Abstract
Foodborne illness is a major public health issue that results in millions of global infections annually. The burden of such illness sits mostly with developing countries, as access to advanced laboratory equipment and skilled lab technicians, as well as consistent power sources, is limited and expensive. Current gold standards in foodborne pathogen screening involve labor-intensive sample enrichment steps, pathogen isolation and purification, and costly readout machinery. Overall, time to detection can take multiple days, excluding the time it takes to ship samples to off-site laboratories. Efforts have been made to simplify the workflow of such tests by integrating multiple steps of foodborne pathogen screening procedures into a singular device, as well as implementing more point-of-need readout methods. In this review, we explore recent advancements in developing point-of-need devices for foodborne pathogen screening. We discuss the detection of surface markers, nucleic acids, and metabolic products using both paper-based and microfluidic devices, focusing primarily on developments that have been made between 2015 and mid-2020.
Keywords
Introduction
Despite existing food safety measures in place, foodborne illness continues to pose a significant threat to the public health and economic growth of developed and developing nations alike. Foodborne illnesses largely result from the consumption of food or beverages contaminated with pathogens, which exert their detrimental effects through colony establishment in host tissues or the production of harmful toxins. While commonly manifesting as mild diarrheal diseases, food poisoning can have life-threatening implications, such as kidney and liver failure, neural disorders, respiratory paralysis, and death. 1 Among the 31 known pathogenic foodborne microorganisms, Norovirus, Salmonel-la, Clostridium perfringens, Campylobacter, Staphylococcus aureus, Clostridium botulinum, Listeria monocytogenes, Escherichia coli O157:H7, and Vibrio have been identified as the leading causative agents responsible for food-related illnesses, hospitalizations, and deaths.2,3 Each year, approximately 600 million individuals globally exhibit symptoms of foodborne illness, and 420,000 of these individuals die. 3 Furthermore, the effects of foodborne pathogens significantly hinder economic development through the burdensome expenditures on medical care, lost productivity during illness, and limited agricultural sales.3,4
Notably, developing nations and remote rural areas systematically bear the largest burden from foodborne illness. According to the World Health Organization in a 2010 report, the highest global burden, as measured by disability-adjusted life-years (DALYs) per 100,000 population, was observed in two African subregions (1300 and 1200 DALYs) followed by Southeast Asian (690 and 1200 DALYs) and Eastern Mediterranean (570 DALYs) subregions. 5 In comparison, the lowest burden was observed in North America (35 DALYs). These developing regions are inherently at higher risk due to insufficient clean water supply, inadequate sanitation measures for food processing and handling, poorly enforced food safety standards, and the absence of electric power and equipment needed for proper food storage. 3 Such regions also lack the healthcare infrastructure for responding to and preventing outbreaks, thus accelerating foodborne illness-related morbidity and mortality. 3
Rapid and accurate testing is essential to mitigating foodborne illness outbreaks and must be utilized at all stages of the food supply chain. As only a few bacterial cells are required for pathogenicity, 6 food safety measures must be highly sensitive to effectively screen and detect contaminated food. Currently, established techniques for foodborne pathogen detection largely rely on microbiological detection and identification. These culturing methods consist of sample homogenization and multistep selective enrichment, followed by various plating steps for colony isolation.7,8 Detection is traditionally performed with phenotypic analysis and biochemical confirmation via Gram staining, metabolic assays, and ribotyping.6–8 Despite the reliability of culture-based techniques, the laborious and time-consuming nature and requirement of trained lab personnel, lab equipment, and sterile environments prevent the universal utilization of such methods.6,9,10 Notably, the 2–3 days required for initial results and 7 days for species confirmation impede timely pathogen detection prior to food consumption. Furthermore, culturing techniques reveal challenges in detecting fastidious and viable but not culturable species, especially among samples with high levels of competing microorganisms.7,11 Recently developed optical sensors based on the analysis of characteristic light diffraction patterns (BARDOT, 12 Scatterometer, 13 converging spherical wave illumination, 14 etc.) have been useful in rapidly differentiating plated bacterial species within a genus and serovars within a species, such as Shiga toxin-producing E. coli (STEC) serogroups. 12 Unfortunately, varying growth conditions, such as incubation time, incubation temperature, and type of nutrient medium, affect colony morphology and, consequently, have been found to interfere with diffraction patterns and limit the accuracy of these light scattering-based methods. 14
With the development of more rapid immunology-based and nucleic acid-based techniques, the conventional screening procedure has transitioned to hybrid techniques utilizing culture-based methods for sample enrichment and molecular techniques for pathogen detection. Immunological assays including enzyme-linked immunosorbent assay (ELISA), enzyme immunoassay (EIA), and enzyme-linked fluorescent assay (ELFA) are commonly utilized for the detection of surface markers on pathogens and pathogen-produced toxins based on antibody–antigen interactions. Polymerase chain reaction (PCR) and its various derivatives, including real-time PCR, reverse transcription PCR (RT-PCR), and multiplex PCR, are commonly employed for the selective amplification and detection of toxin-related genes. 11 Optical techniques including Raman and Fourier transform infrared spectroscopy, flow cytometry, and surface plasmon resonance have also been employed to selectively detect bacterial cells in complex food suspensions.13,15 While time to detection is reduced compared with culture-based methods, these techniques still require numerous operation steps and demand accessibility to expensive equipment, laboratory settings, and trained laboratory personnel. Therefore, these limitations severely restrict the applications of such techniques in developing nations and remote regions.
The lack of food pathogen screening techniques suitable for resource-poor settings has recently warranted increased demand for the development of infrastructure-independent technologies. In particular, efforts have transitioned to focus on point-of-need (PON) food pathogen detection techniques that are affordable, sensitive and specific, user-friendly, rapid and robust, equipment-free, and deliverable to end users, as outlined by the World Health Organization’s ASSURED criteria. 16 Emerging methods commonly incorporate isothermal amplification, lateral flow immunoassay (LFA), functional nucleic acids, immunosensors, or metabolic assays with various detection readouts in paper-based and microfluidic devices. Such hybrid techniques aim to reduce the complexity of food screening processes and increase feasibility for in situ detection.
In this review, we discuss novel PON technologies for foodborne bacterial screening based on their mode of detection: surface marker, nucleic acid, or metabolic-based (
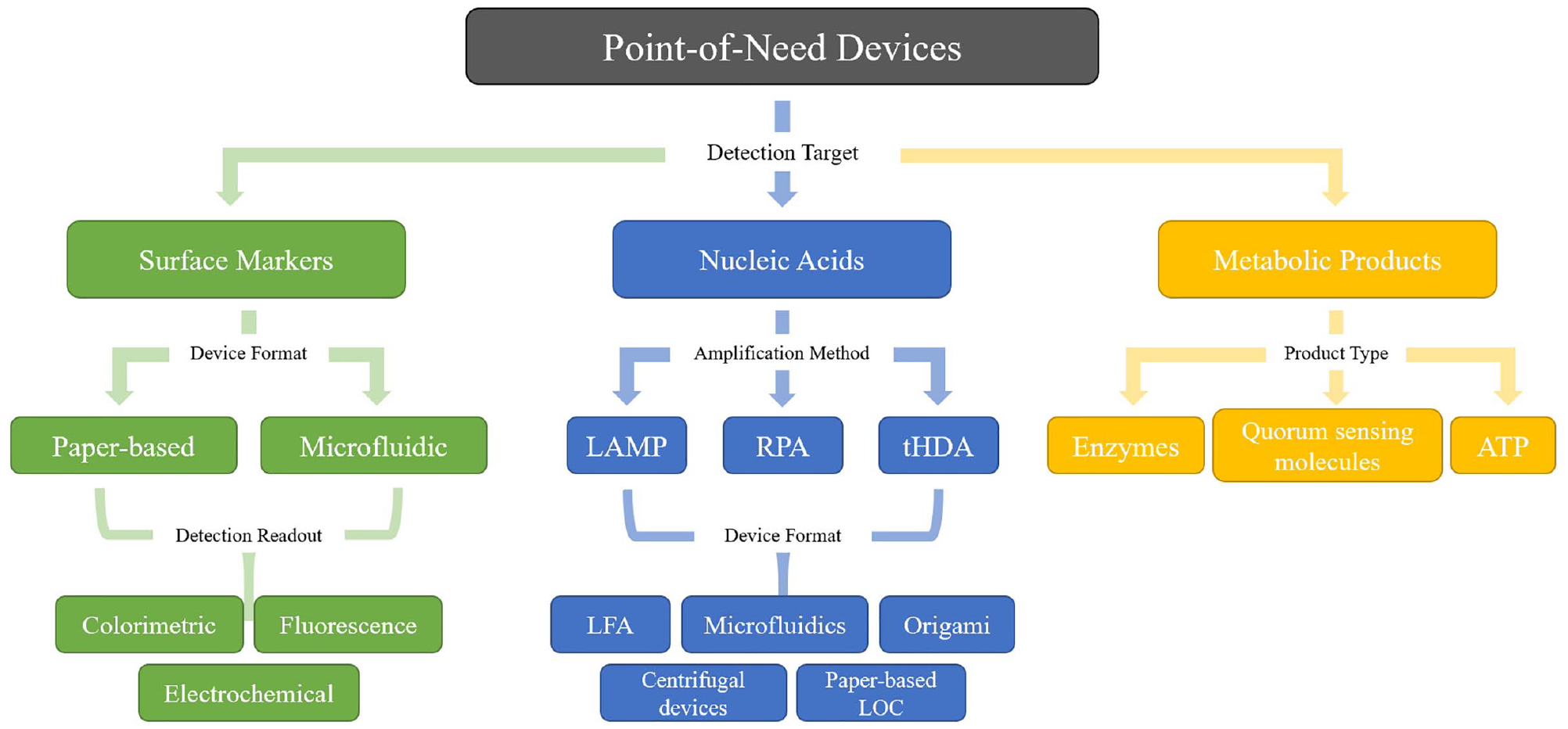
Overview of PON devices for the screening of foodborne pathogens. We give special focus to methods that detect surface markers, nucleic acids, and metabolic products of the target pathogens. The design and readout of devices aimed at detecting each of these targets vary and are discussed in detail in this review.
Detection of Surface Markers
Surface markers have been used extensively in foodborne pathogen testing, as the procedures for producing monoclonal antibodies are well established and facilitate high specificity in binding to the target antigens. 18 Furthermore, the detection of surface markers allows for the analysis of whole cells rather than specific nucleic acid or enzyme targets within the cell, removing the need for additional cell lysis steps. Since PON devices aim to have as few user steps as possible to both minimize time to detection and improve ease of use, this elimination of cell lysis and purification steps is extremely beneficial.
