Abstract
Over the past decade, induced pluripotent stem cells (iPSCs) have become a major focus of stem cell and developmental biology research, offering researchers a clinically relevant source of cells that are amenable to genetic engineering approaches. Though stem cells are promising for both research and commercial endeavors, iPSC-based assays require tedious protocols that include complex treatments, expensive reagents, and specialized equipment that limit their integration into academic curricula and cell biology research groups. Expanding on existing Kit-On-A-Lid-Assay (KOALA) technologies, we have developed a self-contained, injection molded, pipette-less iPSC culture and differentiation platform that significantly reduces associated costs and labor of stem cell maintenance and differentiation. The KOALA kit offers users the full range of iPSC culture necessities, including cell cryopreservation, media exchanges, differentiation, endpoint analysis, and a new capability, cell passaging. Using the KOALA kit, we were able to culture ~20,000 iPSCs per microchannel for at least 7 days, while maintaining stable expression of stemness markers (SSEA4 and Oct4) and normal iPSC phenotype. We also adapted protocols for differentiating iPSCs into neuroepithelial cells, cardiomyocytes, and definitive endodermal cells, a cell type from each germ layer of human development.
Introduction
Cell-based assays are a ubiquitous tool in biotechnology and play a major role in drug development, disease research, and diagnostics. 1 Current cell-based assays, however, typically require specialized equipment, long assay times, and skilled personnel to perform the tasks associated with cell culture and monitoring. 2 Moreover, differences that arise because of the aforementioned variables make it difficult to standardize protocols and create a logistical barrier for collaborating laboratories and medical facilities. Furthermore, schools and education professionals are continually adjusting what they teach to meet the ever-changing science landscape. While cell culture is starting to be taught in some college and high school-level classrooms, the costs and difficulty of doing so are prohibitive to most.3,4 In order to make cell-based assays more accessible and to ensure primary and secondary schools are teaching the most relevant and up-to-date material to students, the costs and complexity of performing mammalian cell culture need to be reduced. One approach researchers have turned to to accomplish this is the use of microfluidics.
Induced pluripotent stem cells (iPSCs) are reprogrammed adult cells that are capable of differentiating into any adult cell type and represent an unprecedented opportunity for improved disease modeling and personalized medicine. 5 Despite the progress in developing iPSC-based assays for drug screening and clinical use, variability and cost continue to hinder further progress. 6 As the use of iPSCs in research and education continues to increase, these problems will become even more important to solve.
Though iPSC microfluidic cell culture is not novel, current platforms still retain a level of complexity that discourages most biologists and teachers from using them. These microfluidic iPSC culture technologies require the use of either a syringe or peristaltic pumps, which cost thousands of U.S. dollars (USD), or consist of complex microfluidic microchannels that require specialized knowledge to fabricate and operate, or both. 7 These factors can make commercialization difficult.8–13 The Kit-On-A-Lid-Assay (KOALA) is a technology capable of facilitating high-throughput, repeatable, low-media-volume cell culture assays, in a simple microfluidic straight channel without the use of liquid handling instruments or micropumps.14–16 This platform is simple and cheap to fabricate, consisting of micromachined polystyrene microchannels, reagent-filled lids, and a cryopreservation apparatus. Once commercialized, KOALA iPSC culture and differentiation kits, consisting of all the required prepackaged cells, culture media, and reagents, could be sold to researchers and educators. Additionally, though these kits would be sold as comprehensive stem cell culture systems, they would also be compatible with custom endpoints such as cell isolation and flow cytometry, drug screening, and enzyme-linked immunosorbent assay (ELISA).
Here we report the use of the KOALA to develop an easy-to-use, self-contained iPSC culture and differentiation system. We take the first step in scaling up the production of KOALA components by using injection molded microchannels. We also develop and characterize a passaging device that allows cells to be passaged from one KOALA microchannel to another microchannel at a user-defined split ratio.
Materials and Methods
Device Fabrication and Preparation
The lids, bases, and passaging adaptors were fabricated by CNC milling (PCNC 770, Tormach, Waunakee, WI, USA), with the lids having standard microscope slide dimensions (25 × 75 mm) ( Fig. 1A ). Bases are composed of three layers: an injection molded and milled post/channel polystyrene layer (Proto Labs, Maple Plain, MN, USA) consisting of 12 sets of posts/channels, a pressure-sensitive adhesive layer (ARcare 90106, Adhesives Research, Glen Rock, PA, USA), and a glass slide layer. The adhesive tape, which covered the entirety of the base except for the microchannels, was cut using a Graphtec Craft Robo Pro plotter cutter. The post/channel layer consists of an inlet and an outlet post connected by a 5 mm long, 2 mm wide, and 0.8 mm deep channel. After assembly, the device is oxygen plasma treated using a Diener Electronic Femto Plasma Surface System.
