Abstract
G-protein-coupled receptors (GPCRs) are modulated by many marketed drugs, and as such, they continue to be key targets for drug discovery and development. Many GPCR targets at Merck Research Laboratories (MRL) are profiled using homogenous time-resolved fluorescence (HTRF) inositol monophosphate (IP-1) cell-based functional assays using adherent cells in 384-well microplates. Due to discrepancies observed across several in vitro assays supporting lead optimization structure–activity relationship (SAR) efforts, different assay paradigms were evaluated for removing growth medium from the assay plates prior to compound addition and determination of IP-1 accumulation. Remarkably, employing the noncontact centrifugation BlueWasher method leads to left-shifted potencies across multiple structural classes and rescues “false negatives” relative to the traditional manual evacuation method. Further, assay performance is improved, with the minimum significant ratio of challenging chemotypes dropping from ~5–6 to <3. While the impact of BlueWasher on a broad range of our GPCR targets remains to be determined, for highly protein-bound small molecules, it provides a path toward improving assay reproducibility across scientists and sites as well as reducing replicates in SAR assay support.
Introduction
Cell-based in vitro pharmacological assays have been routinely used to support medicinal chemistry efforts to identify structurally diverse molecules as potential drug candidates. G-protein-coupled receptors (GPCRs) play a crucial role in drug discovery efforts as one of the most common drug targets for pharmaceutical companies.1,2 There have been numerous high-throughput assay technologies specifically designed to rapidly interrogate GPCR targets, using cell-based assays as the primary screening tool to define a compound as an agonist, antagonist, or inverse agonist and generate detailed information about the potency and efficacy of a compound for functional activity. 3 Assay miniaturization, the opportunity for increased throughput, and the need for a more robust medium removal process have resulted in the introduction of new technologies to efficiently conduct medium exchanges in adherent cell-based assays. Historically, the most common method of manually removing medium has been the “dump-pat,” where the researcher removes medium from the cell plate by inverting it onto an absorbent material and lightly tapping it. This variable method has been replaced by the automated combination plate washer and dispensers and most recently by a noncontact centrifugal washer and dispenser. Assays performed with these different techniques can vary remarkably regarding assay performance, data reproducibility, and predictability of the chemical structure–activity relationship (SAR). Driving chemistry SAR requires highly robust and reproducible assays that can be routinely performed in a high-throughput manner with rapid turnaround time, irrespective of the operator.4,5 During efforts to support the lead optimization (LO) of the Galphaq (Gq)-coupled GPR40 free fatty acid receptor 1 (FFAR1) program, discrepancies were observed in the data obtained from the functional inositol monophosphate (IP-1) accumulation assays. It was speculated that the discrepancies were due to assay support that was divided across sites with a lack of accessibility to identical instrumentation and automation. An alignment of protocols and critical technologies was performed, and it was determined that the methods used for removing the growth media were the source of variability in the assays. Here the underlying factors leading to the selection and implementation of a new technology used for the critical medium exchange step in the assay protocol were reviewed. During this process, a series of compounds were identified that had been inaccurately classified as having weak to no activity in the assays. Using this new assay paradigm, which efficiently and effectively removed all the serum-containing growth medium from the assay plate, it was possible to determine the potency of these compounds more accurately in the assay. In addition, the assay results obtained within Merck Research Laboratories (MRL) as well as externally with our contract research organization (CRO) partnership could be aligned. The implementation of this new technology allowed optimization of assays so that robust, high-quality, and reproducible data were generated routinely across operators and sites for optimal support of the SAR program.
Materials and Methods
Cell Culture