However, this does not mean whole cells are completely exempt from all preparatory steps prior to detection. For bacteria found in complex solid food samples such as meat or vegetables, some type of processing step is required. This typically involves using a machine to homogenize and filter the sample to remove any larger debris that could interfere with detection. 19 These machines are often bulky or expensive, proving to be a major setback in achieving fully PON sample-to-result testing. Even in less complex samples such as milk or juice, the presence of inhibitory food particulates and proteins leads to a decrease in sensitivity; therefore, enrichment protocols are needed to improve the limits of detection (LODs) to an acceptable level. 20 Enrichment culture is typically performed in buffered peptone water or nutrient broth and incubated at 37 °C for several hours, although the incubation time may vary depending on the bacterial strain and sensitivity of the detection device. 21 This preparatory step is not suitable for PON testing because it must be performed in laboratory settings and increases the time to detection significantly. For this reason, groups aiming to achieve entirely PON detection of surface markers on foodborne pathogens must seek non-culture-based isolation and concentration methods. Thus, in the following sections we give special attention to groups that have integrated testing with more PON-friendly enrichment techniques, such as immunomagnetic separation (IMS) and aqueous two-phase systems (ATPSs).
Paper-Based Detection
With growing interest in PON testing, there has been an increase in the development of paper-based devices for the detection of foodborne pathogens. While paper devices with colorimetric readouts, such as LFAs and microfluidic paper-based analytical devices (μPADs), have been around since the early 2000s, it was only recently that these methods have been adapted for use in foodborne pathogen testing. Furthermore, there has been a focus in recent years on addressing the traditional drawbacks of such devices by improving fabrication techniques, LODs, and ease of use. We will discuss the novel developments in paper-based devices utilizing colorimetric, fluorescent, and electrochemical readouts in the following sections, as well as in Table 1 .
Paper-Based Detection of Pathogenic Surface Markers.
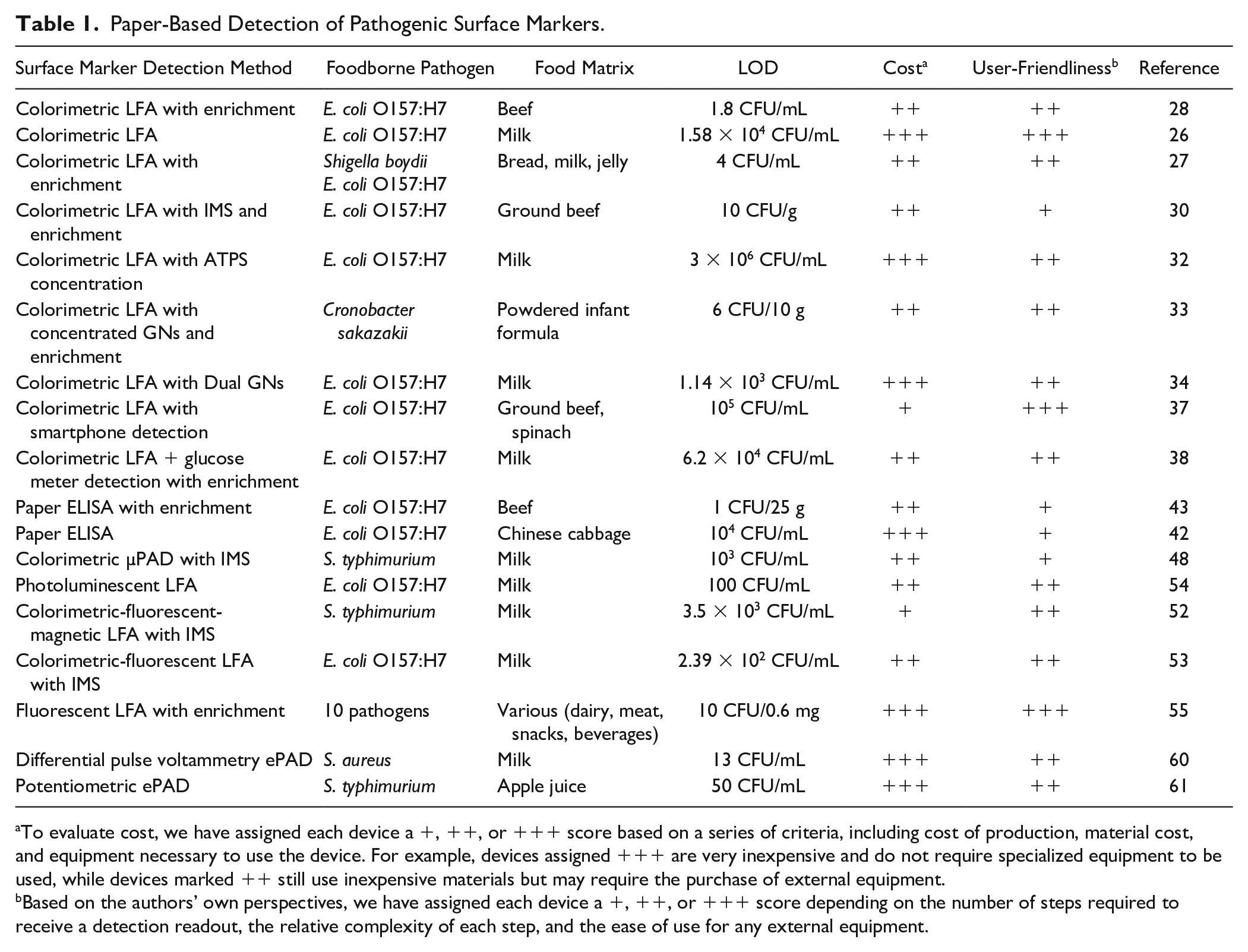
To evaluate cost, we have assigned each device a +, ++, or +++ score based on a series of criteria, including cost of production, material cost, and equipment necessary to use the device. For example, devices assigned +++ are very inexpensive and do not require specialized equipment to be used, while devices marked ++ still use inexpensive materials but may require the purchase of external equipment.
Based on the authors’ own perspectives, we have assigned each device a +, ++, or +++ score depending on the number of steps required to receive a detection readout, the relative complexity of each step, and the ease of use for any external equipment.
Colorimetric Detection
Colorimetric detection is currently the most widely used method in paper-based devices due to its lack of need for external equipment and simplicity of readout for untrained users. 22 LFAs are one such device that have been extremely well characterized and dominate the diagnostic testing market. 23 The sandwich assay form of the LFA is usually assembled on a nitrocellulose membrane with a conjugate pad containing reporter probes, 24 target antibodies printed on the test line, and secondary antibodies immobilized on the control line. 25 When the strip is placed in a positive sample, the fluid flows up the strip via capillary action and the probe-labeled targets bind to the antibodies, resulting in a color change on both the test and control lines. A negative test is indicated by the appearance of only the control line. Early versions of this LFA have proven that the assay can specifically detect the presence of foodborne pathogens in milk,26,27 bread, 27 and raw beef 28 both with and without prior enrichment. Furthermore, the low cost, portability, rapid time to detection, ease of use, and long shelf life make them ideal for use in PON settings.
However, traditional LFAs suffer from a major drawback, as they have relatively low sensitivities of 105 to 106 colony forming units (CFU)/mL unless the sample is enriched for many hours prior to testing.27,28 One of the primary reasons for these poor LODs is the short residence time for the binding of targets to the immobilized antibodies due to the rapid flow of fluid up the strip. 29 Various strategies have been implemented to remedy this issue, such as preconcentration techniques and modification of the gold nanoparticles (GNs). IMS is one such preconcentration method in which antibodies are immobilized on magnetic beads to capture and concentrate the target pathogen from the food sample before it is detected on the LFA. 30 As demonstrated by the Lai group, IMS can increase the sensitivity of LFAs up to 10-fold. ATPSs have also been used for the preconcentration of pathogens. An ATPS is a liquid–liquid extraction technique in which the phase separation of two immiscible solvents results in the preferential partitioning of biomolecules into one phase based on their physical and chemical properties. 31 Our group has previously shown that a UCON–potassium phosphate salt ATPS concentrates E. coli cells in a milk sample by more than 10 times, improving the LOD on a spot test immunoassay. 32 Spot test assays utilize the sandwich LFA format, but the antibodies are immobilized on a spot on the membrane as opposed to a line. Additionally, modifications to the GN reporter probes have been designed to increase binding of the target pathogens in the detection zone. One strategy involved the use of concentrated GNs, which increased the number of reporters captured by the immobilized antibodies on the test line and improved sensitivity 100-fold over traditional LFAs. 33 Another method amplified the LFA detection signal using dual GN complexes, where a secondary antibody-conjugated GN was added to the sample solution and bound to the immobilized anti-E. coli GNs in the conjugate pad as the sample flowed up the LFA. 34 This technique again improved the sensitivity by 100-fold to 1.13 × 103 CFU/mL.
Another issue most current colorimetric LFAs face is the lack of a quantitative readout. Although some studies have suggested the use of a smartphone to measure the intensities of the test bands, they often required analysis using external computer software 35 or had poor accuracy in detecting minute changes. 36 Recently, Bae and coworkers developed a smartphone app that accurately quantified the color intensities on LFAs for E. coli in ground beef and spinach samples, eliminating the need for a computer and making LFA quantitative analysis more PON-friendly. 37 Another study demonstrated the quantitative detection of E. coli using magnetic nanoparticles (MNPs) conjugated to both invertase and antibodies. 38 When the pathogen-bound MNPs reached the detection zone, the invertase catalyzed the conversion of sucrose into glucose that could be read quantitatively using a personal glucose meter. Because MNPs were used rather than GNs to produce the colorimetric change, this method also improved the sensitivity of the test via IMS capture.