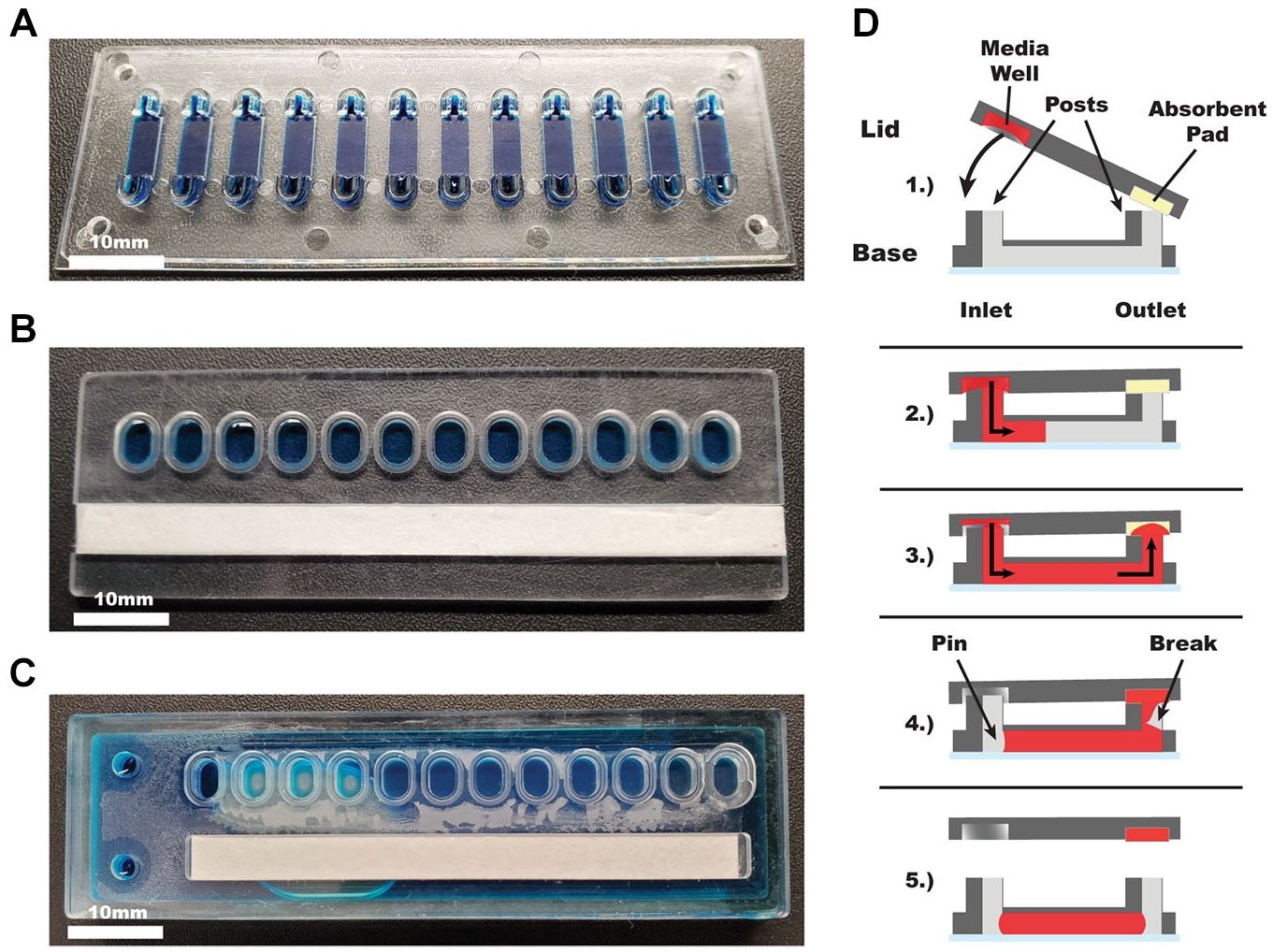
KOALA components and diagram. (
Lids are milled from 2 mm polystyrene (cat. 224-030-01, Goodfellow, Coraopolis, PA, USA). Each lid contains a microwell for each channel and a slot for absorbent pads. Each microwell has an elliptical shape and is spaced to match the pattern of the input posts. The slot for the absorbent pad is 10 mm wide, 72 mm long, and 2 mm deep. The lids are assembled by placing two layers of cellulose fiber absorbent pad (cat. CFSP223000, Millipore Sigma, Burlington, MA, USA) in the designated slot. Reagents or media is added to the wells before each use.
Cryopreservation lids consist of four components, three micromilled polystyrene parts and a nylon membrane filter with a pore size of 0.2 μm (WHA7402009, Millipore Sigma) ( Fig. 1A ). Two of the polystyrene parts fit together to form a large media reservoir with 12 elliptical ports, spaced out to match the spacing of the inlet ports of the KOALA bases and a small media addition port. The nylon filter membrane is placed over the elliptical ports and the last polystyrene piece, consisting of 12 elliptical ports and a slot for an absorbent pad, is aligned on top. All three polystyrene layers are acetonitrile bonded together to form a single device. The bases, lids, and cryopreservation lids are UV disinfected for 30 min, prior to cell culture. For the cryopreservation lid, cells and cryopreservation reagent (10% DMSO) are added to the nylon filter-backed wells, a pressure-sensitive adhesive film (Scotch tape, 3M, Saint Paul, MN, USA) is placed over each well, and the devices are placed in a –80 °C freezer and are frozen at a rate of 1 °C/min.
KOALA passaging devices are milled from 4 mm polystyrene. Each passaging device contains 12 media reservoirs 3.5 mm deep with a half-ellipse shape to fit the inlet post of KOALA microchannels, 12 media transfer channels with 3.5 mm deep, half-ellipse-shaped wells on each end, and a 10 mm wide, 72 mm long, and 3.5 mm deep slot containing twelve 3.5 mm tall wicking posts that align with the outlet posts of KOALA microchannels (see Fig. 4Ai ). Devices are oxygen plasma treated, the media reservoirs and nonfunctional surfaces are manually coated with a thin layer of paraffin wax to prevent liquid adsorption (cat. 22-900-700, Thermo Fisher, Waltham, MA, USA), and two layers of sterile absorbent pads are placed in the slot next to the wicking posts. Devices are UV disinfected for 30 min, prior to cell passaging, and are loaded with reagents prior to each application.
iPSC Culture and Maintenance
IMR90-4 human iPSCs (WiCell Research Institute, Madison, WI, USA) were maintained in feeder-free conditions on Matrigel (cat. 354230, BD Biosciences) in mTeSR1 (WiCell Research Institute) or TeSR-E8 (cat. 05990, STEMCELL Technologies, Vancouver, Canada) media at 37 °C and 5% CO2. All experiments were performed using iPSCs between passages 43 and 60. For routine iPSC maintenance, iPSCs were passaged with Versene (cat. 15040066, Thermo Fisher) every 4–6 days. For passaging iPSCs for differentiation, iPSCs were dissociated with Versene (EDTA solution) and the number of live cells was quantified via a hemocytometer using trypan blue stain (cat. 15250061, Thermo Fisher). iPSCs were cryopreserved in 10% DMSO in liquid nitrogen. iPSCs were then seeded with 10 μM Y27632 (ROCK inhibitor; cat. 1254, Tocris Bioscience, Bristol, UK) for the first 24 h to promote cell attachment. After 24 h, ROCK inhibitor was withdrawn and mTeSR1 or TeSR-E8 was added and replaced every 24 h. These iPSC culture conditions were used in both microdevices and a 96-well plate, with the iPSCs in KOALA microchannels receiving 10 μL of media and the iPSCs in the 96-well plate receiving 200 μL.
iPSC Differentiation
IMR90-4 iPSCs were dissociated and plated overnight in mTeSR1 or TeSR-E8 media at a density of 1.5 × 105 cells per square centimeter on Matrigel as described above. Neuroectoderm, cardiomyocyte, and definitive endoderm differentiation protocols were adapted from published work.17–19 Differentiation was verified using immunofluorescent staining, reverse transcription quantitative PCR (RT-qPCR), and cell phenotype.
Neuroepithelial differentiation: To generate neuroepithelial cells, cells were differentiated for 4 days in TeSR-E6 media (cat. 05946, STEMCELL Technologies).
Cardiomyocyte differentiation: To generate cardiomyocytes, cells were first differentiated overnight in RPMI1640 (Thermo Fisher) and B27 supplement minus insulin (cat. A1895601, Thermo Fisher) with 6 mM CHIR99021 (cat. 72052, STEMCELL Technologies). The following day, the media was replaced with RPMI1640/B27 minus insulin and the cells were allowed to recover for 24 h. On day 2, RPMI1640/B27 minus insulin-containing 4 µM IWP4 (inhibitor of Wnt 4 production; cat. 5214, Tocris) was added. On day 4, the cells were refed RPMI1640/B27.
Definitive endoderm: To generate definitive endoderm, cells were differentiated for 1 day in TeSR-E6 media containing 6 mM CHIR99021 and 100 ng/mL activin A (cat. 338-AC, R&D Systems, Minneapolis, MN, USA), followed by 3 days of activin A alone.
