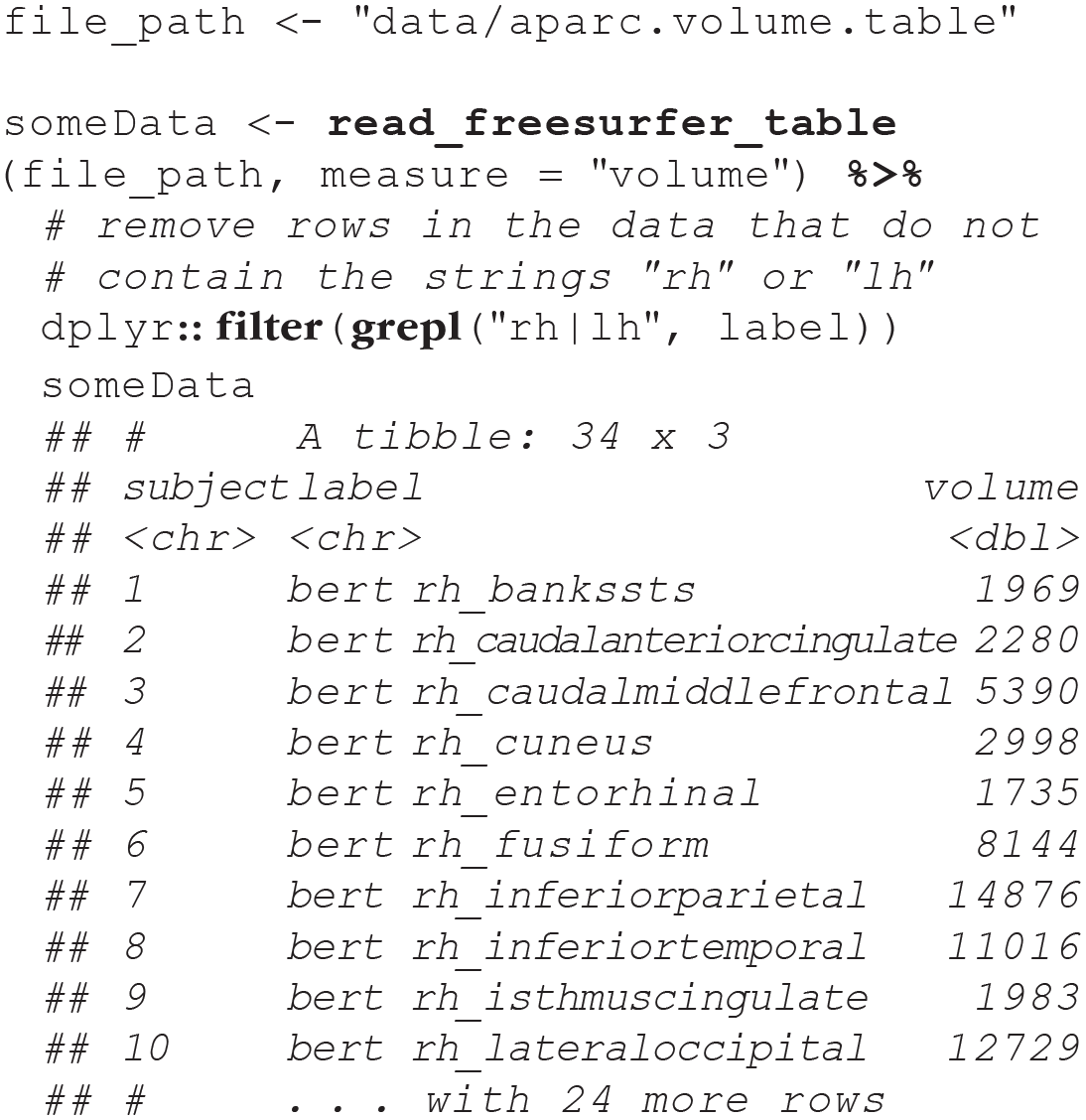
These summary stats files also contain extra metrics, such as intracranial volume and total gray-matter volume. These latter measures do not map onto ggseg, and therefore, one should filter them out of the data returned from
A second option is to read in statistics files directly from the entire subject directory, using regular expression to select the statistics files from the target atlas. The default aparc (FreeSurfer’s name for the dk atlas) has a unique string ending with
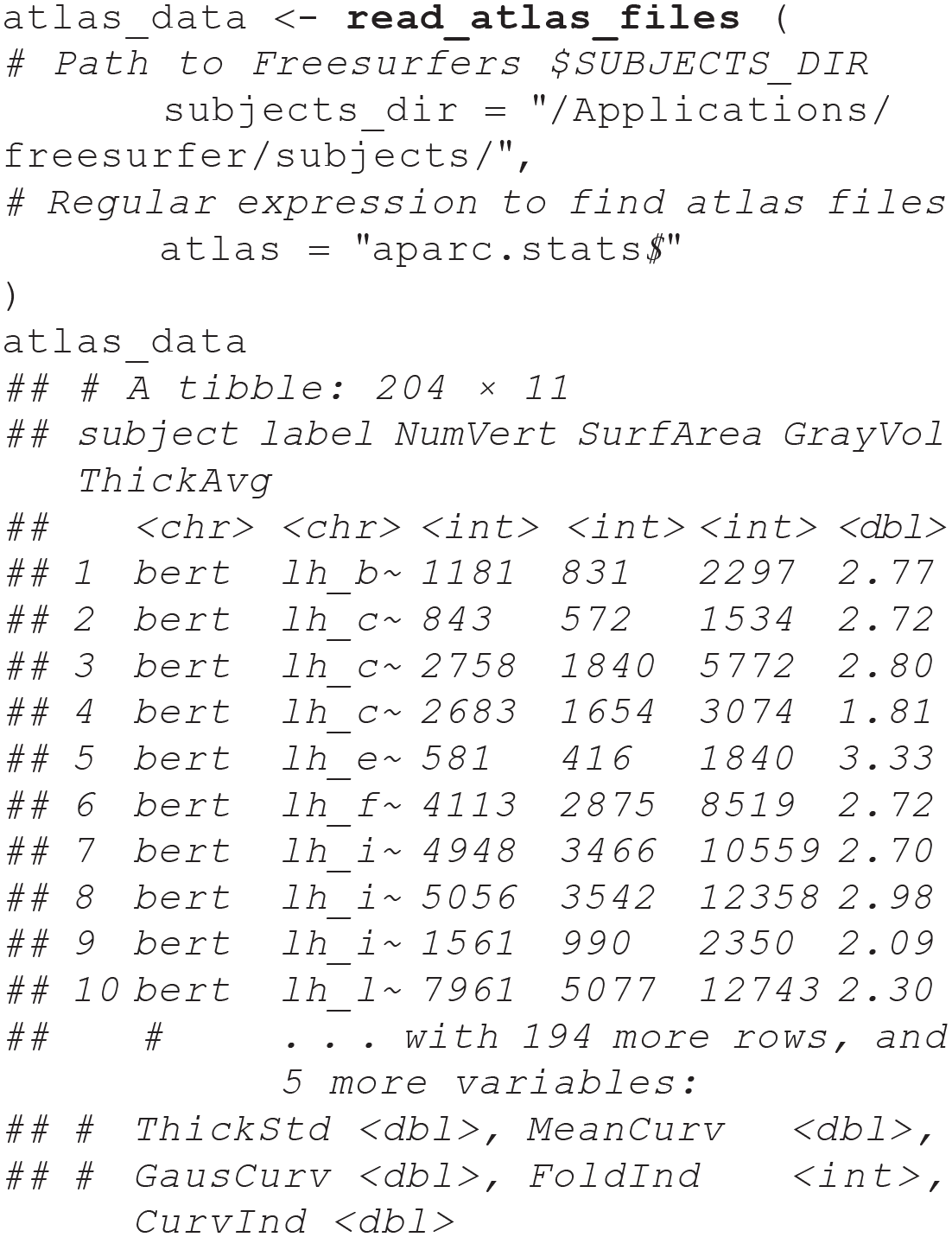
