Abstract
Single-cell cloning is essential in stem cell biology, cancer research, and biotechnology. Regulatory agencies now require an indisputable proof of clonality that current technologies do not readily provide. Here, we report a one-step cloning method using an engineered pipet combined with an impedance-based sensing tip. This technology permits the efficient and traceable isolation of living cells, stem cells, and cancer stem cells that can be individually expanded in culture and transplanted.
Introduction
The ideal single-cell cloning technology must fulfill strict criteria: First, it must allow for the isolation of single cells with high efficiency and avoid the isolation of multiple cells, doublets, and aggregates. Second, it must be disposable to avoid cell cross-contamination from one experiment to another. Third, the procedure must be sterile to prevent bacterial and viral contamination. Fourth, cell viability and functionality must be preserved so that the cells can be either analyzed by state-of-the-art cell biology and omics assays or transplanted. Fifth, the proof of clonality must be recorded in real time to comply with regulatory requirements. 1 Lastly, the technology should be simple, cost-effective, and compatible with standard workflows.
Several technologies are available to isolate single cells, but none meet all the above criteria (Table 1).2–11 Some of them are commercially available, such as the scp single-cell printer (Cytena, Germany)11,12 and the cellenONE (Cellenion, France) based on optical detection to dispense single cells, or the ICell8 13 (Takara Bio, Japan) and the Rhapsody 14 (BD), which allow single-cell trapping in nano- or microwells. Here, we have designed a cell- and user-friendly pipet that allows the versatile isolation of viable single cells together with unambiguous proof of clonality. We engineered a handheld pipet to host an impedance analyzer connected to a computer and a micropump ( Fig. 1a ) coupled with a sterile disposable sensing tip that acts as a cell counter as first described by Wallace Coulter in seminal publications ( Fig. 1b ).15,16 We have previously reported in detail the design and engineering features of the pipet (Bonzon et al., submitted). 17 Briefly, the sensing tip features a gold electrode adherent to its external face and an internal floating electrode made of medical-grade stainless steel ( Fig. 1c ). A 15 µm thick parylene membrane with a 30 µm diameter laser-drilled aperture in its center seals the lower end of the tip ( Fig. 1d ). Altogether, the device senses and records every single particle or cell that flows through the aperture of the tip ( Fig. 1e ). We produced the tips in batch mode and sterilized them by ethylene oxide or gamma irradiation, whereas we sterilized the holding pipet with ethanol. The device is cell-friendly, intuitive, and user-friendly. It easily fits into a laminar flow hood and is compatible with existing workflows.
Advantages and Disadvantages of Existing Single-Cell Isolation Methods.
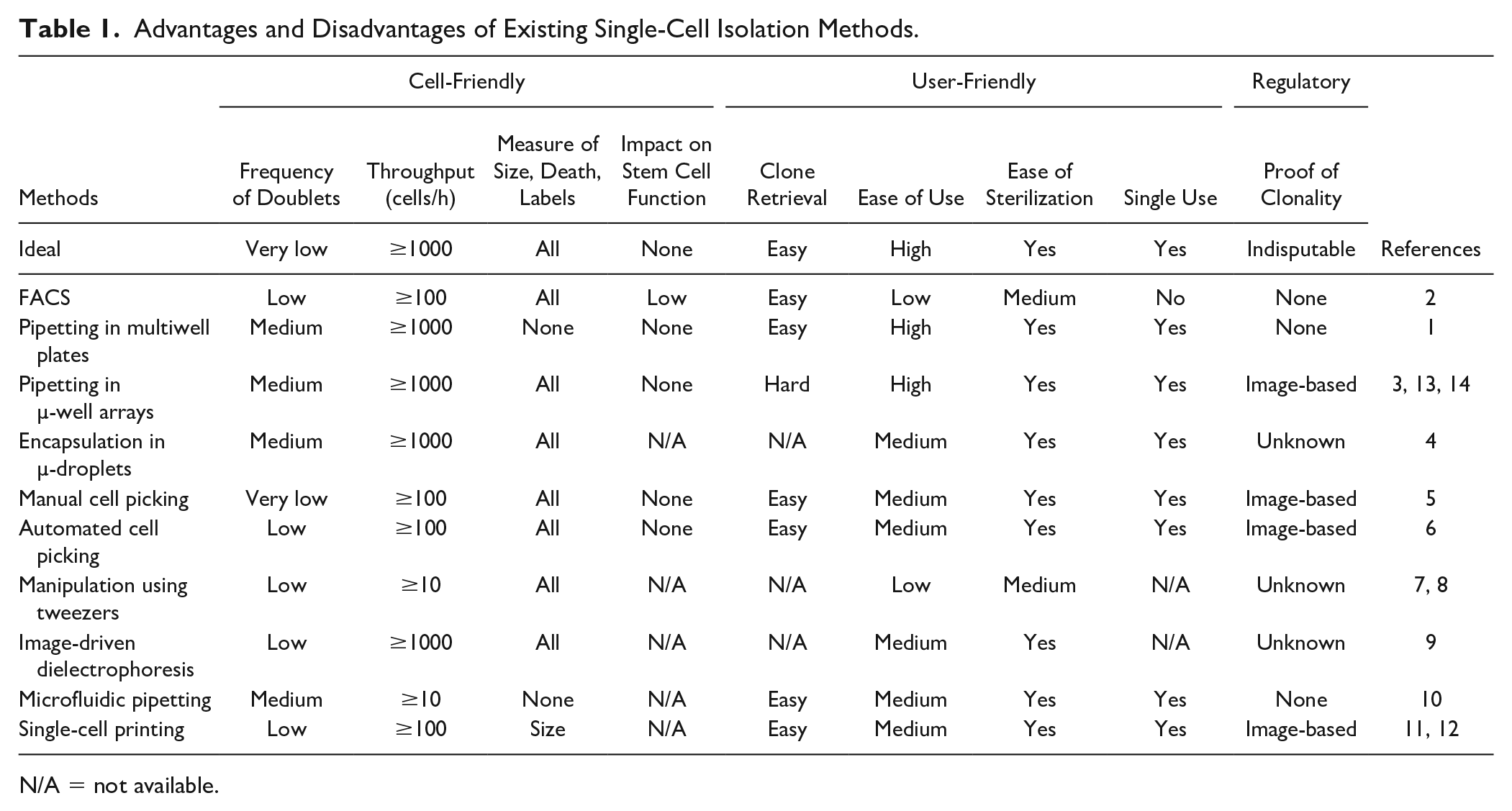
N/A = not available.
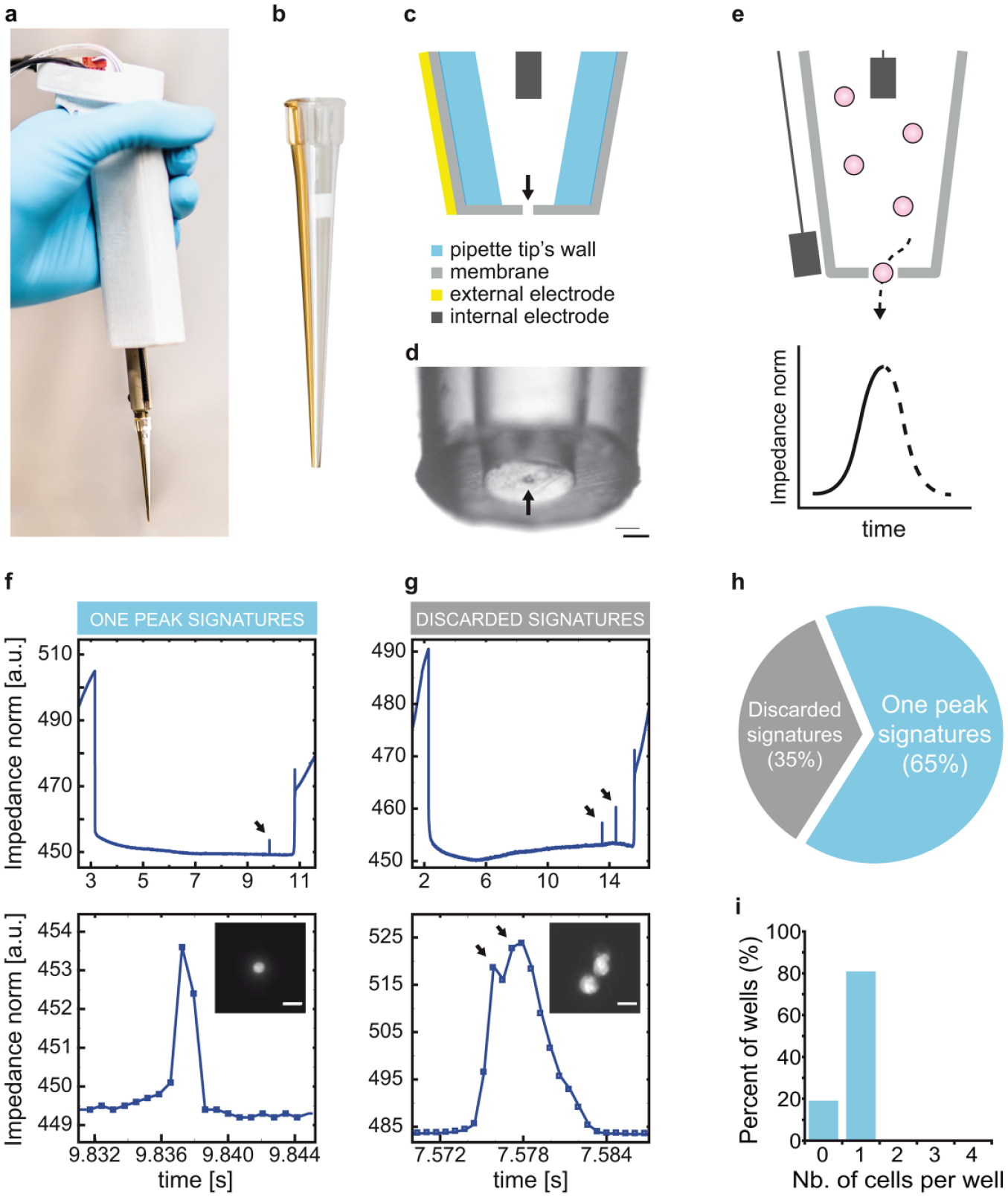