Image Acquisition and Analysis
Bright-field and fluorescent images were obtained using a Nikon Ti Eclipse inverted microscope. Images were processed using Nikon NIS-Elements. Images were analyzed using the open-source software ImageJ.
Viability Assay
Cells were washed with phosphate-buffered saline (PBS) and then stained for 15 min at 37 °C and 5% CO2 with calcein AM (C1340, Thermo Fisher) and ethidium homodimer-1 (E1169, Thermo Fisher). After 15 min, the cells were again washed with PBS and then imaged. Cells staining positive for ethidium homodimer were counted as dead cells, whereas cells staining for only calcein were considered living.
Immunofluorescence
To stain for pluripotency markers, we used the Pluripotent Stem Cell Immunocytochemistry Kit (OCT4, SSEA4; cat. A25526, Thermo Fisher) with all reagents loaded into a KOALA lid. To immunostain for differentiation marker, cells were first washed with 1× PBS and then fixed in 4% paraformaldehyde (cat. 43368, Alfa Aesar, Haverhill, MA, USA) for 15 min. Following fixation, the cells were then permeabilized with 0.1% Triton X-100 (cat. 807426, MP Biomedicals, Irvine, CA, USA) for 30 min and blocked with 1× PBS supplemented with 3% bovine serum albumin for 30 min. The following primary antibodies were used: recombinant anti-islet 1 antibody (EP4182; ab109517, Abcam, Cambridge, UK), anti-cardiac troponin T antibody (1C11; ab8295, Abcam), anti-PAX6 antibody (ab5790, Abcam), anti-N-cadherin antibody (ab76057, Abcam), anti-SOX17 antibody (OTI3B10; ab84990, Abcam), and recombinant anti-FOXA2 antibody (EPR4466; ab108422, Abcam). The following secondary antibodies were used: Alexa Fluor 555 donkey anti-rabbit (A-31572, Thermo Fisher), Alexa Fluor 594 donkey anti-rabbit (A-21207, Thermo Fisher), and Alexa Fluor 488 goat anti-mouse IgG3 (A-21151, Thermo Fisher). Fixed cells were incubated for 4 h at 4 °C with both the primary and secondary antibodies with three wash steps after each incubation.
RT-qPCR
Cells were lysed directly in KOALA microchannels, the cell lysate was pippetted from the devices, and then their mRNA was isolated in 15 µL of 10 mM Tris buffer using Dynabeads mRNA DIRECT Purification Kit (cat. 61011, Invitrogen, Carlsbad, CA, USA) per the manufacturer’s instructions. Immediately following mRNA isolation, a reverse transcription reaction was run using an iScript cDNA Synthesis kit (cat. 170-8891, Bio-Rad) and the resultant cDNA was preamplified with SsoAdvanced PreAmp Supermix (cat. 172-5160, Bio-Rad, Hercules, CA, USA) and primers from Integrated DNA Technologies ( Suppl. Fig. S2 ). Finally, qPCRs were run using iTaq Universal SYBR Green Supermix (cat. 172-5121, Bio-Rad) in a Roche Lightcycler 480 II (Roche Molecular Systems, Pleasanton, CA, USA) and a ΔΔCt analysis was run. GAPDH and ACTB were used for the relative expression analysis. No-template controls were run in parallel for all experiments.
Statistical Analysis
Data were analyzed (Prism 7.0; GraphPad Software, La Jolla, CA) using one-way ANOVA. Tukey’s multiple comparison test with a 95% confidence interval was used when comparing different conditions.
Results
The technology consists of a KOALA base containing 12 microchannels and pairs of input/output posts ( Fig. 1A ), a KOALA lid that contains 12 prefilled wells and a slot for an absorbent pad ( Fig. 1B ), and a KOALA cryopreservation lid consisting of 12 cryopreserved-cell-filled wells attached to a media reservoir, separated by a nylon filter ( Fig. 1C ). The basis of this technology relies on capillary action to perfuse liquid from the KOALA lid, through the KOALA base, and into the absorbent pad of the lid, as well as liquid pinning inside the microchannel, which causes a break in the liquid bridge between the base and the lid ( Fig. 1D ).14,15
iPSC Culture and Cryopreservation Using KOALA
Over time, stem cells cultured in less than optimal conditions will lose their pluripotency and self-renewal capabilities.20,21 Fluctuations in mechanical stimuli, hypoxia, extracellular matrix (ECM) composition, cell density, and small-molecule signaling have all been shown to contribute to this loss.22–25 Thus, we first sought to establish whether human iPSCs could be successfully cultured in KOALA microchannel devices.
Stem cell culture requires optimal cell densities and ECM composition. Using stem cell seeding densities ranging between 250,000 and 2 million cells/cm2, we added IMR90-4 iPSCs into Matrigel-coated KOALA microchannels and cultured them for 2 days, exchanging media twice a day. On the second day, we assessed the viability rate for each density and found that for the lower seeding densities (250,000–500,000 cells/cm2) the average viability rates were less than 33.2%, suggesting that these seeding densities were too low for stem cell culture in the KOALA microchannels ( Fig. 2A ). The average viability rates for the higher seeding densities, 1–2 million cells/cm2, were 75.5% and 90.8%, respectively. Though the higher cell seeding densities correlated with better viability rates, the concentration of cells used to achieve a seeding density of 2 million cells/cm2 was already relatively high and larger seeding densities would be impractical. Thus, we used the 2 million cells/cm2 seeding density for our experiments.