All cell culture reagents and buffers were purchased from Gibco (Gaithersburg, MD) through Thermo Fisher Scientific (Waltham, MA), unless otherwise noted. Human embryonic kidney (HEK)-293 cells stably expressing human GPR40 were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum (FBS), 1× nonessential amino acids (NEAA), 1× penicillin-streptomycin (PS), and 0.5 mg/mL geneticin (G418). Additional GPR40 or GPR120 recombinant cell lines were generated using the Jump-In Cell Engineering Platform6,7 in either a HEK-293 or Chinese hamster ovary (CHO) cell background. The Jump-In cell lines were cultured in DMEM with 10% dialyzed FBS, 1× P-S, 1× NEAA, 200 µg/mL hygromycin B, and 10 µg/mL blasticidin. The INS-1 832/13 rodent pancreatic beta cell line, 8 kindly provided by Dr. Christopher Newgard (Duke University, Durham, NC), was cultured in Roswell Park Memorial Institute (RPMI) 1640 medium supplemented with 10% Hyclone heat-inactivated FBS, 1× penicillin-streptomycin-glutamine (P-S-G), 10 mM HEPES, 1 mM sodium pyruvate, and 55 µM 2-mercaptoethanol (BME). Cells were kept in a 37 °C incubator in a relatively high (90%–95%) humidified atmosphere containing 5% CO2 and passaged twice per week or when the cell replication reached 80% confluency in the flask. Aliquots of assay-ready cryopreserved cell stocks were prepared using Recovery Cell Culture Freezing Medium according to the manufacturer’s instructions using aseptic cell culture procedures. 9
HTRF IP-1 Assay
The IP-1 terbium cryptate (Tb) homogenous time-resolved fluorescence (HTRF) kit was purchased from Cisbio Bioassays (Bedford, MA). DMSO was purchased from Sigma Aldrich (St. Louis, MO). The test compounds of interest were dissolved and serially diluted to 10 concentrations in DMSO ranging from 2 mM to 40 nM at 200× the assay concentration. The reference compound used to determine the maximum response (high control) in the assay was prepared at a single concentration of 1 mM, 200× the final assay concentration. For the minimum response (low control) in the assay, DMSO was prepared at the same final concentration (0.5%) as used for the test compounds. For the adherent cell-based assay, cells were thawed and harvested from a cryopreserved assay-ready frozen aliquot and resuspended in the DMEM complete growth medium as described above at 0.15 million cells/mL. The cells were plated as 7500 cells in 0.05 mL of complete growth medium per well of a 384-well, white, opaque Corning (Corning, NY) BioCoat poly-
For the suspension cell assay, cells were thawed and harvested from a cryopreserved assay-ready frozen aliquot and washed with Hanks’ Balanced Salt Solution (HBSS). The cells were resuspended in IP-1 stimulation buffer at 0.75 million cells/mL and plated as 7500 cells in a 384-well, white, opaque PerkinElmer OptiPlate (cat. 6007290) in a volume of 0.01 mL/well. The assay plate was briefly centrifuged at 500 rpm for 10 s using an Eppendorf (Hauppauge, NY) benchtop centrifuge with a swing bucket rotor. The addition of the test compounds and DMSO controls and the remainder of the protocol were followed as outlined above for the adherent cell-based assay.
Statistical Analysis
The analysis for the dose–response curves was performed using the IDBS ActivityBase for Screening Data Management software (Guildford, UK). Curve fitting was performed using the four-parameter logistic model. The data were reported as the half maximal effective concentration (EC50), as the nanomolar concentration, and as percent activity based on the maximum and minimum controls on each assay plate. Correlation plots of potency values were generated using Microsoft Excel (Redmond, WA). The minimum significant ratio (MSR) decision tool was used to measure the reproducibility and variability of potencies between assay runs.11–14 The MSR is defined as the smallest ratio between the potencies of two compounds that is statistically significant. The mean ratio (MR), the geometric average fold difference in potency between two runs, was also determined. An MR value close to 1 (acceptable range is 0.67–1.5) indicates that there is no statistically significant average difference between the two determinations. MR values outside of this range are due to high variability between assay runs. GraphPad Prism 6 (La Jolla, CA) was used to plot column scatter graphs to compare the assay plate metrics between assay formats.
Fully Integrated Robotic Assay Performed on a HighRes Biosolutions MicroStar System
The need for increased throughput and faster turnaround time for the IP-1 assays supporting the GPR40 program in the LO space demanded a change from the manual assay format that was currently used in the pharmacology labs.4,15–17 The manual assays used the dump-pat method to perform the medium exchange and manual movement of assay plates between peripherals, resulting in inconsistent incubation times and contributing to assay variability and inefficiency. In collaboration with our automation team, a fully integrated robotic assay was designed and validated on the HighRes Biosolutions MicroStar system (Beverly, MA); the system layout and assay workflow are shown in
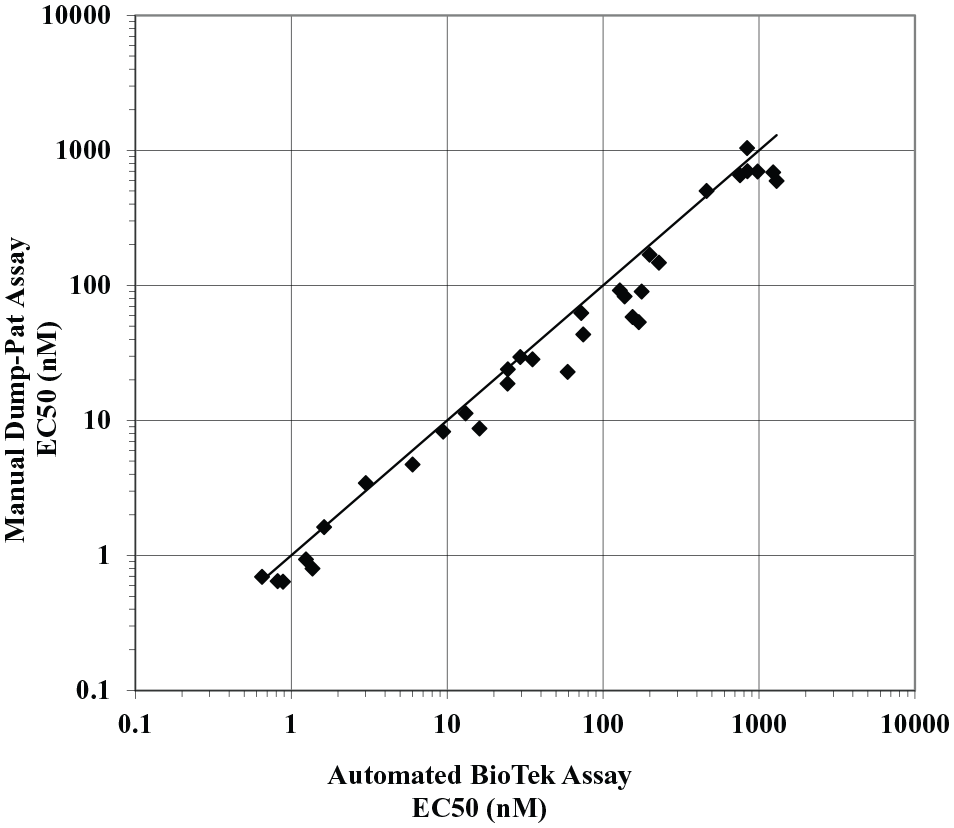
Correlation between the manual dump-pat assay and the automated BioTek assay for the primary GPR40 IP-1 assay. Potency values correlate between the assays using the BioTek EL406 combination plate washer/dispenser from the high-resolution fully integrated system and the manual dump-pat assay method for the removal of growth medium (n = 32; MR, 0.72; MSR, 1.95).
