Abstract
Nomad Technology (Innoprot [Innovative Technologies in Biological Systems], Derio, Spain), a novel tool for multiplexing high-throughput cell-based G protein–coupled receptor (GPCR) assays, is described in this work. This new technology comprises a family of fluorescent biosensors called Nomad Biosensors that allow for the measurement of responses mediated by G proteins through their interactions with second-messenger transduction proteins. GPCRs are one of the largest protein families of receptors in eukaryotes, and their signaling mediates important physiological processes within cells. Thus, GPCRs are associated with a wide variety of diseases, and considered major targets in therapeutic research. Nomad constitutes a novel tool for unraveling the mechanism of GPCR signal transduction by simultaneously tracing different pathways. GPCR activation changes the structural folding of the biosensor and promotes its vesicularization, as well as an increase in the fluorescence intensity. Based on this technology, the MPXNomad cellular model was developed to discriminate between the Ca2+-mediated pathway and the cyclic adenosine monophosphate (cAMP)–mediated pathway. To validate this model, endothelin receptor B (ETBR) was coexpressed into the MPXNomad cell line and assessed with a specific agonist, an antagonist, and a chemical library of compounds. Nomad Technology optimizes the identification of novel GPCR ligands and enables the testing of large numbers of compounds.
Introduction
G protein–coupled receptors (GPCRs) are the largest family of cell membrane receptors. 1 These receptors perceive many extracellular signals, and they have a crucial role in activating and regulating intracellular signal cascades. Consequently, GPCRs have become one of the major therapeutic targets in the pharmaceutical industry.2,3 Indeed, they serve as targets for more than 50% of all modern pharmaceutical drugs. 4 It is widely established that the mechanism of signal transduction of GPCRs is based on the ligand–receptor interaction, which promotes the conformational change of the latter and triggers a cascade of second messengers inside the cells that results in signal amplification via coupling to heterotrimeric G proteins. Thereby, Ca2+ and cyclic adenosine monophosphate (cAMP) are key second messengers involved in amplifying the signal and in modulating different downstream targets. 5
The latest technological advances in microscopy, fluorescent protein (FP) engineering and the emergence of genetically encoded indicators, have led to the design of new tools that enable the measurement of dynamic changes in living cells, promoting a better understanding of the mechanisms that underlie the functional role of GPRCs.6–8 Moreover, thanks to technological advances in the research field, the classic understanding of GPCR signaling has undergone major revisions in recent years with the introduction of the functional selectivity paradigm.9–11 Surely the development of multiplexed assays is necessary to elucidate functional selectivity and, with the utilization of high-content fluorescent techniques, provide spatial and temporal information about second-messenger signaling. 12
The present work describes the development of a new family of genetically encoded biosensors, called Nomad Biosensors (Innoprot [Innovative Technologies in Biological Systems], Derio, Spain), designed to accelerate the discovery of new drugs using high-throughput cell-based GPCR assays.
This new technology comprises a family of fluorescent biosensors that are able to simultaneously detect the activation of different signaling pathways that are triggered by G proteins through their interaction with second-messenger transduction proteins.
Based on this technology, a multiplexed U2OS parental cell line, called the MPXNomad cell line, was created by stably coexpressing the green Ca2+Nomad biosensor with the red cAMPNomad biosensor. Using this approach, we hypothesized that it could be possible to discriminate between any of these two pathways (i.e., Ca2+ and cAMP) in the same living cell using any fluorescence-based equipment. With the aim of validating this hypothesis, the cDNA of the endothelin B receptor (ETBR) was transfected into our multiplexed cell line. ETBR’s ligand, endothelin-1 (ET-1), triggers the activation of both the Ca2+ and cAMP signaling pathways.13,14 This phenotypic cellular-based assay provides a better functional understanding of the post-ligand-binding molecular events than other radiolabeled or label-free biophysical technologies with an acceptable signal-to-noise value.
In this work, we show that the activation of both pathways (i.e., Ca2+ and/or cAMP) can clearly be detected and quantified using the MPXNomad cell line system. Finally, we have further characterized this technology using a library of chemical compounds to validate its utility in compound screening campaigns.
Materials and Methods
Plasmid Construction
The green Ca2+Nomad biosensor plasmid was created by subsequent cloning of the interleukin-2 receptor alpha chain (CD25) (DNA2.0, Newark, CA), turboGFP protein (Evrogen, Moscow, Russia), and a calmodulin (CaM)-binding peptide (ATGGAAAAGAGAAGATGGAAAAAGAATTTCATA GCCGTCTCAGCAGCCAACCGCTTTAAGAAAATCTC ATCCTCCGGGGCACTT) (DNA2.0) fused to a KDEL signal (DNA2.0) into the puromycin-resistant pPuro vector designed in our lab under the control of the cytomegalovirus (CMV) promoter. The red cAMPNomad biosensor plasmid was created by subsequent cloning of the CD25 (DNA2.0), turboFP650 protein (Evrogen), and the Ht31 peptide (GTAGACCTGATTGAGGAGGCCGCCAGCC GAATTGTCGACGCCGTGATTGAGCAGGTGAAAGC AGCAGGCGCCTACGGCGGC) (DNA2.0) fused to a KDEL signal (DNA2.0) into the Geneticin-resistant ptNL vector designed in our lab under the control of the CMV promoter. The ETBR cDNA (cDNA Resource Center, Bloomsburg, PA) was subcloned into a hygromycin-resistant vector, that is, CMV-pHygro, which was designed in our lab using the NheI and XhoI restriction enzymes. The clones were confirmed by DNA sequencing.