Another type of paper-based device that relies on colorimetric readout is a modified version of an ELISA test, the paper-based ELISA (p-ELISA). In a traditional ELISA, the target pathogen binds to primary antibodies immobilized in a well plate, after which an enzyme-conjugated secondary antibody is added and catalyzes a color change that can be measured using a plate reader. 39 ELISA improves upon many of the issues with LFA detection, as it is quantifiable, has higher sensitivities, and allows for high-throughput testing. However, traditional ELISA is not well suited for PON settings due to its long incubation times, high rates of false positives, and need for expensive lab equipment. Thus, there have been efforts to adapt these tests by immobilizing the antibodies on paper substrates, first demonstrated by the Whitesides group using a 96-microzone array format for the detection of HIV. 40 This method has since been adapted for bacterial detection in beef 41 and cabbage 42 samples. The simplest readout for these devices is a colorimetric visual readout, but quantitative measurements are also possible using a smartphone camera and image processing software.41–43 The assay time has also been decreased from around 8 h in traditional ELISA to only 2.5 h, further improving the accessibility for PON testing. 41
μPADs, a concept first introduced by the Whitesides group, have made “lab-on-a-chip” detection possible on paper substrates. 44 The patterning of microchannels and detection reagents on paper using photolithography and printers allows for the use of smaller volumes and the integration of multiple steps, making all-in-one PON pathogen detection more user-friendly. 45 Furthermore, these paper systems are far cheaper than traditional microfluidics chips and eliminate the need for an external power source since fluid flow is driven by capillary action. Most older colorimetric μPADs have similar sensitivity issues to colorimetric LFAs, and thus require many hours of enrichment culture prior to testing.46,47 IMS techniques have been used to address these issues. One such approach detected for Salmonella typhimurium in milk samples incubates the captured magnetic bead–bacteria complexes with streptavidin-β-galactosidase to drive an enzymatic reaction on the μPAD and produces a colorimetric change. 48 This μPAD was also unique in that it utilized a “chemometer” design, which gave a visual quantitative readout based on the length of color development along the capillary flow path and thus eliminated the need for external computer programs. Another problem that limits widespread use of μPADs is the complexity and cost of fabrication. Current techniques revolve around photolithography, which is expensive and requires the use of clean rooms, making it difficult to perform in PON settings. 49 Wax and ink-jet printing have helped reduce the cost of production, but do not provide good precision or reproducibility due to difficulties with printer configuration and material flow control. The Pushpavanam lab recently developed a novel method of fabrication using a laser printer, which is low cost, is easy to use, and yields high resolution, making it much more suitable for PON use. 50
Fluorometric Detection
Recently, there have been efforts to modify these colorimetric paper devices for detection using fluorescence. Fluorometric detection helps resolve some of the major shortcomings of colorimetric devices, as it allows for improved sensitivity and quantitative readouts. 23 The quantum dot (QD) mechanism has been commonly used to integrate fluorescent readouts on paper-based immunoassays. The Merkoçi group created an LFA system that integrated QDs and graphene oxide (GO), a known photoluminescent quencher. 51 QDs conjugated to E. coli antibodies were printed on the test line, and unlabeled QDs were printed on the control line. In a negative test, the flow of GO quenched the fluorescence on both the control and test lines. However, when E. coli was present in the sample, it bound to the antibodies on the test line and blocked the energy transfer from occurring with GO, producing a fluorescent signal that could be measured using a portable lateral flow reader. For E. coli in milk samples, this approach yielded quantitative detection of the pathogen down to a concentration of 100 CFU/mL, demonstrating a significant improvement in LOD compared with colorimetric LFAs. Two studies fabricated fluorescent magnetic nanospheres by integrating QDs with Fe3O4 MNPs.52,53 These particles allowed for lower LODs due to IMS enrichment of the pathogen as well as simultaneous colorimetric, magnetic, and fluorescent readouts. Although the quantification of the fluorescent and magnetic signals required some specialized equipment, the colorimetric change associated with Fe3O4 binding to the test line made PON-friendly qualitative detection possible as well. When testing for S. typhimurium in milk, the device was able to detect a concentration of 2.5 × 103 CFU/mL qualitatively and 2.39 × 102 CFU/mL quantitatively, 2–4 orders of magnitude lower than the LODs of traditional LFAs. 53
Upconversion nanoparticles (UCNPs) have also been integrated with LFAs for fluorescence-based detection of foodborne pathogens. UCNPs are different from traditional fluorophores in that they are excited by near-infrared light rather than ultraviolet light, which eliminates the hazards associated with UV-based detection, helps reduce background fluorescence, and improves detection sensitivity. 54 Zhou and coworkers used this method to develop a multiplex LFA for the simultaneous detection of 10 different types of foodborne pathogens in 279 different food samples. 55 The device consisted of a disc with 10 detection channels, each with a sandwich immunoassay with a conjugate pad containing the antibody-conjugated UCNPs. The tests yielded high specificity of all target pathogens even in the presence of complex food matrices. Although the test yielded a low LOD of around 10 CFU/0.6 mg, enrichment culture was needed to achieve this. In addition, detection required the use of a specialized reader, so smartphone-based detection must be developed for this device to be fully adaptable for PON use.
Electrochemical Detection
Electrochemical detection offers the advantages of rapid detection with greater sensitivity and specificity than traditional methods, but electrochemical biosensors were previously incompatible with resource-poor testing due to their complicated fabrication processes and expensive materials.56–58 There has been a recent shift toward integration with paper-based devices to create electrochemical paper-based analytical devices (ePADs), which significantly decrease the cost of production of these biosensors and allow for the miniaturization of complex processes to increase portability and ease of use. 23 These devices utilize a three-electrode system containing a reference, counter, and working electrode. 59 The working electrode is typically coated with enzymes or antibodies that catalyze an electron transfer reaction in the presence of the target pathogen, which produces a current correlating with the concentration of the target in the sample. One such study functionalized S. aureus antibodies to carbon nanotubes on the working electrode of their device, which induced an electrical current upon binding to bacterial antigens that could be measured using a potentiometer. 60 Although the actual mechanism behind this increase in current is not yet fully understood, this technique was able to achieve an LOD of 13 CFU/mL for pathogens in a milk sample in under 30 min. Another study tested two different configurations of immobilizing the antibodies on the paper electrodes. 61 The first was a direct conjugation of the anti-Salmonella antibody to the membrane, and the second relied on an intermediate polyamidoamine dendrimer layer that increased the available area for the antibodies to be immobilized. Although this second configuration was designed to increase antibody concentration and improve sensitivity, it was found to have poor reproducibility and was thus unsuitable for testing food samples. The direct-conjugation setup was used to measure the electromotive force (EMF) change that resulted when the target bacteria from apple juice samples bound to the antibodies and achieved an LOD as low as 50 CFU/mL. These recent developments in devices relying on electrochemical detection have demonstrated a significant improvement in sensitivity over the devices discussed in the previous sections.
Microfluidic-Based Detection
Recent efforts have sought to leverage microfluidic devices to develop lab-on-a-chip systems to detect foodborne pathogens. Microfluidic chips offer a number of advantages for PON testing, including storage of reagents, control of fluid flow, and integration of multiple steps of the diagnostic workflow. Furthermore, these integrated steps in the chip workflow also have the potential to be automated, allowing for the detection of foodborne pathogens to be achieved with the push of a single button. Traditional chip fabrication methods include soft lithography and polydimethylsiloxane (PDMS) glass bonding, but clean rooms, complicated SU-8 mold fabrication protocols, and difficult reagent handling create expensive and inefficient limitations for scaling up to mass production of these chips.62,63 Alternative approaches have emerged, including 3D printing, laser jets, print and peel methods, and micromilling.62,63 Although these methods still require machinery, they use technology that is more widely accessible and feasible for large-scale production.62,63 In this section, colorimetric-, fluorescence-, and electrochemical-based detection of surface epitopes of foodborne pathogens with microfluidic devices will be discussed. A summary of these devices is also presented in Table 2 .
Microfluidic-Based Detection of Pathogenic Surface Markers.
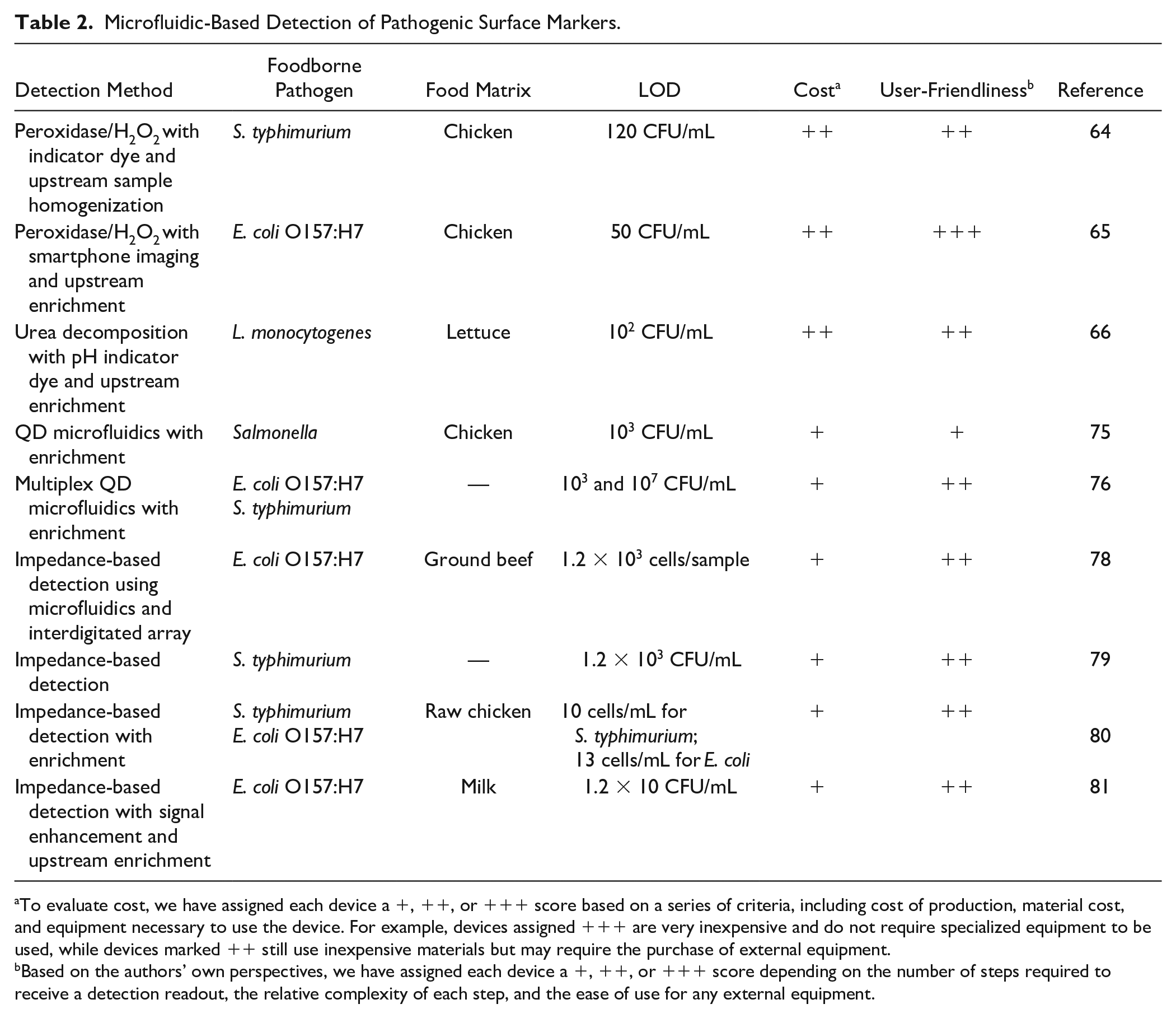
To evaluate cost, we have assigned each device a +, ++, or +++ score based on a series of criteria, including cost of production, material cost, and equipment necessary to use the device. For example, devices assigned +++ are very inexpensive and do not require specialized equipment to be used, while devices marked ++ still use inexpensive materials but may require the purchase of external equipment.
