Abstract
A digital assay is one in which the sample is partitioned into many small containers such that each partition contains a discrete number of biological entities (0, 1, 2, 3, …). A powerful technique in the biologist’s toolkit, digital assays bring a new level of precision in quantifying nucleic acids, measuring proteins and their enzymatic activity, and probing single-cell genotypes and phenotypes. Part I of this review begins with the benefits and Poisson statistics of partitioning, including sources of error. The remainder focuses on digital PCR (dPCR) for quantification of nucleic acids. We discuss five commercial instruments that partition samples into physically isolated chambers (cdPCR) or droplet emulsions (ddPCR). We compare the strengths of dPCR (absolute quantitation, precision, and ability to detect rare or mutant targets) with those of its predecessor, quantitative real-time PCR (dynamic range, larger sample volumes, and throughput). Lastly, we describe several promising applications of dPCR, including copy number variation, quantitation of circulating tumor DNA and viral load, RNA/miRNA quantitation with reverse transcription dPCR, and library preparation for next-generation sequencing. This review is intended to give a broad perspective to scientists interested in adopting digital assays into their workflows. Part II focuses on digital protein and cell assays.
Introduction
For the purposes of this article, we define a biological assay as the quantification of the concentration or activity of a biological entity in a sample container. A digital assay is one in which the sample is first partitioned into many small containers such that each partition contains a discrete number of biological entities (0, 1, 2, 3, …). The term
Digital versus Analog Assays
Figure 1
compares a conventional analog assay with a digital assay. Analog assays, typically performed in a vial or a microplate, provide a readout signal
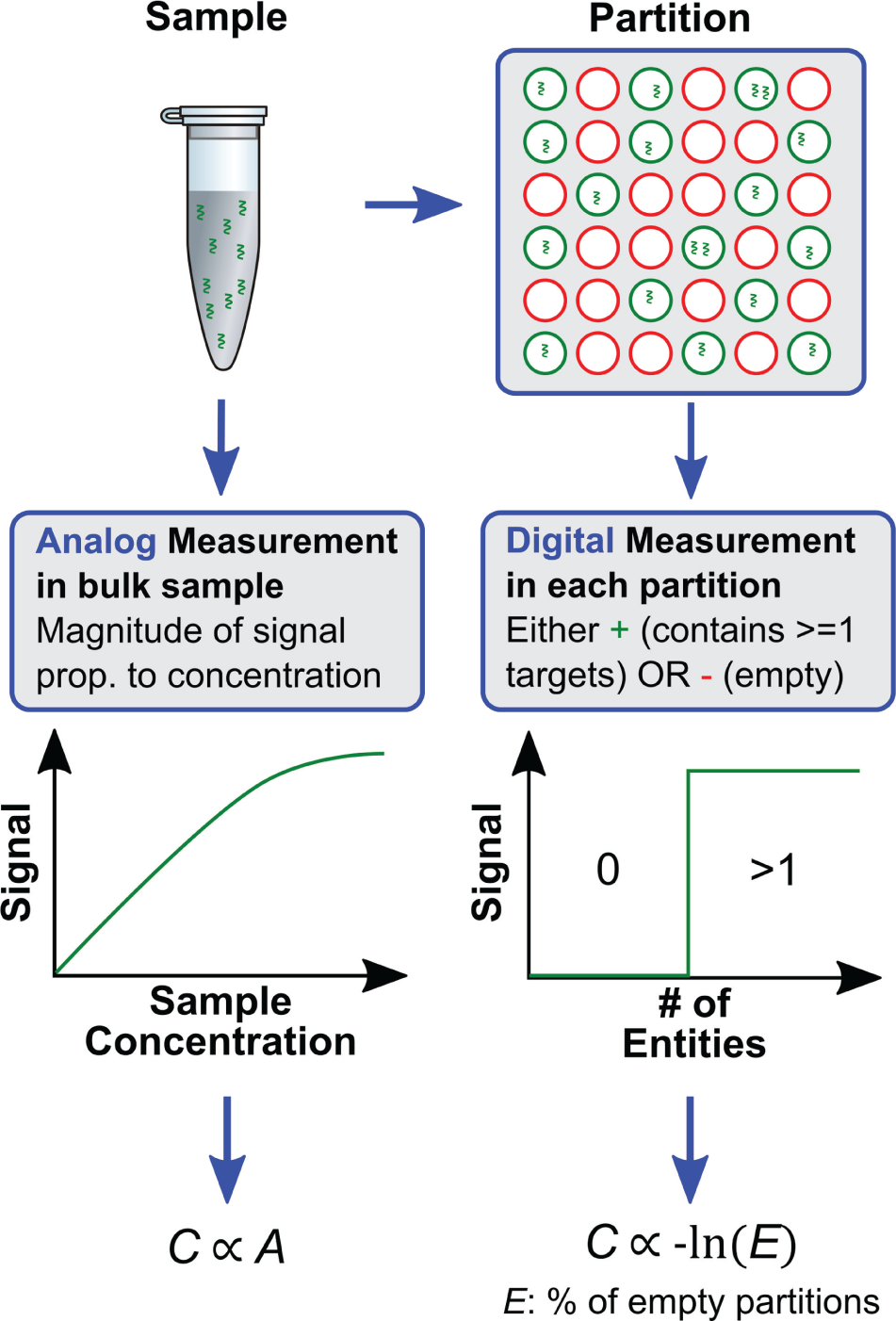
Comparison of digital assays vs. conventional analog biological assays.
The term
Benefits of Partitioning
Partitioning provides several benefits to digital assays over their conventional counterparts, particularly as the number of partitions increases and the partition volumes become small. Two notable effects of partitioning are illustrated in Figure 2 . The first of these is the concentration effect: the limit of detection (LOD) is improved because a small reaction volume increases the effective concentration of the target molecule. 5 This makes it possible, for example, to amplify and detect single molecules of DNA6–8 and single-cell secretions to the extracellular environment.9–11 The second of these, the enrichment effect, improves the analysis of complex mixtures by purifying the target of interest from interfering compounds. More specifically, it increases the ratio of the target of interest versus the background. In an illustrative example ( Fig. 2B ), partitioning results in 8× purification in one replicate, and complete purification in another replicate. In digital PCR (dPCR), the enrichment effect improves amplification efficiency of low-abundance mutant nucleotides against a wild-type background, which is useful for applications like cancer diagnostics. 12 The concentration effect scales inversely with sample volume, while the enrichment effect scales with the number of partitions. A third benefit of partitioning is that a digital assay provides superior precision and linearity when counting copies of targets, simply because it measures individual molecules rather than an ensemble concentration. Precision improves with the number of partitions, since more single entities can be resolved within a large population. Finally, increasing the number of partitions extends the dynamic range, that is, the range of sample concentration that can be measured.