Cell Culture
The U2OS human bone osteosarcoma cell line (DSMZ, Braunschweig Germany) was obtained from ATCC (cat. HTB-96) and grown in Dulbecco’s modified Eagle’s medium nutrient mixture F-12 HAM (Sigma-Aldrich, St. Louis, MO) supplemented with 10% fetal bovine serum (Sigma-Aldrich), MEM nonessential amino acids (Sigma-Aldrich), and gentamicin (Sigma-Aldrich) at 37 °C in a humidified atmosphere supplemented with 5% CO2.
Generation of the Recombinant Stable Cell Lines
The plasmid encoding the green Ca2+Nomad biosensor was transfected into U2OS cells using Lipofectamine LTX (Thermo Fisher Scientific, Waltham, MA). Resistant clones were obtained by limit dilution, and positively transfected cells were selected using 10 µg/mL puromycin (Thermo Fisher Scientific). Once this stable cell line was ready, a positive clone was transfected with the plasmid encoding the red cAMPNomad biosensor, and the positive clones were selected by limit dilution using 10 µg/mL puromycin (Thermo Fisher Scientific) and 500 µg/mL Geneticin (Sigma-Aldrich). The cell line coexpressing both biosensors, named the MPXNomad cell line, was subsequently transfected with the ETBR-CMV-pHygro construction, and resistant clones were obtained by limit dilution and selected using 500 µg/mL Geneticin (Sigma-Aldrich), 10 µg/mL puromycin (Thermo Fisher Scientific), and 100 µg/mL hygromycin B (Thermo Fisher Scientific). These cells constitutively expressed green Ca2+Nomad biosensor, red cAMPNomad biosensor, and the ETBR, and this triple stable cell line was denominated the MPXNomad ETBR cell line. The MPXNomad ETBR cells were then tested using human ET-1 peptide (Tocris Bioscience, Bristol, UK) for functional Ca2+ and cAMP responses. Finally, a ready-to-use frozen batch of the MPXNomad ETBR cell line was placed into vials for use in each experiment. Likewise, the MPXNomad neurokinin 2 receptor (NK2R) cell line was engineered cloning the NK2R-CMV-pHygro construct into the cell line expressing both biosensors.
Cell Treatments
Cells were seeded at 20,000 cells/well in 96-well black, clear-bottom imaging plates (BD, Franklin Lakes, NJ) for agonism, antagonism, and screening assays. The Hamilton’s (Reno, NV) Microlab Star automated liquid handling workstation was used to dispense the liquid media containing the cells and compounds.
Agonism Assay
The MPXNomad ETBR cell line was treated with serial logarithmic dilutions of human ET-1 (Tocris Bioscience), endothelin-2 (ET-2; Sigma-Aldrich), and endothelin-3 (ET-3; Sigma-Aldrich) at concentrations ranging from 10 pM to 1 µM in Optimem media (Thermo Fisher Scientific) for 24 h at 37 °C, 5% CO2, and 95% relative humidity before data acquisition.
Antagonism Assay
The MPXNomad ETBR cell line was cotreated with ET-1 at a constant concentration of 10 nM, and serial logarithmic dilutions of BQ-788 (Bachem, Bubendorf, Switzerland) ranging from 30 µM to 30 nM in Optimem media (Thermo Fisher Scientific) were applied for 24 h at 37 °C, 5% CO2, and 95% relative humidity before data acquisition.
Specificity/Versatility Assay
The MPXNomad, the MPXNomad ETBR, and the MPXNomad NK2R cell lines were treated with human ET-1 (Tocris Bioscience) 100 nM, neurokinin A (NKA) (Sigma-Aldrich) 1 µM, and the vehicle (H2O) in Optimem media (Thermo Fisher Scientific) for 24 h at 37 °C, 5% CO2, and 95% relative humidity before data acquisition.
Screening Assay
A chemical library comprising 1200 compounds was sourced from the Prestwick Chemical Library (Illkirch, France), and based on the compounds’ chemical and pharmacological diversity and their bioavailability and safety for humans, these compounds were used for the screening assay. The reference compound ET-1 was purchased from Tocris Bioscience. All compounds from the library were dissolved in DMSO (Sigma-Aldrich) at 5 mM and the reference compound (i.e., ET-1) in water at 100 µM. The MPXNomad ETBR cells were treated with ET-1 (positive control) at 1 μM, whereas the compounds of the library were tested at 10 μM for 24 h in Optimem media (Thermo Fisher Scientific) before image acquisition.