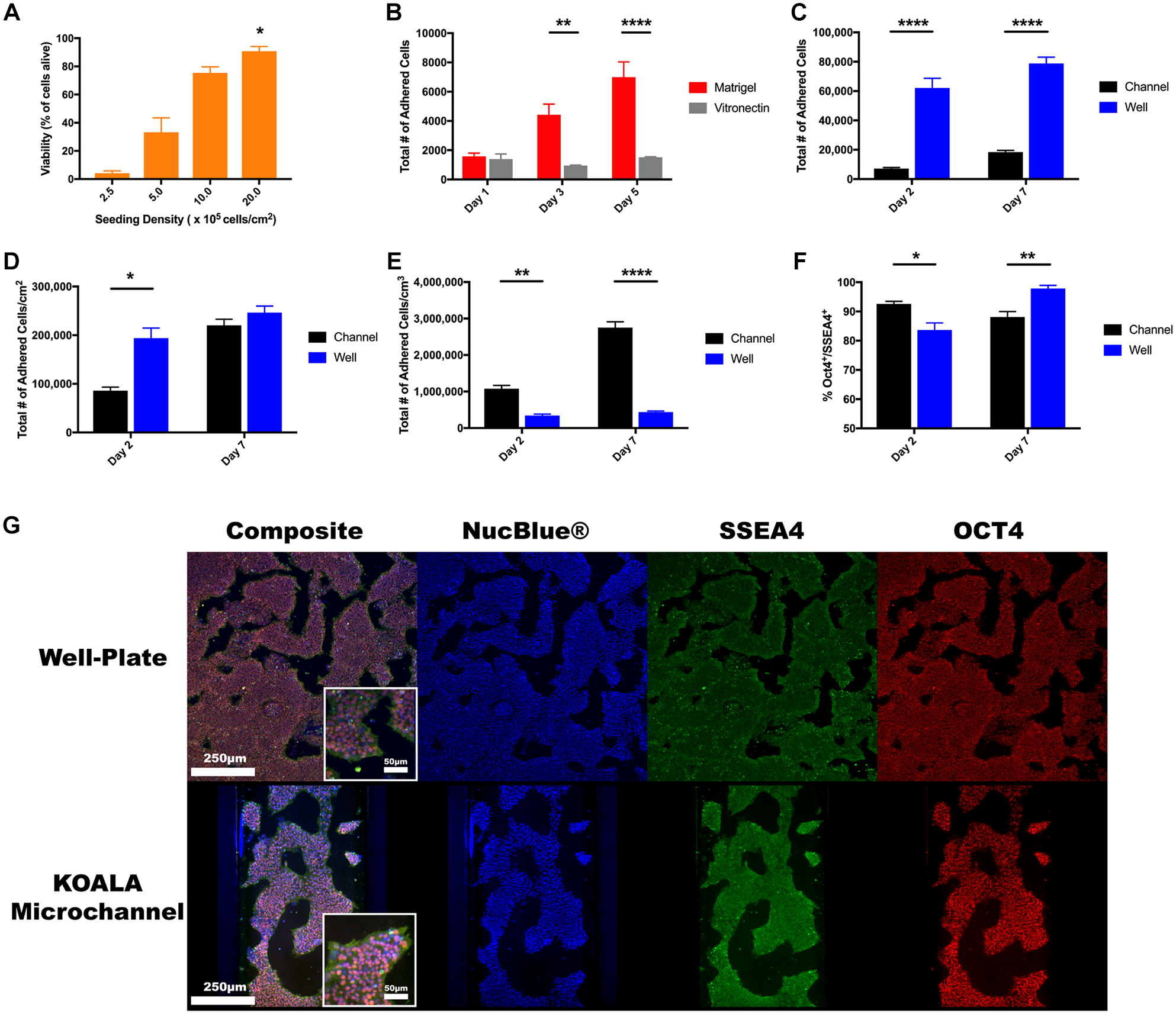
iPSC culture in KOALA versus well plate. (
Stem cell maintenance requires appropriate ECM conditions to support cellular self-renewal and to reduce spontaneous differentiation. 26 Classically, mouse embryonic fibroblasts and undefined basement membrane extracts such as Matrigel have been used for this purpose. Though Matrigel excels at facilitating stem cell culture, batch-to-batch variability has prompted researchers to turn to more defined ECM options. 27 In order to make the KOALA platform more appealing to cell biology researchers, we tested vitronectin, a fully defined protein substrate that has been shown to be amenable to stem cell culture. 28 To test vitronectin’s performance as an ECM substrate in KOALA, we compared its performance to Matrigel and analyzed the number of IMR90-4 iPSCs over 5 days. After 24 h of cell culture, there was no statistical difference between the number of adhered cells for both substrates with an average of 1581 ± 227 and 1399 ± 346 cells adhered to the Matrigel-coated and vitronectin-coated channels, respectively ( Fig. 2B ). However, at 72 and 120 h, the number of stem cells adhered to the Matrigel-coated microchannel was 4418 ± 739 and 6994 ± 1045, triple and quintuple that of the 24 h measurement, while the number of cells adhered to the vitronectin-coated microchannels remained similar to the 24 h measurement at 957 ± 28 and 1519 ± 36, respectively. For the vitronectin condition, we also noted a large number of cell detachments after each media change, suggesting that the iPSCs might not have adhered to the vitronectin as strongly as to Matrigel, or might have died and detached from the substrate. These results indicated that vitronectin did not perform as well as the widely used Matrigel and is not a suitable culture substrate for iPSCs in the KOALA platform. Thus, we decided that Matrigel would be used for future experiments.
In order to evaluate whether iPSCs can be cultured using KOALA, we compared stem cell culture in KOALA to cell culture in a 96-well plate, a conventional method of culturing and differentiating stem cells. We first seeded IMR90-4 iPSCs into KOALA microchannels and a 96-well plate and then counted the number of adhered cells at days 2 and 7 ( Fig. 2C ). At both time points, 2 and 7 days, the average number of adhered cells in the 96-well plate significantly outnumbered the number of cells adhered to the microchannels: 62,084 ± 6608 and 78,818 ± 4314 for the 96-well plate versus 7243 ± 606 and 18,491 ± 1070 for the microchannels. Taking into account differences in surface area, we found that the average number of adhered cells per surface area was significantly higher in the 96-well plate (194,014 ± 20,651 cells/cm2) compared with the microchannels (86,226 ± 7219 cells/cm2) for the day 2 time point, but no differences remained by day 7 (246,307 ± 13,480 cells/cm2 in the 96-well plates and 220,134 ± 12,735 cells/cm2 in the microchannels). This result suggests that there is an equivalent cell culturing capacity between the two platforms ( Fig. 2D ). One of the advantages of microfluidic cell culture platforms is their high cell number-to-volume ratios, which enable stronger cell–cell signaling.29–31 However, due to the high number of cells in a reduced volume, there is also an increased potential for nutrient depletion and cell starvation. When we look at the number of adhered cells per volume of media, we see a significant difference between the two culture platforms for both days, with the cells per volume ratio for the KOALA microchannels being six times higher than those for that of the 96-well plate ( Fig. 2E ).
To test if the increased cell-to-volume ratio affected the cells’ pluripotency, we immunostained IMR90-4 iPSCs in both KOALA microchannels and a 96-well plate after 2 and 7 days of culture and quantified the number of SSEA4+/Oct4+ cells, markers of pluripotency (
One of the main technological innovations of KOALA is the cryopreservation lid, which allows cells to be frozen and thawed and the cytotoxic cryopreservation media to be removed all within one microfluidic device. Briefly, the KOALA cryopreservation lid works similarly to a normal KOALA lid; however, there is an extra compartment attached to the back of the cryopreservation lid that is fluidically connected to the media wells by filter membrane. Cells are frozen in the wells of the cryopreservation lid, and when it is time to thaw and add them to microchannels, warm media is added to the rear compartment and the toxic cryopreservation reagents are dialyzed out. Once the cryopreservation reagents are diluted, the cells can be added to a KOALA microchannel, the same manner way a normal KOALA lid operates. 14 To test whether this device could be used to cryopreserve stem cells, we compared its performance to traditional cryopreservation methods. IMR90-4 iPSCs from the same culture were dissociated and frozen in media containing 10% DMSO in either a KOALA cryopreservation lid, 200,000 cells/well, or a cryovial. Both samples were then thawed using previously described protocols and the cells were seeded, either with the KOALA cryopreservation lid or by manually pipetting, into a Matrigel-coated KOALA microchannel. 14 Total adhered cells were then quantified and assayed for gene expression of pluripotency markers after 2 and 5 days of culture ( Fig. 3A ). After both 2 and 5 days of cell culture postthaw, there was no statistical difference in the total number of adhered cells. Additionally, there was no difference in the gene expression of POU5F1 and SOX2, markers for pluripotency, for both cryopreservation methods after 5 days of culture in a KOALA microchannel after cell seeding ( Fig. 3B ). These results indicate that the KOALA cryopreservation lid performs as well as traditional iPSC cryopreservation methods in preserving viability and pluripotency.
