Abstract
A reverse transcription loop-mediated isothermal amplification (RT-LAMP) assay was developed for the rapid identification of Infectious bursal disease virus (IBDV). The RT-LAMP assay used a set of 4 primers to amplify the viral protein 2 gene of IBDV for the detection of IBDV, showing not only high efficiency but also analytic specificity. The data demonstrated that the RT-LAMP assay detected 30 different IBDV isolates, had no cross-reaction with 3 other avian viruses (Infectious bronchitis virus, Newcastle disease virus, and Avian influenza virus), and obtained a 95.45% sensitivity in 22 positive clinical samples in reference to virus isolation. Therefore, this rapid, specific, sensitive, and convenient RT-LAMP assay could be applicable to the identification of IBDV in less-equipped laboratories as well as in the field.
Infectious bursal disease virus (IBDV) is the cause of infectious bursal disease, which is a highly contagious disease of young chickens and is a serious threat to the poultry industry worldwide. 2 Conventional methods for detection of IBDV include virus isolation, virus neutralization, reverse transcription polymerase chain reaction (RT-PCR), agar gel precipitation test, and antigen-capture enzyme-linked immunosorbent assay. However, all of these conventional methods are considered to be time consuming and unsuitable for rapid and routine detection of IBDV. Thus, a rapid, simple, and convenient detection method is needed.
A novel nucleic acid amplification method termed loop-mediated isothermal amplification (LAMP) was developed. 8,9 LAMP relies on autocycling strand displacement DNA synthesis performed by using a polymerase enzyme and is able to amplify nucleic acid under isothermal conditions and low reaction temperature. Compared with all PCR-based assays, LAMP does not require any expensive or special equipment. Therefore, LAMP-based detection assay would be suitable for detection-on-spot field situations or primitive laboratories in developing countries. LAMP-based detection assays have been used to successfully detect animal viruses (e.g., Avian influenza virus [AIV], Newcastle disease virus [NDV], Foot-and-mouth disease virus, and Porcine reproductive and respiratory syndrome virus). 3,4,5,10 The aim of the current study was to develop a novel method for the detection of IBDV in a simple, rapid, and cost-effective manner.
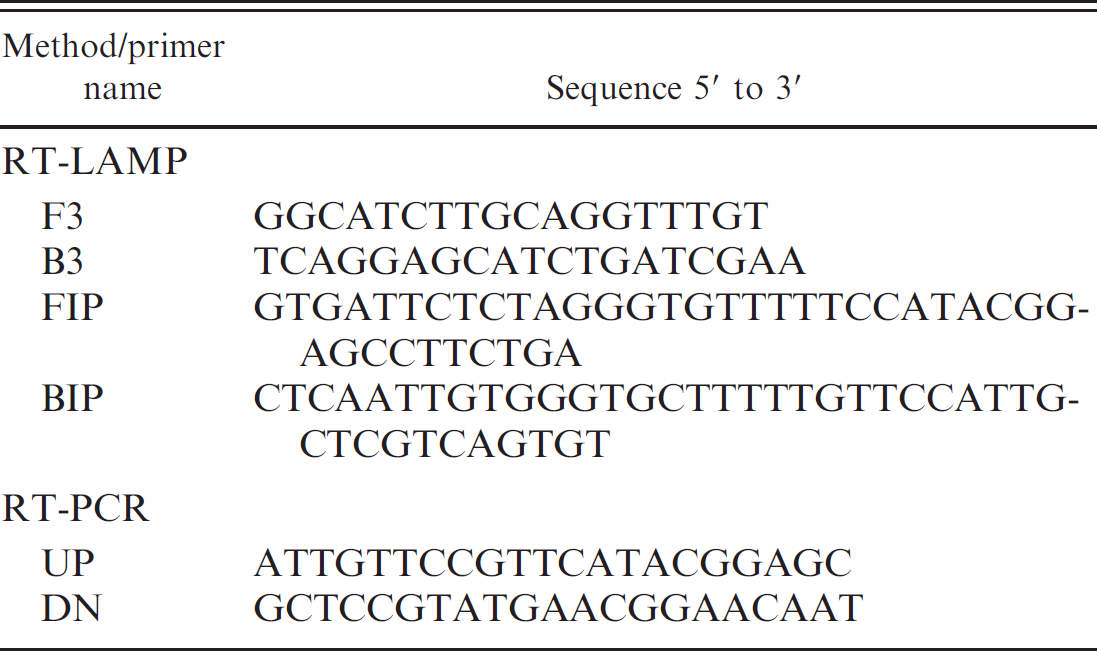
After alignment of the viral protein (VP)2 sequences of 98 IBDV isolates published in GenBank, a highly conserved region was chosen as a target sequence for the primer design. A set of 4 primers composed of an outer pair (F3 and B3) and an inner pair (FIP and BIP) were designed using Primer Explorer version 3. a The primers used in RT-PCR were designed by Primer 5.0. The nucleotide sequences of the primers are shown in Table 1.