Based on the authors’ own perspectives, we have assigned each device a +, ++, or +++ score depending on the number of steps required to receive a detection readout, the relative complexity of each step, and the ease of use for any external equipment.
Colorimetric detection
Colorimetric-based detection is especially advantageous, as qualitative detection can be performed with the naked eye. One such device combined IMS and enzyme catalysis to isolate, concentrate, and detect S. typhimurium. 64 MNPs isolated and concentrated the pathogen, and polystyrene microspheres were subsequently incubated with the bacteria–nanoparticle conjugate. Two mixing channels intertwined with one another in a looped fashion to allow for the sample to thoroughly combine with H2O2. The decomposition of H2O2, facilitated by the catalase proteins on polystyrene microspheres, produced oxygen gas that forced an indicator dye in a downstream channel to move along a ruler. The distance traveled by the indicator dye was then used to determine the concentration of S. typhimurium in the original sample, as the amount of oxygen gas produced was related to the amount of bacterial present. The reaction mechanism of H2O2 decomposition by catalase-decorated polystyrene microspheres has also been combined with smartphone imaging technology to provide quantitative detection of E. coli. 65 In a study by Zheng and coworkers, a change in color hue was observed due to the aggregation of GNs. If E. coli was absent in the sample, the GNs would aggregate and change the color of the solution from red to blue. If E. coli was present in the sample, the catalase from the MNP–bacteria–polystyrene–catalase complex would decompose the H2O2 in the sample and the solution would remain red. While the change in color hue could be detected with the naked eye, a smartphone app was developed in parallel to quantify the degree of color change.
While the peroxidase/H2O2 mechanism is most commonly used in biosensor assays, the instability of H2O2 makes for a short shelf life. 66 Furthermore, the ubiquity of peroxidases in a variety of cells increases the difficulty of achieving specific detection of a target pathogen. To address this issue, recent efforts have been made to leverage the urease-catalyzed decomposition of urea in such assays. Urease is inexpensive, highly catalytic, and involves nontoxic substrates. IMS was first used to separate and concentrate the pathogen; then antibody- and urease-decorated GNs were conjugated to the MNP–pathogen complex. The sample was introduced to a well plate containing urea and a pH indicator, and as the urease on the GNs decomposed the urea into ammonium carbonate, the pH of the solution increased and a color change was observed. The color change was proportional to the concentration of pathogen present in the well. Although this mechanism has yet to be incorporated into a microfluidic setup, the similarity between this configuration and that of the peroxidase/H2O2 mechanism suggests that it has potential for application in microfluidic foodborne pathogen detection.
Fluorometric Detection
In addition to paper-based detection methods, QDs have been pursued in conjunction with microfluidic IMS to isolate and detect S. typhimurium. 67 QD nanoparticles and polyclonal antibodies sandwiched the Salmonella on the QD surface, and a portable fluorometer was then used to measure the signal produced by the sandwich complex. Using this setup, Salmonella could be detected at 103 CFU/mL in chicken extract and borate buffer. In addition to producing a quantitative readout for detection, fluorometric methods are also advantageous due to their multiplexing capabilities by using different probes with different excitation and emission wavelengths. The Paknikar group explored multiplexing for the detection of a variety of waterborne pathogens using circular microfluidic channels. 68 By integrating copper wire, magnets, and QDs into the microfluidic device, they achieved immunomagnetic concentration and fluorometric detection of both E. coli and S. typhimurium between 103 and 107 CFU/mL in phosphate-buffered saline (PBS). While the pathogens were concentrated and captured within the device, detection was performed using an upright fluorescence microscope. Future work suggests incorporating a portable or compact fluorometer into the workflow, as well as testing samples with greater complexity, allowing for more PON-friendly detection that still contains multiplexing capabilities.
Electrochemical Detection
Within the realm of foodborne pathogen screening, impedance microbiology has grown as an increasingly popular technique. Impedance biosensors have been integrated with microfluidic chips to create PON detection platforms that require minimal power and produce a quantitative readout. 69 Impedance-based detection relies on differences in electrical impedance resulting from different concentrations of bacteria in the solution. When introduced in a microfluidic setup, the pathogen can be captured and concentrated by immobilized antibodies in the fluid channels, and electrodes can then measure the difference in impedance across the length of the channel. For example, the Tung lab incorporated a gold interdigitated array microelectrode (IDAM) and MNPs into a microfluidic setup that achieved an LOD of 1.2 × 103 E. coli O157:H7 cells in ground beef in 35 min. 70 The change in impedance was correlated to the concentration of E. coli present on the IDAM surface within the microfluidic chip. This work is especially noteworthy because the authors did not use any upstream sample enrichment prior to introducing the sample to the microfluidic chip, as the MNPs were able to sufficiently isolate and concentrate the target pathogen. Almasri and coworkers compared a similar impedance setup to a nonmicrofluidic biosensor and found that the microfluidic setup resulted in a 10-fold improvement in the LOD, as microfluidic integration reduced the risk of sample loss due to multiple liquid handling steps. 71 Almasri and coworkers also achieved LOD values of 10 cells/mL of S. typhimurium and E. coli O157:H7 in raw chicken product in less than 1 h using a glass substrate etched via micromachining. 72 However, the manufacturing of such devices involved multiple complex steps inside a clean room, and the impedance analyzer used to quantify the impedance changes within the device was not portable and required a power source.70–72 While the sample processing step of the workflow was greatly simplified and pathogen concentration and detection were fully integrated, the diagnostic device still requires simplified fabrication and the use of more portable equipment to be considered PON.
In addition to using the pathogens themselves to change the impedance of the solution in the microfluidic channels, additional signal enhancement can be performed by introducing ions into the solution. The Lin research group successfully advanced the aforementioned microfluidic impedance-based detection method by introducing GNPs decorated with urease and anti-E. coli antibodies. 73 The urease on the GNPs catalyzed the hydrolysis of urea produced by the bacteria into ammonium carbonate, which decreased the impedance within the device even further. With this setup, the LOD was found to be 12 CFU/mL with a time to detection of 2 h. 66
Detection of Nucleic Acids
In an effort to replace traditional culture techniques to grow detectable amounts of foodborne pathogens, there has been a movement toward nucleic acid-based detection methods. These methods include amplification-based techniques, in which specific nucleic acid sequences within the genome of the target are amplified to detectable levels, as well as oligonucleotide probes, in which synthetic sequences of nucleic acids are hybridized with the target sequence. While this paper will mainly focus on amplification-based techniques that are highly suitable for PON settings, several oligonucleotide probing methods will be briefly mentioned here. Fluorescence in situ hybridization (FISH) has seen a dramatic increase in use in recent years for foodborne pathogen screening. Initially developed by the Pace and Stahl laboratories, FISH involves fluorescent oligonucleotide probes that target ribosomal RNA (rRNA) within intact pathogen cells.74–76 In contrast to FISH, where hybridization and fluorescence can be observed in whole cells, microarrays have also emerged as a popular detection technique. These microarrays contain fluorescent oligonucleotide probes immobilized onto a glass slide, allowing the target sequence to hybridize with the probe and be sequestered onto the slide for imaging.77,78 In addition, molecular beacons are unique, hairpin-structured probes that are labeled with a fluorescent reporter dye at one end and a quencher at another; upon binding the target, the quencher separates from the probe and the hybridized target fluoresces.79,80 Although many versions of oligonucleotide probes are single-stranded, double-stranded probes do exist, in which the quencher and fluorophore exist on opposite strands. As the target hybridizes with the fluorophore-labeled strand in a thermodynamically driven fashion, the quencher strand separates from the fluorophore strand and fluorescence can be observed. 81 These oligonucleotide probing mechanisms have become increasingly popular in the past decade. However, multiple liquid handling and washing steps currently limit their applicability at the PON.
In the field of amplification-based detection methods, PCR has emerged as the new gold standard. While PCR is robust, rapid, and specific, it requires the use of expensive, power-consuming thermocyclers to denature, anneal, and elongate the DNA. 82 The use of thermocyclers and the need for specialized laboratory personnel make PCR unsuitable for use outside the laboratory, especially in resource-poor settings. Furthermore, additional laboratory equipment and trained technicians are required to perform upstream sample processing steps to lyse the pathogen and purify the genetic material for amplification. For these reasons, the use of PCR outside the traditional laboratory has been replaced by isothermal nucleic acid amplification tests (NAATs). Because thermocyclers are not necessary for isothermal techniques, both the timescale and cost of amplification is reduced.
Among isothermal NAATs, loop-mediated isothermal amplification (LAMP) has become the most widely researched and applied isothermal amplification technique since its development in 2000 by Hase and coworkers. 83 LAMP is characterized by its use of four to six primers to generate hairpin loops that recognize six to eight different regions on the target gene, and it generally operates at temperatures between 60 and 65 °C. Depending on the subsequent detection method, this amplification scheme may be modified to incorporate a labeled probe.84,85 This amplification scheme is highly efficient, capable of amplifying a few copies of genetic material to 109 copies within 1 h. Furthermore, additional primers increase the specificity of this technique. Among isothermal amplification techniques, 78% of recent publications for food-based analysis are LAMP-based. 86 Another widely used technique is recombinase polymerase amplification (RPA), developed by Piepenburg and researchers. Instead of relying on heat-based denaturation, RPA leverages recombinase proteins to unravel the DNA and provide space for polymerization, allowing the oligonucleotide primers to find the homologous sequence and bind to the strands. Operating between 37 and 42 °C, RPA is especially advantageous because it does not require significant heat, 82 and it has even been shown to amplify at room temperature. 87 The last isothermal amplification technique we will discuss in this review is thermophilic helicase-dependent amplification (tHDA), which operates at an optimal temperature of 65 °C. 88 In this mechanism, the protein helicase unwinds and unravels the duplex to allow for the insertion of the amplification machinery, including oligonucleotide primers and, depending on the detection method, a labeled probe.89,90 While LAMP, RPA, and tHDA are the most commonly used isothermal mechanisms in the foodborne pathogen NAAT field, other isothermal techniques exist, including rolling circle amplification (RCA), nicking enzyme amplification reaction (NEAR), strand displacement amplification (SDA), transcription-mediated amplification, self-sustained sequence replication (3SR), and nucleic acid sequence-based amplification (NASBA).91–93 As such, the following section will mainly focus on devices that utilize LAMP, RPA, and tHDA as their amplification techniques. A summary of such devices is also provided in Table 3 .
Detection of Pathogenic Nucleic Acids via LAMP, RPA, and tHDA Amplification Methods.