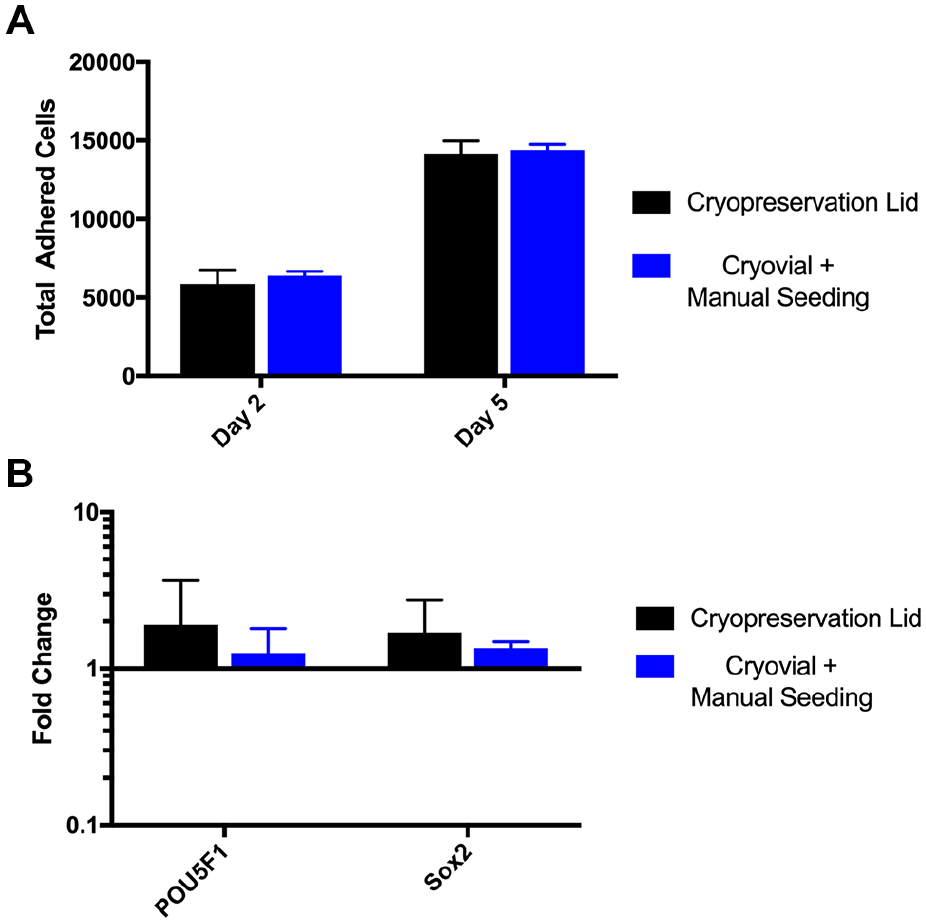
Cryopreservation lid characterization. (
Pipette-less Cell Passaging between Microchannels
We have demonstrated that the KOALA cell culture platform is capable of maintaining iPSCs with no significant effects on the cells’ pluripotency and self-renewal. We have also shown that KOALA performs comparably to traditional stem cell culture in a well plate. However, for KOALA to be a self-contained platform, a method of passaging cells to other devices for basic cell maintenance or downstream applications is needed.
We designed a microfluidic passaging device that allows cells to be transferred into new KOALA microchannels at a user-defined split ratio without the use of micropipettes ( Fig. 4A ). The device, which is similar to a KOALA lid, consists of a micromachined polystyrene plate with 12 media reservoirs, 12 sets of transfer channels/wells, and 12 wicking posts ( Fig. 4B ). The mechanism for how the device achieves unidirectional flow through the donor KOALA base, the passaging device, and into the new KOALA base relies on differences in surface hydrophobicity/hydrophilicity, and capillary action. To prepare the passaging device, it was first oxygen plasma treated to render the media transfer channels and wicking posts hydrophilic. Next, the media reservoirs and nonfunctional surfaces of the passaging device were coated with paraffin wax to make those surfaces hydrophobic. Lastly, an absorbent pad was placed next to the wicking initiation posts.
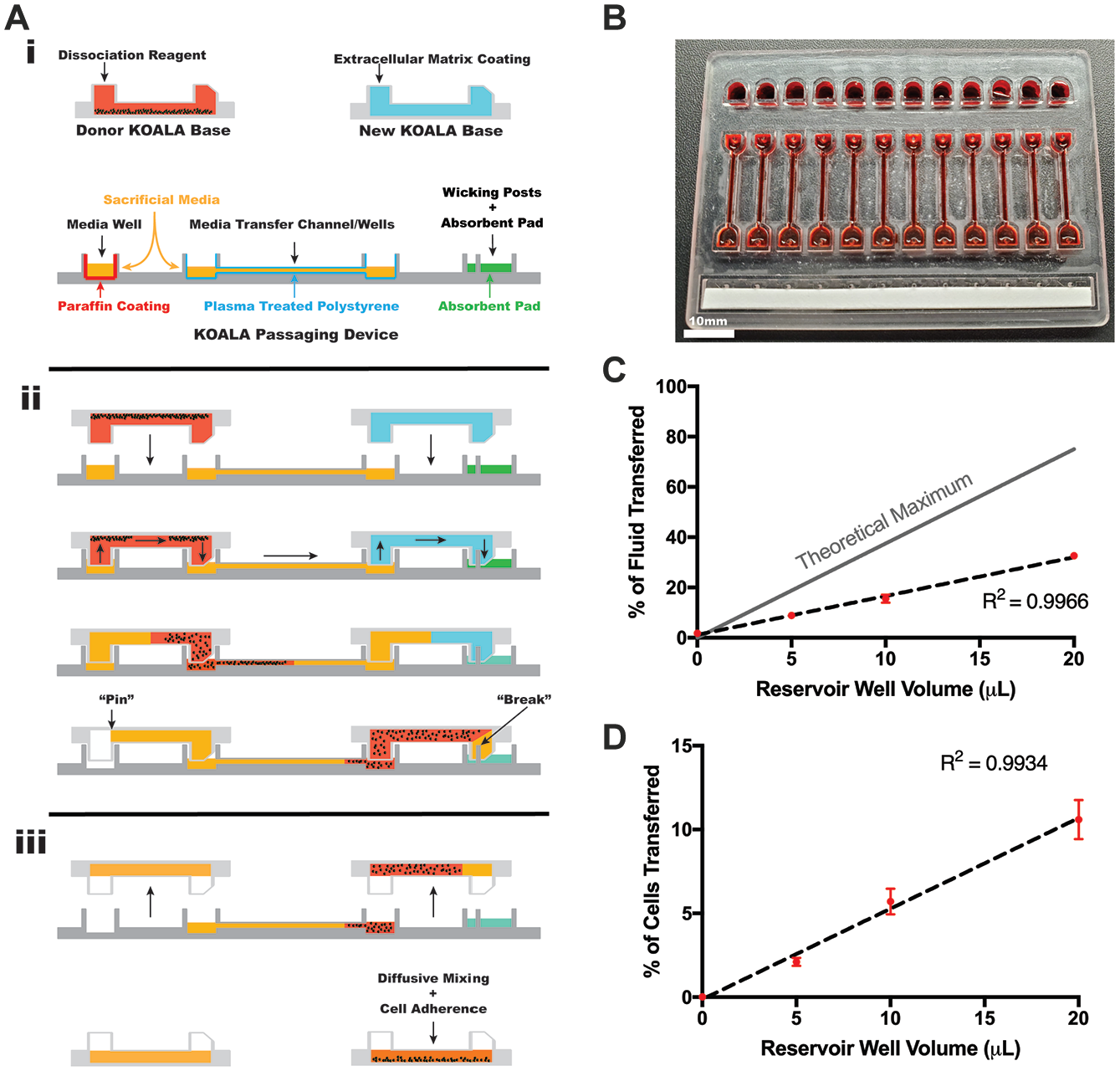
KOALA passaging device characterization. (
To operate the passaging device, a user-defined volume of cell culture media is added to the media reservoirs, along with 15 µL of sacrificial media to the transfer channels/wells ( Fig. 4Ai ). Cell dissociation reagent is added to the donor KOALA device to detach adherent cells, and ECM solution is added to the new KOALA device to coat it with the necessary ECM. Once the cells become detached, the donor KOALA device is placed upside down onto the passaging device so that the input posts of the donor KOALA device align and fit into the media reservoir wells and the output posts of the donor KOALA device fit into the wells of the transfer channels ( Fig. 4Aii ). This action creates a continuous liquid interface from the media reservoirs, through the donor KOALA device, and into the transfer channels. Next, the ECM-containing KOALA device is placed upside down onto the passaging device so that the input posts of the KOALA device fit into the wells of the transfer channel, opposite those of the donor KOALA device, and the output posts of the new KOALA device align with the wicking posts of the passaging device and fit into the output posts. Once fluidic contact is made between the transfer channels, the ECM-coated KOALA device, and the wicking posts/absorbent pad, capillary action, driven by the absorbent pad, will induce laminar flow from the media reservoirs, through the donor KOALA device, into the transfer channels, through the new KOALA device, and into the absorbent pad. Flow of liquid will continue until all of the cell culture media in the media reservoir is transferred through the devices, after which, due to the design and dimensions of the KOALA posts, the media will “pin” in the input post of the donor KOALA device, which will stop the flow of media into the ECM-coated KOALA device. 14 Once the liquid transfer is complete, both KOALA devices can be separated from the passaging device ( Fig. 4Aiii ). By adding more or less liquid to the media reservoirs of the passaging device initially, one can control the amount of fluid transferred from the donor device to the receiving device, and thus the split ratio. Additionally, homogenous cell seeding in the ECM-coated KOALA device can be achieved by placing the device, with the unadhered cells, immediately onto a rocker table to allow the cells to roll and adhere throughout the microchannel.