Total RNA was extracted from IBDV isolates using an RNA extraction kit a according to the manufacturer's protocol. The RT-LAMP reaction was carried out in a 25-μl reaction mixture containing 2 M each of the inner primers FIP and BIP, 0.2 M each of the outer primers F3 and B3, 1.4 mM deoxyribonucleotide triphosphate (dNTP) mix, b 5 mM MgSO4, 16 units of Bacillus stearothermophilus (Bst) DNA polymerase, c 1 × the supplied Bst DNA polymerase buffer, 0.125 units of AMV reverse transcriptase, d and 2 μl of template RNA. The RT-LAMP reaction mixtures were incubated at the optimal reaction temperature (65°C) for the optimal reaction time (50 min) and were finally heat inactivated at 80°C for 5 min to terminate the reaction. To prevent cross-contamination, 20 μl of mineral oil e was added in the reaction before the incubation period. The amplified DNA products from the RT-LAMP reaction mixture were analyzed on 1% agarose gel stained with ethidium bromide, where the positive reaction mixtures showed a characteristic ladder of multiple bands. The RT-LAMP assay successfully amplified the target sequence of the VP2 gene of IBDV, as observed by agarose gel electrophoresis. The amplified product was observed as a ladder-like pattern on the gel due to the formation of a mixture of stem-loop DNA with various stem lengths and cauliflower-like structures with multiple loops formed by annealing between alternative inverted repeats of the target sequence in the same strand.
The reverse transcription loop-mediated isothermal amplification (RT-LAMP) and reverse transcription polymerase chain reaction (RT-PCR) primer sets used in the current study.
Temperature and reaction duration are 2 parameters critical in RT-LAMP reaction. To determine the optimal reaction temperature, the RT-LAMP reactions were carried out for 60 min at 61°C, 62°C, 63°C, 64°C, and 65°C. The DNA products from all reactions showed an obvious ladderlike pattern on the gel. However, the intensities of the DNA products at 64°C and 65°C were apparently higher than those at other temperatures, and the intensity of the DNA product at 65°C was the highest. Therefore, 65°C was considered the optimal temperature for the RT-LAMP reaction for detecting IBDV. To determine the optimal reaction time, the total reaction mixture was incubated at 65°C for 10, 20, 30, 40, and 50 min. The DNA products from the reaction with a duration of 50 min showed the highest intensity; thus, 50 min was considered the optimal reaction duration for the RT-LAMP reaction for detecting IBDV.
To ascertain the sensitivity of the RT-LAMP assay, 10-fold serial dilutions of RNA templates extracted from the samples were tested and compared to that of the conventional RT-PCR. The full-length complementary DNA of VP2 was synthesized by a commercial RT-PCR kit f according to the manufacturer's protocol. Briefly, a 50-μl reaction mixture contained 0.5 μl of RTase, 40 units of R Nase inhibitor, 20 mM dNTP mixture, 5 μl of 10 × 1-step RT-PCR buffer, 1 μl of 1-step enhancer solution (each of which were included in the commercial kit), 0.4 μM each of the UP and DN primers, 5 units of hot-start DNA polymerase, and 4 μl of RNA template. The thermal profile for PCR was 50°C for 30 min and 94°C for 2 min, followed by 35 cycles of 94°C for 30 sec, 56°C for 30 sec, and 72°C for 40 sec, with a final extension cycle at 72°C for 10 min. The amplified products were electrophoresed on 1% agarose gel stained with ethidium bromide. The result showed that the RT-LAMP assay was able to detect the RNA templates at 105-fold dilution, whereas RT-PCR was able to detect at 103-fold dilution (Fig. 1). Thus, the sensitivity of the RT-LAMP reaction for detecting IBDV is about 100 times higher than that of the RT-PCR reaction. In addition, the time for RT-LAMP reaction for the detection of IBDV was faster, taking only 50 min, compared with 2.5 hr for RT-PCR.
