Abstract
The drug discovery process is dependent on the ability of screening efforts to identify and optimize lead compounds with therapeutic potential. Because G-protein–coupled receptors (GPCRs) account for the most “druggable” targets, the development of high-throughput, low-cost, and high-density GPCR assays to accommodate the increasing size of compound collections is significant. In this study, we report the application of an advanced LEADseeker robotic platform equipped with customized IT solutions for rapid data transfer to reduce assay cycles times for support of GPCR panel screening. The advantages of assay throughput, format, automation design, data management flow, and data reproducibility are discussed in terms of gains in productivity for lead optimization. The GPCR robotic platform demonstrates how automation technology can leverage traditional drug discovery processes by providing consistent and reliable data packages to expedite lead optimization efforts.
The major challenge confronting the pharmaceutical research area of lead optimization is to screen more compounds against an increasing number of targets using more robust methods while conserving time, cost, and resources. With the onset of parallel combinatorial synthesis, medicinal chemists are providing more compounds for evaluation in primary structure-activity relationship (SAR) screens and selectivity assays.1,2 In addition to chemistry discovery efforts, the fields of macromolecular modeling, database mining, and computer-based de novo design of druglike molecules contribute additional screening sets of compounds to identify new lead molecules.3,4
Given the many drug targets and the quantity of compounds synthesized on a regular basis by pharmaceutical chemistry teams, lead optimization has become more and more dependent on employing high-throughput screening (HTS) technology to efficiently and rapidly enable the characterization of novel chemotypes.5–7 To increase efficiency of the lead evaluation process, screening methods must be faster, cost-effective, and highly reproducible to enable quicker result turnaround time to monitor SAR efforts. To achieve this objective, assay miniaturization with precise, automated execution and rapid data delivery has become the industry goal. 8 To most efficiently adapt screening strategies to automation, assay formats should be validated with the goal being ease of automation. For example, converting assays to homogeneous formats requiring no separation steps or multiple incubations can be very amenable to HTS robotic platforms. Therefore, assay design should be streamlined to combine simplicity, speed, and robustness for automation adaptation. The enhanced productivity and data consistency of employing automation and smart assay design allow for gaining a quicker read on SAR to allow for more informed decisions for chemical synthesis sooner in the development process. 5
G-protein–coupled receptors (GPCRs) are the largest class of targets for drug discovery.9–13 GPCRs are a family of seven transmembrane proteins that mediate intracellular signaling responses that trigger physiological processes such as neurotransmission, metabolism, secretion, chemotaxis, inflammation, proliferation, smooth muscle contractility, and visual and chemosensory perception.14,15 Despite the vast diversity of ligand types, GPCR function is very modular whereby ligand-activated GPCRs undergo confirmational change and couple to a subset of the heterotrimeric G-protein subtypes, which are functionally grouped into four families (Gs, Gi/o, Gq, and G12). 14 These G-proteins in turn regulate intracellular effectors such as second messengers (i.e., cAMP, calcium, IP3), second messenger–dependent protein kinases, and ion channels to produce a biological outcome. 14
Many diseases and disorders have been linked to GPCRs, and >25% of the top 200 marketed drugs act to block ligand interaction on the extracellular domain of GPCRs to modulate intracellular signaling pathways.9–11,16–22 With the majority of therapeutic areas being served by GPCR-regulating drugs, the industry continues with rigorous efforts to search for new leads or more efficacious compounds for known and orphan GPCRs. In addition to assaying activity against the primary GPCR target, potential drug candidates must also be simultaneously examined in multiple GPCR panel screens targeting the central nervous system and the periphery to monitor off-target selectivity (cross-reactivity) and potential liability (side effect) issues before advancing compounds into in vivo pharmacokinetic and pharmacodynamic studies for efficacy determination. 23 The amount of resources, time, and money devoted to such lead evaluation efforts necessitates the need for lab automation and new technologies to expedite the process of GPCR preclinical drug development.
Both biochemical and functional cell-based GPCR assays have been successfully automated for HTS hit identification studies, and these same technologies are now being successfully applied to lead optimization for the benefit of faster data turn-around times for each program compound.11,24,25 In the present study, we describe the creation of a novel, custom-tailored robotic platform that harnesses commercially available instrumentation and scheduling package but is ultimately operated by internally derived informatics components that facilitate efficient data transfer. We use the CCR1 and CCR5 chemokine GPCR ligand-binding screens as a template for assay miniaturization and automation to achieve gains in throughput and productivity. Application of the GPCR LEADseeker scintillation proximity assay (SPA) robot allows for consistent, high-quality, and reproducible results in decreased timelines for chemists to monitor SAR trends for design of next-generation molecules.