These long-format data can be used directly with the

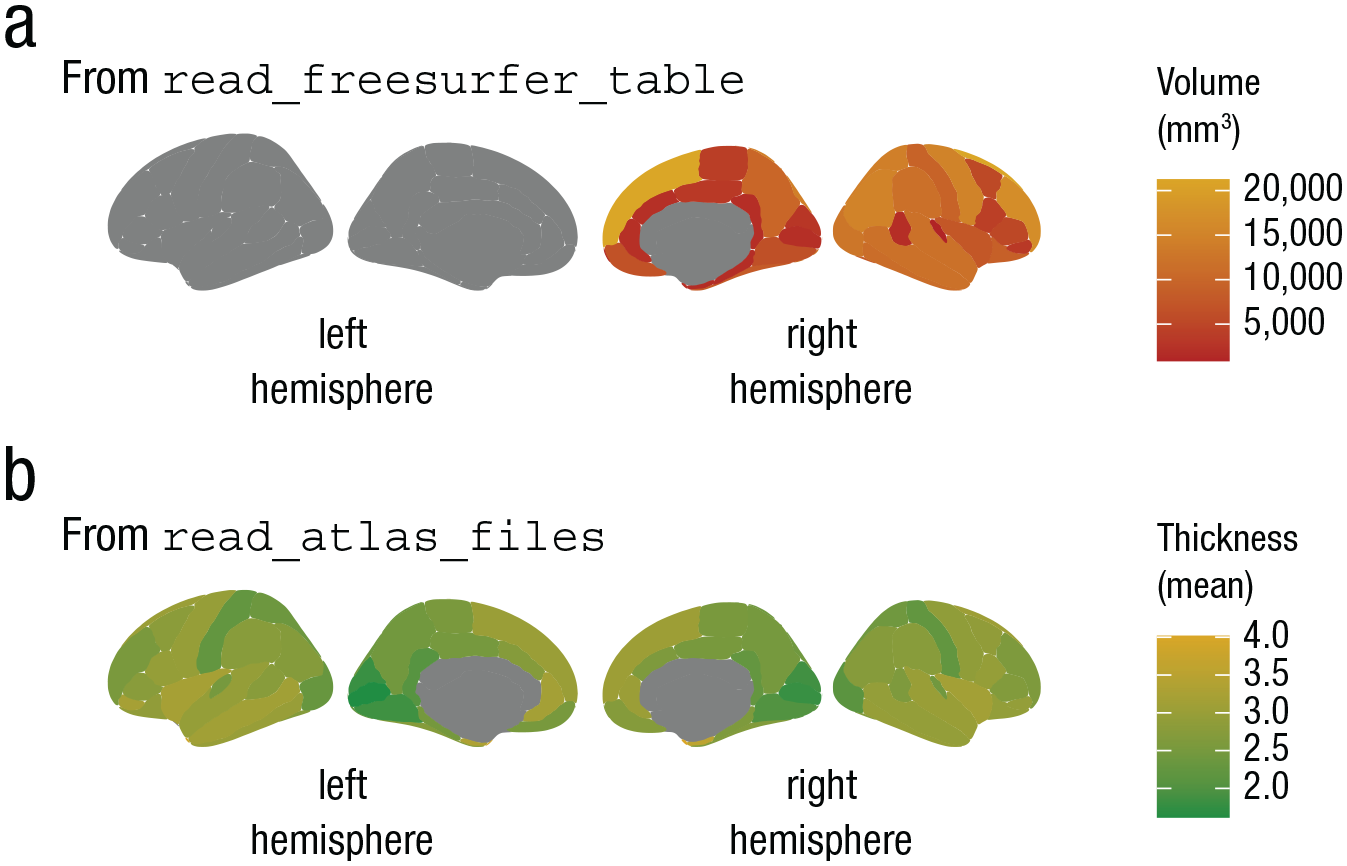
Examples of outputs obtained using
If the results are only in one hemisphere, but you still want to plot both of them, make sure your
Creating subplots
There is often the need to plot a statistic of interest in different groups (e.g., to plot thickness or brain-cognition relationships separately in young and older adults). This may be accomplished with
Convert the data to long format with a column indexing which group each row corresponds to (group data should appear in separate rows, not in separate columns).