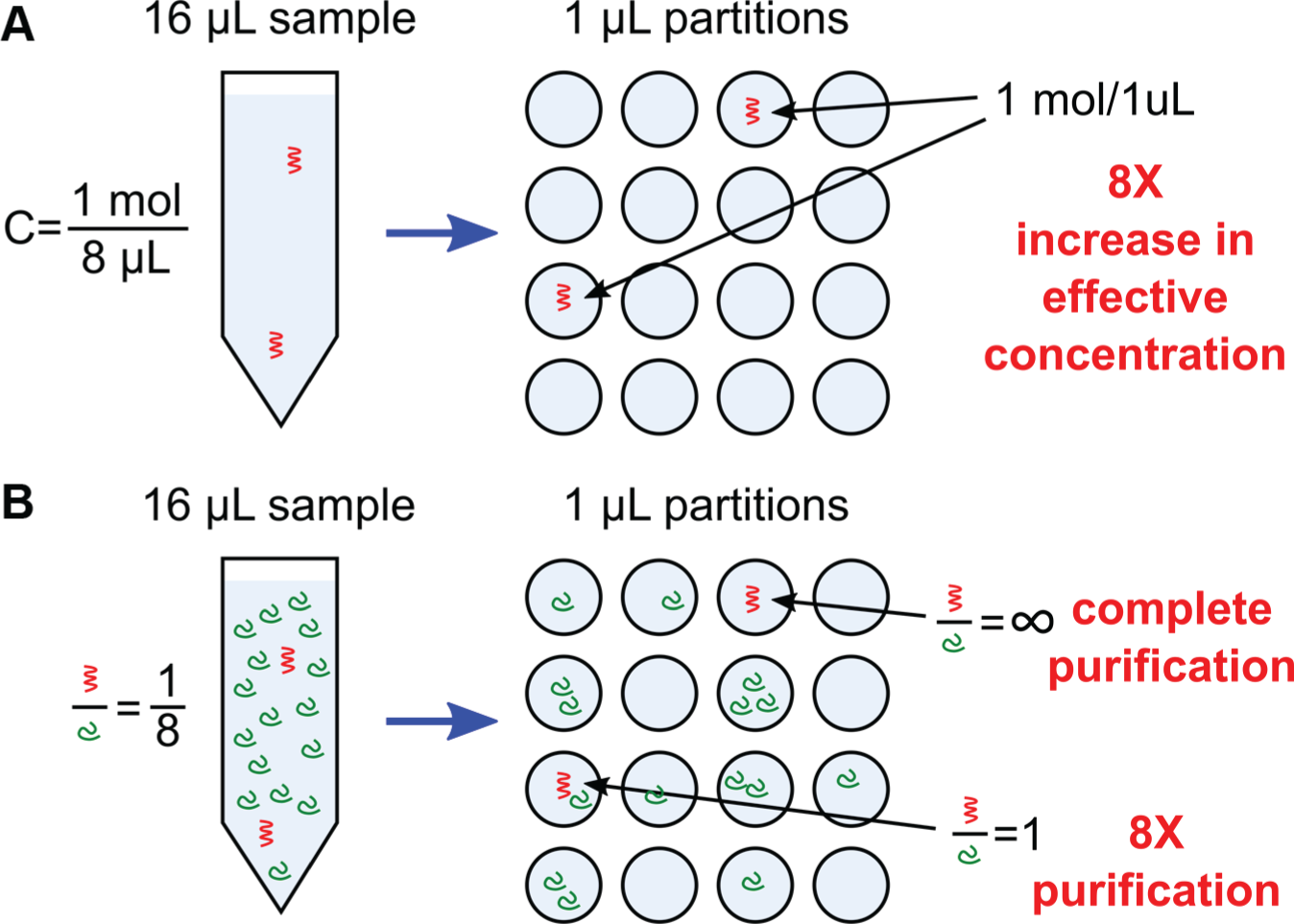
Benefits of partitioning in digital assays. Shown are two illustrative examples: (
Partitioning Methods: Chambers and Droplets
Partitioning is accomplished using two general approaches: (1) arrays of physically isolated chambers or wells or (2) droplet emulsions where the reaction is conducted in water droplets separated by a continuous oil phase. Either approach can provide partition volumes ranging from femtoliters to almost microliter volumes.
In the case of chamber arrays, digital assays have been conducted in commercially available 96-, 384-, or 1536-well plates. However, as described above, scaling to more wells improves the dynamic range and the ability to enrich mutant samples. Scaling can be achieved by miniaturizing the wells using photolithographic and microfabrication processes. For example, photolithography and silicon etching have been used to fabricate 50,000 wells for digital enzyme-linked immunosorbent assay (ELISA), each with a 50 fL volume. 13 Similar platforms for performing digital enzyme assays, with wells designed to accommodate 1.4 fL volumes, have also been fabricated by micromolding a flexible elastomer (PDMS) against a 1 µm silicon structure. 14 Chamber arrays can also be made in plastics using micromolding 15 or imprint lithography,16,17 and in other materials using variants of soft lithography. 18 The resolution of the manufacturing process determines volume variability. Using photolithography, tolerances can be as small as hundreds of nanometers, and with nanoscale imprint lithography, they can be <25 nm. Volume variability contributes to uncertainty in digital counting assays, as described in the next section.
The second approach for partitioning utilizes droplet emulsions in oil. Monodisperse droplets can be made using microfluidic t-junctions where an aqueous stream is combined with a stream of oil, 19 or by flow-focusing cross-geometries where the aqueous stream is combined with two orthogonal streams of oil. 20 The oil should be nonreactive and have low solubility to prevent the diffusion of reactants between reactors; often, fluorocarbon oils are chosen for this purpose. Stable droplet generation requires the addition of surfactants to stabilize the water oil interface and prevent droplet coalescence.21–24 The surfactant must be optimized so that it does not interfere with the assay, and keeps small biomolecules from diffusing into the oil. For more details on droplet microfluidics, the reader is referred to several excellent reviews on the subject, including a comprehensive 2012 review by Guo et al. 25 and a 2010 review by Theberge et al. 26 Tran et al.’s 2013 27 and Teh et al.’s 2008 28 reviews cover technical developments in droplet microfluidics technology, including droplet processing components and their integration in laboratory workflows. Lagu et al.’s 2013 29 and Zagnoni and Cooper’s 2011 30 reviews focus on the application of droplet microfluidics for single-cell biology, highlighting the physics of droplet formation and manipulation, cell encapsulation methods, a variety of sensing and sorting methods, and the use of confinement in single-cell analysis.
Each of the two approaches (chambers or droplets) has its respective benefits. In general, the advantage of droplet systems is that they can generate as many partitions as needed. Droplet dPCR (ddPCR) systems, for example, can provide as many as 10 million partitions. However, droplets are generated serially, which requires minutes to hours to generate sufficient partitions. Step emulsification systems address this issue by massive parallelization, allowing them to generate tens of thousands of droplets in minutes while maintaining relatively uniform droplet size.31,32 The volume uncertainty in droplet systems is due to the polydispersity of droplet generators, usually <5%, while volume uncertainty for lithographically defined chambers is determined by the manufacturing tolerances, as described above. For example, commercial dPCR systems using chamber and droplet implementations typically have <3% uncertainty. 33 In either implementation, care must be taken to avoid adsorption of biomolecules to the interfaces, which can lead to sample loss or enzyme deactivation. Physical chambers can be coated with blocking agents (e.g., bovine serum albumin) to prevent the adsorption and deactivation of biomolecules. 14 Similarly, droplet systems can utilize amphiphilic surfactants, which present a blocking group such as polyethylene glycol (PEG) facing the aqueous side, and a hydrophobic tail facing the oil side.22,24,34
Organization and Scope of This Review
This two-part review article will cover the most popular types of digital assays, including nucleic acids (dPCR), proteins (digital ELISA and enzymatic activity), and cells (single-cell gene expression, proteomics, and transcriptomics). The large variety of assays is accompanied by an equally wide range of applications, each of which will be covered in the respective sections. Part I covers partitioning statistics, and then moves on to dPCR concepts, systems, and applications. Part II 35 discusses digital protein assays, digital cell assays, and their respective applications. Given the readership of SLAS journals, the intended audience is the scientific community using and developing laboratory automation. Accordingly, the article covers fundamental concepts, the methods and types of digital assays reported in the literature, commercial products, and future trends.