Outsourcing Capabilities Support
Building collaborative relationships outside of our internal organization is essential. To that end, partnerships have been established with several CROs to utilize their unique chemistry capabilities and screening expertise for many programs, including the GPR40 program.18,19 For this program, the data turnaround time was reduced by carrying out the IP-1 assays at the same site where a substantial portion of the chemistry was being performed. The screening capabilities at the CRO did not align with the fully automated process we had established internally for these assays. The assay protocol used by the CRO was more aligned with the manual dump-pat assays initially used in the internal pharmacology group, as previously mentioned. The assays were validated and successfully established externally using the manual dump-pat method; however, over time the variability in the assays led to a struggle to consistently support the SAR for the program. An extensive evaluation of the data generated by the CRO and the internal pharmacology group revealed an unacceptable correlation between the two assays. The data from the CRO showed a significant right shift in potency for compounds in the submicromolar range and a left shift in potency for weaker compounds; in some cases, compounds that had no activity in the internal assay exhibited weak activity in the CRO’s assay (
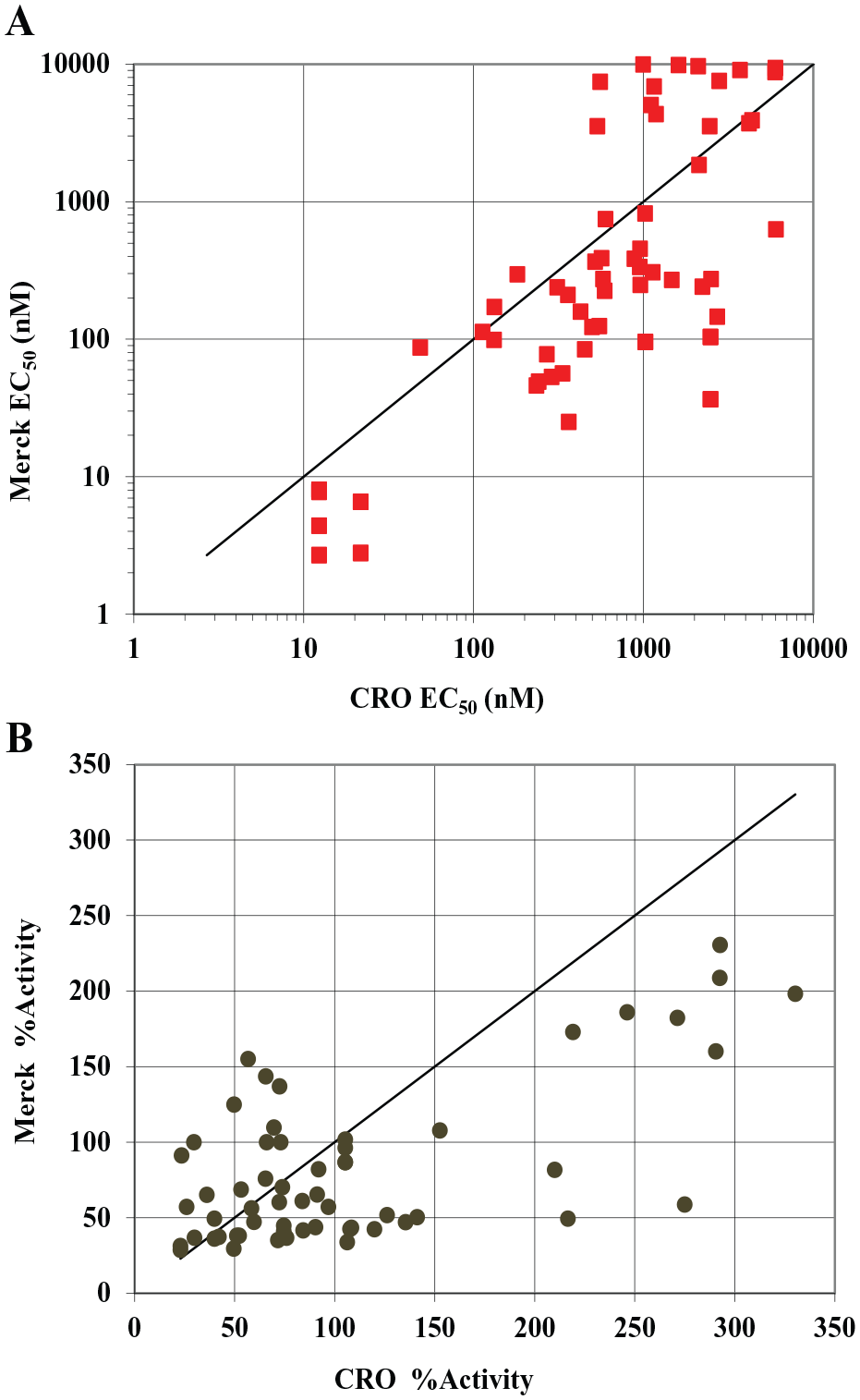
Struggle to meet criteria for SAR support. Internal (Merck) and external (CRO) variability for potency (
Methods for Medium Exchange
GPR40 is from the class of GPCRs that signal through the Gq pathway.20–22 Therefore, the highly robust Cisbio HTRF IP-1 Gq cell-based assay kit was employed that detects the accumulation of IP-1, a stable downstream metabolite of IP-3 induced by activation of a phospholipase C (PLC) cascade.3,23–27 According to the Cisbio IP-1 instructions, the kit allows direct characterization of all types of compounds acting on Gq-coupled receptors in either adherent or suspension cell assays. After further review of the assay protocols used both internally at MRL and externally at the CRO, it was determined that the protocols were fully aligned except for the method used to remove the growth medium after the overnight incubation. For this step, the CRO researcher would manually remove the growth medium by inverting the assay plate and quickly shake or flick the plate several times to discard the growth medium. The inverted plate would then be tapped multiple times on a stack of paper towels to remove any remaining medium from the wells and the top of the plate would be wiped clean with a paper towel to remove residual medium on the outside of the wells. Internally, this method is referred to as the dump-pat method and is extremely user dependent and can vary significantly across assay plates, even in the hands of the same researcher. Upon further evaluation, it was determined that the force exerted to shake or flick the plate and the number of times to shake or blot the plate