In order to confirm the function of the device, we first tested its liquid transfer capabilities using fluorescein dye. To do this, we filled donor KOALA devices with a solution of fluorescein and then used passaging devices filled with different volumes of water in the media reservoir wells to transfer the fluorescein to a new, water-filled KOALA device. By measuring fluorescein fluorescence in the water-filled KOALA device, we were able to calculate the percentage of fluid, from the donor device, that was transferred to the new device ( Fig. 4C ). We found a linear relationship between the initial media reservoir volume and the percent of liquid transferred to the new channel (R2 = 0.9966). Additionally, the passaging device was consistent in its operation across all media reservoir volumes tested, demonstrating low standard deviations for each volume. Though the passaging device worked with high consistency, the percent of media transferred was significantly lower than the calculated theoretical maximum.
Next, we tested the passaging device with stem cells. We first seeded KOALA devices with IMR90-4 iPSCs and grew them to confluency. We then coated new KOALA microchannels with Matrigel. Using Versene and mechanical tapping of the device to detach and dissociate the iPSCs, we used the passaging device to transfer iPSCs to the Matrigel-coated channels at different volumes ( Fig. 4D ). Though we achieved a similar level of consistency, low standard deviations across multiple replicates, across all of the media reservoir volumes used, the percentage of cells that were transferred to the new device did not compare directly with the results from the previous experiment. For the 20 μL condition, 10.6% of the cells were transferred to the KOALA device compared with 32.6% of the fluorescein solution. As a whole, these data show that the passaging device is capable of passaging iPSCs with high precision and consistency.
iPSC Directed Differentiation in Microchannels
Stem cell differentiation is an important functionality for KOALA’s broader application as a tool for stem cell research. To demonstrate this, we used directed differentiation to differentiate IMR90-4 iPSCs into cell types from each germ layer: neuroepithelium (ectoderm), cardiomyocytes (mesoderm), and definitive endoderm (endoderm).
Neuroepithelial cells are the first precursor cells that form during the development of the nervous system. 32 Using a differentiation protocol developed by Lippmann et al., we were able to differentiate IMR90-4 iPSCs into neuroepithelial cells after 7 days of directed differentiation in KOALA. 17 IMR90-4 iPSCs were seeded into Matrigel-coated KOALA channels, allowed to adhere, and then cultured in mTeSR-E6 for 7 days, according to the established protocol. Cells from KOALA channels were collected at 2, 5, and 7 days after the start of differentiation and the gene expression of PAX6, SOX2, and POU5F1 was quantified using a qPCR analysis. During neuroectoderm differentiation, gene expression of PAX6 should increase, SOX2 levels should remain stable, and POU5F1 expression should decrease. After 2 days of differentiation, we observed a 38,838-fold increase in PAX6, a 0.93-fold reduction in SOX2, and a 0.38-fold reduction in POU5F1 gene expression, consistent with the published protocol ( Fig. 5Ai ). These gene expression levels remained consistent through 7 days. Another indicator of neuroectoderm differentiation is the formation of polarized rosette structures that can be visualized by immunofluorescent staining for Pax-6 and N-cadherin. 33 After 7 days of differentiation, we fixed and stained cells within KOALA microchannels for these two proteins. While we observed strong and consistent staining for both Pax6 and N-cadherin, additional markers of neuroepithelial cells, we were not able to detect robust rosettes structures ( Fig. 5Aii ). However, the qPCR results and the strong Pax6 and N-cadherin staining indicate we were able to successfully differentiate iPSCs into neuroepithelial cells in the KOALA device, though further optimization will be needed to differentiate these cells down the neural lineage.