Image Acquisition and Analysis
After 24 h of treatment, the cells were fixed with phosphate-buffered saline (PBS; Sigma-Aldrich) supplemented with 3.7% formaldehyde (FA; Sigma-Aldrich) for 10 min at room temperature (RT) and permeabilized with 0.3% Triton-X100 (Sigma-Aldrich) in PBS for 3 min at RT. The cell nuclei were then stained with DAPI (Sigma-Aldrich) for 5 min at RT. Fluorescent images were acquired in the BD Pathway 855 High-Content automated image platform with a 20× dry objective. The changes in the fluorescence patterns of the Ca2+Nomad and cAMPNomad biosensors were processed and quantified using image analysis algorithms with the Attovision bioimaging software (BD) in 3 × 3 subfields of each well (500–1000 cells analyzed per well). The excitation and emission filters used were the following: for DAPI, 380/10 and 450/10; for green fluorescent protein (GFP), 488/10 and 540/20; and for rhodamine, 548/20 and 570LP. Ca2+ activity was measured by summing the green fluorescent intensity of every pixel contained within a cell. cAMP activity was measured by counting the red fluorescent granules and normalizing the result to the cell number. The data were normalized as percentages of activity compared with the positive control (ET-1) after subtracting the value of the vehicle control. Cellular viability was determined by counting the nuclei (DAPI). For each condition, five replicates were used. The data were processed using Excel and plotted in SigmaPlot 11.0. The results are expressed as the mean ± SD. EC50 determination curves were fitted to a four-parameter logistic regression. The Z′ factor was calculated using the following formula: Z′ = 1 – [(3σc+ + 3σc–)/(µc+ – µc–)], where µ and σ are the average and standard deviation, respectively.
Results
Currently, there is a growing necessity to develop novel tools to study membrane receptors. Existing approaches do not consider the capacity of the ligands to stabilize different conformations of a receptor, which leads to a preferential activation of one signaling pathway over the others. Besides, these approaches are normally not useful for testing a large number of compounds. 15 In an attempt to respond to this need, Nomad Technology has optimized multiplexed high-throughput cell-based GPCR assays.