Materials and Methods
Cell Culture
The human monocytic leukemic cell line, which expresses endogenous CCR1, was cultured in RPMI 1640 without HEPES (Invitrogen, Grand Island, NY; Cat. No. 111875-085) supplemented with 10% fetal bovine serum (Invitrogen; Cat. No.26140-095), 1% penicillin/streptomycin (Invitrogen; Cat. No. 15140-122) and 1x 2-mercaptoethanol (Invitrogen; Cat. No. 21985-023). An Athersys HT1080 human fibrosarcoma cell line stably expressing a RAGE-activated CCR5 receptor was grown in alpha MEM (Mediatech, Herndon, VA; Cat. No. 15-012-CV) supplemented with 10% dialyzed fetal bovine serum, 2% penicillin/streptomycin/glutamine (Invitrogen; Cat. No. 10378-016), and 500 µg/mL hygromycin B (Invitrogen; Cat. No. 25300-062). For CCR5 membrane preparation, 1 × 108 cells/pellet were suspended in buffer (50 mM HEPES, 5 mM MgCl2, 1 mM CaCl2), homogenized on ice with a Polytron homogenizer, and centrifuged at 48,000
Competitive Ligand Binding Assays
Filtration format
Radioligand competitive binding assays were used for the assessment of binding affinities of various compounds to the CCR1 and CCR5 receptors. For ligand-binding filtration assays, 2.5 × 105 THP-1 cells/well (in buffer containing RPMI 1640 without phenol red [Invitrogen, Cat. No. 11835], 0.1% bovine serum albumin [BSA; Sigma, St. Louis, MO; Cat. No. A2153], 20 mM HEPES [Invitrogen; Cat. No. 15630]) for the CCR1 assay or 15 µg/well membrane suspension (in 50 mM HEPES, 5 mM MgCl2, 1mM CaCl2, 0.1% BSA) for the CCR5 assay were added to 96-well assay plates containing test compounds in concentration-response curves (threefold, 11-point serial dilutions) with final concentrations ranging from 10−10 to 10−5 M. Threefold serial dilutions represent half-log increments for concentration-response curves and allow coverage of a broader dynamic range of IC50 values. Unlabeled ligands MIP-1α or MIP-1β were used in 100-fold excess for determining nonspecific binding for CCR1 and CCR5, respectively. Subsequently, the radioligand 125I-MIP-1α for CCR1 or 125I-MIP-1β for CCR5 was added to the reaction at a final concentration equal to the Kd. Based on binding kinetic experiments, equilibrium binding is achieved at approximately 60 min. Therefore, a 60-min incubation period was used for ligand-binding filtration methods. After a 60-min incubation period at room temperature, the binding reaction was terminated by harvesting onto GF/C filter plates (PerkinElmer, Waltham, MA; Cat. No. 6005174) using a Packard Cell Harvester followed by washing with an ice-cold HEPES-based wash buffer to remove unbound ligand. The plates were subsequently analyzed by the Packard TopCount reader after scintillant addition.
The competition data of the test compound over a range of concentrations was plotted as percentage inhibition of radioligand-specific bound in the absence of test compound (percentage of total signal). After correcting for nonspecific binding, IC50 values were determined. The IC50 is defined as the concentration of competing ligand needed to reduce specific binding by 50%. The concentration of test compound that inhibited 50% of radioligand bound (IC50) was quantified using the four-parameter logistic equation to fit the normalized data. The Ki values were calculated by application of the Cheng-Prusoff equation to the IC50 values, where Ki = IC50/(1+ ligand concentration/Kd).
The Z′ value as a criterion for data quality was calculated according to Zhang et al., 26 with the following equation Z′= 1 – (3*SDhigh control + 3*SDlow control)/(|Meanhigh control – Meanlow control|), where SD = standard deviation. A Z′ value of 1 represents an ideal assay, whereas a Z′ value between 0.5 and 1 indicates a good assay quality. The coefficient of variation (%CV), defined as the percentage ratio of standard deviation to signal magnitude, was calculated as 100*SDcontrol counts/Meancontrol counts.
SPA format
For LEADseeker SPA assays, the scintillant is incorporated into red-shifted SPA imaging beads, which are constructed to bind specific molecules, in this case protein. When the radioligand is bound to the SPA bead through interaction with the capture protein, the β-particle released is capable of exciting the scintillant contained in the bead, and detectable light is emitted. Emission energy from a radioligand not bound close to the SPA bead is dissipated in the aqueous medium and is not detected. 27 For competition studies, 384-well assay plates containing various test compounds at concentration-response curves with final concentrations (threefold, 11-point serial dilutions) ranging from 10–10 to 10–5 M were prepared containing respective standard compounds for determination of nonspecific binding. 1.0 × 105 THP-1 cells/well (CCR1 assay) or 5 µg protein/well (CCR5 assay) and 100 µg/well LS-WGA-PS beads (Amersham; Cat. No. RPNQ0260) were mixed and then added to each well of the assay plate. Subsequently, the radioligand 125I-MIP-1α for CCR1 or 125I-MIP-1β for CCR5 was added to the reaction at a final concentration equal to the Kd. The assay plates were incubated for 8 h at room temperature, followed by analysis using the LEADseeker. The competition data of the test compound over a range of concentrations were plotted as described above to determine IC50 and Ki values.
LEADseeker SPA Robotic System Instrument Design
GPCR target assays offer some inherent challenges that can be addressed by automation. These challenges include the need to have very tightly and consistently scheduled assay plates, to incorporate long incubation times, and to screen large compound collections through parallel multiple target receptors. A solution to address these challenges was designed and developed while maintaining a flexible platform that can be used to screen a wide variety of target classes. The system incorporates commercially available robotics, instrumentation, and scheduling package but is ultimately tied together by internally-derived informatics components that facilitate the smooth transfer of data.