A method for the traceable isolation of living single cells. (
Methods
Pipet Setup
The cloning pipet was composed of a fluidic pump (MFCS-EZ; Fluigent, France), an impedance analyzer set at a sampling rate of 1444 Hz (SI-QSD; Sensima, Switzerland), and a control unit (in-house-developed printed circuit board). We described the technical characterization of the setup in a previous paper. 17 The data sent by the impedance analyzer were analyzed using a custom-made software (LabVIEW; National Instruments, Switzerland) that in turn controlled the micropump so that one cell was dispensed at a time. More details on the data analysis are also available for the reader in our precedent paper. 17 In parallel, we recorded the impedance signal at each dispensing procedure and analyzed it using MATLAB (R2014a; MathWorks, Natick, MA).
Fabrication of Sensing Tips
We modified the pipet tips (200 µL colorless; RatioLab, Germany) as described previously. 17 Briefly, we closed the extremity of each tip with a 15 µm thick membrane of medical-grade Parylene-C using chemical vapor deposition (Comelec C-30 S; Comelec, Switzerland). We opened a 30 µm aperture through the membrane by laser ablation (LightShot; Optec, Belgium). The inner electrode was made of a medical-grade stainless steel wire (316L; Sadevinox, France). We coated a 150 nm thick gold electrode on the external surface of the tips using evaporation through a shadow (EVA760; Alliance Concept, France). The tips were then assembled under laminar flow and sterilized by gamma irradiation (30K Gy).