Use dplyr’s
Finally, apply
Figure 5b provides an example of separate subplots created following these three steps. In this example, ggseg was applied to a mock data set including summary statistics for groups labeled “Young” and “Old.”
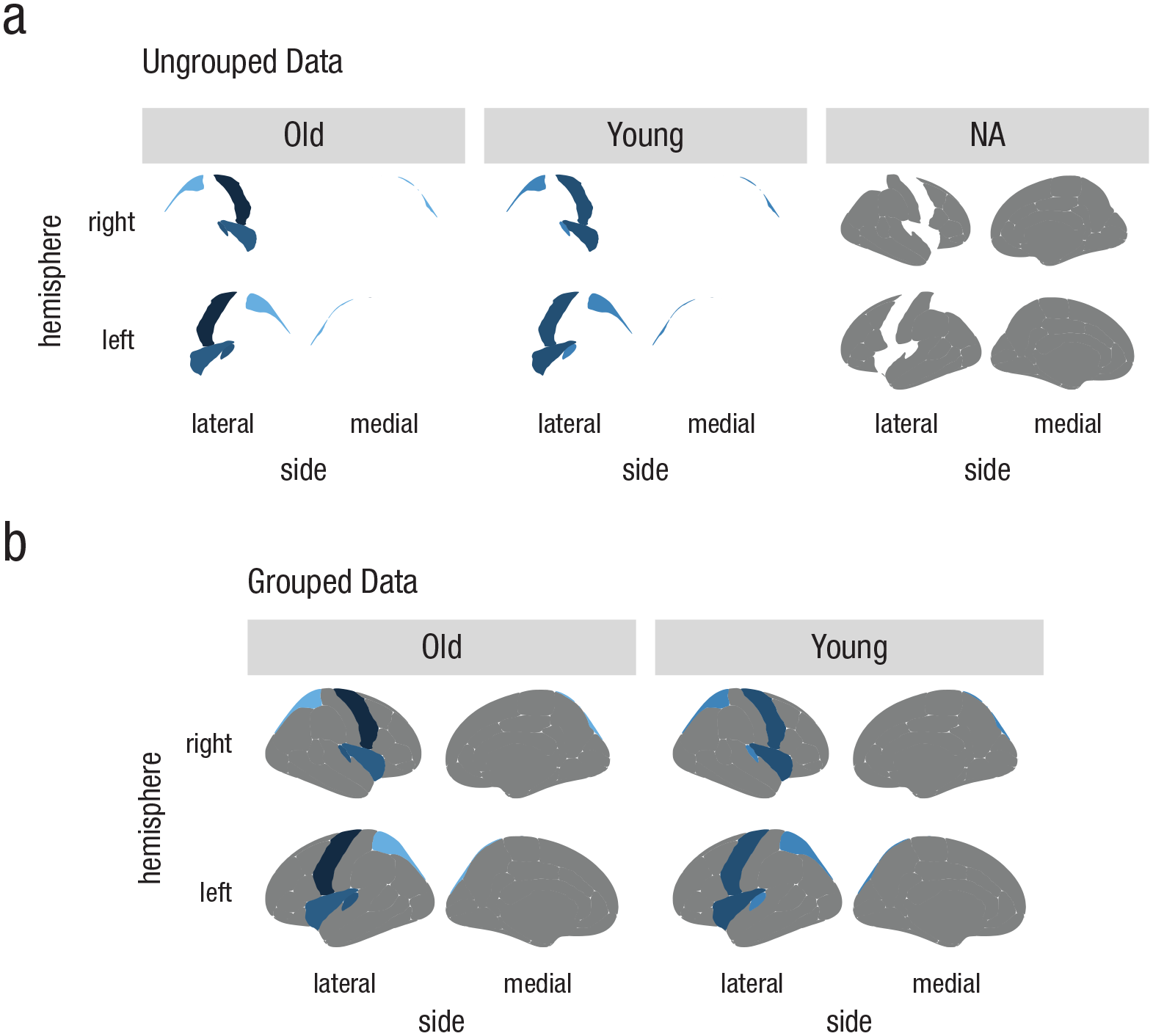
Examples of incorrect and correct faceting with ggseg. The plots in (a) were created using ungrouped data. The data were faceted incorrectly because they were not merged correctly with the atlas. In contrast, when the data were grouped (b), they merged in the expected way, and correctly faceted plots were created. NA = not applicable or not represented in the data.