will vary dramatically between researchers, as well as with the same researcher from day to day and across the assay plates within a single experiment. This disparity results in variable amounts of residual growth medium in the wells of the assay plate. Due to the inconsistency of this manual medium exchange step, it was determined that the protocol needed to be less user dependent and capable of generating consistent data irrespective of operator or location. Alternative methods to simplify and standardize the manual medium exchange process were explored that would be sustainable for both internally and externally run assays.
The BioTek EL406 washer/dispenser (Winooski, VT) is considered by many as the industry standard method for liquid aspiration and dispensing in high-throughput, fully integrated automation processes. 16 The BioTek uses an automated, programmable method resulting in a standardized medium exchange in assay plates and has been successfully used within the pharmacology group’s fully automated high-throughput screening process. However, the BioTek was not routinely used in the lower-throughput pharmacology processes due to the additional programming and monitoring required for each cell type, the risk of disturbing the cell monolayer and loss of cells, and the inherent residual volume remaining following the wash procedure. Additionally, the external partner supporting the IP-1 assays for the GPR40 project did not use the BioTek due to complications previously experienced with adherent cell-based assays. Concurrently, during this time the pharmacology group acquired a Blue Cat Bio BlueWasher that was evaluated as an alternative method to the manual dump-pat technique. The BlueWasher uses a noncontact, centrifugal process to remove liquid from assay plates and comes preprogrammed with various settings specific for adherent cell-based assays, leaving little to no residual liquid in the wells. Additional settings include a decant step to mimic the manual plate flick motion and the ability to add medium or buffer back to the assay wells (not tested in this evaluation).
The GPR40 IP-1 assays were developed using an adherent cell-based protocol; however, the Cisbio IP-1 assay kit is designed to be used equally well for adherent or suspension cell-based assays. The GPR40 IP-1 assay was evaluated in suspension cell mode to eliminate the need to remove the growth medium from the assay plate and align the assays with other IP-1 HTRF protocols used within the pharmacology group. Like the manual dump-pat method, the suspension cell assay protocol does not require the use of specialized instrumentation but requires a medium exchange using centrifugation prior to plating for the assay. The suspension cell protocol also eliminates the need for overnight incubation, allowing the assay to be performed within a single day. 23 With the exception of the initial cell preparation, the remainder of this protocol is automation friendly and can be used in low- to ultra-high-throughput processes with minimal protocol changes.