Cell Preparation
We cultured murine fibroblasts (3T3-J2), human epidermal stem cells (strain YF29), human squamous carcinoma cancer stem cells (SCC1318), and rat hair follicle multipotent stem cells (clone YR219P3-cl719) as described elsewhere.5,20 We detached adherent cultured cells with trypsin/EDTA and filtered the cell suspension through a 40 µm cell strainer (BD) to avoid aggregates, which may clog the sensing tip. Then, we diluted the single-cell suspension to a final cell density of 104 cells/mL. We incubated 3T3-J2 cells with 1 µM calcein AM for 30 min (Thermo Fisher, Waltham, MA). The medium used in latter experiments was a Hank’s balanced salt solution (HBSS; Gibco, Thermo Fisher) supplemented with 3% bovine serum (BS; Gibco).
Recombinant Protein Production Assay
As a model, we used a CHO cell line engineered to express tumor necrosis factor receptor as an Fc fusion protein (TNFR:Fc). 21 We measured the concentration of the protein expression of the culture medium by sandwich enzyme-linked immunosorbent assay (ELISA) using a defined protocol. 21 We used a goat anti-human Fcγ antibody (Jackson ImmunoResearch Laboratories, Inc., West Grove, PA) for coating. Then, we used alkaline phosphatase (AP)-conjugated goat anti-human gamma chain antibody (Invitrogen Corp., Thermo Fisher) for detection. Finally, we used the enzyme p-nitrophenyl phosphate (New England Biolabs) as the substrate and measured the absorption at 490 nm on a microplate reader (SPECTRAmax 340; Molecular Devices, Palo Alto, CA). As a standard for quantification, we used a commercially available affinity-purified recombinant human TNFR:Fc fusion protein (Enbrel; Amgen, Zug, Switzerland).
Holoclone Assay
We performed holoclone assays as previously described. 5 Briefly, single cells were isolated individually into culture dishes. After 7 days of culture, we transferred each growing clone to two indicator dishes (one-fifth of each clone per dish). Twelve days later, we fixed and stained the dishes for classification of clonal types. When less than 0%−5% of the colonies on the indicator dishes was terminal, we classified the clone as holoclone. When more than 95% of the colonies was terminal, we scored the clone as paraclone; otherwise, it was classified as meroclone.
Long-Term Morphogenetic Assay
We performed animal experiments according to the Swiss legislation (authorization no. 2855; Canton de Vaud, Switzerland). Long-term skin reconstitution assays were performed as previously described. 19 Briefly, dissociated cultured cells from green fluorescent protein (GFP) rat whisker were injected at the dermoepidermal junction of a newborn mouse skin, forming a transplant. This transplant was then engrafted onto the back of a nude mouse and left for skin development and hair cycling for more than 100 days.
Tumorigenic Assays
Tumorigenic assays were performed as previously described. 18 Dissociated cells from human squamous cell carcinoma (SCC13) were injected subcutaneously into nude mice and left for development for more than 100 days.
Histology and Immunohistochemistry
Skin samples were harvested and fixed with 4% paraformaldehyde (PFA; Thermo Fisher) before being dehydrated and included into paraffin. Five-micrometer sections were realized with a Leica 2065 microtome (Leica, Germany). Staining of the SCC13 samples was performed according to Goldner’s trichrome protocol. 22 Enhance GFP (EGFP) immunohistochemistry was performed using chicken anti-GFP (ab13970; Abcam, Cambridge, UK) and goat anti-chicken Alexa488 (A11039; Molecular Probes) antibodies.