This GPCR LEADseeker SPA robotic platform contains instruments capable of reagent dispensing and plate reading. As shown in Figure 1 , the physical system is based on a Thermo (Waltham, MA) CRS 465 fixed-base robotic arm controlled by Polara software version 2.3. A Generation 3 GE LEADseeker imager is employed as the detector for the SPA assays conducted on the system. This plate reader is capable of processing 384-well assay formats with a scan time of less than 5 min per plate with its charged coupled device (CCD) camera, which detects the red-shifted SPA imaging beads emitting at a 615 nM wavelength. For these assays, test compounds are predispensed into assay-ready plates, so all liquid handling on the system can be accommodated with bulk dispensers; in this case, five Thermo Multidrops are used. Additional peripheral components include a Thermo CRS automated carousel, Thermo Cytomat incubator, Velocity11 PlateLoc sealer (Agilent Technologies, Santa Clara, CA), Velocity11 Vspin centrifuge, Thermo/CRS regrip turn table, a magnetic spinner for mixing reagents for addition, and a Keyence (Elmwood Park, NJ) BL-601 barcode reader. These components are all mounted on a custom-built modular table system consisting of extruded aluminum frames, supporting solid aluminum tops, and a custom locking mechanism. The modular table system works equally well with both fixed base and track-based robotic systems. In either case, the design is such that tables can be interchanged between systems, thereby further enhancing the flexibility of the overall design. When a table with associated equipment is returned to a previously taught position, it is done so with a repeatability within 1 mm. All components of this system are fully integrated and require no manual intervention, for completely walk-away operation. Figure 2 demonstrates the protocol that the software initiates for the hardware to execute its functioning.
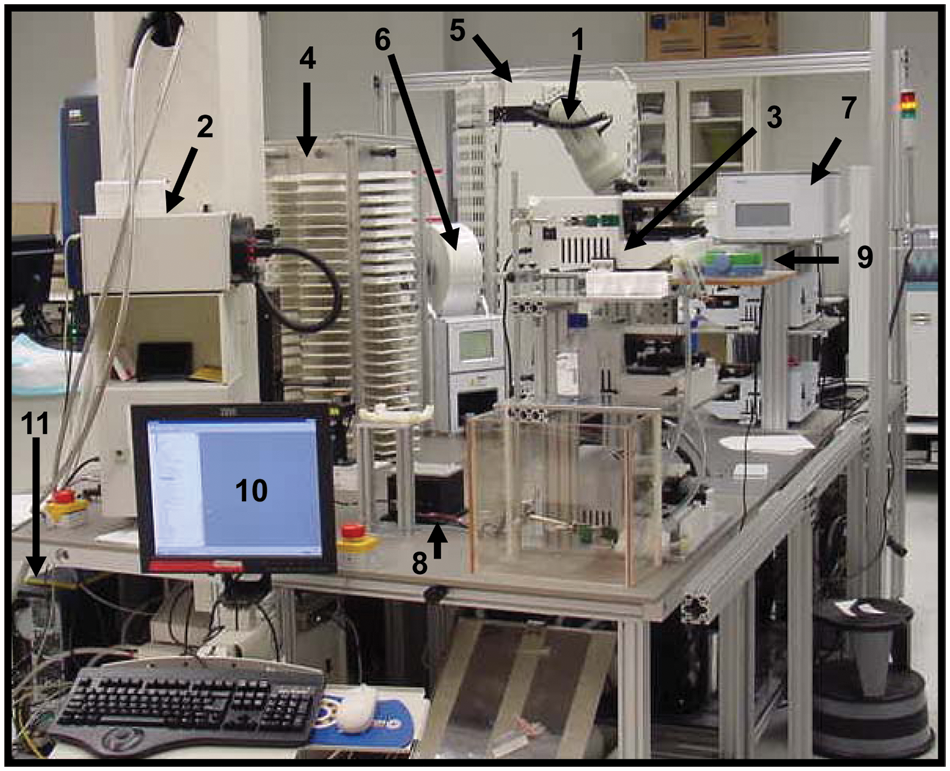

The automated G-protein–coupled receptor (GPCR) LEADseeker scintillation proximity assay robot at Bristol-Myers Squibb. Integral components of the system include Thermo CRS 465 fixed base arm controlled by Polara software version 2.3 (1), Generation 3 GE LEADseeker (2), and five stacked bulk dispense liquid-handling Thermo Multidrops (3). Additional peripheral components include Thermo CRS automated carousel (4), Thermo Cytomat incubator (5), Velocity11 PlateLoc sealer (6), Velocity11 Vspin centrifuge (7), Thermo/CRS Regrip turn table (8), magnetic spinner to prevent reagent settling (9), IBM computer with Polara and in-house software for robot execution (10), and a second computer driving the LEADseeker imager (11).