Partitioning Statistics
In digital assays, partitioning statistics are used to determine sample concentrations and confidence intervals. Partitioning the sample into small containers results in a statistical distribution of targets (DNA molecules, proteins, or cells) among the partitions (
Fig. 1
). Partitioning is similar to aliquotting, a common step in a laboratory workflow; however, the latter typically involves such a large number of targets per aliquot that the statistical variation between samples is ignored. By contrast, in digital assays, the number of targets in the sample (m) is typically less than or on the order of the number of partitions (
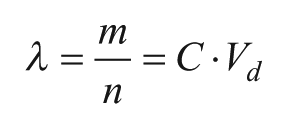
Here, m is the number of targets in the sample and Vd is the partition volume.
The probability that a partition will contain
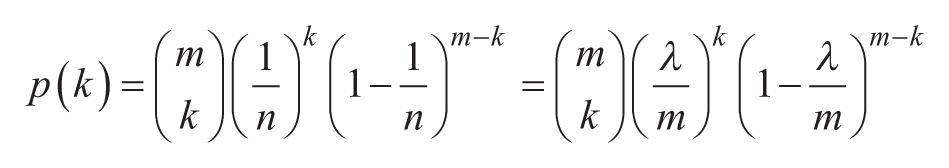
Digital assays typically employ a large number of partitions. When
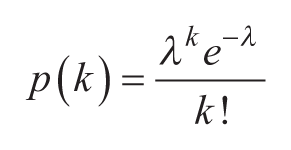
The mean and variance of the Poisson distribution are both equal to
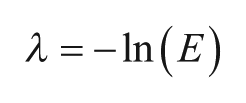
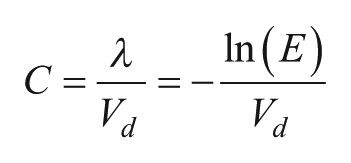
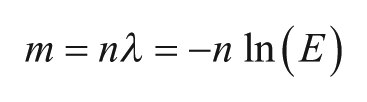
Figure 3
shows calculated Poisson distributions for various values of sample concentration l. As l increases, the proportion of empty partitions falls to zero, making quantitation impossible. To statistically have at least one empty partition (nE > 1), the number of targets must be less than
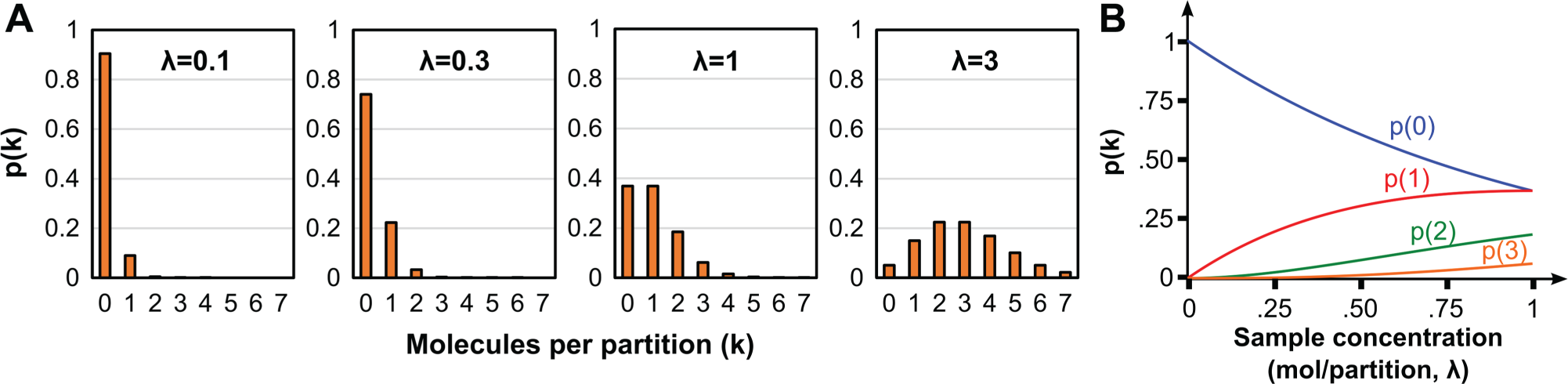
Poisson encapsulation statistics. (
Sources of Error
In conventional analog assays, measurement uncertainty is typically limited by the resolution of the detection instrument ( Fig. 1 ), for example, the ability of a fluorimeter to resolve small differences in emitted fluorescence. In digital assays like dPCR, the detection instrument needs only to identify whether a partition has 0 or >1 targets, which makes data analysis less dependent on the properties of the detector or assay chemistry. Digital assays have two sources of error: subsampling error and partitioning error. 5 Subsampling error sets the lower detection limit at low concentrations and is independent of the instrument, while partitioning error dominates at high concentrations and may depend on the sampling and partitioning instrument.
Subsampling errors arise in any biological assay (digital or analog), which does not analyze the full volume of the sample, but rather a subsample of it, resulting in statistical variation between replicate tests. For example, a diagnostic sample like blood serum could be several milliliters, while a typical dPCR system can handle only 20 µL samples. The subsampling process introduces an unavoidable source of error, particularly when there are few targets within the original sample to detect. When subsampling a fraction of a larger sample, the standard deviation of the targets per subsample is
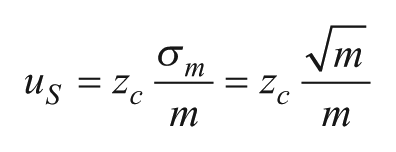
Partitioning errors, which are specific to digital assays, occur because the distribution of targets among partitions may differ from one experiment to the next. In a set of replicate experiments, there is a variance in the number of empty partitions E that propagates to a corresponding variance in the calculated concentration l.