The molecular structures of the Nomad Biosensors comprise a membrane localization peptide, an FP, a vesicularization signal, and a second-messenger transduction protein–binding peptide ( Fig. 1A ). The latter can be replaced depending on the specific cascade triggered after GPCR activation. In this manner, the Ca2+Nomad biosensor contains a CaM-binding domain and the cAMPNomad biosensor, that is, a protein kinase A (PKA)–interacting Ht31 peptide. Nomad Biosensors are normally localized in the plasma membrane in basal conditions ( Fig. 1B ). Upon ligand binding, the activation of the GPCR leads to an increase in second-messenger levels (Ca2+ and/or cAMP) that, in turn, will activate the transduction proteins, CaM, or PKA. The interaction of the CaM or the PKA with the Nomad Biosensors produces changes in their folding and promotes an increase in the fluorescence and vesicularization patterns.
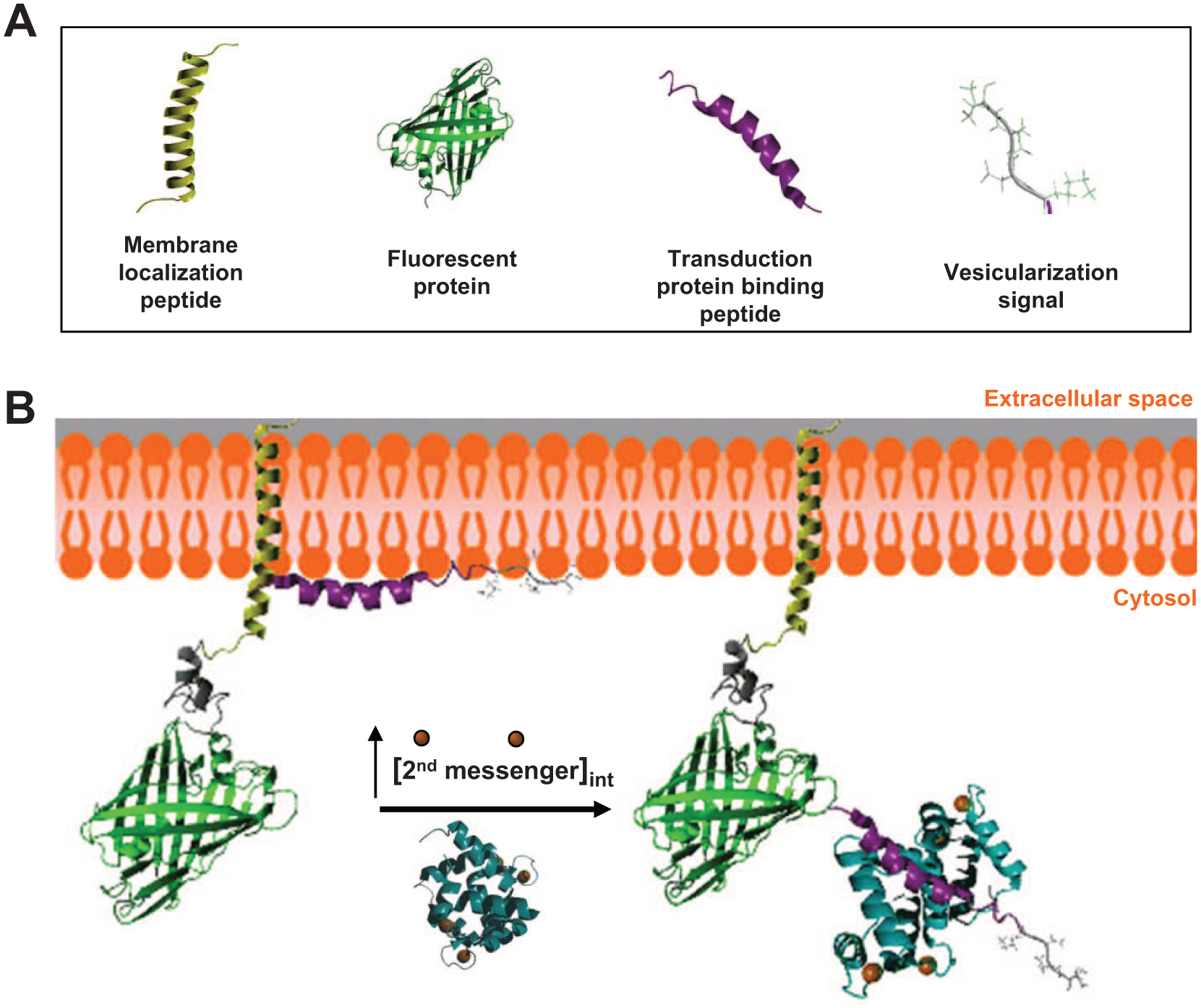
(
Despite the second-messenger responses usually taking few seconds or minutes, 16 the conformational change and internalization of the biosensors could take longer. In order to elucidate the best time point to acquire the data, we first performed a time-course experiment. The MPXNomad ETBR cell line was treated with the natural agonist ET-1 (1 µM), and the increase in activity was measured at different time points for 24 h ( Fig. 2 ). Notwithstanding that the internalization of the biosensor could already be observed between 6 and 9 h after the agonist addition, the time to obtain an appropriate dynamic range was around 18–24 h. The same tendency was observed with other cell lines expressing unrelated receptors (data not shown). So, an incubation time of 24 h was set for the rest of the experiments.
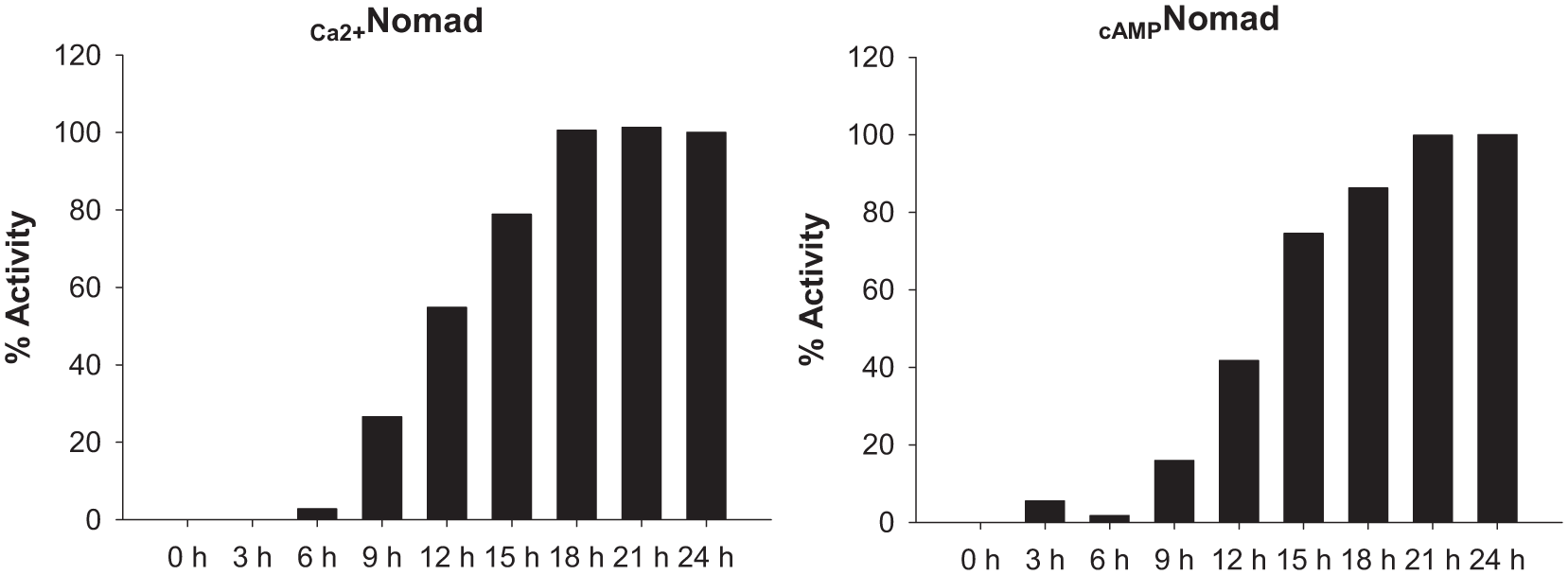
Time-course experiment of Nomad Biosensors signaling. MPXNomad-ETBR cell line was treated with ET-1 1 µM, and measurements were taken every 3 h during 24 h. After 6 h of incubation, the signal can be detected, but the best dynamic range, common for both biosensors, is reached after 18–20 h.