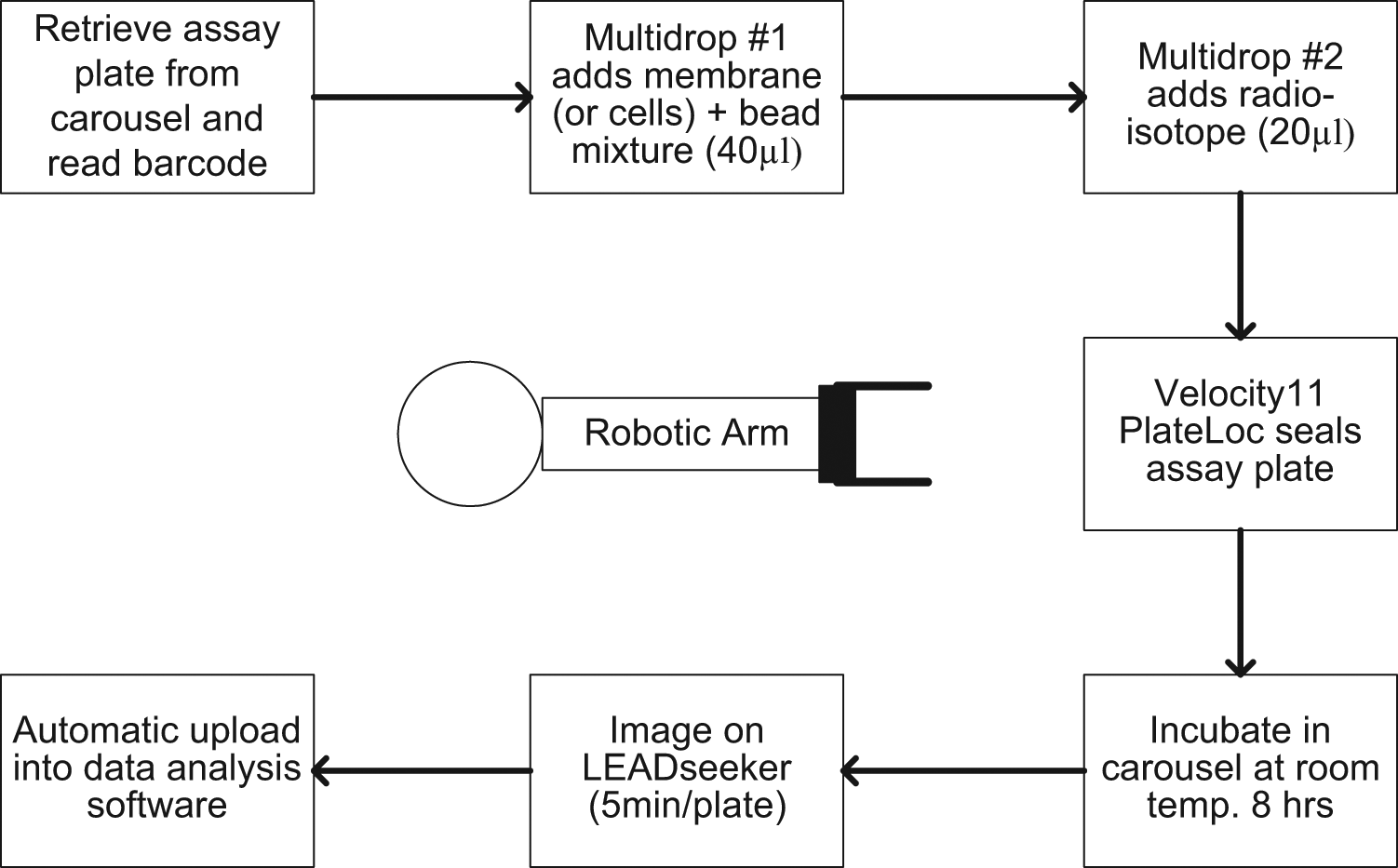
Flowchart of the robotic scintillation proximity assay protocol for evaluating ligand-binding affinity for G-protein–coupled receptor targets.
Informatics Process for Bristol-Myers Squibb Automated Systems
A key component of the robots employed at Bristol-Myers Squibb is that the systems not only automate the plate manipulation processes but also manage the data as well. Numerous different plate readers and detection devices are used on these systems; however, the methodology used to acquire and transition the data is the same ( Fig. 3 ). For the GPCR LEADseeker SPA robot, the plate reader operates on a separate computer from the computer that manages the Polara scheduling and robot control software. At the completion of each assay plate read, the data are saved by the plate reader’s software on its computer, and a copy is also saved within the Polara log file, which results in an immediate duplicate backup copy of the data. In real time, a data preprocessor extracts the data from the Polara log file and writes it to a separate text file for each assay plate. The generation of this text file triggers the QCMAD processor, which parses the data into a file consistent with database tables in the HTS Toolset Hub. Subsequently, the data are uploaded into the Toolset database. In Toolset, the raw data are merged with compound information that was previously entered into the database through PlateMaker at the time of compound plate creation as well as assay parameters that had been entered directly into Toolset prior to assay initiation. Because this sequence happens in real time, the CurveMaster utility can be used to review data sets on a plate-by-plate basis as they are generated or in a batch mode after completion of the run. CurveMaster is also the utility that is used to verify the data and publish it to the Central Database Repository (CDR), where it can be viewed by the chemists and biologists of the therapeutic area teams. This approach, whereby both the physical assay and data manipulations are automated and closely tied together, has provided a significant reduction in the turn-around time from compound submission to data reporting.