5
An estimate of partitioning error can be found based on the analysis of Dube et al.,
39
which models the partitioning as a binomial process. The standard deviation in the proportion of negative partitions E is
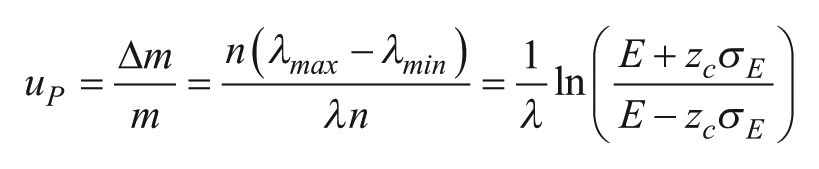
This variation becomes significant at high concentrations (large l), when very few partitions are empty (
Figure 4
compares the relative contributions of subsampling and partitioning errors versus sample concentrations at 20,000 and 1 million partitions. At large l, partitioning error limits the maximum concentration that can be measured, while both errors increase at l < 1. In practice, the sample concentration is chosen to minimize error. For example, in dPCR, the sample concentration is typically adjusted such that l ranges from 0.001 to 65,38 (and in some cases, up to 8
40
). At
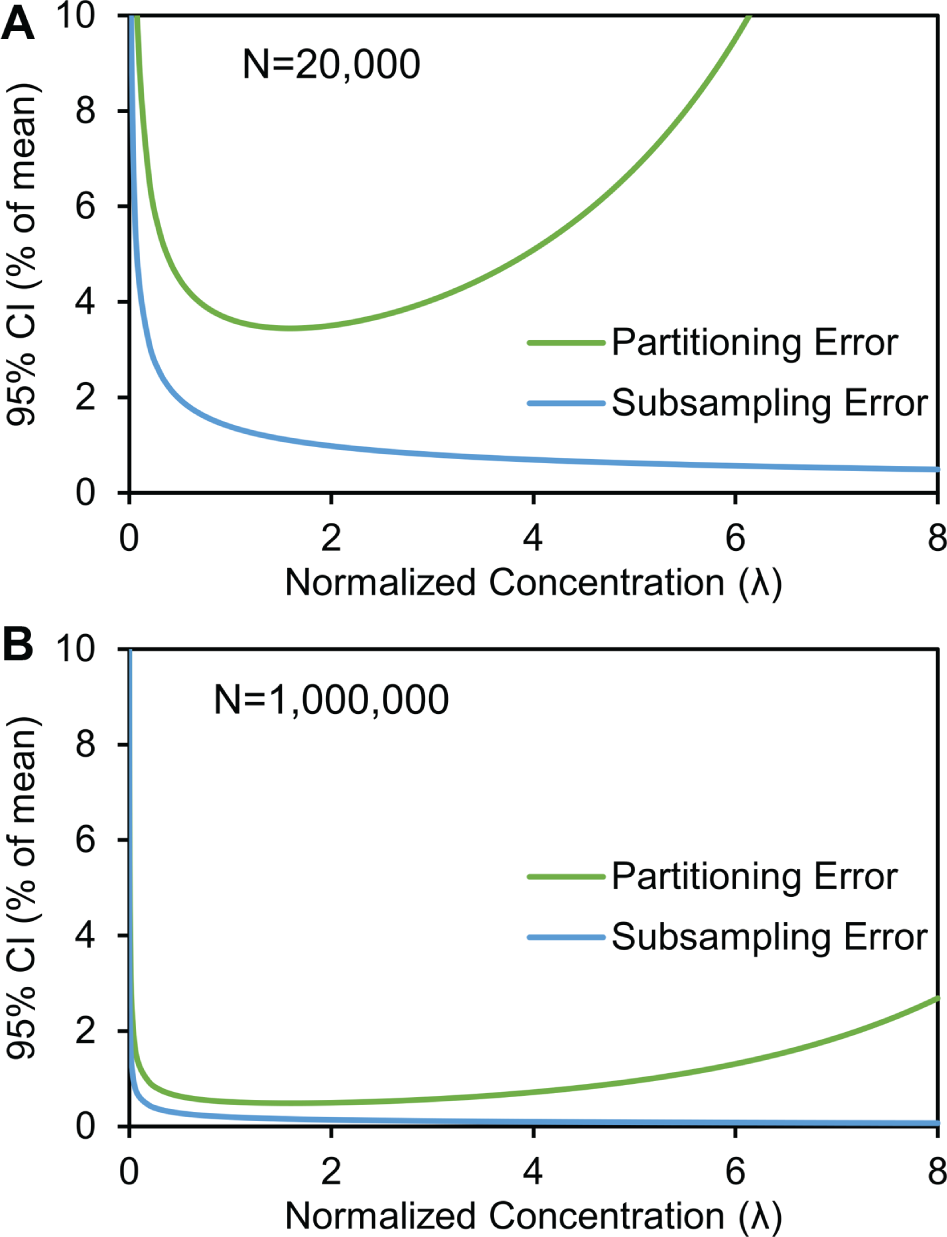
Subsampling errors and partitioning errors in digital assays assuming (
The Poisson model does not take into account variation in partition volume, which can skew the distribution of targets, particularly at high concentrations.12,41 As described by Pinheiro et al.,
42
the volume uncertainty uVd
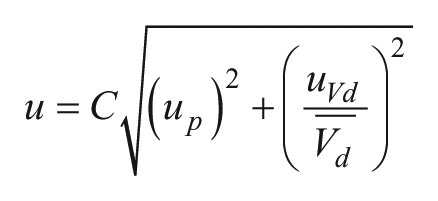
Here,
Digital PCR
The most mature digital assay, digital PCR (dPCR) is a method for absolute quantification of a nucleotide sequence.
5
Although the first paper dates back to 1999,
43
dPCR did not gain widespread use until the first commercial instruments became available in 2007.
44
dPCR is based on the same biochemical principles as quantitative real-time (qPCR), but differs in how quantitation is performed. In qPCR, a sample containing target DNA, PCR reagents, and fluorescent probes is thermally cycled, while monitoring fluorescence at each cycle. During each replication cycle, hydrolysis probes (e.g., Taqman) or intercalating DNA binding dyes (Evagreen or SYBRgreen) exponentially increase fluorescence emission at each cycle, with a rate dependent on amplification efficiency (
Fig. 5
). Hydrolysis probes release fluorescence when cleaved during each replication cycle, while DNA intercalating dyes increase fluorescence emission when bound to double-stranded DNA. Hydrolysis probes typically offer higher specificity, sensitivity, and quantitation. In all cases, the number of cycles required to achieve a detectable fluorescence,
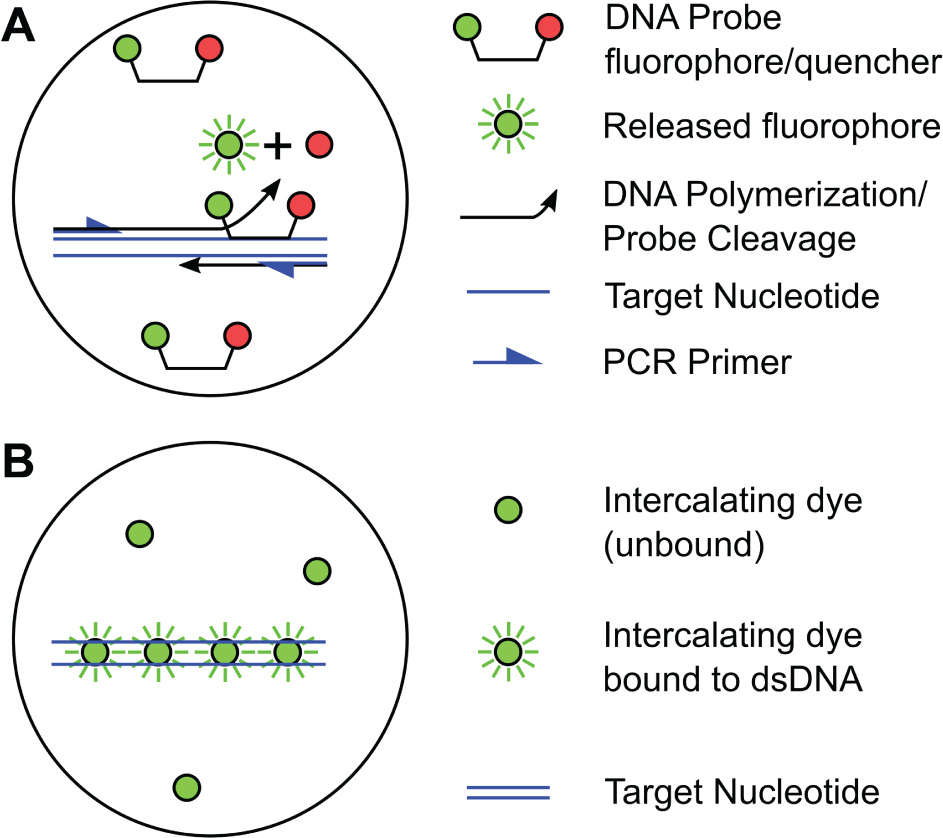
Amplification and detection chemistry in dPCR (same for qPCR). (
In dPCR, the detection chemistry is essentially identical; however, the sample, reagents, and probes are first partitioned into many replicate reactions (
Fig. 1
), with commercial systems generating between 1000 and 10 million partitions.