The second step is needed because the
The following code was used to create the plots in Figure 5:

Using other atlases
Although the dk atlas is the default atlas for the

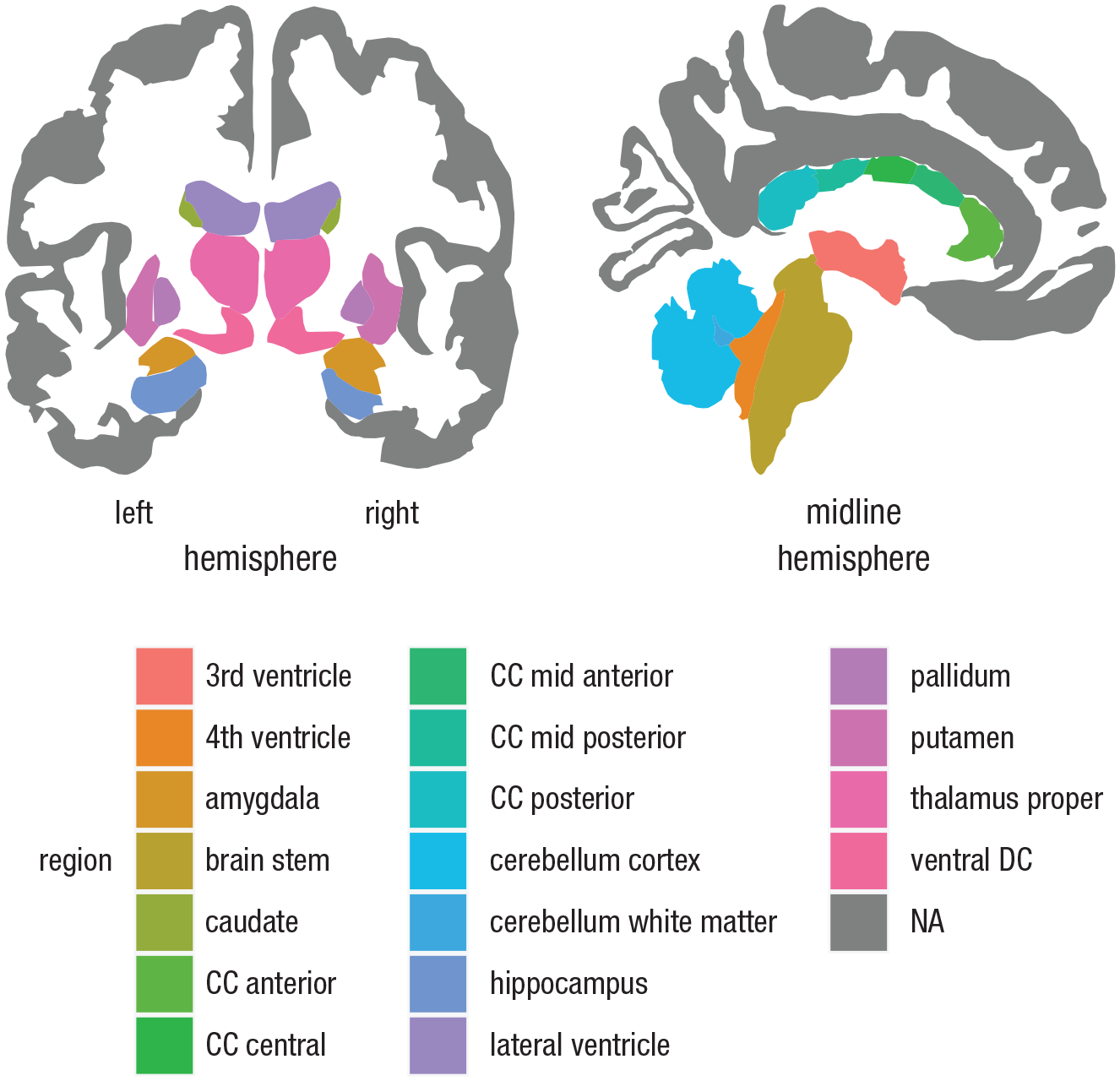
Plotting subcortical atlases using ggseg. Atlases for subcortical structures have some distinct differences from the cortical atlases, such as the Desikan-Killiany atlas (Desikan et al., 2006). For instance, there is no option to show only a single hemisphere. Furthermore, rather than showing lateral and medial surfaces, they show axial, coronal, or sagittal slices. To produce the plot in this figure, the Automatic Segmentation of Subcortical Structures (aseg atlas; Fischl et al., 2002) was supplied to the
Plotting 3D Mesh Data
Representing brains as 2D polygons is a good solution for fast, efficient, and flexible plotting, and these plots can be easily incorporated in interactive apps such as shiny (Chang, Cheng, Allaire, Xie, & McPherson, 2019). Yet brains are intrinsically 3D, and it can be challenging to recognize the location of a region in a flattened image. This problem is exacerbated in atlases that represent subcortical features because they cannot be flattened into two dimensions, as cortical features can. Hence, we have also provided the ggseg3d package for plotting, viewing, and printing 3D atlases in R. The functionality of ggseg3d is based on tri-surface mesh plots through plotly (Sievert, 2019). The data structure is more complex than that of the ggplot2 polygons, and the package includes additional options, such as for brain inflation, glass brains, and camera locations. Because ggseg3d is based on plotly, the resulting brain atlases are interactive, which is useful for interpretation and public dissemination. We recommend that users familiarize themselves with plotly (Sievert, 2019) before using ggseg3d.
Out of the box,
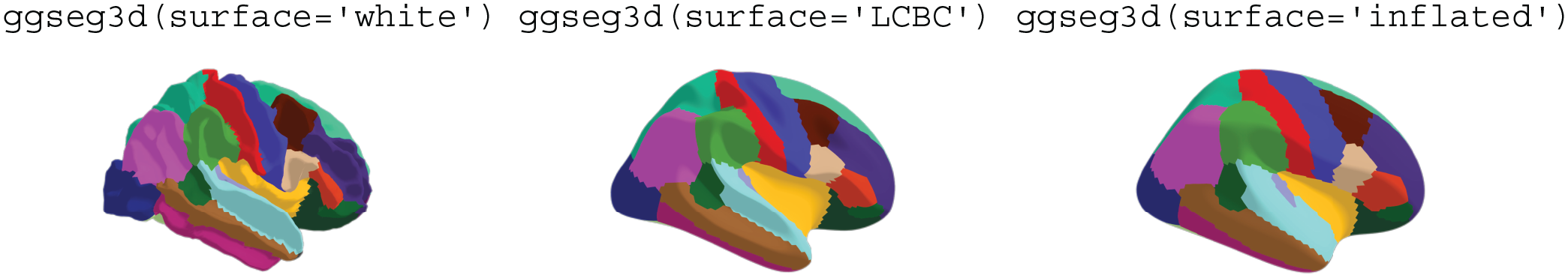


The three surface options provided in ggseg3d atlases. The white surface (left) is the white-matter surface, the LCBC (Center for Lifespan Changes in Brain and Cognition) surface is a semi-inflated (inflated over 10 iterations) white-matter surface (center), and the inflated surface (right) is a fully inflated gray-matter cortical surface provided by the FreeSurfer software. The label above each plot is the literal code used to create it.