Accordingly, the MPXNomad ETBR cell line was validated by measuring the redistribution of the Nomad Biosensors with fluorescence image acquisition after treatment with the vehicle (H2O) and the natural agonist ET-1 (1 µM) for 24 h ( Fig. 3A ). The negative control produced a characteristic membrane distribution of the biosensor, whereas the ET-1-treated cells presented an increase in the fluorescence intensity for the green Ca2+Nomad and a pattern of vesicle retention throughout the cytoplasm for the red cAMPNomad biosensor. This increase in fluorescence in the Nomad Biosensors was verified using flow cytometry as an additional measurement method ( Suppl. Fig. S1 ).
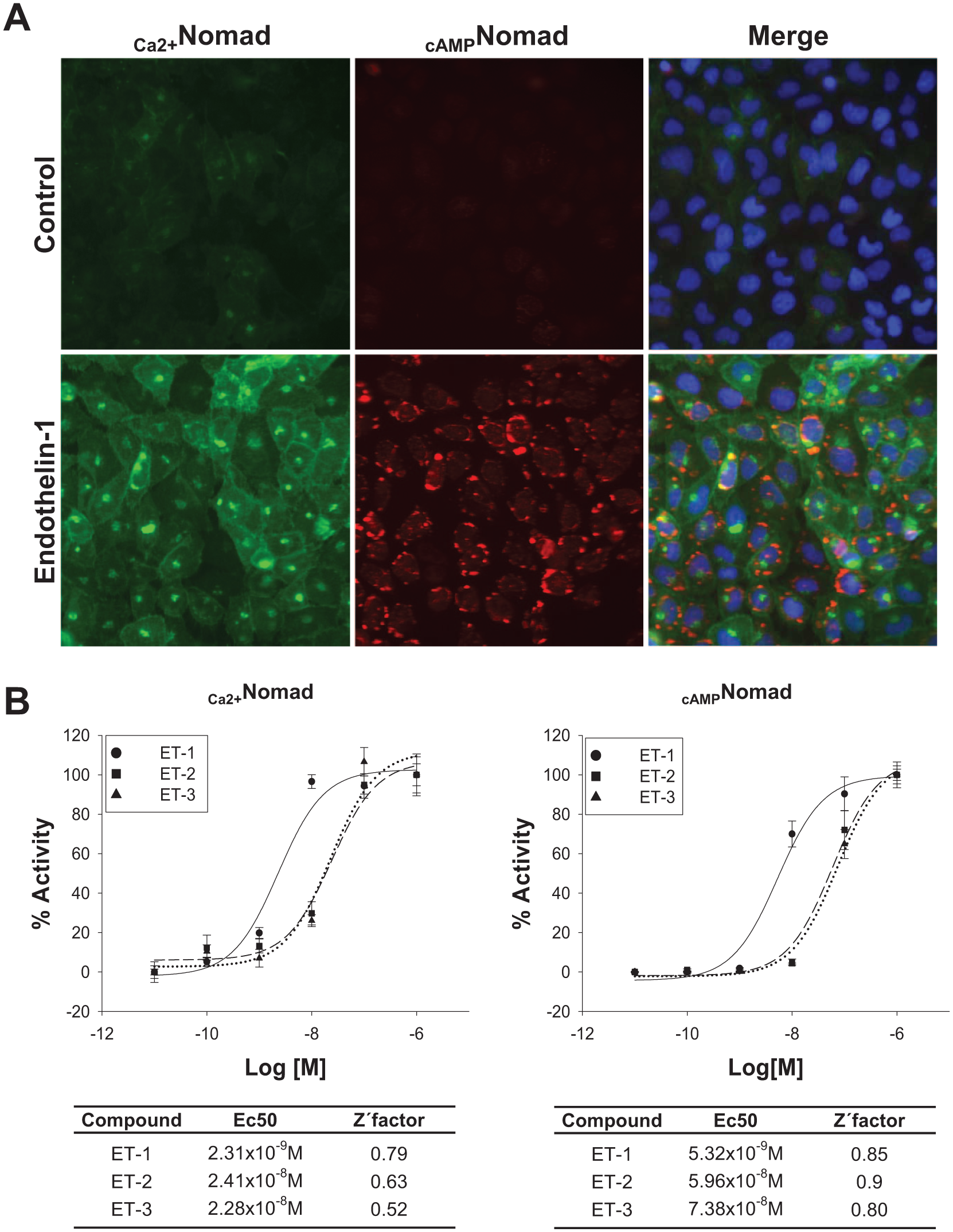
MPXNomad ETBR cellular fluorescence redistribution upon ET-1 treatment in U2OS cells. (
In addition to ET-1, there are two other vasoactive peptides of the endothelin peptide family, that is, ET-2 and ET-3, that can also activate ETBR. 17 To explore the robustness and reliability of our multiplexed model, dose–response curves were created for each endothelin peptide (i.e., ET-1, ET-2, and ET-3; Fig. 3B ). Although it is thought that ETBR is activated with similar affinities by the ET-1, ET-2, and ET-3 peptides, the MPXNomad model showed that ET-1 promotes the mobilization of Ca2+ and cAMP in a more efficient manner than ET-2 and ET-3, even though they also properly activate the receptor. Finally, to check the assay quality, Z′ factors were determined for all of the dose–response curves ( Fig. 3B ). It is widely accepted that a Z′ factor > 0.5 confirms that an assay is robust.18,19
Nomad Biosensor distribution can alternatively be quantified with different laboratory equipment. The results revealed very similar EC50 values for ET-1, ET-2, and ET-3 for both the calcium and the cAMP biosensors when analyzed in a conventional microplate reader. The Z′ factors for each ligand and biosensor were also calculated and resulted in values > 0.6, which confirmed the robustness of the assay ( Suppl. Fig. S2 ).