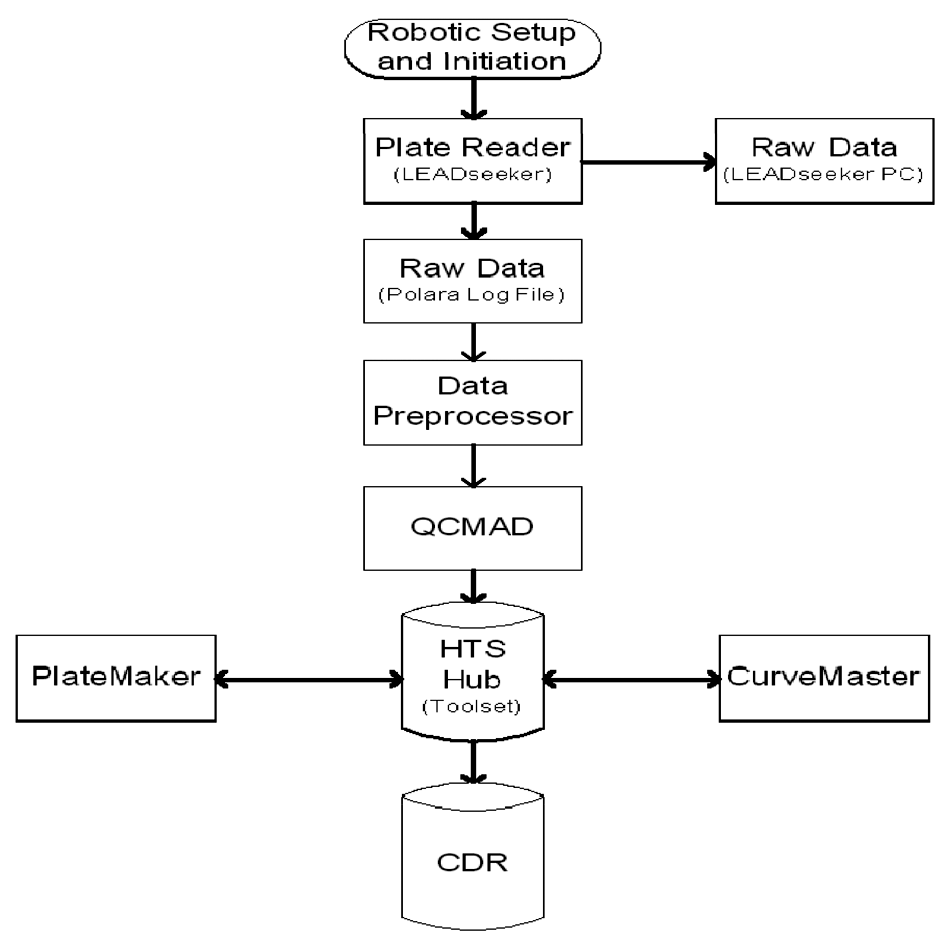
Informatics data processing and flow from the G-protein–coupled receptor LEADseeker SPA robot to the main database (refer to text for details).
The described system has been used to automate multiple SPA assays, which have been run in production mode very successfully. The capacity and efficiency of the SPA assays would not have been possible without the use of some type of automation. This platform not only facilitated the assays but also allowed for further streamlining of the process.
Results and Discussion
By miniaturizing assay formats and using automation technology, it is possible to increase assay throughput and efficiency for lead optimization as well as decrease operational costs associated with reagent consumption and employee time. However, it is first important to ensure the assay is suitable for automation and that conversion to an amenable assay format remains robust and sensitive and yields comparable pharmacological results. The CCR1 and CCR5 96-well ligand-binding filtration assays were converted to the homogeneous 384-well LEADseeker SPA assay through a series of experiments to determine cell density, bead titration, radioligand concentration, and incubation time (data not shown). For example, CCR1 assay performance was not sacrificed with conversion from the manually intensive gold standard filtration assay (
Fig. 4A
) to the mix-and-read SPA format, which eliminates harvesting separation and wash steps (
Fig. 4B
). The Z′ factor serves as an indicator of assay quality and takes into account the variability in sample data as well as the dynamic range between the high (total counts) and low (background counts) data populations.
26
Typically, biochemical assays have a high Z′ value (>0.7), whereas for cell-based assays, a Z′ value of 0.5 is an acceptable measure of favorable assay quality. The %CV, defined as the percentage ratio of standard deviation to signal magnitude, is typically used to quantify data variability and the precision available in the assay to identify hits with confidence.