The 3D-atlas data are stored in nested tibbles. Each cortical atlas has data sets for the three different surfaces (see Fig. 7) and the two hemispheres. Only one surface is available for subcortical atlases, as inflation procedures are irrelevant. The
Supplying external data
The

Note the
Customizing colors and the color bar
Custom color palettes can be created either in Hex or in R names. Colors will be evenly spaced in the color scale. A palette may also be supplied as a named numeric vector, in which the vector names are the colors that the user wishes to use and the numeric values are the break points for the colors (e.g.,

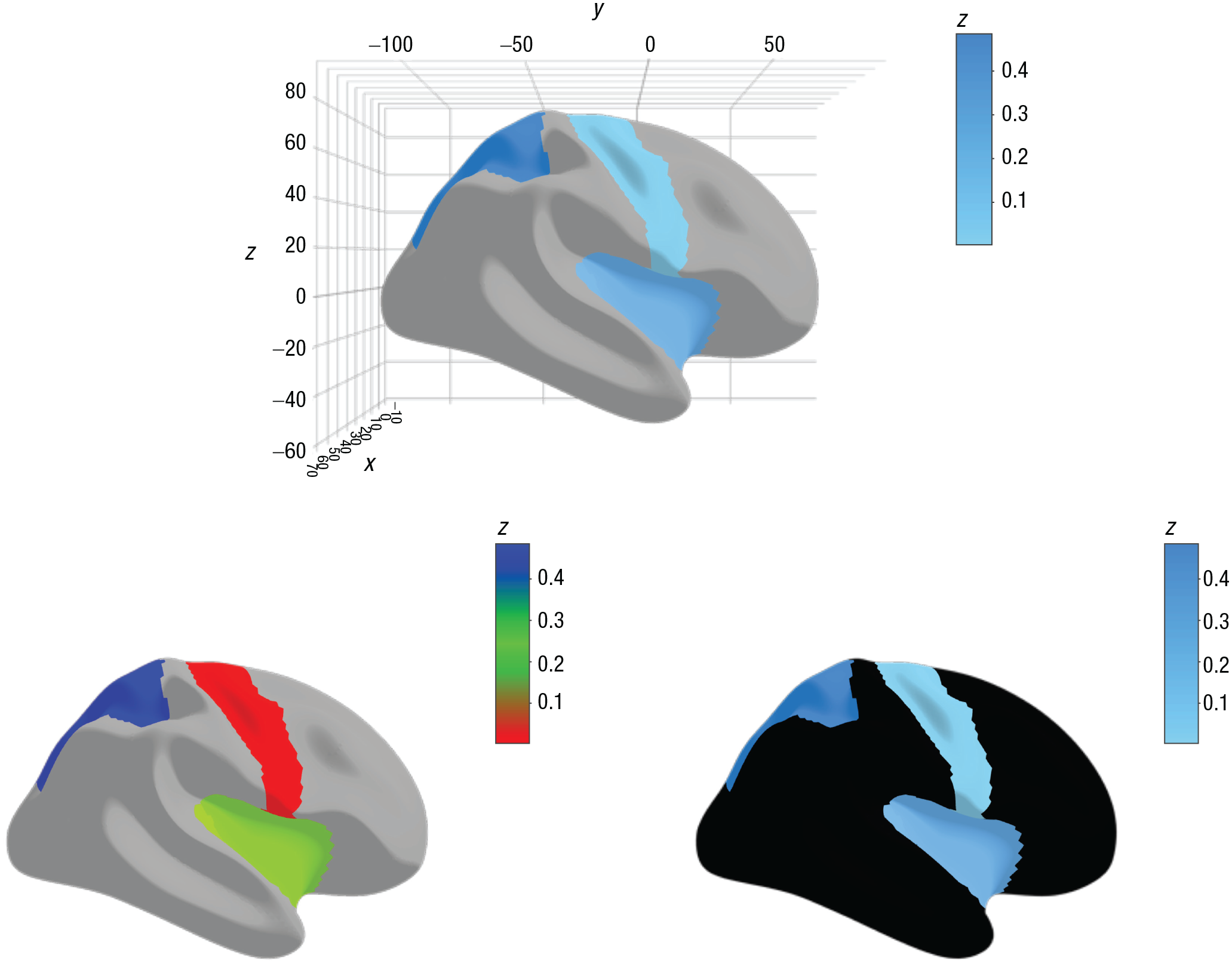
Illustration of the use of color palettes in ggseg3d. Supplying data to
Adding a glass brain
A glass brain is a translucent representation of the brain that can provide a frame of reference, particularly in the case of subcortical parcellations, and it serves only as a visual aid. A cortical glass brain can be added to ggseg3d plots through the function

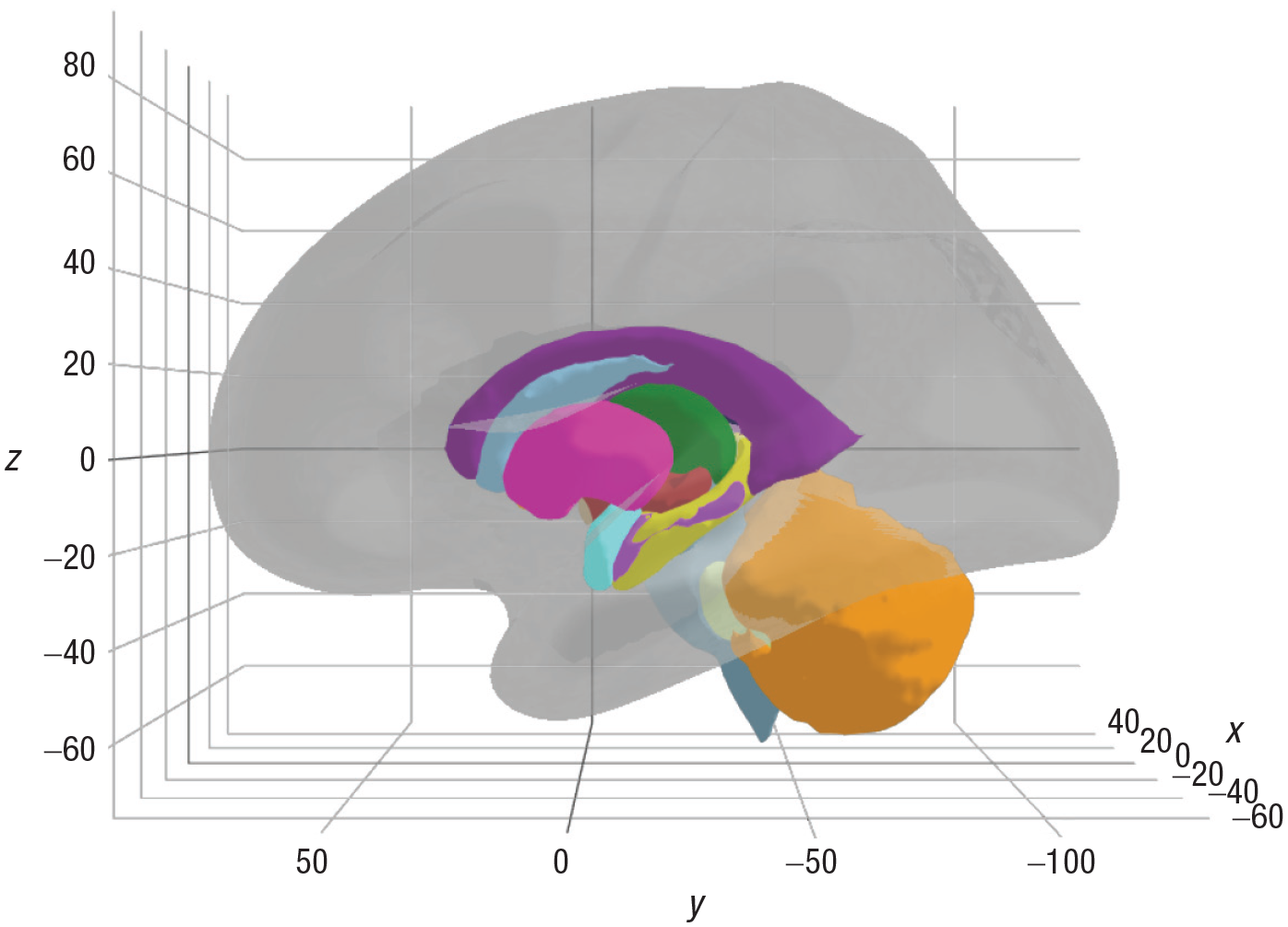
Example of a glass brain added to the visualization of subcortical structure. The plot of the glass brain is controlled by three arguments: opacity, hemisphere, and color. In this example, the glass brain was added for the left hemisphere, and the remaining two arguments were left as their default values. A similar, interactive version of this plot can be found in the online documentation (https://lcbc-uio.github.io/ggseg3d//articles/ggseg3d.html#colours).