Results and Discussion
The procedure to isolate single cells is straightforward. The pipet is programmed to aspirate 20 µL of cloning medium (HBSS supplemented with 8% BS) containing 104 cells/mL. Next, the tip of the pipet is rinsed in medium and then gently immersed into the center of a cloning culture dish filled with cell-type-specific culture medium. The user then clicks on the user interface to command the isolation of a single cell and the recording of the impedance. As a single cell passes through the Coulter aperture to flow into the culture dish, it leaves an electrical signature that appears as a unique peak on the computer screen ( Fig. 1f ). Simultaneously, a sound is emitted to inform the user that a cell has been isolated. The recording of the impedance stops and the data are saved for traceability. The procedure is then repeated as many times as needed. At the end of the experiment, each impedance profile is magnified and examined in detail. A single and sharp peak is the signature of a single cell ( Fig. 1f ), whereas multiple peaks result from doublets, multiple cells, and aggregates ( Fig. 1g ). A small amplitude peak is the mark of debris. Culture dishes that do not meet the quality control criteria are rejected. In future projects, we will perform an in-depth characterization of the differentiating signals obtained for single cells versus cell aggregates, as clusters could be specifically crucial for key applications.
We tested the pipet performances using mouse embryonic fibroblasts (3T3-J2 cells) stained with a green fluorescent dye (1 µM, calcein AM). We repeated the cloning procedure described above 144 times during a 50 min period using 96-well plates. Ninety-four wells (65%) were determined to contain a single cell and 50 wells were eliminated because their impedance profile did not match the one peak signature ( Fig. 1h ). We then examined each positive well under an inverted fluorescent microscope for the presence of a fluorescent cell. A single cell was contained in 75 wells, whereas the other wells were empty ( Fig. 1i ). With this experiment, we demonstrated the practicality of the technology and the robustness of the impedance profile for quality control. In future experiments, we will specifically assess the impact of cell concentration on the efficiency of single-cell dispensing.
When it comes to the production of biotherapeutics, regulatory bodies require demonstrating that each new cell line derives from a single cell to ensure product safety and quality. 1 Because single-cell isolation methods are not fully traceable yet, companies waste up to 70 weeks in subcloning steps to ensure acceptable probabilities of monoclonality. 2 Hence, we used the cloning pipet to isolate impedance-based traceable clones from a cell line (CHO TNFR:Fc, GFP +) engineered to produce the GPF as well as a therapeutic protein. 21 We isolated 36 single cells, each of them traced by an impedance profile ( Fig. 2a ). In parallel, we isolated single cells into wells using limiting dilution. We examined these wells under a fluorescent microscope to identify 35 wells hosting a single GFP+ cell. Growing clones ( Fig. 2b ) were obtained with both procedures (9 clones vs 14 clones). Six clones of each group were inoculated at equal density and further expanded in culture. All clones showed high cell density and viability ( Suppl. Fig. S1 ) and produced the TNFR:Fc protein within the same range ( Fig. 2c ). Most importantly, this experiment showed that the cloning pipet provides an immediate proof of monoclonality that can be used to clone cells in one single round.