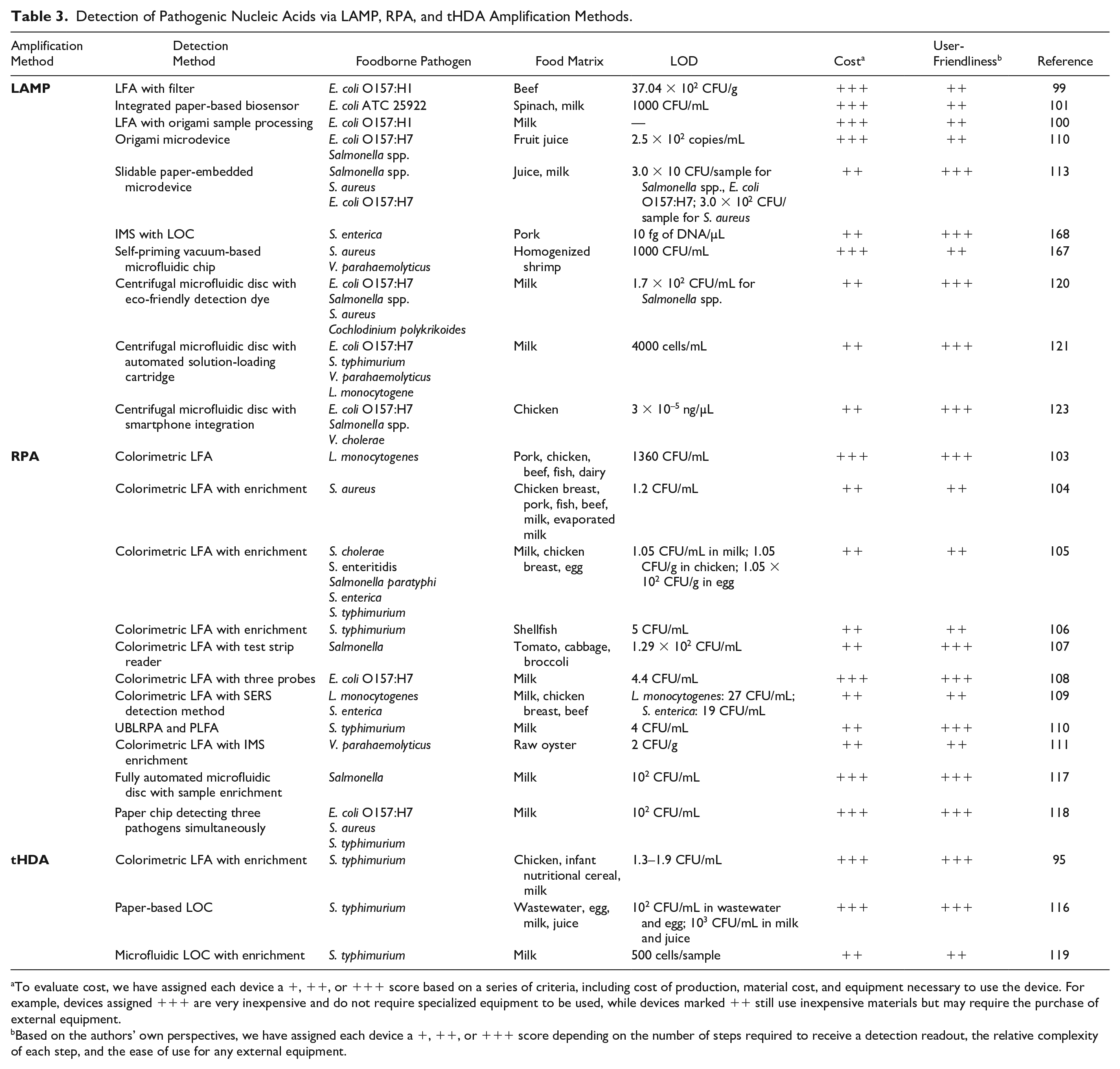
To evaluate cost, we have assigned each device a +, ++, or +++ score based on a series of criteria, including cost of production, material cost, and equipment necessary to use the device. For example, devices assigned +++ are very inexpensive and do not require specialized equipment to be used, while devices marked ++ still use inexpensive materials but may require the purchase of external equipment.
Based on the authors’ own perspectives, we have assigned each device a +, ++, or +++ score depending on the number of steps required to receive a detection readout, the relative complexity of each step, and the ease of use for any external equipment.
Although isothermal amplification techniques have improved upon the NAAT and increased its feasibility for PON settings, each technique poses its own set of limitations. For example, LAMP requires multiple primers, which introduces additional liquid handling steps that must be performed by the user. This also further complicates primer design to avoid dimerization and hairpin structures. Additionally, both tHDA and LAMP operate at relatively high temperatures, requiring a more powerful heat source to maintain the constant temperature. Finally, RPA and tHDA are prone to developing primer artifacts and easily succumb to contamination. 94 Since most PON settings are not sterile environments and lack experienced lab technicians, improvements must be made to these amplification strategies in order to achieve successful implementation in resource-poor settings. These issues, along with sample processing challenges, limit the widespread application of these tests. However, significant strides have been made to develop isothermal NAAT devices suitable for PON use and will be further discussed in the following sections.
Paper-Based Detection Methods
Paper-based diagnostics have received much attention as a viable substitution for quantitative PCR, which is currently considered the gold standard for detection in the NAAT workflow. These devices are inexpensive, can store reagents, and can transport fluid via capillary action, removing the need for pumps. Devices such as the LFA, paper-based lab-on-a-chip devices, and origami-inspired paper folding devices have all been combined with isothermal amplification techniques to create a PON NAAT platform. We will discuss these devices in detail in the following sections.
Lateral Flow Immunoassay
As the most widely used paper-based device, the LFA can provide a qualitative colorimetric readout that can be perceived by the naked eye. In addition to visual readout, imaging technology can also be applied in conjunction with the LFA to quantify the intensity of the test line. One of the most common versions of the LFA involves sandwiching the target biomarker between two antibody complexes on the test line. These sandwich LFAs can be utilized to capture dual-labeled amplicons on the test line, where during the amplification cycle each end of the amplicon is tagged with a different label. Antibodies targeting these labels can then sandwich the amplicon on the test line. This method highlights the universality of the LFA format, as this technique can also be used to capture surface markers of pathogens as discussed previously. As long as the amplicon is labeled at both the 5′ and 3′ ends, it can be sandwiched by antibodies on the GNs and antibodies on the test line. Labeling the amplicon is important, as antibodies cannot sandwich DNA alone. This extra labeling step occurs concurrently with amplification and is not required for surface marker detection via the LFA.
Several LFAs compatible with LAMP have been developed to allow for the detection of foodborne pathogens in various food matrices, including meats, produce, milk, and seafood.95–98 A number of studies have attempted to integrate the LFA with more PON sample processing steps.99,100 For example, the Kim group utilized a cellulose nitrate filter to concentrate and isolate E. coli in beef samples before downstream LAMP and LFA detection. An LOD of 10 CFU/g was demonstrated, a noteworthy 100-fold improvement over LODs determined in unfiltered samples. 99 While the group was able to apply a PON method for sample processing, a Stomacher was still required to remove pathogens from the filter before subsequent detection. To further move toward integration of the NAAT workflow, Luo and coworkers developed a multilayered paper-based biosensor that integrated nucleic acid extraction, amplification, and LFA detection. 101 The different layers contained zones for each step of the NAAT workflow and were separated by hydrophobic layers to allow for controlled fluid flow between each layer. The researchers also developed a handheld battery-powered heating device, eliminating the need for power-intensive heaters. Smartphone imaging was performed to allow the results to be quantified. Utilizing a similar setup, the Xu lab was able to achieve an LOD of 1000 CFU/mL in spinach and 10 CFU/mL in milk within 70 min. While the biosensor integrated all of NAAT steps into one device, the setup still required multiple user handling steps, which increases the risk of contamination and requires the test to be performed by trained personnel.
In addition to LAMP, RPA has also been integrated with the LFA for colorimetric readout after isothermal amplification. Combined with RPA, the LFA has been shown to be highly sensitive, with LODs ranging as low as 1.05 CFU/mL with enrichment of Salmonella in milk to 1360 CFU/mL without enrichment of L. monocytogenes in generalized food samples.102–106 However, enrichment times vary between the types of sample and pathogen tested. Improvements upon the traditional RPA + LFA setup include work by the Wang research group, which applied surface-enhanced Raman scattering (SERS) to an LFA to simultaneously detect L. monocytogenes and Salmonella enteritidis serotype Enteritidis following RPA. 107 SERS enhances the Raman scattering of molecules adsorbed to the structures. Analysis of the Raman scattering through spectroscopy allows for highly sensitive and quantitative analysis of the bacteria. After amplification, the product was purified and applied to the SERS-LFA. Concentrations as low as 27 CFU/mL for S. enteritidis and 19 CFU/mL for L. monocytogenes were detected in various food samples. While the technique above is highly specific and allows for quantitative analysis, there are a few drawbacks that prevent application in resource-poor settings. Primarily, the analysis of the Raman signals required specialized equipment, a nonportable confocal micro-Raman spectroscopic system. Additionally, the process to develop the specialized GNs was time-consuming and multiple purification steps were needed to prevent nonspecific binding.
To improve the sensitivity of the LFA, Luo and coworkers utilized universal blocking linker recombinase polymerase amplification (UBLRPA) to incorporate a peptide nucleic acid lateral flow device (PLFD) to facilitate stronger binding. 108 UBLRPA uses the same amplification mechanism as RPA, with the exception that ssDNA is produced instead of dsDNA. The LOD for the UBLRPA-PLFD device for S. typhimurium bacteria in milk was 4 CFU/mL, which was a significant improvement compared with other PON NAAT devices that detect pathogens in complex food matrices. The Wang lab further explored sensitivity improvements by using IMS to capture and isolate Vibrio parahaemolyticus in contaminated raw oyster samples. 109 IMS decreased the LOD to 2 CFU/g after 4 h of enrichment from 2 × 106 CFU/g in the oyster samples. However, the increased sensitivity of RPA-LFA compromises assay time; though it exhibits lower time to detection than traditional methods like PCR, this hybrid technique is much less rapid than RPA or LFA alone.
Finally, tests combining upstream tHDA and downstream LFA detection have been developed and have detected pathogens in food samples ranging from milk to meat products. For example, Wang and coworkers developed a workflow with tHDA and an encased LFA that resulted in LODs as low as 1.3–1.9 CFU/mL in contaminated chicken products with a time to detection of less than 90 min. 94 However, although the LFA and tHDA techniques employed made these NAAT tests more PON, the protocols that led to these low LODs still relied on upstream sample enrichment and purification steps. Furthermore, icing steps were required immediately after amplification to minimize the production of primer artifacts. Therefore, steps must be taken to develop sample processing methods that can be performed at the PON setting and do not depend on laboratory technology or trained personnel to perform. Primer design must also be performed carefully to minimize the formation of hairpin structures and dimers, which both removes primers available for amplification and increases the risk of false positives.