26
The lower the %CV, the higher the assay precision. Both Z′ and %CV were improved with the LEADseeker SPA CCR1 assay over the filtration assay (
Fig. 4A
,
B
). The enhanced assay quality is due to the homogeneity of the SPA assay as opposed to the more variable nature of the filtration assay with its harvesting step and numerous wash steps involved. The signal-to-noise ratio (S:N) is reduced in the case of the LEADseeker readout, which is measured in integrated optical density compared with counts per minute obtained by TopCount measurements. An S:N of 4 to 8 for the LEADseeker readout is considered optimal because of the much-improved precision of the imager over TopCount photomultiplier technology (PMT). By design, a PMT-based instrument will yield greater counts than a CCD-based instrument. The LEADseeker, with its deeply cooled CCD camera, special lenses, and adjustable filters, will eliminate more background for an overall decreased signal, which is compensated by reduced noise and less well-to-well variability from rapidly flat-field imaging all 384 wells at once.28,29 When comparing CCR1 SPA and filtration binding assays, Ki assessments covering a range of test compounds with different potencies showed a significant correlation, with an average
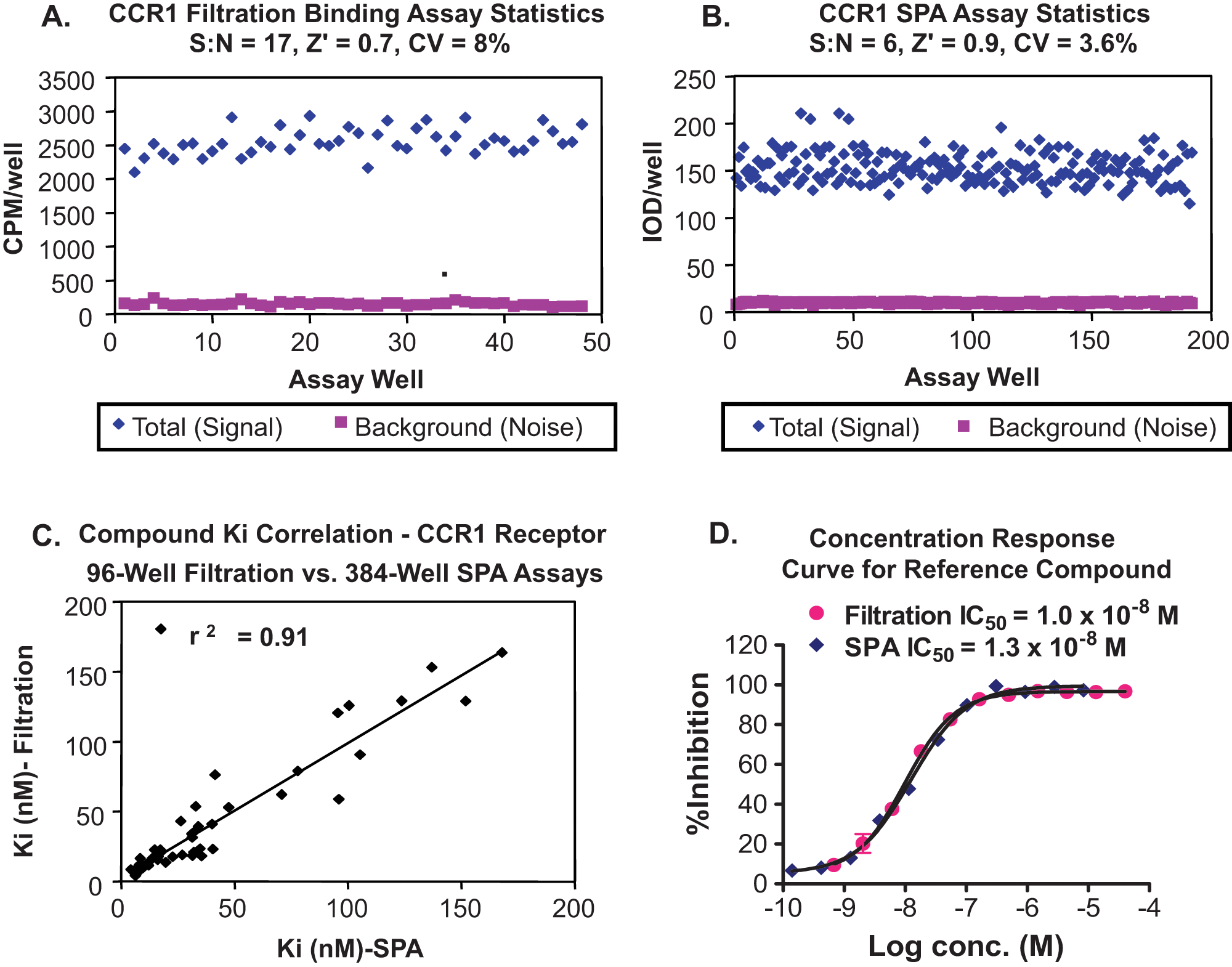
Assay format comparison between CCR1 ligand binding filtration and scintillation proximity assay (SPA) assays. (
GPCR LEADseeker SPA robot validation with the CCR1 and CCR5 assays yielded consistent Z′ values between 0.8 and 0.9 for both assays with plates in a given run and across runs (
Fig. 5A
,
B
). Compound Ki value comparisons between historical data and fully automated SPA experiments were in strong correlation, displaying
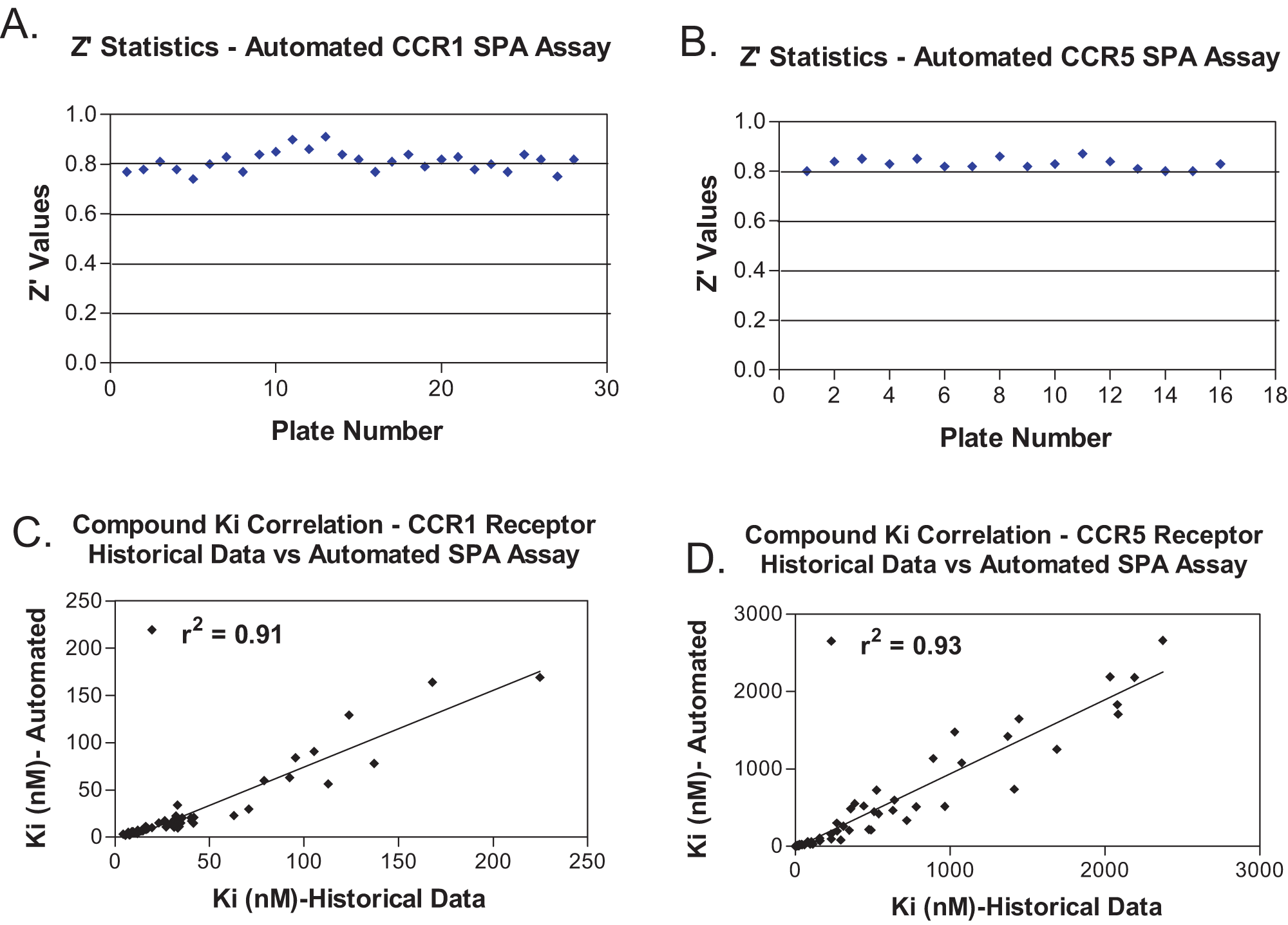
Automation validation: CCR1 and CCR5 automated scintillation proximity assay (SPA) performance on the G-protein–coupled receptor LEADseeker SPA robot. Interplate Z′ values determined by measuring total and background counts for separate assay plates in the CCR1 (
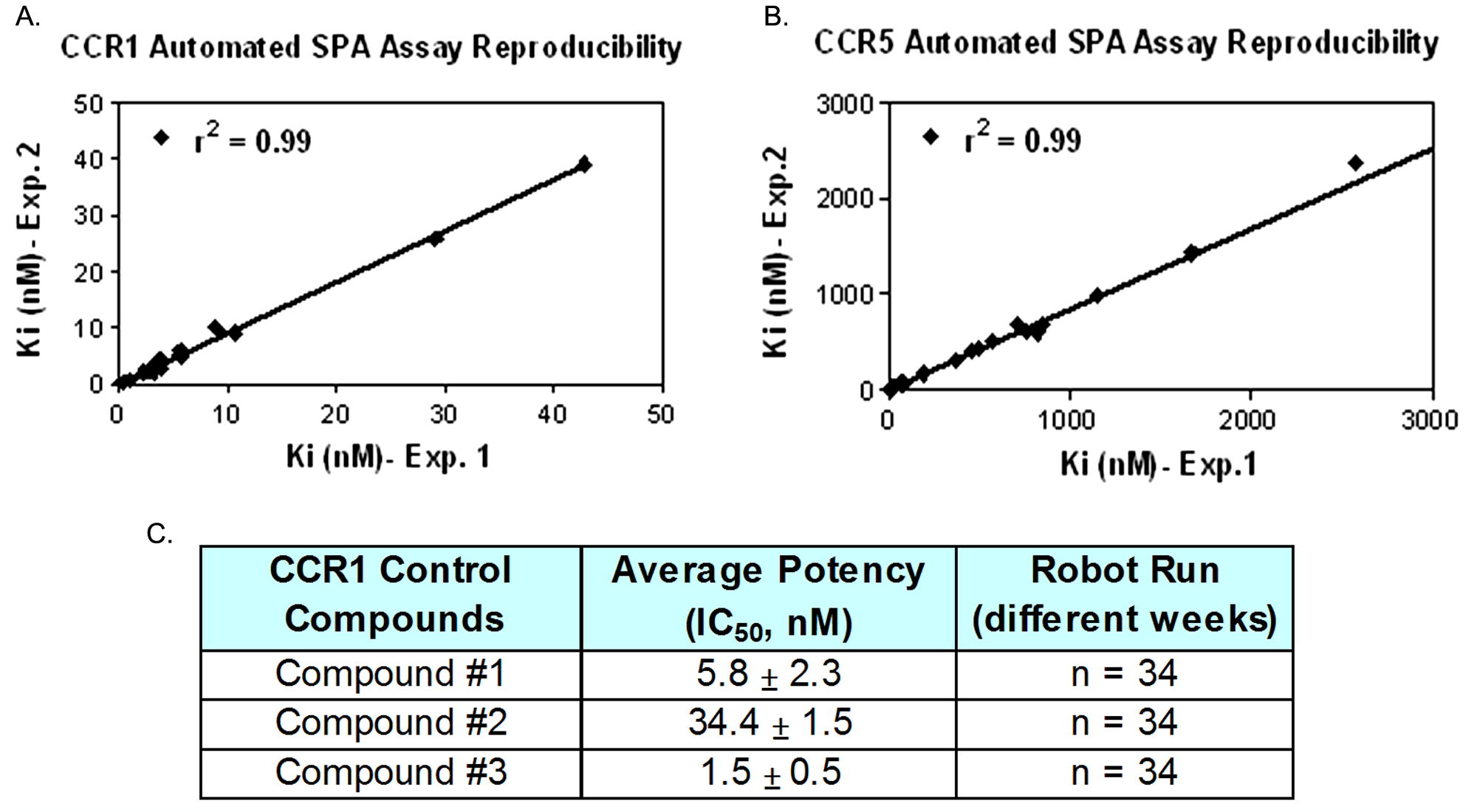
Assay reproducibility on the G-protein–coupled receptor LEADseeker scintillation proximity assay (SPA) robot. Run-to-run compound Ki comparison for fully automated CCR1 (