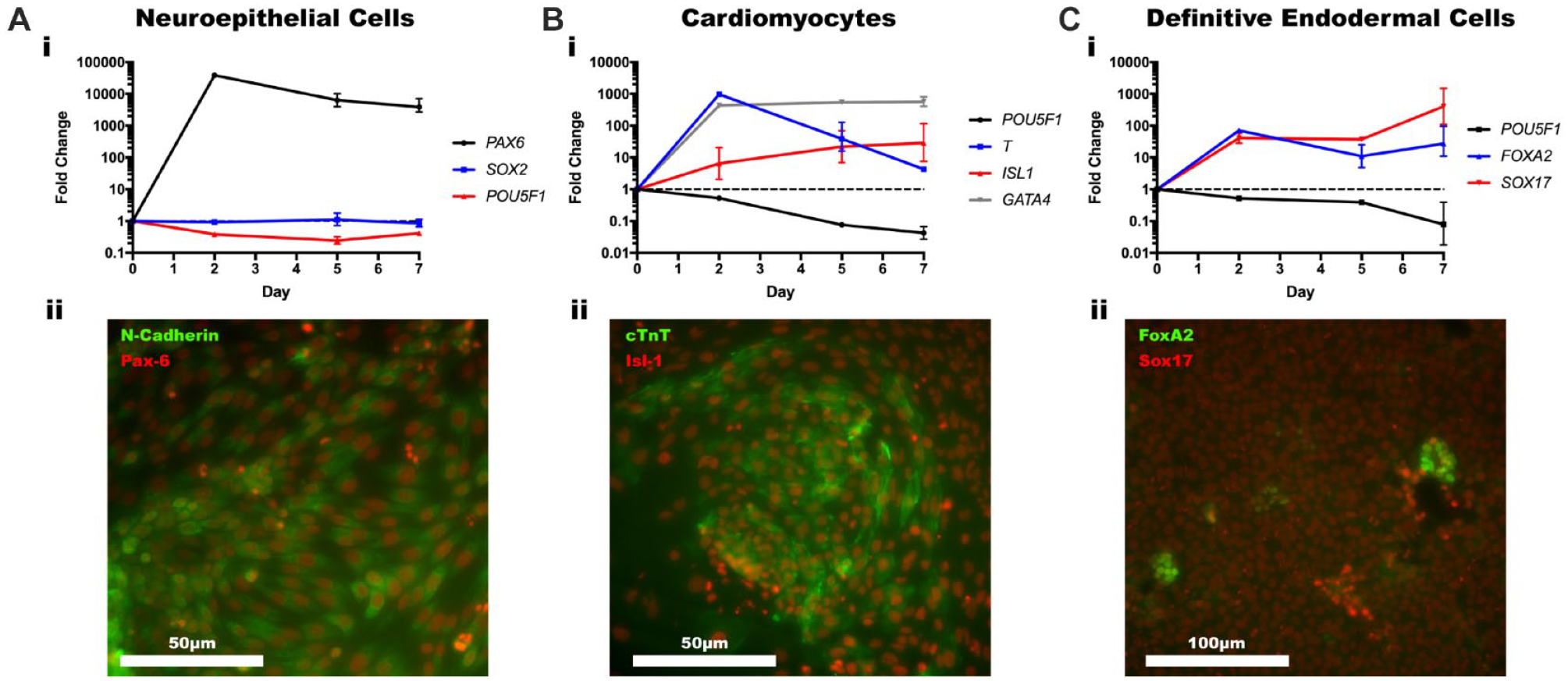
Directed differentiation in KOALA. (
Cardiomyocytes are the muscle cells that make up cardiac muscle and are derived from the mesodermal germ layer. 34 Using a differentiation protocol developed by Lian et al., we were able to differentiate IMR90-4 iPSCs into cardiomyocytes cells after 7 days of directed differentiation in KOALA. 18 IMR90-4 iPSCs were seeded into Matrigel-coated KOALA channels, allowed to adhere, and then differentiated according to the established protocol. Cells from KOALA channels were collected at 2, 5, and 7 days after the start of differentiation and a qPCR analysis was run to measure gene expression of POU5F1, T, ISL1, and GATA4 ( Fig. 5Bi ). During cardiomyocyte differentiation, gene expression of POU5F1 will steadily decrease, expression of T will initially increase significantly as the cells take on a mesoderm-like phenotype and then decrease as the cells differentiate further down the cardiomyocyte lineage, and expression of ISL1 and GATA4 will increase. As expected, gene expression of POU5F1 decreased by 0.53-fold after 2 days of differentiation and continued to decrease further, expression of T increased 975-fold by day 2 and then only by 4.27-fold by day 7, expression of ISL1 steadily increased by 28.61-fold over 7 days, and expression of GATA4 increased by 434.53-fold after 2 days of differentiation and remained steady for the next 5 days. On the seventh day of differentiation, we also fixed and immunostained cells in KOALA microchannels for Isl1 and cTnT (cardiac troponin T) ( Fig. 5Bii ). We observed uniform staining for both proteins, which further suggested successful cardiomyocyte differentiation. However, cardiomyocytes in 2D cell culture will begin rhythmically contracting without any external signals. While we only observed two instances of contracting cells ( Suppl. Movie 1 ), combined with the qPCR and immunostaining results, we conclude that we were able to successfully differentiate iPSCs into cardiomyocytes using the KOALA platform.
Lastly, the definitive endoderm gives rise to the epithelial lining of the digestive and respiratory tracts, and to the thyroid, thymus, lungs, liver, and pancreas.35,36 Using a differentiation protocol developed by Kunisada et al., we were able to differentiate IMR90-4 iPSCs into definitive endodermal cells after 7 days of directed differentiation in KOALA. 19 IMR90-4 iPSCs were seeded into Matrigel-coated KOALA channels, allowed to adhere, and then differentiated according to the established protocol. Cells from KOALA channels were collected at 2, 5, and 7 days after the start of differentiation and the expression of definitive endoderm markers SOX17 and FOXA2 as well as POU5F1 was quantified using qPCR ( Fig. 5Ci ). After 7 days of differentiation, expression of SOX17 and FOXA2 increased 403.1- and 27.5-fold, respectively, while POU5F1 expression decreased 0.08-fold, indicative of a definitive endoderm phenotype. We also immunostained for Sox17 and FoxA2, and while Sox17 stained uniformly, we only found small patches of cells stained for FoxA2 ( Fig. 5Cii ). According to the gene expression, FOXA2 was less upregulated than SOX17 and, consequently, more time may have been needed for higher levels of FoxA2 protein to be translated. Also, compared with the neuroepithelial and cardiomyocyte differentiation protocols, the protocol followed here resulted in a much lower differentiation efficiency rate. Regardless, based on the high gene expression for SOX17 and FOXA2, we successfully differentiated iPSCs into definitive endodermal cells using KOALA.