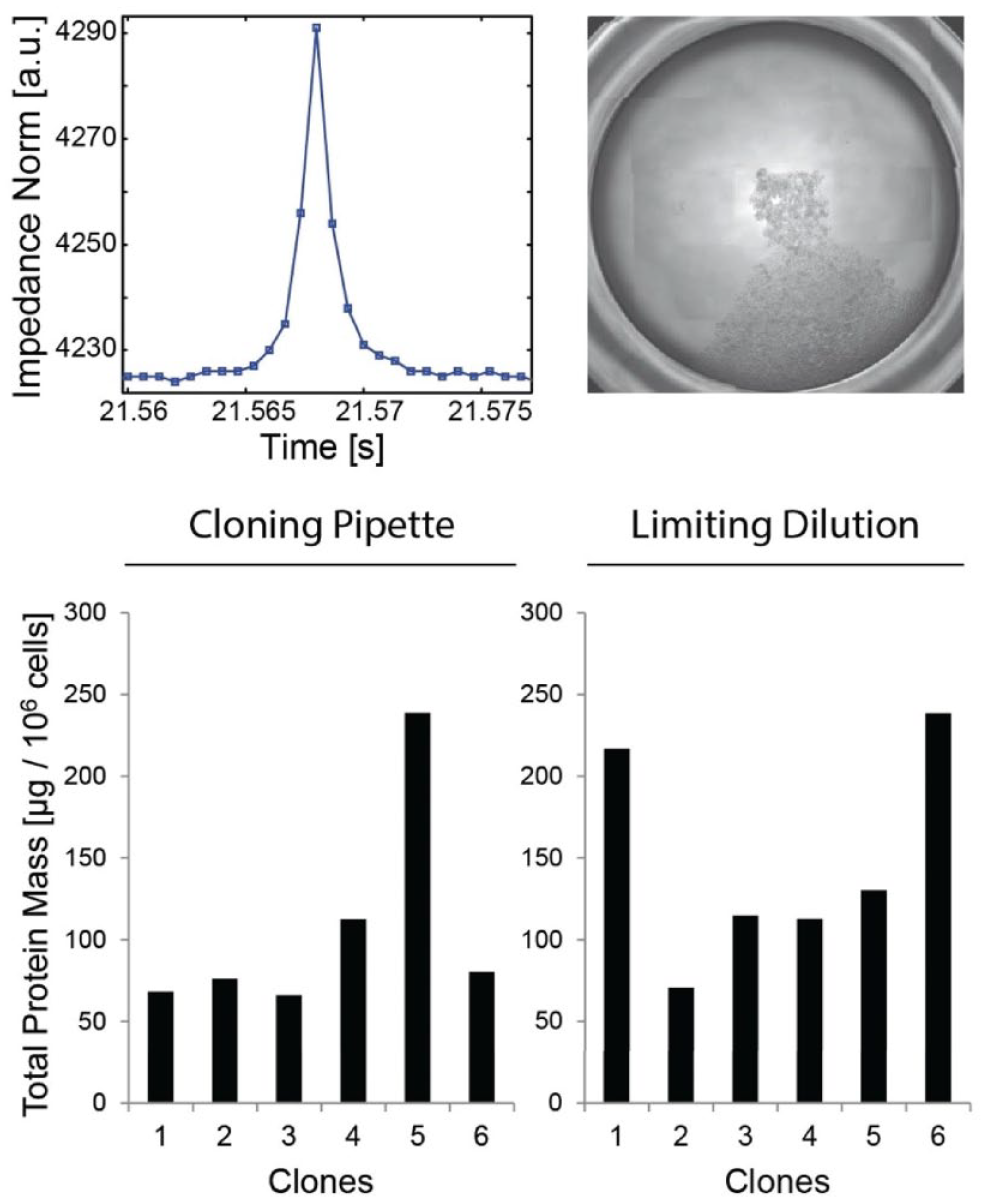
Single CHO cells isolated with the cloning pipet retain their growth and protein production capabilities. The performances of protein-expressing cells (cell line CHO TNFR:fc, GFP +) isolated with the cloning pipet are maintained. (
Human epidermal stem cells are used in regenerative medicine.23–25 Recently, we proposed a novel single epidermal stem cell strategy safe gene therapy. 26 When it comes to regenerative medicine, one of the main challenges is the handling of human epidermal stem cells in vitro as they are very sensitive to stress, to which they respond by loss of stemness, premature senescence, and differentiation.5,20 A holoclone is a stem cell and has enough growth capability to reconstitute the entire epidermis of an adult human.26,27 As of today, the holoclone assay is the best quality control to evaluate the presence of epidermal stem cells in vitro.22,28 This assay measures the growth capacity of a single epidermal cell. 5 In this assay, each single cell is individually isolated under direct microscopic vision using an elongated Pasteur pipet and individually cultured for 7 days before the clones are passaged. The entire cloning procedure from isolation to cultivation by mouth pipetting lasts less than 30 s for each cell and is extremely gentle. Consequently, high colony-forming efficiency is usually much better than that achieved by fluorescence-activated cell sorting (FACS). Hence, we challenged the cloning pipet against mouth pipetting in a holoclone assay to measure the stress of the cloning pipet on human epidermal stem cells. We isolated human epidermal stem cells (YF29) in parallel by mouth pipetting (72 cells) and with a cloning pipet (58 cells), with each cloning procedure being performed by independent operators. The three clonal types (holoclone, meroclone, and paraclone) were obtained with both procedures ( Fig. 3a ), even if the total number of clones was higher by mouth pipetting (26 clones vs 7 clones), most likely because of the mouth pipetting skills of the operator. Next, we evaluated the capabilities of single cells isolated with the cloning pipet in a transplantation assay developed for the formation of hair follicles using multipotent hair follicle stem cells of the rat (YR219P3-cl7). Seven clones were obtained out of 33 isolated single cells, and five were expanded in culture before being transplanted into developing mouse skin as described. 19 Two of the clones contributed to the generation of cycling hair follicles and sebaceous glands for months, as expected ( Fig. 3b ). Altogether, these experiments demonstrate that the cloning pipet does not impact stem cell properties.