Results
Evaluating Alternative Methods for Medium Exchange
To simplify, standardize, and validate the GPR40 IP-1 assays, enabling efficient assay transfer across operators or sites, a series of structurally diverse chemical matter (107 unique compounds) were evaluated using the medium exchange methods mentioned above, with the exception of the BioTek, as this was not a feasible alternative for our external partner. For each method, correlation plots were used to assess the compound potency (EC50) and efficacy (percent activity) for the assays performed on subsequent days by a single operator. The results were averaged and then plotted against the averaged results from the standard assay using the manual dump-pat technique. MSR statistics were utilized to determine if there was a significant difference (>3-fold) or a collective shift in the results either across days for a single method or between two methods tested.
Suspension Cell Assay
The GPR40 IP-1 assay was performed using the suspension cell protocol previously mentioned. The reproducibility of this method shows a modest correlation with respect to potency and efficacy (
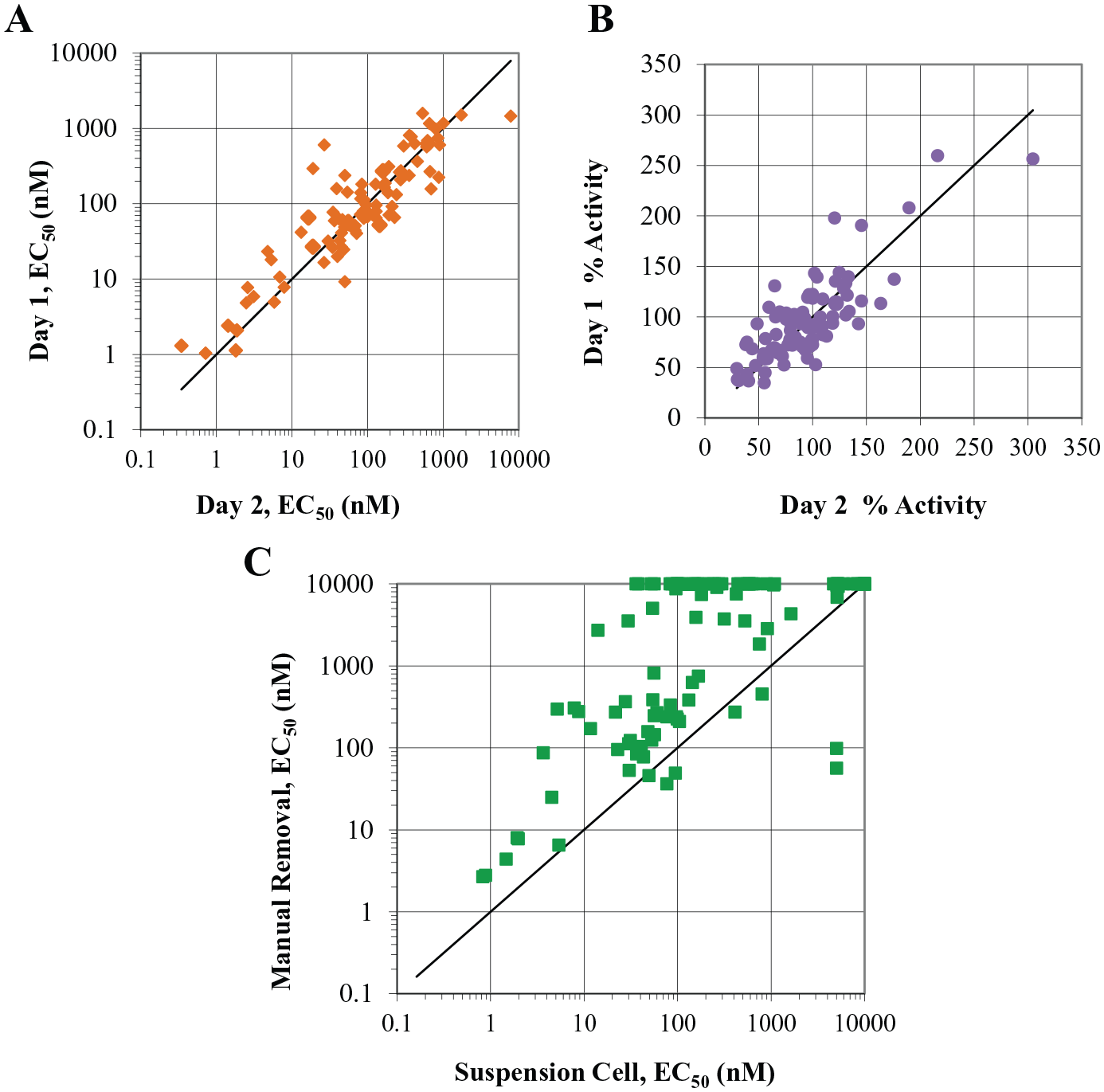
Alternative methods for medium exchange in assay plates: suspension cell assay. The IP-1 assay performed using cells in suspension mode correlates in potency (
BlueWasher Assay
The GPR40 IP-1 assay was performed using adherent cells that were incubated in the assay plate overnight. The following day, we used the BlueWasher to remove the growth medium from the assay plate. The BlueWasher protocol demonstrated highly reproducible results across days for potency and efficacy (
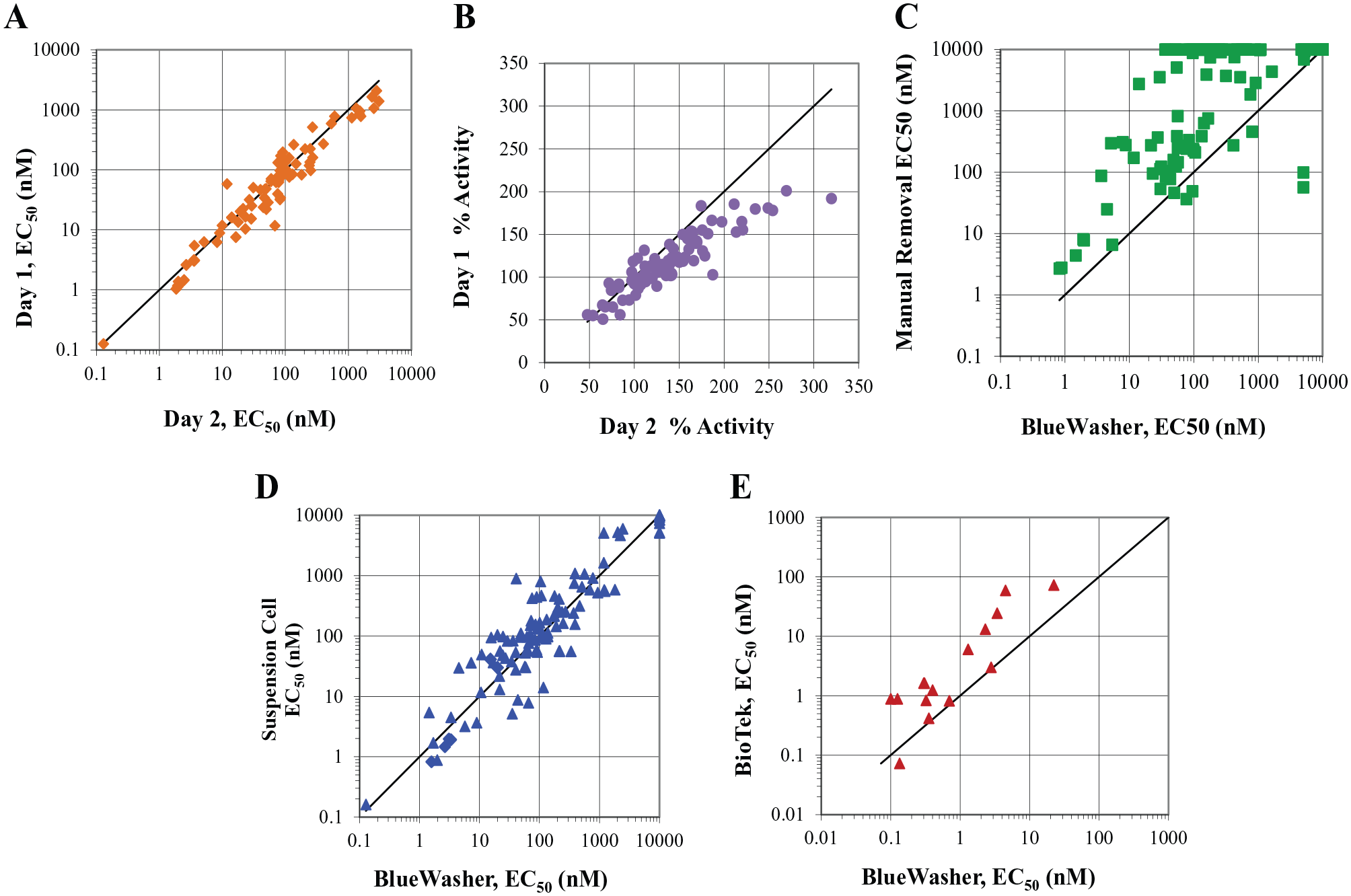