Origami-Based Devices
Origami-based devices present another promising paper-based platform that has been utilized for microbiological testing of food.110,111 Origami is the traditional Japanese art of paper folding, which has been utilized for several centuries to construct 3D geometries using a flat piece of paper. 112 These foldable devices can be fabricated on a single sheet of paper and assembled via simple folding, which reduces production costs and complexity. Paper folding can occur directly after fabrication or during the detection scheme, which allows for control of the timing between sequential steps of the detection workflow. Furthermore, these devices allow for the integration of multiple NAAT steps into a single platform, reducing the amount of storage space and number of liquid handling steps required for detection workflows.
Lee and coworkers developed a foldable microdevice composed of a thin polycarbonate film that contained multiple reaction and detection zones to allow for multiplex detection. 110 To improve the PON applicability of their device, they developed a thin graphene-based heater that was powered with a low-voltage, handheld power bank. The device was able to successfully detect Salmonella and E. coli O157:H7 in spiked juice samples after a separate thermal lysis and purification step utilizing polydopamine-coated paper. Furthermore, a notable LOD of 2.5 × 102 copies/mL was achieved when detecting E. coli O157:H7 in juice samples. While this device demonstrates promise for use in PON settings, some modifications must be considered. The colorimetric detection method required a UV irradiation step prior to visualization with the naked eye, which required additional user steps and the use of a hazardous UV light source. Furthermore, integration of sample lysis and purification steps with the foldable platform would be necessary to reduce the number of liquid handling steps required.
Paper-Based LOC
To integrate the entire NAAT workflow into one single platform, as well as leverage the capillary action of paper-based devices, paper LOC devices have been developed to detect foodborne pathogens using isothermal amplification. For example, Lee and coworkers developed a slidable, multilayered microdevice that integrated DNA extraction, purification, LAMP-based amplification, and colorimetric detection. 113 A handle allowed for the middle layer of the device, which contained paper discs containing reagents required for amplification and detection, to slide in a stepwise fashion. To perform each step, deionized water was utilized to rehydrate the paper discs containing reagents. This aqueous solution with rehydrated components then ended up in the bottom layer of the device, where the sample was located. Through this setup, the authors were able to successfully screen for Salmonella, S. aureus, and E. coli O157:H7 in artificially spiked juice and milk samples within 75 min. Additionally, the authors were able to demonstrate an LOD of 30 CFU/sample for Salmonella spp. and E. coli O157:H7 and 3.0 × 102 CFU/sample for S. aureus.
In addition to LAMP LOC devices, tHDA has also been leveraged in this paper-based platform. The Tang research group developed a particularly noteworthy device that contained paper-based modules for DNA extraction, amplification, and detection. 114 Using sponges to store lysis and washing buffers, the sample was introduced to the sample pad, and the user pressed a button to release the buffers from their respective sponge reservoirs. The lysate was filtered through paper and the isolated DNA was then transferred to amplification and detection zones by moving the paper on a copper strip between zones. Detection of S. typhimurium was achieved in less than an hour with LODs of 102 CFU/mL in egg and 103 CFU/mL in milk and juice. Although some simple user manipulation was required to move the sample through the extraction, amplification, and detection modules within the device, the only liquid handling step involved was the initial sample introduction step. This fully integrated device shows promise not only that the NAAT workflow can be concentrated into a single device, but also that these devices have the potential to be automated into single-step, battery-operated diagnostic tests.
Chip-Based Detection Methods
Microfluidic chip-based methods of detection are another promising field of study for PON testing of food pathogens due to their ability to integrate cell lysis, DNA extraction, amplification, and detection into a single, portable device. In a wireless laser diode-controlled device developed by the Cho laboratory, laser-based cell lysis and RPA were performed in a polycarbonate disc. 115 The sample was detected qualitatively via colorimetric readout on an embedded sandwich lateral flow device. The increased versatility and portability of the single-step chip was attributed to the simpler system using the laser diodes and localized heating. The automation of many steps used to detect food pathogens achieved an LOD of 102 CFU/mL in milk. Another chip incorporating RPA developed by the Kim group allowed for the simultaneous testing of up to three pathogens with a single sample introduction step. 116 Lateral flow through the asymmetric membrane allowed the sample to reach each corner of the chip containing dehydrated RPA reagents. Primers specific to each pathogen were segregated in the different regions to decrease the chance of nonspecific bacterial amplification. The pathogens were then detected via fluorescent readout using a nonportable fluorometer. The LODs of E. coli, S. aureus, and S. typhimurium with this method were found to be 102 CFU/mL. While this chip made great strides toward fully PON RPA-based diagnostics, complex food samples cannot be injected into the chip as they interfere with the fluorescence. Further research will need to be applied to increase the user-friendliness of the device as sample introduction via injection could potentially be dangerous for untrained personnel due to the use of needles. The fluorescence also needs to be measured with a fluorometric reader, increasing potential up-front costs for the device.
Microfluidic devices have also been employed in conjunction with tHDA in an effort to integrate the NAAT workflow into a single device. Tsougeni and coworkers developed three microfluidic platforms for cell capture and lysis, DNA purification, and DNA amplification, respectively. 117 Cell capture and the lysis device were designed with a high surface area, allowing for nearly 100% capture efficiency by antibodies immobilized within the channels. Amplification and detection were performed on a separate chip, where a liquid tHDA reaction was introduced to the inlet and amplification took place as the fluid flowed through the microfluidic channels. The group successfully screened for S. typhimurium spiked in milk at levels below 500 cells/sample, making the device a promising alternative to the current gold standard NAAT. However, several modifications must be considered in order to make the device more suitable for PON settings. For example, multiple liquid handling steps need to take place to move the sample from module to module. Furthermore, the device relied on gel electrophoresis as a detection output. Finally, liquid tHDA reagents were mixed with the sample obtained from the purification module, prior to addition to the amplification module. Dehydration of tHDA reagents into the inlet of the amplification module would increase the suitability of this device for PON settings by removing unnecessary liquid handling steps and supporting a more continuous workflow.
Centrifugal Microfluidics
Centrifugal microfluidics, or lab on a disc, presents a promising platform for PON diagnostics. These devices consist of embedding complex assays in fluidic networks on a disc. A single, compact motor is used to generate continuous centrifugal force to transport fluid unlike traditional microfluidics, which generally require bulky external instrumentation such as external pumps. 118 Using different spin profiles, this platform allows for the automation and integration of multiple microfluidic fluidic processes, such as mixing reagents, valving, or metering samples, into one device. 119 Additionally, multiple assays can be simultaneously performed on the same disc. 118 This section will discuss some of the current progress in the development of centrifugal microfluidic devices for PON foodborne pathogen screening.
A number of studies have utilized the centrifugal platform to develop automated devices for PON microbiological testing in food samples.120–126 For example, Thong and coworkers developed a centrifugal device that employed multiple microfluidic operations to integrate amplification and fluorometric detection into one device. 123 The device contained multiple units that allowed 30 reactions to be performed simultaneously. Each unit consisted of loading chambers to load the LAMP assay (including fluorescent dye) and primers separately, channels and chambers to mix the LAMP assay, metering chambers to aliquot the LAMP assay into equal volumes, and an amplification chamber that contained the loaded sample. Sealing materials were loaded into reaction chambers to enclose the amplification chamber to prevent evaporation during the reaction before subsequent detection was performed. To create a simple endpoint analysis system and improve portability, the researchers created a chamber that housed important electronics such as the stepper motor and microcontroller for fluid transport as well as a UV emitter and photodiode for detection. A smartphone connected via Bluetooth was utilized to control microfluidic operations and analyze results. Utilizing this setup, the authors were able to achieve an LOD of 3 × 10–5 ng/µL DNA to detect for samples containing E. coli, Salmonella spp., and Vibrio cholera spiked with chicken DNA within 60 min. However, the use of fluorometric indicators has the limitation of requiring an extra UV illuminator for detection, making it less suitable for PON settings. In another example, Oh and Seo developed a centrifugal device that integrated microbead-assisted DNA extraction, purification, amplification, and colorimetric detection into a single device to detect pathogens within 65 min. 121 Samples containing pathogens could be visualized with the naked eye by observing a color change from purple to sky blue of the solution in the reaction chamber. Additionally, the authors employed a 3D-printed solution-loading cartridge to preload sample, washing solution, elution solution, and a LAMP cocktail, therefore eliminating the need for laborious sample loading steps. Utilizing this device, the researchers were able to achieve multiplex detection of E. coli O157:H7, S. typhimurium, V. parahaemolyticus, and L. monocytogenes in milk samples within 65 min with an LOD of 4 × 103 cells/mL. Further iterations of the device improved on the design by integrating a super absorbent polymer into the waste chamber so that large sample sizes up to 1 mL could be tested. 124 Furthermore, to allow for real-time analysis, an LED UV-Vis detector, a safer alternative to conventional fluorometers, was integrated into a portable chamber that also contained the motor for centrifugal control and heaters for the reaction.
Detection of Metabolites
In addition to targeting unique pathogen surface markers and nucleic acid sequences, food screening methods have also exploited bacterial metabolic pathways. Enzymatic metabolic assays for microbe detection, identification, and quantification are considered standard procedure in the laboratory following culture enrichment and typically utilize chromogenic media substrates for spectrophotometric analysis.127–129 Other metabolic targets of common food screening techniques include adenosine triphosphate (ATP) and quorum-sensing molecules (QSMs). However, gold standard techniques for metabolic detection face the same drawbacks as surface marker-based and nucleic acid-based detection methods, including extensive enrichment steps, sophisticated lab-based equipment, and complex operation.
Emerging detection methods commonly aim to improve the usability of these metabolic-based techniques in resource-poor settings by integration in inexpensive, portable devices requiring minimal user intervention. Unfortunately, due to the universality of bacterial enzymes and metabolites, metabolic-based detection methods face inherent limitations involving low specificity. Techniques involving optimized enrichment procedures or capture antibodies have been utilized to enhance the selectivity of metabolic tests; however, many challenges still exist in accurate species-specific identification and quantification. In this section, we cover the development of PON techniques that rely on the detection of (1) enzymatic activity of the target species, (2) ATP, and (3) QSMs that aim to address the aforementioned limitations for more accurate on-site foodborne pathogen detection. Key metabolic-based devices are summarized in Table 4 .
Methods of Targeting Metabolic Products to Detect Pathogenic Enzymes, ATP, and QSMs.