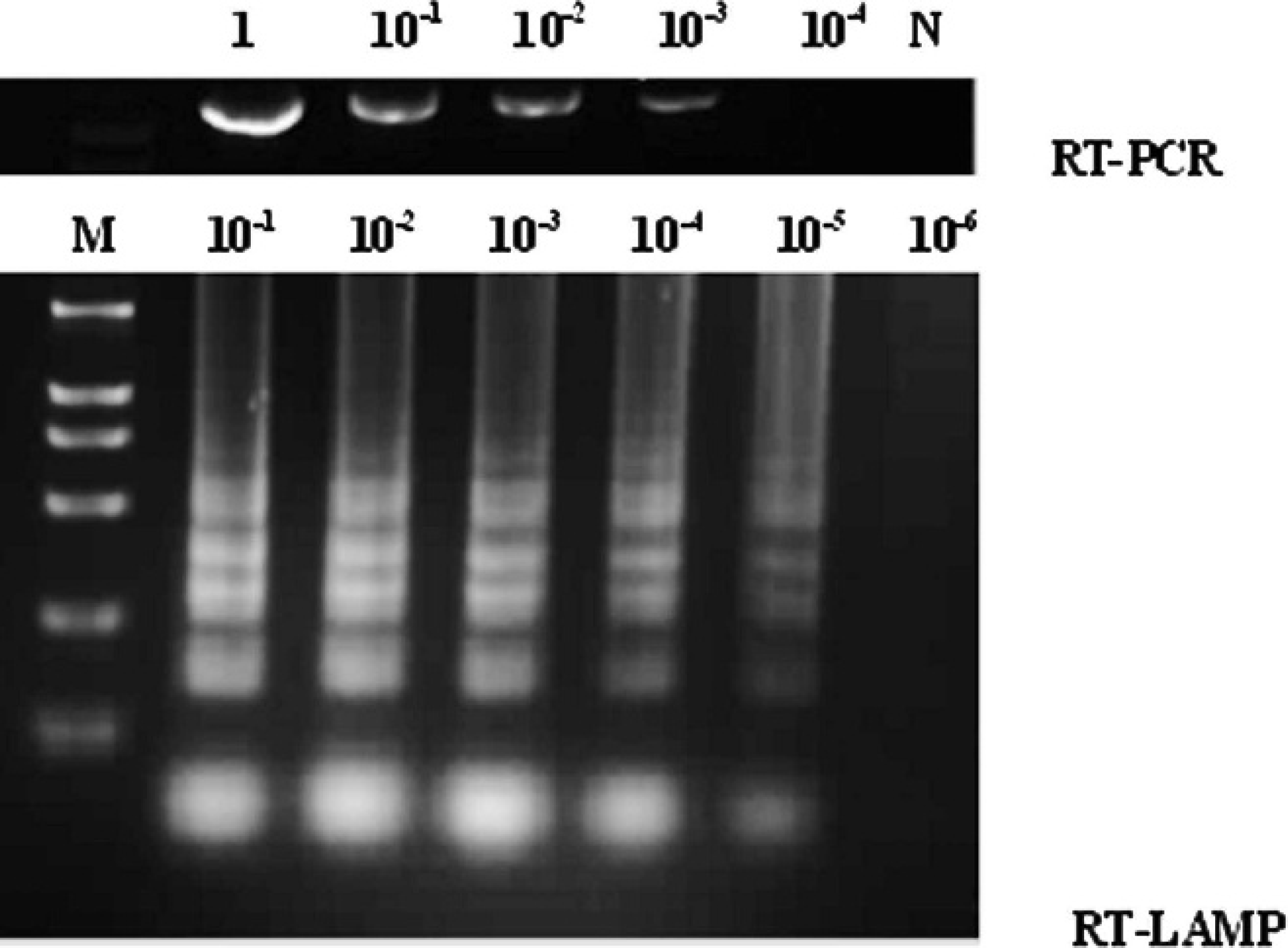
Comparison of sensitivity of reverse transcription polymerase chain reaction (RT-PCR) and reverse transcription loop-mediated isothermal amplification (RT-LAMP). Lane M: 2,000-bp ladder DNA marker; lanes 1–10−4 and 1–10−6: the corresponding dilutions of RNA template; and lane N: negative control, using water as template.