More Resources
The
The online documentation for ggseg3d also includes interactive versions of many of the figures in this Tutorial and several more options not covered here (see https://lcbc-uio.github.io/ggseg3d/articles/ggseg3d.html). The online documentation will be updated as new features are required and as we improve user functionality and options.
Additional Atlases
The ggseg and ggseg3d packages have two atlases each: 2D and 3D variations, respectively, of the dk (Desikan et al., 2006) and aseg (Fischl et al., 2002) atlases. These are, however, only two among many meaningful ways of segmenting the brain into different regions. Thus, we have developed the ggsegExtra package, a library with two main functionalities: (a) installing additional data sets from other repositories for plotting with the ggseg and ggseg3d packages and (b) creating custom atlases for ggseg and ggseg3d.
Verified existing atlases
The number of new atlases being created is ever increasing, as research and methods in neuroimaging analysis progress. The ggsegExtra package is intended to be expanded as a community effort, as new and informative atlases are published. A sample of the atlases currently listed in the ggsegExtra package is illustrated in Figure 10 and explored in our shiny demonstration (https://athanasiamo.shinyapps.io/ggsegDemo/). The following code shows how you can look for ggseg-compatible atlases using functions in the ggsegExtra package:
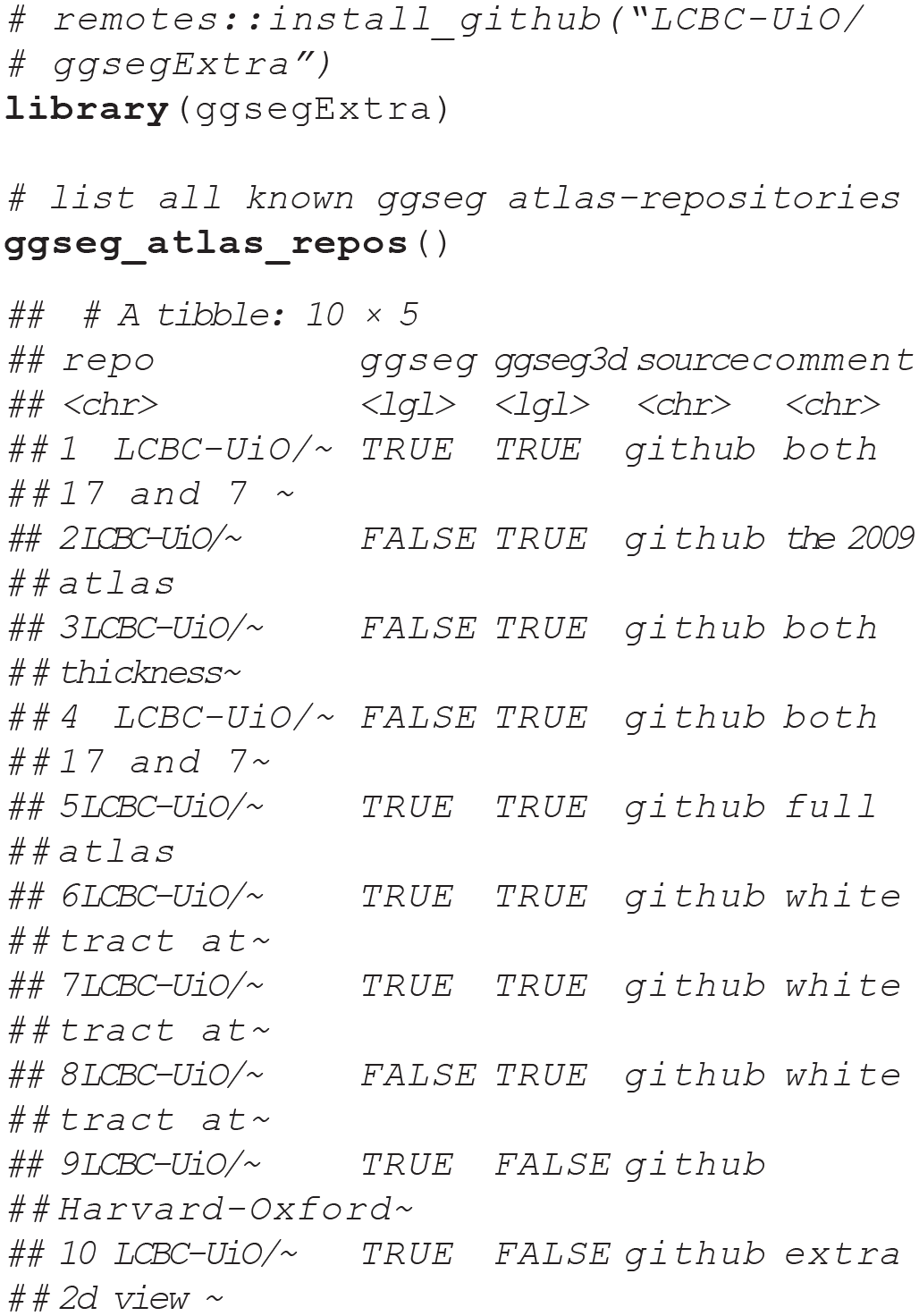