Alternative methods for medium exchange in assay plates: BlueWasher technology. The IP-1 assay performed using the BlueWasher technology to remove the growth medium from the assay plate correlates well regarding potency (
Cross-Assay Metrics and Validation
Based on the significant shift in potency observed in the GPR40 IP-1 assays using the various methods for media exchange, it was necessary to establish a highly robust and reproducible assay protocol that could be standardized across cell lines, operators, and sites, generating reproducible and reliable data to support the SAR needs of the program. As part of the assay validation, statistical metrics such as signal/background (S/B) and Z′ factor5,28 were included for each paradigm used. The S/B is the analysis for the assay window and is calculated as the fold difference between the averaged high and low controls. Ideally, this parameter should be greater than 2 to minimize the effect of background noise on the assay. However, S/B does not take into account the variability within the control wells. The Z′ factor encompasses the assay window and the variability within the high and low controls to measure the robustness of assay performance. Z′ factor values range from –1 to 1, with acceptable assay performance ranging from 0.5 to 1. For the HEK human GPR40 IP-1 assay, the S/B was highest for the dump-pat assay and lowest for the suspension cell assay (
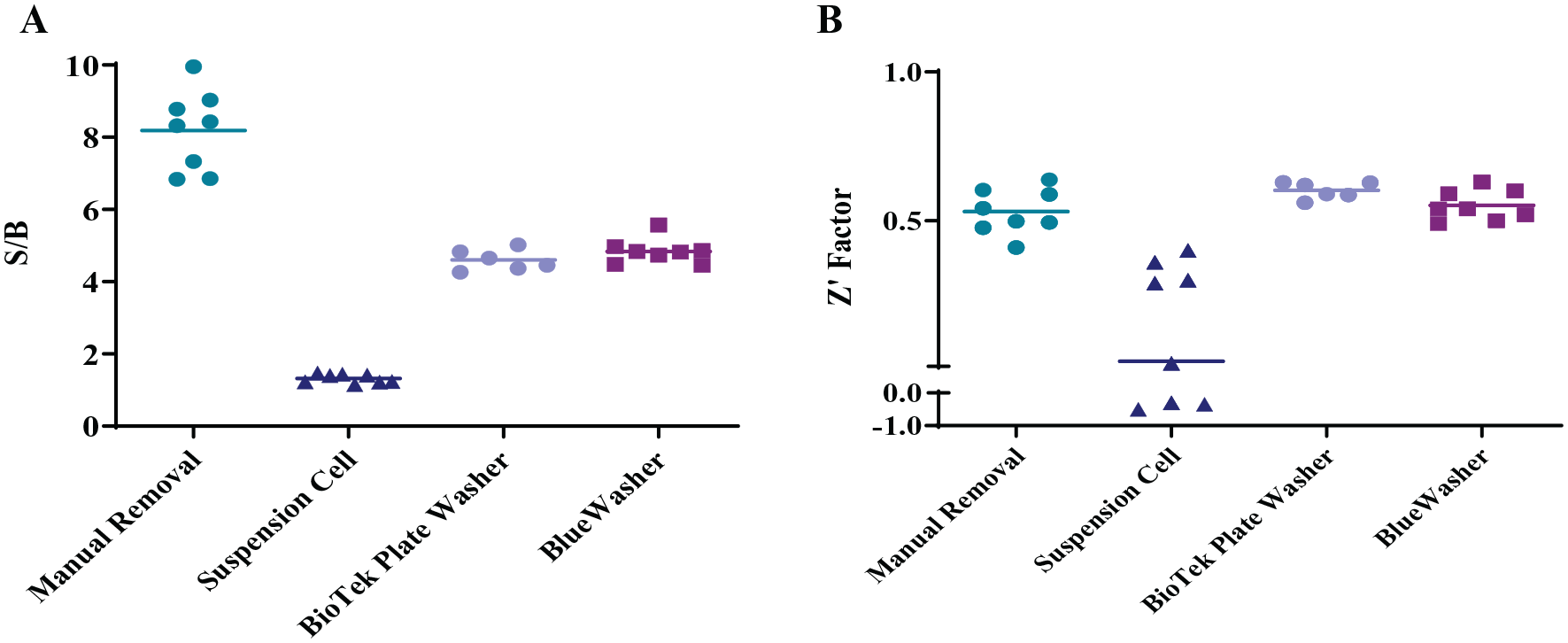
HEK GPR40 IP-1 assay metrics for the different medium exchange paradigms. (
To better understand and predict the in vitro to in vivo correlation, the IP-1 assay was used to evaluate the cross-species assays that involve varying cell line backgrounds, as well as the biologically relevant INS-1 cell line. These cell lines were validated in the different assay paradigms (excluding the BioTek) and evaluated the same statistical metrics for each (