Through assay miniaturization, the fully automated 384-well CCR1 and CCR5 LEADseeker SPA assays reduced assay cost and test compound consumption by fourfold. By using the robotic technology platform, assay productivity and compound capacity were enhanced fourfold through automation and IT implementation, which expedited high-quality data packages to chemistry teams. The walk-away system also allowed significant time savings (threefold) to allow release of staff scientist time. Furthermore, with automation, compound capacity can increase without affecting staff time or labor because the robot executes the time-consuming and routine aspects of screening, which enables the staff to devote more efforts for assay development experimentation and mechanism studies. Thus, the GPCR LEADseeker SPA robotic platform permits the capabilities of screening large compound collections such as combinatorial libraries or data-mining sets without process limitations and enhances the flexibility of when these assays can be performed, as the robot can run throughout the night and on weekends.
The main advantage of the LEADseeker SPA system for evaluating GPCRs is the ability to batch runs together for simultaneous execution of multiple receptor screening on one robotic platform. The LEADseeker SPA robot can facilitate the running of 10 different target GPCRs simultaneously and has the capacity to handle a maximum of two hundred 384-well assay plates while precisely timing reagent addition steps and incubation times for each assay plate. This capability can enable parallel multi-GPCR target panel screening for off-target selectivity analysis to provide more comprehensive data packages and allow chemists to compare compounds against a multiparameter matrix. In addition, the marriage of the robotic system software components with the internal database allows for instantaneous uploading of LEADseeker raw data into the data analysis package software. Real-time analysis of the data not only allows for faster data turn-around times but also enables the ability to pinpoint any potential problems with the assay quality at an earlier time point. The utility of the LEADseeker robotic platform can be extended to target classes beyond GPCRs. For example, a panel of six enzyme selectivity assays using SPA beads to capture tritiated reaction products was optimized on the robotic system. Full automation of the enzyme assay suite enabled a 5-h time savings for the scientist and resulted in improved and consistent assay quality on the robotic platform (Z′ = 0.82) compared with assay execution in manual mode (Z′ = 0.63; data not shown).