Additionally, to determine whether Nomad Technology represents a valid strategy for searching for other antagonists, a well-known ETBR antagonist, BQ-788
20
(
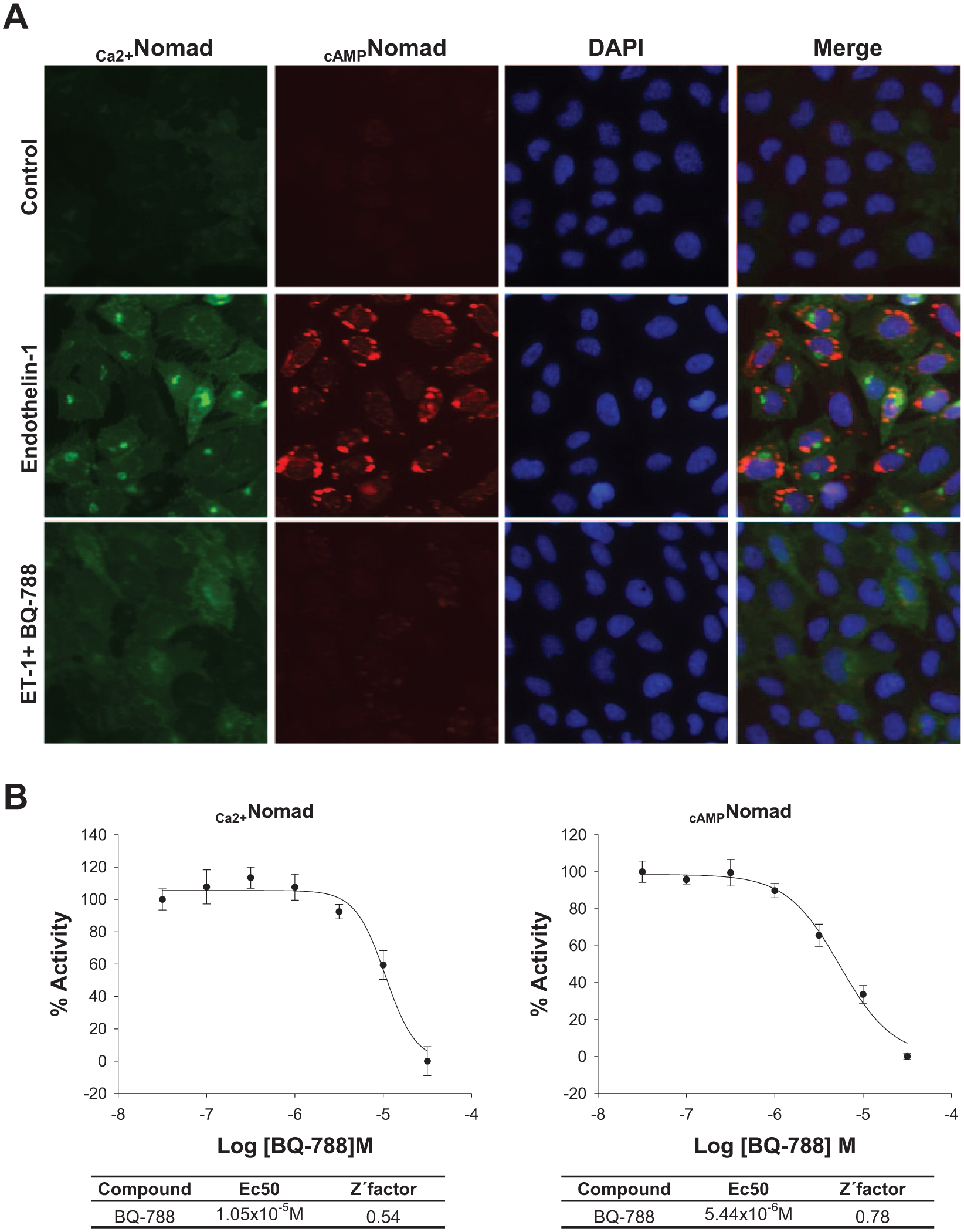
MPXNomad ETBR cellular fluorescence redistribution upon BQ-788 treatment in U2OS cells. (
In order to prove the specificity of the Nomad Biosensor for the receptor of interest, a cell line expressing both biosensors but lacking the receptor was engineered (MPXNomad). The MPXNomad cell line was treated with the vehicle (H2O) or ET-1 100 nM for 24 h. Neither the vehicle nor the agonist produced an increase in the biosensor signal, showing that the increase observed with the MPXNomad-ETBR cell line is specific for the ETBR. The versatility of the technology was analyzed using a cell line expressing an unrelated GPCR, the NK2R. The treatment of the MPXNomad-NK2R cell line with the NKA agonist produced an increase in the biosensor signal that was specific for the receptor, as the treatment of the MPXNomad cell line with NKA did not produce any change. Furthermore, the MPXNomad-ETBR and MPXNomad-NK2R cell lines only responded to their cognate ligand, further proving the specificity of the technology ( Table 1 ).