43
The partitions are then subjected to excess rounds of thermal cycling (>30), so that any partition that contains one or more DNA targets becomes fluorescent, regardless of amplification efficiency. Then, the proportion of nonfluorescent partitions, through Poisson statistics described above, is used to calculate the target concentration within a well-defined confidence interval.5,39,45 To have a sufficient number of empty partitions, high-concentration samples must first be diluted so that
Digital PCR Systems
Figure 6
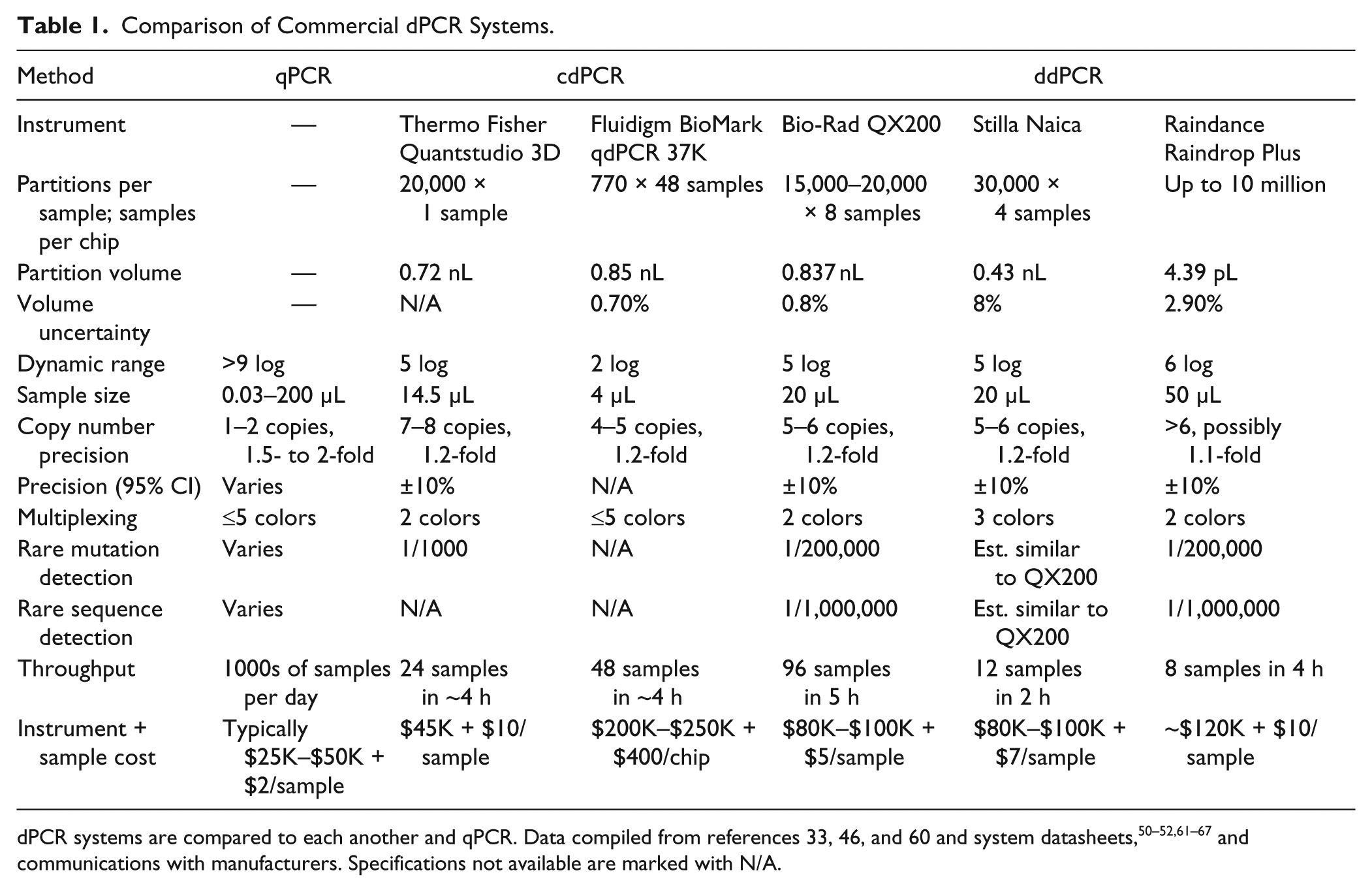
and
Table 1
compare five commercially available dPCR systems, which fall into two categories, depending on the way they partition and detect samples.