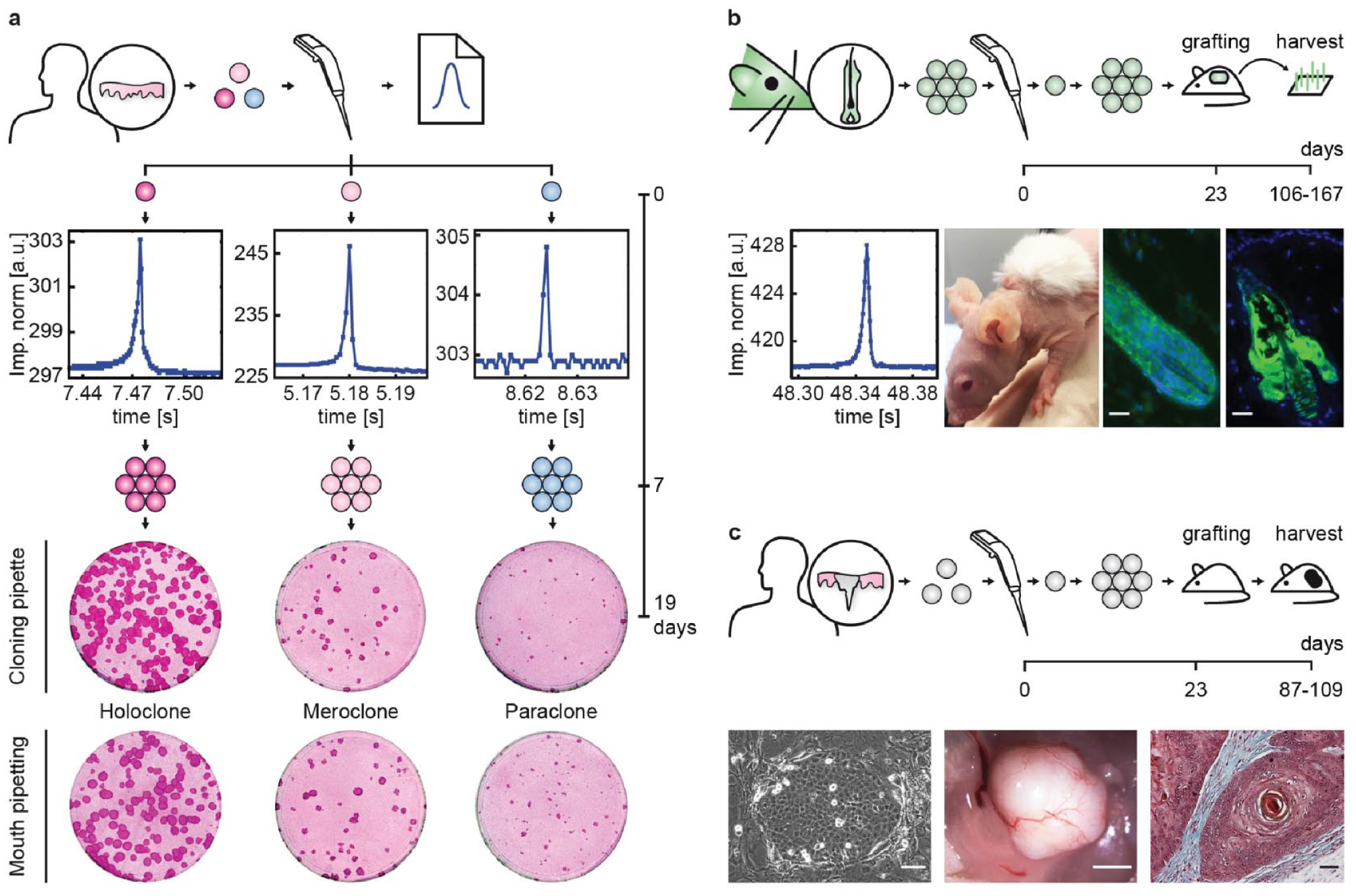
Single stem cells isolated with the cloning pipet retain their fundamentals. (
Single-cell cloning is also widely used to study cancer stem cells. Hence, we used the cloning pipet to clone single human squamous carcinoma cells from a cell line (SCC13) known to generate tumors when injected into immunodeficient mice. 18 Five clones were obtained out of 21 isolated single cells and three were expanded before being injected subcutaneously in athymic mice and observed for 109 days. All clones produced growing tumors as for the original cell line. The histology of the tumors was identical with the presence of keratin pearls ( Fig. 3c ). Collectively, these experiments demonstrate that single cells isolated by the cloning pipet maintained their fundamental properties, including stemness or tumorigenicity. This opens applications for use with patient-derived xenografts to better understand cancer heterogeneity and the metastasis process. Such studies would benefit from the isolation and cloning of single cancer cells, either from tumor chunks or from circulating tumor cells.29,30
In conclusion, we developed and validated an impedance-based procedure to isolate single stem cells using an engineered cloning pipet. The procedure is efficient, cell-friendly, and user-friendly. Above all, this technology is compliant with regulatory guidelines as it is disposable and provides a traceable record of the cell isolation, which both represent key features over other cloning procedures. For the next steps, we will focus on its validation for various applications, such as stem cell biology, biotechnology, and single-cell omics, among others.
Supplemental Material
Supplemental_Material_for_Traceable_Impedance-Based_Dispensing_by_Muller_et_al – Supplemental material for Traceable Impedance-Based Dispensing and Cloning of Living Single Cells