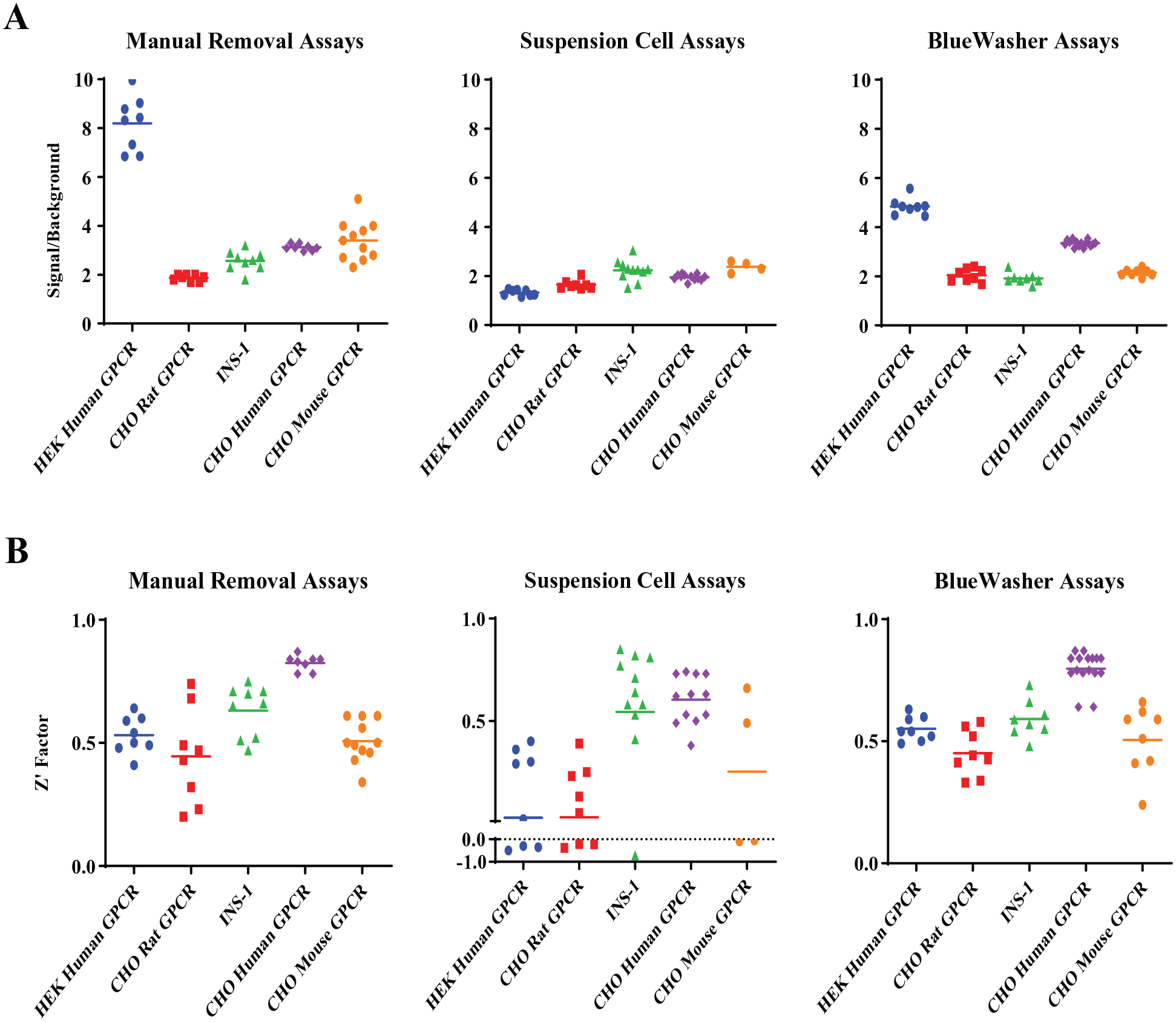
Assay metrics for additional cell lines for the different medium exchange paradigms. Most cell lines are affected by the assay paradigm used to remove the growth medium; however, the BlueWasher paradigm resulted in the most highly consistent robust assays compared with the others. (
Discussion
One of the critical components in supporting LO medicinal chemistry efforts is to have high-quality, robust, and reproducible assays in place for routine screening.1,2 For the GPR40 program discussed here, an HTRF IP-1 adherent cell-based assay was used as the primary functional assay, for cross-species evaluations and for off-target characterization. The lead chemical series in these programs included chemical structures that were highly plasma protein bound (>99%) (data not shown) and the potency of these compounds was extensively altered by the presence of serum (FBS) (data not shown) in the assay. Unaware of the significant impact of minor residual volumes of complete growth medium in the wells of the assay plate, the pharmacology group routinely supported the programs using the manual dump-pat method for removing the growth medium from assay plates. Using this technique, the amount of complete growth medium remaining in the wells would vary from plate to plate and across operators and days. Even with the assay performed on the fully automated platform using the BioTek EL406 washer/dispenser, there was an inherent residual amount of complete growth medium that remained in the wells after the medium exchange. It was not realized until the assay was performed using a method that completely eliminated