46
The first category, chamber-based digital PCR (cdPCR) systems, utilize physical structures as partitions. Vogelstein, who coined the term
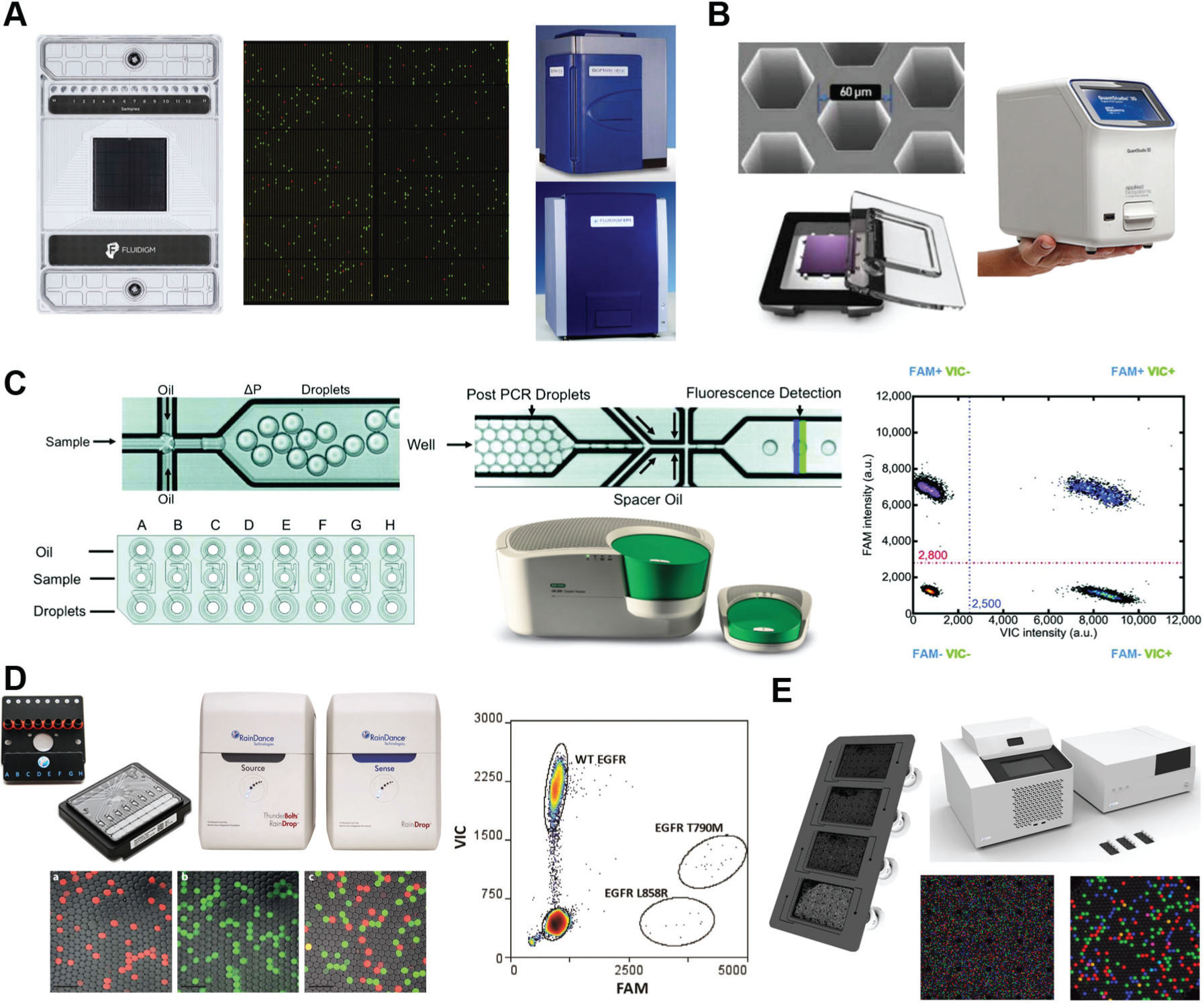

Commercial dPCR systems. (
The second category, ddPCR systems, performs PCRs in water-in-oil droplet emulsions, as described in the introduction section. The PCR mix in each droplet must also include PCR-compatible surfactants, often proprietary, to stabilize the emulsion and ensure chemical isolation during amplification. 22 The first report of ddPCR in 2007 49 demonstrated single-copy detection limits using Taqman hydrolysis probes in 10 pL droplets within 18 cycles. This technology was spun out from Lawrence Livermore Labs into QuantaLife, a start-up that developed the first commercial ddPCR system and eventually sold it to Bio-Rad. Bio-Rad’s current offering ( Fig. 6C ) is the QX200 ddPCR system, which generates up to 20,000 droplets with volumes of ~1 nL.40,50 Raindance Technology’s Raindrop system ( Fig. 6D ) 51 generates up to 10 million droplets with volumes of 5 pL, utilizing an array of eight parallel drop generators to increase throughput. Raindance was acquired by Bio-Rad in January 2017. The Stilla Naica System for Crystal Digital PCR 52 ( Fig. 6E ) utilizes step emulsion droplet generators,32,53,54 which can be massively parallelized to hundreds of nozzles; however, Stilla’s current product focuses on a moderate number of droplets (30,000) with short workflow times (2 h) and increased readout capabilities (three fluorescence channels). A unique feature of step emulsion generators is that they do not require the flow of oil; hence, in the Stilla system, the oil is preloaded into the Sapphire chip, simplifying operation and reducing potential contamination.
It is evident that compared with cdPCR systems, ddPCR can readily scale to more partitions in a cost-effective manner. As mentioned earlier, increasing the number of partitions provides several benefits: First, adding partitions proportionally increases dynamic range, so a larger range of samples can be accommodated without dilution. Second, it improves the ability to detect rare targets in the presence of similar nucleotide sequences or inhibitors, due to the enrichment effect ( Fig. 2B ). This is helpful in detecting single-nucleotide polymorphisms (SNPs) and other rare alleles in a largely wild-type population, such as circulating tumor DNA (ctDNA) (to be discussed later). Third, it is better at resolving copy number variations (CNVs), particularly when the target concentration is low. 55
The commercial systems also differ in how they monitor fluorescence during PCR, either by fluorescence imaging or laser-induced fluorescence (LIF) cytometry. The former approach is used in the Fluidigm BioMark system, which is unique in that it can measure fluorescence at each partition after each cycle. The BioMark system can therefore operate in qPCR or dPCR mode. Thermo Fisher’s Quantstudio 12K, a predecessor to the Quantstudio 3D, also provides both capabilities. Real-time, intracycle monitoring on all partitions may help to eliminate false positives in dPCR, but limits the maximum number of supported partitions. Quantstudio 3D and Naica 52 also use fluorescent imaging, but perform only a single-endpoint fluorescence measurement upon the conclusion of all cycles. Fluorescence imaging, when extended to a wide field of view, can analyze as many as 1 million droplets simultaneously. 56 The second approach, LIF droplet flow cytometry, 57 is used by the ddPCR systems from Bio-Rad and Raindance, which serially analyze droplet fluorescence using flow cytometers operating at hundreds or thousands of drops per second. LIF can achieve higher sensitivity and dynamic range than imaging, but its serial nature reading requires more time.
Multiplexing capabilities, offered on all systems to varying degrees, are useful for oncology and food analysis, where multiple targets are quantified simultaneously. The QX200, Raindrop, and Quantstudio 3D support two fluorescence channels at nonoverlapping spectral wavelengths, and Naica provides three. One way to increase the number of fluorophores is amplitude multiplexing, where probes of substantially different intensities can be simultaneously used on the same fluorescence channel. 58 In traditional flow cytometry, amplitude multiplexing comes at the expense of dynamic range, but in digital assays the trade-off is less severe because the fluorescence is used only for counting and not for direct analog quantification. The Raindance system claims to multiplex up to 10 different probes by using 5 amplitude multiplexed probes on two fluorescence channels. For example, a 7-plex assay was used to quantify common mutations in oncogenes. 59 Although not officially offered, other systems in principle should also be able to achieve amplitude multiplexing to various degrees.