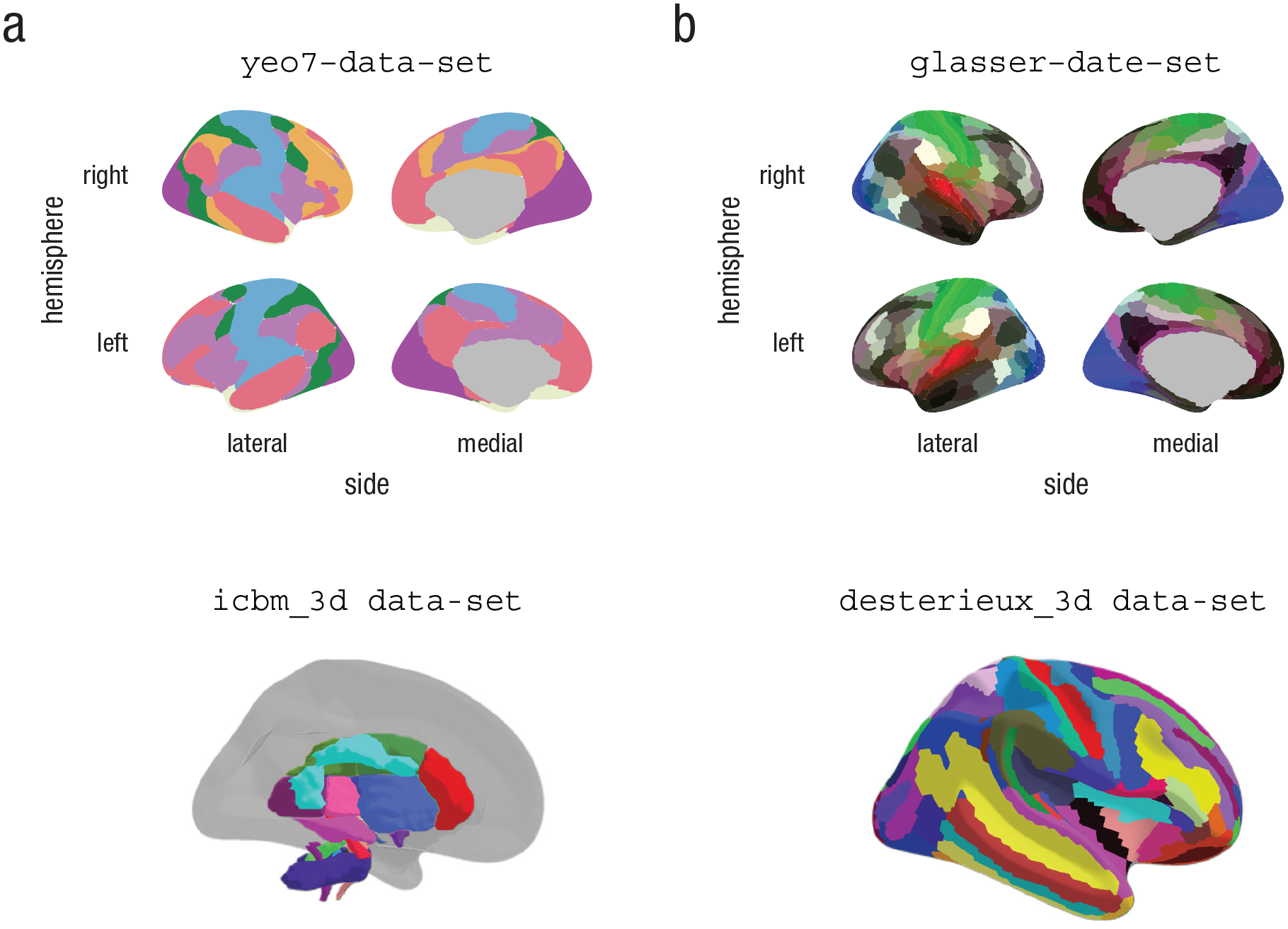
Four example data sets from the ggsegExtra library, plotted with the
Having identified an atlas you wish to use, you will need to install it as you would an R library. There is a convenience function in ggsegExtra that will help you do that. Alternatively, you can install all the existing atlases. This function will take time to execute and will consume quite some space, as some of the atlases (particularly the ggseg3d atlases) are large. Once the atlases are installed, the atlas packages can be loaded in the same way as any other R package, using the

Creating custom atlases
In many cases, users would like to create their own atlases based on existing parcellations or their own imaging results. There are several functions in ggsegExtra to aid users in creating atlases compatible with either ggseg or ggseg3d. The procedures depend on the origin of the segmentation (surface based vs. volume based) and what type of system the user is working on. In addition, users need other software dependencies to create new ggseg-compatible atlases. Therefore, we do not describe the procedure here, but refer readers to the “Articles” sections of the online documentation (https://lcbc-uio.github.io/ggsegExtra/index.html) for further explanations and guides. We encourage users who successfully create new atlases to contribute to the ggsegExtra brain-atlas repository by adding the repositories containing their brain atlases to the repository list.