Supplemental material, Supplemental_Material_for_Traceable_Impedance-Based_Dispensing_by_Muller_et_al for Traceable Impedance-Based Dispensing and Cloning of Living Single Cells by Georges Muller, David Bonzon, Stéphanie Claudinot, Ariane Rochat, Philippe Renaud and Yann Barrandon in SLAS Technology
Footnotes
Acknowledgements
We are grateful to J. B. Bureau for his help on the microfabrication of the sensing tip and to N. Uffer and N. Beuchat for their help with the design of the software. We are thankful to D. Hacker from the Protein Production Core Facility at the EPFL for providing us with the expertise and materials to perform the recombinant protein production assay. We thank A. Bertsch and D. Forchelet for their advice and EPFL students C. Dayer, B. Charrez, C. Broenimann, N. Krischer, and N. Van Neghem for helpful discussions. We are grateful to A. Catana of the EPFL Technology Transfer Office for his continuous support and to the ENABLE program for funding interns F. Mattey, S. Béguin, N. Beuchat, and N. Uffer. We also thank M. Lany for providing us with state-of-the art ASIC and the staff from the Center of Micronanotechnology for its help with microfabrication of the tips.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: G.M., D.B., P.R., and Y.B. have financial interests in SEED Biosciences SA, a company commercializing the technology as well as intellectual property described herein.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work was supported by grants from the Ecole Polytechnique Fédérale de Lausanne to P.R. and Y.B. and from the Centre Hospitalier Universitaire Vaudois and the Fondation Enfants Papillons Switzerland to Y.B.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