Comparison of dPCR with qPCR
Since the transformational invention of PCR in 1983, nucleic acid quantitation has evolved from a qualitative technique (PCR + electrophoresis), to semiquantitative (qPCR), and now, with the advent of dPCR, to absolute quantitation. Compared with its most recent predecessor, qPCR, dPCR provides benefits in three areas. 46
On the other hand, qPCR still wins in other areas.
Other aspects of qPCR and dPCR systems to consider include the following:
Applications of Digital PCR
Based on the comparisons above, dPCR is best suited in applications where the key requirements are precise and accurate quantification and the ability to detect rare mutants or sequences. On the other hand, qPCR is preferable when dynamic range or throughput is the driver. Examples of dPCR applications ( Fig. 7 ) include CNV for genotyping genetic diversity and disease processes, detection of ctDNA, quantification of viral load and low-level pathogens, and quantification of NGS libraries. Each is discussed in turn below.
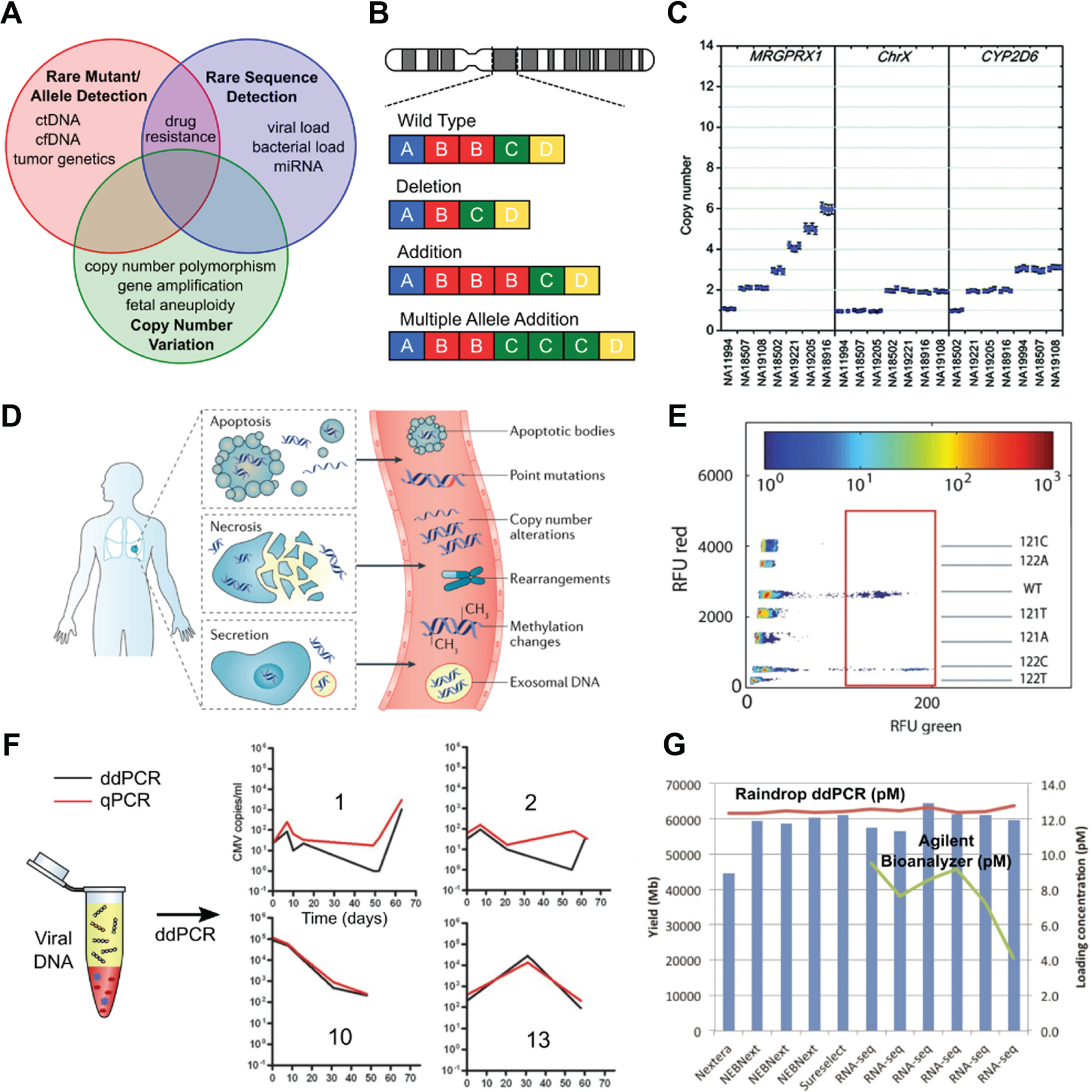
Applications of dPCR. (
Copy Number Variation Genotyping
CNVs (
Fig. 7B
) are sequences of >1 kb pairs that are present in variable copies in as much as 6%–19% of the human genome,75–77 as well as plants.
78
CNVs are considered structural alterations of the genome, and may include the deletion or addition of entire genes, along with their regulatory regions. A rich source of genetic diversity among individuals, CNVs can alter gene dosage, modify protein coding regions, unmask recessive alleles, and influence a variety of complex disease phenotypes,
75
including susceptibility to HIV, autism, and schizophrenia, as well as the process of cancer metastasis.
79
For example, fewer copies of the
Fluidigm’s BioMark system (
Liquid Biopsy: Detecting Circulating Tumor DNA
dPCR can be used for liquid biopsy,
91
a cancer diagnostic that measures ctDNA released into a patient’s bloodstream by tumor apoptosis, necrosis, or secretion
92
(
Fig. 7D
). Less invasive than direct biopsy, liquid biopsy is particularly attractive when the tumor is not accessible. Several somatic mutations in ctDNA (
A challenge in liquid biopsy is that ctDNA is typically <1% of fragmented DNA in the bloodstream. 71 Moreover, only a fraction of the ctDNA (<10%) is mutated, depending on how advanced the cancer may be. 71 A hallmark of dPCR is its ability to detect rare mutants by partitioning, which enriches the mutant relative to the wild type. ddPCR systems from Raindance and Bio-Rad can resolve rare mutants (RMD) at frequencies as low as 0.0005%,72,73 compared with 1% for qPCR and 2% for NGS. 91 Furthermore, while NGS can only detect if a mutation is present, dPCR can quantify it as well, enabling one to monitor the progression of the disease. When measuring ctDNA, the fact that tumor DNA is as little as 5% makes it proportionally more challenging to resolve subtle changes. The high-copy-number resolution of dPCR (as little as 1.1) is an added benefit in diagnostics that require longitudinal monitoring, such as viral load.
Several studies have shown promising results, beginning with a clinical breast cancer study that detected ctDNA using the Fluidigm BioMark system.