Discussion
The main aim of the ggseg, ggseg3d, and ggsegExtra packages is to ease and streamline visualization of brain-atlas data in R, by gathering a collection of atlases from several scientific sources and providing customized plotting functions. In this Tutorial, we have introduced the packages by presenting some use examples and highlighting the main functions and options that are available. As visualization tools, these packages add complementary functionality to other packages, such as ggBrain (Fisher, 2019) and ggneuro (Muschelli, 2017) in R and software-specific image viewers such as FSLeyes (McCarthy, 2019) and Freeview (Dale et al., 1999). We do not aim to compete with software-specific visualizations or advocate for the superiority of the ggseg packages as visualization tools. After all, flattened 2D polygons do not rely on a meaningful brain coordinate system, and the units of information in 3D meshes are limited to the number of parcellations. On the contrary, we believe that ggseg’s niche among visualization tools resides in its simplicity and its ability to be combined with statistical-analysis pipelines. The possibility for the ggseg packages to serve as interactive tools for dissemination and reproducibility purposes when combined with other technologies, such as Binder (Project Jupyter et al., 2018) or shiny (Chang et al., 2019), is an added benefit (for an example, see the Supplementary Material in Vidal-Piñeiro et al., 2019).
The functions in the ggseg packages provide users with the possibility of adapting the plots to their wishes, and also make it possible to create and contribute to the atlas repository in ggsegExtra. The three ggseg packages offer the following main features:
A collection of 2D-polygon and 3D-mesh brain-parcellation atlases: The atlas data include the necessary coordinates for plotting and include other helpful information for easier user access.
Functions for visualization: The
Complementary features, like color scales, and functions for converting compatible data frames to atlas classes, reading in likely data sources (e.g., FreeSurfer analysis output), and creating new atlases for use with ggseg and ggseg3d.
The foundations of the ggseg packages trace back to the necessity of visualizing and exploring the life-span trajectories of cortical thickness across different brain regions (see the Supplementary Material in Vidal-Piñeiro et al., 2019). That is, ggseg arose from the need to inspect and display brain information over time (i.e., to include a spatial dimension and a time-varying factor) in a way that overcomes the constraints of print journals and classical 2D plots (e.g., bar plots). The current state of science requires researches to share the results of studies in both high detail and an intuitive manner, so as to permit communication to wide audiences and facilitate reproducibility. Hence, we believe these tools conform to the essence of open science and invite users to improve the code, provide examples or tutorials, and contribute to the atlas collection according to their own interest and needs via the public ggseg GitHub repository (https://github.com/LCBC-UiO/ggseg), ggseg3d GitHub repository (https://github.com/LCBC-UiO/ggseg3d), and ggsegExtra GitHub repository (https://github.com/LCBC-UiO/ggsegExtra).
The ggseg packages are based on ggplot2 and plotly, which are only two of several visualization tools in R, in addition to R base plotting. Although the functionality demonstrated here could also be integrated into packages such as lattice (Sarkar, 2020), both ggplot2 and plotly have become established tools in the R community and have extensive online documentation. This makes them great packages to base further extensions and wrappers on.
Finally, although the ggseg packages are limited to brain parcellations, we believe that the structure and functions of the packages can be easily applied to any scientific field that benefits from data being displayed across the spatial dimension. We encourage readers to borrow the package functionalities and adapt them to their respective fields and structures of interest, such as has already been done with the gganatogram package (Maag, 2018).
Planned Package Improvements
For ggseg, we are working on making a proper
In the case of ggseg3d, we hope to make it possible to color each individual mesh triangle, and thereby provide users with more flexibility in the plotting. This should also improve the speed of
Conclusion
Visualization is a fundamental aspect of neuroimaging, used to explore and understand data, guide interpretation, and communicate with colleagues and the general audience. In this Tutorial, we have introduced the ggseg packages, tools for visualizing brain statistics through brain-parcellation atlases in R. This visualization tool easily combines with interactive routines as well as with diverse statistical-analysis pipelines. We hope this tool and Tutorial prove useful to neuroscientists and inspire other researchers to apply the functions in a wide variety of fields and structures.
Footnotes
Acknowledgements
We thank John Muschelli for comments on the package code and its adaptation to Neuroconductor (Muschelli et al., 2019), and Richard Beare for the first community-created atlas and comments on the base code. We are indebted to A. M. Winkler for providing scripts for converting neuroimaging data (FreeSurfer) into .ply files, a necessary step for conversion to mesh plots, and to Inge Amlien for the LCBC surface files and helpful contributions to the ggseg3d pipeline. We are thankful for those users who actively contributed to the package and all the individuals who worked on the framework on which the package is sustained (e.g., R Project, neuroimaging software). Finally, we thank Anders M. Fjell and Kristine B. Walhovd for their encouragement and financial support.
Transparency
Action Editor: Alex O. Holcombe
Editor: Daniel J. Simons
Author Contributions
D. Vidal-Piñeiro generated the idea for the tool and the initial scripts for plot visualization. He has also been responsible for converting neuroimaging data to ggseg-like data (e.g., polygons and mesh data). A. M. Mowinckel adapted the initial scripts and converted the functions into package format and has continued developing the functions with the aim of increasing user friendliness. She is also responsible for conceiving the idea of adding the mesh-plot functionality through plotly and developing the pipeline for making that possible. A. M. Mowinckel wrote the first draft of the manuscript, and both authors critically edited it.
Prior Versions
Some of the content presented here also appears in the package vignettes of ggseg and ggseg3d, which may be accessed through R or the package websites (Mowinckel & Vidal-Piñeiro, 2019a, 2019b). The first author’s blog also has several tutorials on ggseg’s creation and functionality (Mowinckel, 2018a, ![]() ).
).