the growth medium as to how significantly the residual complete growth medium can affect these compound potencies. This phenomenon was further evaluated by assessing a series of known highly plasma protein-bound compounds, using both the standard dump-pat assay protocol and the BlueWasher method, and observed the same trend as shown in
Figure 4C
. Since the pharmacology group heavily relied on a panel of IP-1 assays to support the GPR40 program and similar GPCR targets, the assay paradigm could not be shifted based on the improved results of a single assay. It was necessary to align and standardize the assay method across all cell lines used to support the programs. Five key cell lines were tested in the assay paradigm that has been shown to fully remove the growth media from the assay plates and the results were compared with those of the dump-pat assay method, evaluating the S/B and Z′ factors for each assay paradigm (
Supplemental Material
Thomas-Fowlkes_et_al_Supplemental_Figures – Supplemental material for Cell-Based In Vitro Assay Automation: Balancing Technology and Data Reproducibility/Predictability
Supplemental material, Thomas-Fowlkes_et_al_Supplemental_Figures for Cell-Based In Vitro Assay Automation: Balancing Technology and Data Reproducibility/Predictability by Brande Thomas-Fowlkes, Steven Cifelli, Sarah Souza, Richard Visconti, Alice Struck, Adam Weinglass and Mary Jo Wildey in SLAS Technology
Footnotes
Acknowledgements
The authors thank Blue Cat Bio for their help and support with the BlueWasher instrument. We thank Scott Greene for helping with the BioTek automation process and Jessica Liu for helping with the production of the cell lines used for these studies.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