In summary, the GPCR LEADseeker SPA robot described here has demonstrated its utility as an HTS tool for accelerating the lead evaluation compound selection process through IC50 profiling and selectivity monitoring. With novel approaches to assay design, assay format standardization, and enhanced informatics tools, integration of automated platforms such as the GPCR LEADseeker SPA robot can therefore improve the lead optimization process by reducing compound characterization times and accelerating delivery of high-quality, comprehensive data packages to increase knowledge applied to future compound design. The major advantage of technology enablement is that parallel processing of compounds through multiple assays is now being performed more quickly, with higher throughput for an overall reduced cost, which is the definition of automation justification. Especially for lead optimization, data quality and assay reproducibility are essential to closely monitor SAR trends and to benchmark reference compounds on a weekly basis. By achieving shorter timelines between compound synthesis and in vitro data delivery of on-target and off-target compound characterization, earlier and more informed decisions on which compounds to synthesize and progress into in vivo efficacy tests will significantly help to improve success rates in providing drug candidates to the clinic.
Footnotes
Acknowledgements
The authors would like to acknowledge James Gill, Nelly Masias, Daniel Maloney, Benjamin Carvelho, and Michael Beluch for development and support of the IT technology for data analysis and data extraction and processing for the robotic system. We are appreciative of the efforts of Michael Thibeualt, Edward Pineda, and Joe Yacobucci on the robot engineering and the support of James Myslik for robotic development. In addition, we recognize the efforts of the Cellular Resource group including Jonathan Lippy, Bert Ribeiro, and Suki Jayachandra as well as Reshma Panemangalore for enzymatic studies. Thanks also to the compound management team of Chad Johnson, Rob Hutchins, Henry Foster, Scott Manolopoulous, and Keith Wilson for creation of assay-ready compound plates for experimentation. We thank Percy Carter, John Hynes, and the BMS Immunology Discovery Chemistry Teams for supplying compounds for this study. We express our gratitude to Ge Zhang for critical review of the manuscript and figure formatting.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