96
The prevalence of the
Infectious Disease: Rare Sequence Quantification of Viral or Bacterial Load
Precise quantification of circulating viral or bacterial DNA is critical not only for early diagnosis, but also to monitor the progression of infection and response to treatment. 99 HIV patients undergoing retroviral therapy are regularly monitored for residual infection, and immunosuppressed organ transplant patients are proactively screened for cytomegalovirus (CMV), Epstein–Barr virus, adenovirus, and others. Early diagnostic approaches for identifying pathogens relied on light microscopy; however, this approach was laborious and could not identify small cells like viruses. Since the 1990s, the gold standard has become qPCR, which quantifies viral or bacterial load by measuring circulating DNA. However, in qPCR, the construction of calibration curves and amplification efficiency can vary greatly from lab to lab, resulting in as much as 20%–30% CV at low template concentrations. 99 dPCR provides (1) absolute quantification of target sequences, (2) in some cases, lower CVs at low concentration, and (3) tolerance to PCR inhibitors like sodium dodecyl sulfate (SDS) and heparin.69,100 Sedlak and Gerome’s 2013 and 2014 reviews99,100 outline the use of dPCR in viral and microbial diagnostics, including the measurement of occult RNA, HIV-1, and adenovirus using commercial dPCR products.
In 2008, an early variant of the Raindance platform was used to amplify and detect spiked-in adenovirus solutions with a LOD as low as 1 template per 167 droplets, or 92 per microliter.
101
This chip used continuous-flow ddPCR where droplets shuttle between hot and cold zones during amplification, and cross “neck-down” regions where their fluorescence is detected. In 2013, Strain et al.
102
used the Bio-Rad QX100 system to quantify HIV
Quantitation and Preamplification of Next-Generation Sequencing Libraries
NGS begins with a high-quality DNA library in precise quantities. Constructing an NGS library (e.g., Illumina) includes several steps: fragmentation of the genomic sample, size selection, end repair, adapter ligation, PCR enrichment, and quantitation. Of these, dPCR has proved to be useful in evaluating the integrity of size selection, enrichment, and quantitation. Using the Raindance system, Didelot et al. 106 used ddPCR to quantify the size distribution and integrity of DNA fragments. They performed multiplex ddPCR with four primer probes, each targeted to different lengths of DNA. In doing so, the approximate distribution of nucleotide sequence lengths could be digitally quantified by counting the number of amplified species resulting from each probe. When the same distribution is analyzed using qPCR, it has CVs on the order of 10%–50%. After library construction, a portion of the genomic sample is selected for sequencing. Targeted sequencing often requires unbiased amplification of predesignated loci, usually by PCR with a selected primer library. The common issue of biased amplification can be addressed by dPCR. Partitioning fragmented DNA into small containers effectively enriches the target sequences against background, thus improving the amplification of rare sequences or those difficult to amplify. Using a variant of the Raindance system, Tewhey et al. 107 amplified nearly 4000 targets simultaneously using a highly multiplexed primer library. First, multiple droplet libraries containing each of the primers were generated and pooled. Then, the primer library drops were serially merged with droplets containing the genomic sample. After PCR, the droplet emulsion was broken, and the amplified DNA was enriched and sequenced with high specificity and sensitivity. The last but important use of dPCR is quantitation. To achieve high-yield and high-quality sequencing, NGS sequencers require a relatively narrow range of DNA loading capacity (e.g., Illumina suggests 60 gigabases per lane 108 ). Too little DNA will result in poor coverage, low read depth, and failure to detect SNPs or rare sequences. Too much will result in overclustering, where all sequences may not be resolved. DNA quantitation with spectrophotometry or fluorescence is qualitative and has limited sensitivity, requiring more than 1000× more sample than the sequencer itself, and thus wasting precious sample. 109 Using the Fluidigm dPCR chip with universal adapter primers, White et al. demonstrated absolute quantitation of as few as 100 template molecules, and was able to sequence with nanogram-scale libraries rather than microgram ones. 109 In another example, quantitation by the Raindance ddPCR system correlated more closely with sequencing yield compared than the Agilent Bioanalyzer, which uses capillary electrophoresis and fluorescence 108 ( Fig. 7G ). In this study, 11 samples were sequenced using an Illumina HiSeq 2500. Prior to loading, the DNA was quantified using both methods, and 60 GB was loaded onto each lane based on the ddPCR quantitation. The ddPCR provides more accurate and precise quantitation, which correlates more closely to yield than the Bioanalyzer; however, it should be noted that the Bioanalyzer also provides sizing data. Postsequencing, ddPCR can be used to validate any discovered target sequences. Due to the excellent quantitation and amplification capabilities of dPCR, it is rapidly becoming a standard tool in NGS.
Quantifying mRNA through Reverse Transcriptase dPCR
By adding a reverse transcription step, dPCR can quantify RNA levels with virtually all the benefits of digital partitioning. Of particular interest is the detection of circulating microRNAs (miRNAs), which are known to be biomarkers for cancer, endocrine dysfunction, and other diseases.
7
Hindson et al. demonstrated reverse transcriptase (RT) dPCR by performing a reverse transcription step manually prior to dPCR in the Bio-Rad QX100 system.
7
RT-dPCR had 37%–86% lower variability and 7× better day-to-day reproducibility than its qPCR counterpart, and with a similar detection limit. In serum samples, RT-dPCR could better distinguish between samples that were positive or negative for
Food and Agricultural Testing
dPCR has also found use in the testing of environmental samples (seeds, plant material, soil, and wastewater), where inhibitors from the DNA isolation process can decrease the efficiency and reproducibility of qPCR.
115
Morisset et al.
116
demonstrated the utility of ddPCR for quantifying genetically modified organisms (GMOs) in maize feed samples.
117
A duplex ddPCR assay compared levels of the
Summary and Conclusions: Part I
Digital assays partition samples into thousands of wells or droplets, and utilize Poisson statistics to calculate the sample concentration with absolute precision. The most mature digital assay is dPCR, evidenced by several commercial instruments currently used by researchers for a wide variety of applications. With the advent of dPCR, nucleic acid amplification has evolved from qualitative (PCR + electrophoresis) to semiquantitative (qPCR), and now to absolute quantitation. dPCR substantially improves precision in counting single molecules and resolving a small number of copies in the presence of inhibitors or wild-type populations. However, it does so at the expense of throughput. The applications of dPCR center primarily around three key capabilities: rare mutation detection (ctDNA), RSD (viral load, bacterial load, and circulating miRNA), and precise copy number quantitation (CNV and NGS libraries). Looking to the future, the use of dPCR in quantifying circulating nucleic acids is gaining widespread acceptance and holds considerable promise in improving detection limits and lab-to-lab reproducibility. To further standardize methodologies and results, Huggett et al. published the dMIQE (Minimum Information for Publication of Quantitative Digital PCR Experiments). 41 The use of dPCR is also quickly becoming a standard technique in NGS library prep. 109 Having covered digital assays with nucleic acids on Part I of this review, Part II will focus on digital approaches in proteins and cells.
Footnotes
Acknowledgements
The author thanks Dr. Rémi Dangla for insightful comments and discussions on the manuscript, and Priyan Weerappuli for a detailed manuscript review.
Declaration of Conflicting Interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The author gratefully acknowledges support from the National Science Foundation Division of Chemical, Bioengineering, Environmental, and Transport Systems (CBET), award numbers 1236764 and 1512544, and from the Division of Electrical, Communications and Cyber Systems (ECCS), award number 1232226.