The specificity of the RT-LAMP assay was determined with 30 IBDV isolates and 3 other avian viruses (H9N2 AIV, NDV, and Infectious bronchitis virus g ). Twenty-six IBDV isolates were obtained from China in 2006–2007 by standard isolation methods 7 in 9-day-old, specific pathogen-free (SPF) chicken embryonated eggs. Standard IBDV strains 2512 and D78 were provided by commercial sources. h Strains HK46 (vvIBDV 1,6 ) and GZ902 (variant strain of IBDV 1 ) were isolated from South China and stored in the authors' laboratory. All 30 IBDV strains were positive, whereas all 3 of the other avian viruses were negative. This demonstrated that the RT-LAMP assay was specific, with no cross-reaction with the 3 other avian RNA viruses and, more importantly, that the RT-LAMP assay was applicable to diverse strains of IBDV, strongly suggesting the broad applications of RT-LAMP detection of IBDV in clinical samples.
To evaluate the application of RT-LAMP to detect IBDV in clinical samples, the test was performed on 25 field clinical bursal samples that were obtained from chickens suspected of infection with IBDV. All samples were placed in isotonic phosphate-buffered saline (pH 7.4) containing antibiotics and stored at −80°C until total RNA was extracted. All of the samples were subjected to virus isolation in SPF embryonated eggs, which yielded 22 positive and 3 negative. All 3 negative samples were also negative by both RT-LAMP and RT-PCR assays. For the RT-PCR method, 3 VI-positive samples were negative, resulting in a sensitivity of 86.36%. For the RT-LAMP method, 1 VI-positive sample was negative, resulting in a sensitivity of 95.45%. The RT-LAMP assay missed 1 infected sample, the reasons for which are still unclear. One possible reason is that the amount of IBDV in the 1 missed sample was so low that the virus might have been lost during the extraction process, or the amount of extracted viral RNA was not enough to trigger the LAMP reaction. To solve the problem, different extraction protocols and kits should be tried for improving the extraction efficacy. The second possibility is that IBDV in the 1 missed sample might contain mutations that might cause mismatches between the viral RNA and primers. This could be ruled out by sequencing the VP2 gene of IBDV isolated from the 1 sample.
In summary, the current study presented the rapid detection of IBDV by RT-LAMP with high sensitivity and analytic specificity. The RT-LAMP assay is feasible for use in less-equipped laboratories.
Acknowledgements. This work was supported by the grants from State Public Industry Scientific Research Programs (nyhyzx07-038) and Major Programs of Science Technology Strategic Plan (2007A020400006) of Guangdong, People's Republic of China. The authors would like to thank Dr. Qingfeng Zhou, Guangdong Wen's Co. Ltd., and Merial Animal Health Co. Ltd. for supplying clinical samples. Chunyi Xue and Yun Zhang contributed equally to this study.
Footnotes
b.
TaKaRa Biotechnology, Dalian, China.
c.
New England Biolabs Inc., Ipswich, MA.
d.
TaKaRa Bio Inc., Otsu, Japan.
e.
Sigma-Aldrich, St. Louis, MO.
f.
Primescript™, Ex Taq™ HS DNA Polymerase; TaKaRa Bio Inc., Otsu, Japan.
g.
Kindly provided by Guangdong Wen's Food Co. Ltd., Xinxing, China.
h.
Kindly provided by Merial Animal Health Co. Ltd., Shanghai, China, and Schering-Plough Co. Ltd., Shanghai, China, respectively.