Discussion
iPSCs are sensitive cells whose pluripotency and self-renewal are highly influenced by slight changes in their environment. This poses technical and economic barriers for many researchers or educators who lack experience and setup. KOALA is a pipette-less, self-contained cell culture platform that can ameliorate some of the inconsistencies in stem cell culture, provide a more customizable microenvironment, and decrease overall costs. In this paper, we described the optimization and characterization of iPSC culture in the KOALA platform.
Most microfluidic cell culture devices fail to be adopted by the science community. Either their complexity or specific application, or both, limits their scalability and use by scientists. For example, Kamei et al. and Gómez-Sjöberg et al. both use intricate microfluidic designs, which include on- and off-chip valves and pumps, and small channel dimensions to culture stem cells.8,10 While useful, their operation and fabrication require microfluidic expertise that most biology researchers and educators do not have. Even more straightforward microfluidic stem cell culture technologies like the ones used by Giobbe et al. and Liu et al. require expensive syringe/peristaltic pumps to operate, essentially making these stem cell culturing platforms an inaccessible option for most scientists.9,11 The KOALA platform contains everything a user needs to culture mammalian cells except for incubators and biosafety cabinets. The mechanism off which the KOALA technology is based is simple to perform and requires very little expertise. Moreover, the components used for KOALA are simple in design and disposable.
In terms of cost, all of the KOALA components were fabricated using cheap materials. Besides the KOALA base, which was composed of an injection molded polystyrene piece, tape, and a glass slide, the rest of the KOALA components (i.e., lid, cryopreservation lid, and passaging device) were made from micromachined polystyrene sheets. Without factoring in reagents and labor, the cost of a single KOALA kit, consisting of two KOALA bases, a cryopreservation lid, a passaging device, and 20× KOALA lids, would be less than $10 USD. While stem cell culture reagents are generally expensive, the lower reagent volumes used in KOALA could help reduce the cost per kit. Additionally, though the fabrication of KOALA components outlined in this paper required nontrivial amounts of labor, because of the low complexity of KOALA, we have taken steps toward commercialization and have begun to use injection molded devices. In this paper, all of the KOALA bases, which contain the microchannels, were injection molded, a step very few microfluidic devices achieve. Once designs are finalized for the KOALA lids, cryopreservation lids, and passaging devices, we will have those components injection molded as well, which will significantly reduce the associated labor costs.
We first optimized basic iPSC maintenance in KOALA microchannels and compared its performance with that in traditional well plates and found that there were no significant differences between the two methods in terms of cell viability and cell density. Additionally, we noticed no microorganism contamination in either platform over 7 days. However, when comparing the number of cells expressing pluripotency markers between KOALA microchannels and well plates, we noticed that the expression of these markers was higher after 2 days of culture in microchannels but then decreased by 7 days, whereas the reverse was true for well plates. The higher cell-to-fluid volume ratios at the microscale leads to a faster accumulation of secreted factors and could explain the higher numbers of cells expressing pluripotency markers in microchannels versus well plates. 30 However, waste products and metabolites also accumulate faster and, when exposed to these detrimental factors for long enough, could also explain the decrease in pluripotency markers for iPSCs cultured in KOALA microchannels. By better understanding this dynamic in terms of stem cells, an improved design or protocol (e.g., timing of media changes) could be implemented to help mitigate the detrimental effects observed here and could produce a more stable iPSC culture platform.
In this paper, we introduced a new component to KOALA that facilitates cell passaging between two devices at a user-defined split ratio. The device operates similarly to a KOALA lid and was able to passage cells with a high level of precision between technical and biological replicates. One challenge with the device was the lower than expected cell passaging rate. When we inspected the passaging device after passaging cells, we observed clumps of cells remaining in the wells at the ends of the cell transfer channels, indicating either that there is nonlaminar flow through portions of these wells or that the cells are adhering to the plasma-treated surface. Regardless of this loss of cells, the passaging device performed consistently and future use of the device will use media volumes that account for these differences. Cell passaging was the last cell culture procedure that KOALA was not able to perform. The addition of the passaging device to the KOALA platform means that all of the steps involved with iPSC culture can be carried out within the KOALA platform.
In terms of functionality, we demonstrated how KOALA can support directed differentiation of iPSCs down cell lineages from all three germ layers, neuroepithelium (ectoderm), cardiomyocytes (mesoderm), and definitive endoderm (endoderm). For all three directed differentiations, we were able to achieve robust gene expression of the proper lineage markers for each cell type. Additionally, we observed strong expression of lineage markers for all but definitive endodermal cells. However, even though we attained uniform immunostaining for the neuroepithelial and cardiomyocyte differentiations, we failed to observe rosette-like structures or consistent spontaneous contractions, hallmarks for each cell type. This lack of phenotype could stem back from our previous point that while growth factors accumulate quicker in microdevices, so do metabolites, and these metabolites could be having a detrimental effect on the cells we were attempting to differentiate. The directed differentiation protocols used in this paper most likely did not account for issues like this. Though they worked reasonably well in KOALA, future work will include optimization of differentiation protocols that take into account parameters that affect cell culture at the microscale.
Together this work demonstrates the utility of KOALA in stem cell research and opens up the possibility for the creation and commercialization of microfluidic stem cell differentiation kits. Compared with traditional stem cell culture, reagent volumes used in KOALA are significantly reduced, 10 µL reagent/media/microchannel versus 200 µL/well. Overall, KOALA iPSC differentiation kits could be a cheaper and easier alternative for researchers and educators to use.
Future work will focus on both improving differentiation protocols and expanding the functionality of the device. Most differentiation protocols are optimized for use in well plates. From our results, we found that these protocols do not translate directly to microchannels, most likely due to the higher cell-to-volume ratio found in microfluidic devices. Additional work is needed to optimize differentiation protocols for KOALA microchannels. We have shown that KOALA is amenable to many endpoints, such as most cell staining and imaging techniques, and RNA/DNA isolation. One of the attractive applications of stem cells is their ability to be genetically altered and then differentiated down multiple cell lineages, allowing researchers to better model and study genetic diseases. Adding functionality for gene editing to KOALA could be a useful function for it to have.
Supplemental Material
Supplementary_Information – Supplemental material for Induced Pluripotent Stem Cells on a Chip: A Self-Contained, Accessible, Pipette-less iPSC Culturing and Differentiation Kit
Supplemental material, Supplementary_Information for Induced Pluripotent Stem Cells on a Chip: A Self-Contained, Accessible, Pipette-less iPSC Culturing and Differentiation Kit by Patrick McMinn, David J. Guckenberger and David J. Beebe in SLAS Technology
Footnotes
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: David J. Beebe holds equity in Bellbrook Labs LLC, Tasso Inc., Salus Discovery LLC, LynxBiosciences Inc., Turba LLC, Stacks to the Future LLC, and Onexio Biosystems LLC. David J. Guckenberger holds equity in Tasso Inc. and Salus Discovery LLC.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by the NIH: National Institutes of Allergy and Infectious Diseases through R01 AI134749, the State Economic Engagement & Development (SEED) Research Program, and the University of Wisconsin Carbone Cancer Center Support Grant P30 CA014520.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