Specificity and Versatility Assays.
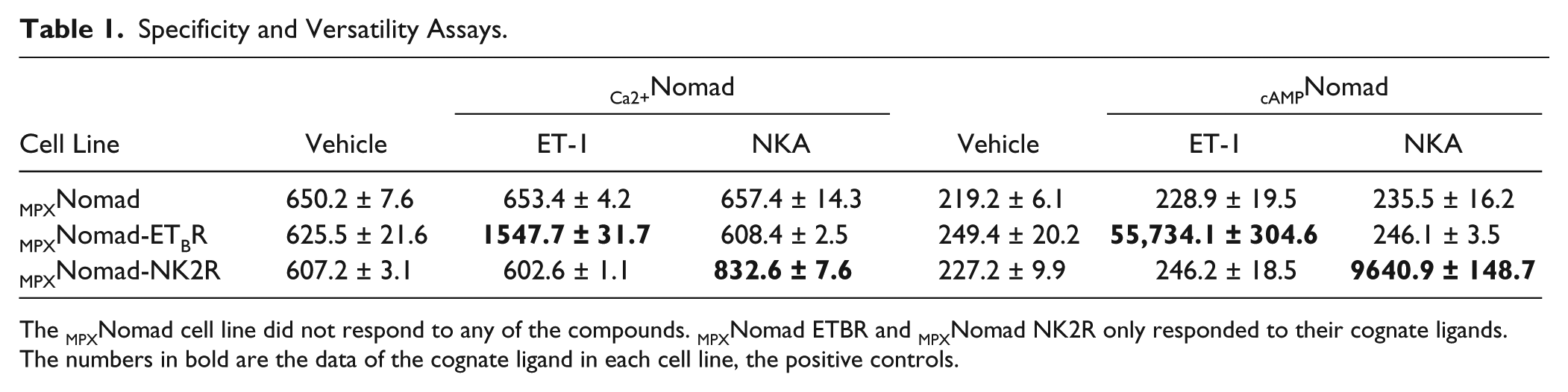
The MPXNomad cell line did not respond to any of the compounds. MPXNomad ETBR and MPXNomad NK2R only responded to their cognate ligands. The numbers in bold are the data of the cognate ligand in each cell line, the positive controls.
The main objective of the development of Nomad Technology is its use in compound screening platforms. High-throughput screening (HTS) and high-content screening (HCS) are the most useful tools for drug screening. However, library screening campaigns usually measure only one parameter and obviate other signaling pathways that might be activated through the same receptor.