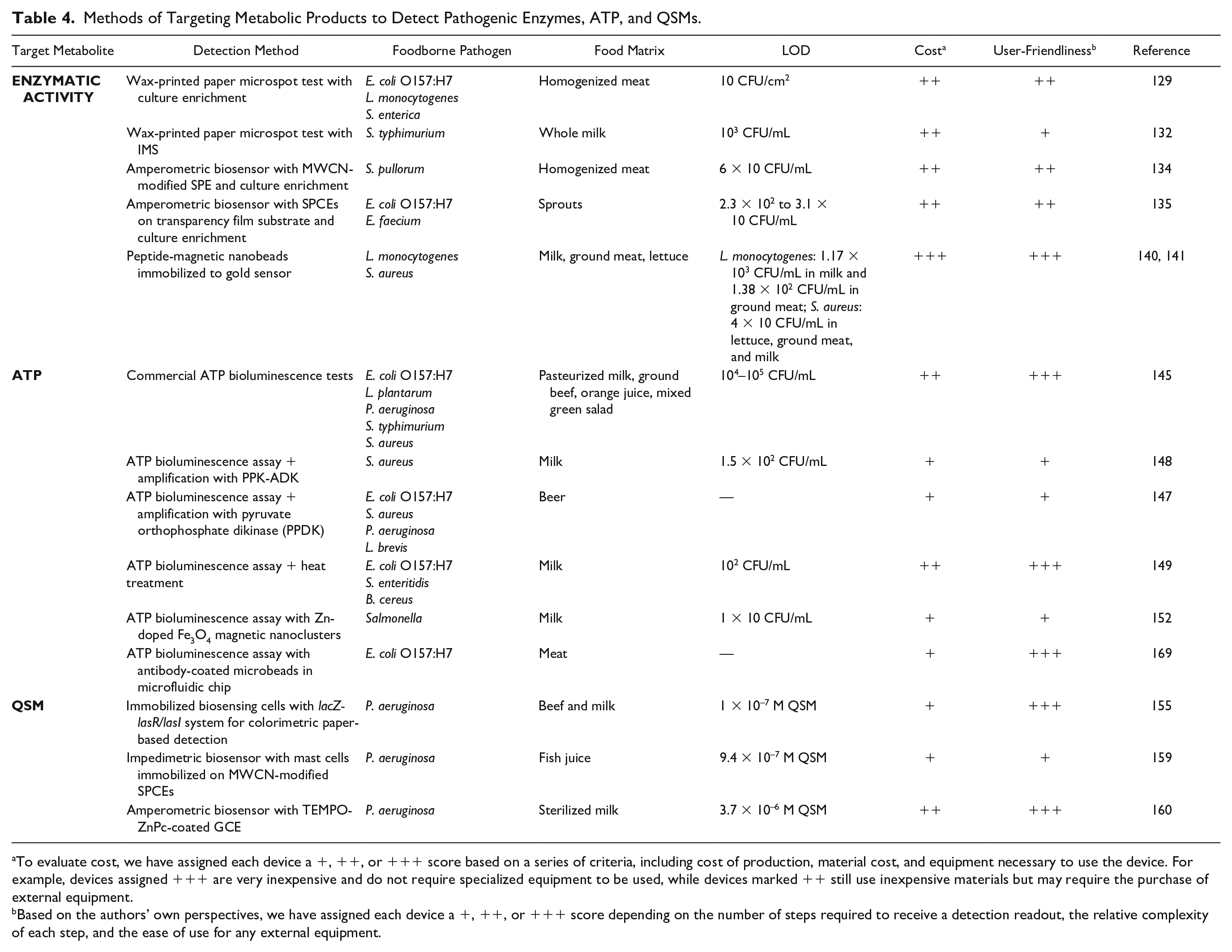
To evaluate cost, we have assigned each device a +, ++, or +++ score based on a series of criteria, including cost of production, material cost, and equipment necessary to use the device. For example, devices assigned +++ are very inexpensive and do not require specialized equipment to be used, while devices marked ++ still use inexpensive materials but may require the purchase of external equipment.
Based on the authors’ own perspectives, we have assigned each device a +, ++, or +++ score depending on the number of steps required to receive a detection readout, the relative complexity of each step, and the ease of use for any external equipment.
Detection of Enzymatic Activity
Species-specific enzyme metabolism has been measured as an indication of bacterial concentration in food samples using paper-based colorimetric detection and electrochemical detection. Most commonly, colorimetric assays detect metabolic activity by measuring the color change occurring due to the chemical reaction between a species-specific enzyme and a chromogenic substrate. The use of chromogenic substrates for colorimetric detection of enzymatic activity in food samples is not a novel concept; however, its integration in µPADs is a relatively new development that has increased the viability of on-site food screening.
For example, a wax-printed paper microspot test was developed by the Henry group for the detection of E. coli, L. monocytogenes, and Salmonella enterica in ready-to-eat meat samples through use of the three species-specific enzyme–substrate pairs. 127 Due to the scaling down of culture enrichment volumes, a more rapid detection time ranging from 8 to 12 h was achieved for the three species. This is a significant improvement upon the 2–3 days required for traditional culturing and metabolic test protocols. A recent study by the same group further improved upon time to detection by pioneering the integration of IMS with a µPAD for the detection of S. typhimurium in milk samples. 130 In the presence of β-galactosidase, chlorophenol red β-galactopyranoside (CPRG) substrate in the paper well array exhibited a color change detectable by the naked eye. When compared with culturing methods, IMS allowed for a drastic decrease in the overall time for enrichment and detection to 90 min.
Unfortunately, due to their dependence on visual inspection, colorimetric assays are subject to external factors affecting device accuracy such as environmental lighting and human bias.131,132 Evaporation of the substrate over time also results in progressive signal degradation and variability of test results based on environmental temperature and buffer pH. 132 As discussed before, novel ePADs utilize handheld potentiostats with electrodes attached to electrochemically active paper or film substrates and have increasingly been employed for the detection of bacterial enzymes to address the sensitivity issues of colorimetric assays. 131
Chai and coworkers developed a portable amperometric biosensor connected to a screen-printed electrode (SPE) modified with multiwall carbon nanotubes (MWCNs) conjugated to chitosan peroxidase that monitored H2O2 consumption catalyzed by Salmonella pollorum-specific catalase. 133 Increased sensitivity for the target bacterium was achieved through use of anti-Salmonella polyclonal antibodies. Detection in homogenized chicken samples occurred in 2 h with an LOD of 60 CFU/mL with enrichment. While demonstrating relatively high sensitivity and rapid detection, the test is still prone to false positives due to the low specificity of polyclonal antibodies. Furthermore, although the reaction rate is drastically increased in the presence of bacterial catalase, H2O2 degradation can occur spontaneously and lead to inconsistent test results. 134 The Henry group later reported the use of electrochemically active and chromogenic substrates for dual electrochemical and colorimetric detection of enzyme activity in sprout samples. 132 In this study, β-galactosidase, β-glucuronidase, and β-glucosidase activity associated with E. coli and Enterococcus spp. were monitored with the detection of oxidized o-nitrophenol (ONP) and p-nitrophenol (PNP) in paper-based wells and transparency film-based electrochemical cells. While both methods had similar detection times of 4 h, including enrichment, electrochemical detection resulted in a low LOD comparable to that of a conventional plate reader.
Many challenges of enzyme detection involve the low sensitivity of tests stemming from difficulties in detecting intracellular enzyme activity. Upstream sample processing and purification steps are oftentimes employed to extract and isolate bacterial enzymes prior to detection in order to achieve clinically relevant sensitivity. In addition, bacterial cell damage during food processing can alter cellular metabolism and inhibit enzymatic activity; thus, detection following these procedures has a higher likelihood of false negatives or inadequate quantitative results. 135 Second, enzyme-based detection lacks high specificity due to the confounding production of target enzymes by nontarget strains. For example, while the enzyme–substrate pair β-galactosidase–CRPG is commonly used for the detection of E. coli O157:H7, many other nonpathogenic strains of E. coli have also been shown to produce varying amounts of β-galactosidase. 136 Inhibition of competitive nontarget detection and reduction of false positives can be accomplished with the use of target-specific antibodies during detection steps, such as in the study by the Chai group. While increasing selectivity, the integration of antibodies simultaneously increases cost and decreases assay stability. Selective preenrichment with culturing or IMS also increases the specificity of enzymatic assays, yet further motivates the nonideal sample processing requirement.
Novel technologies must be developed to overcome this critical limitation by offering in-device sample processing or improved sensitivity without the need for such techniques. Inducers have been commonly proposed to increase enzyme expression as a simple technique for improving LODs.
137
For example, isopropyl-β-
Detection of ATP
Although a less common method, the detection of ATP has also been utilized in food safety analysis. ATP is an energy carrier present in all living cells for the regulation and maintenance of cellular metabolic processes such as biosynthesis, energy storage, and transport mechanisms. Upon cell death, ATP synthesis is immediately arrested and the remaining intracellular ATP is hydrolyzed within minutes. 140 Intracellular ATP content thus serves as a promising indication of the number of viable cells in a sample. ATP bioluminescence assays are most frequently utilized for their relatively high sensitivity, specificity, and detection speed when compared with other ATP-based methods. In the presence of firefly luciferase, intracellular ATP released from bacterial cells reacts with luciferin and oxygen to generate a bioluminescence signal measurable with a luminometer. The intensity of bioluminescence is proportional to the ATP content, which in turn has a linear relationship with the total bacterial concentration.140–142 ATP bioluminescence assays are particularly advantageous for their ability to produce on-site food screening results within 5–20 min, including sample processing steps. 140
Commercial ATP monitoring systems such as the Hygiena SystemSURE Plus + UltraSnap/Supersnap, Neogen AccuPoint, 3M UniLite NG + CleanTrace and Biocontrol MVP Lighting + Surface Test, and Charm Science Novalum + Pocketswab Plus are battery-powered devices that have been utilized for the on-site detection of foodborne pathogens. 143 These devices include disposable swab tips with premoistened ATP extractant for sampling and ATP collection. In the Charm and 3M devices, the sample is mixed and reacted with luciferin–luciferase reagents by puncturing the swab tip through foil seals. The Hygiena, Neogen, and Biocontrol devices do not require foil puncturing. The bioluminescence signal is subsequently detected by either a photodiode or photomultiplier tube in a handheld luminometer. Rapid detection readout is available within 5–20 s after insertion of the swabbed sample. 144 While offering convenience and speed, these ATP tests are insufficient at detecting clinically relevant concentrations due to high LODs ranging from 104 to 105 CFU/mL.
A number of studies have attempted to address sensitivity issues in bioluminescence assays by incorporating enzymes that catalyze the conversion of adenosine diphosphate (ADP) or adenosine monophosphate (AMP) to ATP.145–148 For example, Kuroda and coworkers utilized the fusion protein adenylate kinase–polyphosphate kinase (PPK-ADK) for ATP signal amplification by converting AMP and ATP to two molecules of ADP, and then converting ADP back to ATP. This method achieved a significant improvement in assay sensitivity to detect 1.5 × 102 CFU/mL in milk samples. 147 However, the long incubation times and numerous purification steps involved in preparing the enzyme-based enhancement method limit PON feasibility. The Park lab developed an alternative signal enhancement protocol involving simple heat treatment. 146 Above an organism’s optimal growth temperature, the rate of ATP synthesis increases until a maximum temperature is reached, at which cells die and ATP production ceases. Accordingly, this study found that sample incubation at a moderately elevated temperature of 60 °C for 10 min resulted in a decrease in the LOD in milk samples to 102 CFU/mL, an order of magnitude lower than the conventional ATP bioluminescence assay. Unfortunately, heat-based signal enhancement methods are only effective for mesophilic and thermophilic bacteria and would result in a severe loss of signal and false-negative results if psychrotrophs were present in the food sample.
Currently, many emerging PON tests and commercial ATP detection kits measure total ATP content in the sample for simple and rapid detection. As a result of the ubiquity of ATP, ATP-based testing is oftentimes nonspecific and unable to discriminate between bacterial species in mixed samples. Furthermore, ATP-based tests are relatively inaccurate due to the wide range of intracellular ATP content in different bacterial cells, which varies by 4–5 orders of magnitude. 149 Higher test accuracy, sensitivity, and selectivity of target species have been accomplished by utilizing enrichment steps involving selective nutrient broth and IMS prior to detection. For example, the Jeon research group developed a system using anti-Salmonella functionalized Zn-doped Fe3O4 magnetic nanoclusters for bacterial concentration before assessment with a commercial ATP kit and portable luminometer. 150 This study achieved a noteworthy LOD of 10 CFU/mL for Salmonella in milk samples.
In addition, a number of groups have incorporated microbead-based IMS with ATP detection within a microfluidic chip for enhanced bacterial preconcentration.151,152 Zhang and coworkers incorporated microbead-based IMS with ATP detection within a microfluidic chip for enhanced antigen capture. 155 Anti-E. coli O157:H7 antibodies were immobilized onto the glass microbeads, which were placed in the concentration zone of a microfluidic chip. Assay reagent was injected into the inlet of the chip and flowed over the E. coli caught by the microbeads. The luminescent signal produced by the assay was then measured by a photomultiplier tube placed at the outlet reservoir of the chip. Pathogen capture efficiency was calculated to be 75.49% ± 15.48% for E. coli-spiked meat. However, the LOD was only reported for E. coli samples suspended in PBS with a value of 3.2 × 10 CFU/mL within 20 min. While the capture efficiency of E. coli spiked in meat was extremely similar between the microfluidic chip and traditional culture methods, more work needs to be done in reporting an LOD for complex samples using the microbead-in-a-chip detection platform.
Even with improved species-specific testing, however, significant variability between tests can still occur due to the dependence of ATP content on the cellular metabolic state. Variable factors such as bacterial phase of growth, nutrient availability, and external environmental conditions in food samples influence ATP production and can result in the unreliable analysis of cell concentration. 149 On this note, damage to the bacterial cell during sample processing procedures can alter metabolic state and ATP production rates. In addition, the presence of nonbacterial ATP can amplify bioluminescence intensity and overquantify bacterial concentration. This is a particularly significant issue in fresh produce such as meat and vegetables that contain high levels of somatic ATP. Nontarget ATP detection can typically be prevented by sample pretreatment consisting of selective lysis of somatic cells followed by filtration for the removal of extracellular ATP. However, the time-consuming and laborious nature of these procedures limits their use in commercial products and justifies their exclusion from PON tests. Therefore, while current PON tests are attractive alternatives to conventional methods for their rapidity, convenience, and on-site applicability, they primarily offer general indications of bacterial presence rather than reliable quantifications and are used to flag samples for further analysis.
Detection of QSMs
Another indirect approach is the detection of extracellular QSMs or autoinducers. Many pathogenic bacteria are known to utilize quorum sensing (QS) as a cell–cell communication mechanism to coordinate metabolism in response to stimuli. 153 QS bacteria share the ability to synthesize, secrete, and respond to QSMs, which are diffusible signaling molecules that are relatively specific to a species or class of microbes. 154 For this reason, QSM levels can be associated with a specific bacterial species and cell density. Once extracellular QSMs reach a critical threshold concentration, they bind regulatory proteins to control the expression of specialized genes for modulation of cell metabolism, cell phenotype, and population behavior. Pathogenic features such as virulence factor and antibiotic production, biofilm formation, sporulation, and motility are associated with QS signaling. Furthermore, QS systems are largely implicated in the process of food spoilage. Spoiled food is oftentimes observed to house an accumulation of QSMs due to their key role in the expression of food-degrading enzymes. 155 Due to their prevalence among bacterial species and role in facilitating pathogenicity and food spoilage, QSMs are a promising biomarker for the detection of bacteria in food samples.
Detection of QSMs has been achieved using methods such as matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS), gas chromatography–mass spectrometry, high-performance liquid chromatography, and high-performance mass spectrometry. 156 These systems require expensive instrumentation and tedious sample processing steps, making them nonconducive to on-site testing. Simplified methods have recently employed cell-based biosensors with genetically engineered reporter strains coupled with colorimetric and electrochemical detection. Daunert and coworkers developed one such cell-based biosensor technology to quantitatively measure the concentration of the QSMs N-acyl homoserine lactone (AHL) and autoinducer-2 (AI-2) in food samples. 153 Their portable paper-based microdot test utilized E. coli reporter strains containing the plasmid pSD908, which encodes the lacZ gene under the control of the las recognition/regulatory system. Filter paper strips were incubated at 37 °C with the immobilized biosensing cells and β-galactosidase substrate. In the presence of QSM N-3-oxododecanoyl homoserine lactone (3OC12-HSL), the lacZ-controlled production of β-galactosidase resulted in a visible color change in the substrate. Detectable levels of inoculated Pseudomonas aeruginosa whole cells in beef and milk samples were achieved within 1 h after inoculation for a total time of 2.5 h, including incubation steps.
The use of mammalian immune cells as biorecognition elements has also been reported for QSM detection in food samples. The Fang group developed a novel electrochemical impedance sensor that utilized alginate/GO hydrogels for the encapsulation of rat basophilic leukemia mast cells (RBL-2H3) on MWCNs. 157 Binding of P. aeruginosa-produced 3OC12-HSLs to the immobilized mast cells triggered degranulation of secretory vesicles containing inflammatory mediators, which resulted in a significant decrease in cell viability and an increased electron transfer rate at the working electrode. Notably, the use of MWCN-modified screen-printed carbon electrodes (SPCEs) over traditional electrodes warranted a lower-cost, disposable device with improved electrochemical sensitivity and stability. This technique was applied for the detection of P. aeruginosa-produced AHLs in fish juice and achieved an LOD of 0.094 μM 3OC12-HSL. Prior to electrochemical analysis, however, the sample required multiple steps for AHL extraction.
Notably, the aforementioned cell-based biosensors do not report tests confirming assay selectivity for 3OC12-HSLs among other AHL molecules or substances found in the sample, which limits their use in species-level diagnostics. Furthermore, the use of immobilized bioreceptors increases manufacturing costs and limits chemical and thermal stability. 158 Issues regarding storage and shelf life with the use of cell-based biosensors may also arise due to the specific requirements for maintaining live cells. To address these challenges, Sesal and coworkers proposed a 2,2,6,6-tetramethyl-1 piperidinyloxy zinc(II) phthalocyanine (TEMPO-ZnPc)-coated glassy carbon electrode (GCE) as an amperometric biosensor with high selectivity for 3OC12-HSLs. 158 In this device, the TEMPO-ZnPc coating effectively replaced the use of biomimetic receptor molecules in electrochemical biosensors. Upon reaction between the radical groups of TEMPO and the amino group of AHL molecules, the redox activity of TEMPO groups was decreased and resulted in measurable electrochemical responses. Due to the variability in structure between different AHL molecules, current changes measured at the GCE/TEMPO-ZnPc electrode drastically differed upon the binding of each AHL. When compared with other AHLs, square wave voltammetry measurements for 3OC12-HSL were the only AHL to exhibit both a decrease in peak current and a shift in peak potential with increasing concentration. This method was employed to rapidly detect 3OC12-HSL within 5 s in milk samples with an equivalent LOD of 107 CFU/mL. Thus, while achieving rapid and selective detection, the GCE/TEMPO-ZNPc technology exhibited low sensitivity when compared with gold standard techniques. In addition, the reactivity of the TEMPO electrode was found to induce long-term instability under atmospheric conditions and required vacuum storage.
Overall, QSM-based techniques have garnered increased interest in the detection of foodborne pathogens. Many developed methods are rapid, simple, and amenable to miniaturization for portable detection. However, due to the density-dependent nature of QSM production, QSMs are typically present in food samples at nanomolar concentrations, which drives the necessity for increasingly sensitive detection methods. Furthermore, methods relying on QSM detection alone are insufficient for ensuring the absence of bacterial contamination. Some prevalent pathogenic bacteria, such as E. coli O157:H7, S. typhimurium, and S. aureus ATCC 29213, do not produce QSMs 156 and will go undetected by such devices. Even in devices that achieve selective detection of specific QSMs, species-specific identification remains a challenge due to the production of the same QSMs by multiple bacterial species.
Conclusions
In this review, we present recent developments in PON diagnostic tests that screen for foodborne pathogens. Special emphasis was placed on devices reported between 2015 and 2020 that involved detection in complex food and beverage matrices such as ground meat, milk, and juice. Devices were categorized by their method of detection, by either capture via antibody–surface marker interactions, nucleic acid amplification, or detection of metabolic products. These detection methods were further organized into paper-based and microfluidic devices, as well as several hybrid assay techniques.
While research groups have made significant advancements to improve the accessibility of these diagnostic devices in resource-poor settings, several obstacles remain that limit the widespread applicability of such tests in field settings. Many techniques still require extensive pretreatment (sample homogenization, cell lysis, and preconcentration) that necessitates external bulky and expensive equipment and reliable power sources. Furthermore, the need for numerous user handling steps and a controlled setting limit the feasibility for use by an untrained user without access to a laboratory. Even for devices with reagent-storing capabilities, researchers face challenges involving short shelf lives and strict storage requirements.159,160 Additionally, PON devices generally suffer from lower sensitivity when compared with conventional PCR and culturing techniques.159,160 Furthermore, accurate target quantification, multiplexing capabilities, and multifunctionality are critical components of PON devices that require further improvement.159,160 Recent research efforts have reduced the costs on the detection end of the device workflow by introducing paper-based detection methods with simple colorimetric readouts or integrating multiple liquid handling steps onto a single platform. In comparison, few improvements have been made in upstream sample processing. In the future, more focus must be placed on sample extraction, purification, and concentration from complex food and beverage matrices if these diagnostic devices are to achieve widespread use in the food industry. Several groups have shown promise in this regard, with the development of microfluidic ELISA platforms, 161 fluorometric 162 or LFA-based aptasensors, 163 electricity-free NAAT methods, 164 and metabolic-based devices that harness bacteria-mediated chemistry between functionalized nanoparticles, 165 or the detection of additional metabolic products such as cytochrome oxidase. 166 Future directions for these projects should involve testing for foodborne pathogens in complicated food samples to determine the feasibility of field testing using these devices.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