21
To demonstrate that the Nomad multiplexed cell line could be useful in the identification of possible biased hits that may be positive for the Ca2+ pathway or for the cAMP pathway, a high-content assay was implemented in 96-well plates with the MPXNomad ETBR U2OS cells. The optimized cell-based assay was used to screen a library of 1200 compounds that belong to the Prestwick Chemical Library at 10 µM for 24 h. The hit compounds for the different second-messenger pathways revealed an increase in the Nomad Biosensor fluorescence intensities and vesicularization as expressed as Ca2+ or cAMP activity, respectively. The library screening results are provided in
Figure 5A
. Setting a hit threshold of 3 standard deviations (SD) over the mean (<33.64% for Ca2+ activity and <12.04% for cAMP activity) resulted in the acquisition of 17 hits (1.41% hit rate). The Z′ factors calculated from the positive and negative control wells on each plate produced an average Z′ of 0.75 ± 0.07 for Ca2+ activity and a Z′ of 0.80 ± 0.06 for cAMP activity (
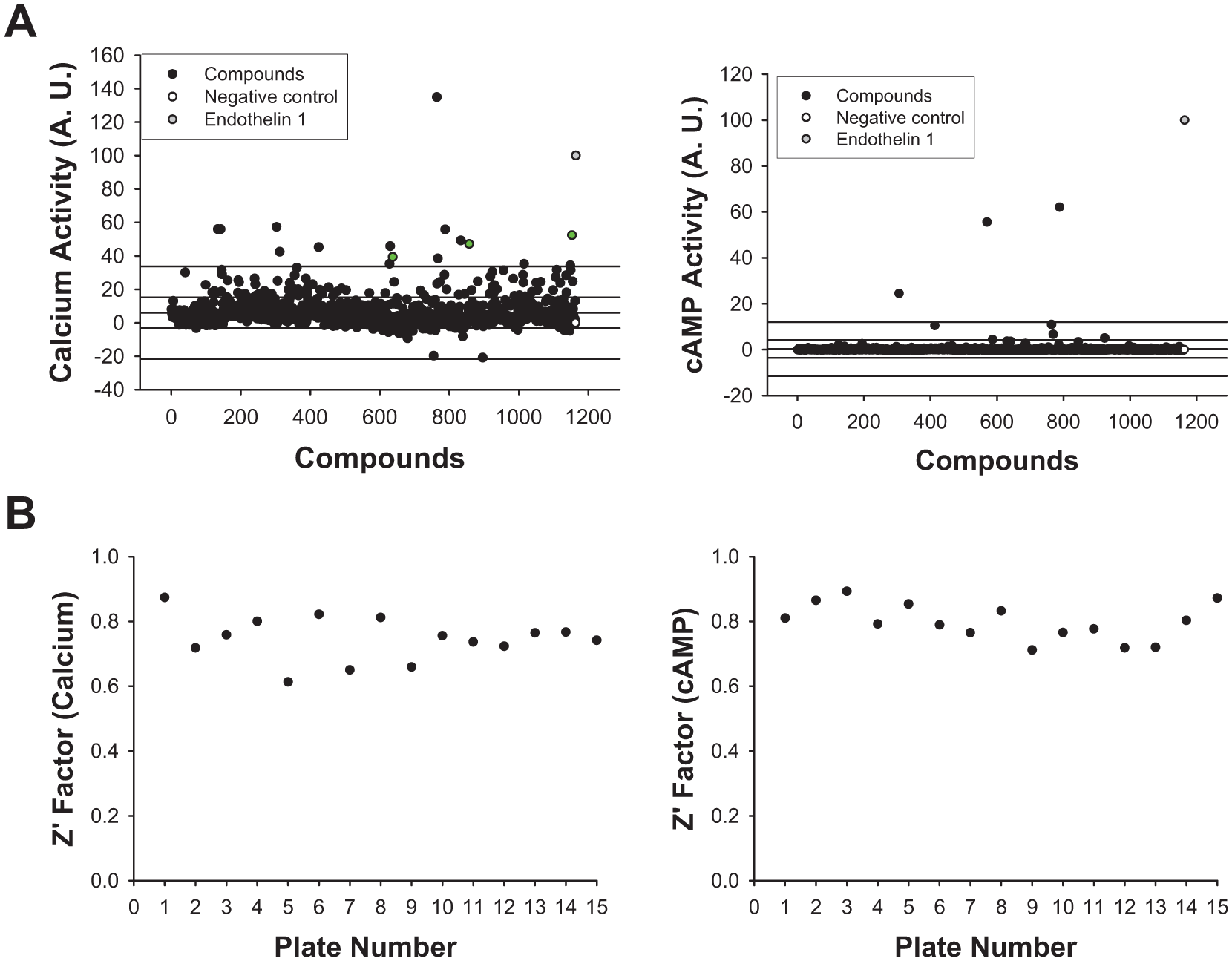
HCS of the Prestwick Chemical Library of 1200 compounds. (
Subsequently, those 17 hits were tested at different concentrations, and only the ones that presented activation in all these concentrations were selected for further analyses: lanatoside C, imatinib, and digoxigenin (green dots in 4A). The three positive compounds were demonstrated to increase the green fluorescence produced by the Ca2+Nomad biosensor, while only imatinib caused an increase in the vesicularity pattern of the cAMPNomad biosensor ( Fig. 6A ). To confirm the previous results, dose–response curves were created using these three compounds to test them for functional Ca2+ and/or cAMP responses. With this purpose, cells were treated with a range of concentrations comprising dilution points spanning from 3 nM to 3 μM for lanatoside C, from 1 nM to 3 μM for imatinib, and from 1 nM to 10 μM for digoxigenin over 24 h in 96-well plates ( Fig. 6B ). The error bars represent the standard deviations from four replicates per well. The EC50 values for lanatoside C, imatinib, and digoxigenin in the Ca2+ response assay were 3.86 × 10−7 M, 4.00 × 10−8 M, and 1.16 × 10−6 M, respectively. The corresponding Z′ factors calculated for these compounds in the Ca2+ response assay were 0.66, 0.64, and 0.67. No dose–response curves were obtained from the cAMP response assay with the exception of imatinib, for which the EC50 was 5.02 × 10−8 M, and the Z′ factor was 0.74. None of the positive compounds performed better than the natural ET1 agonist.
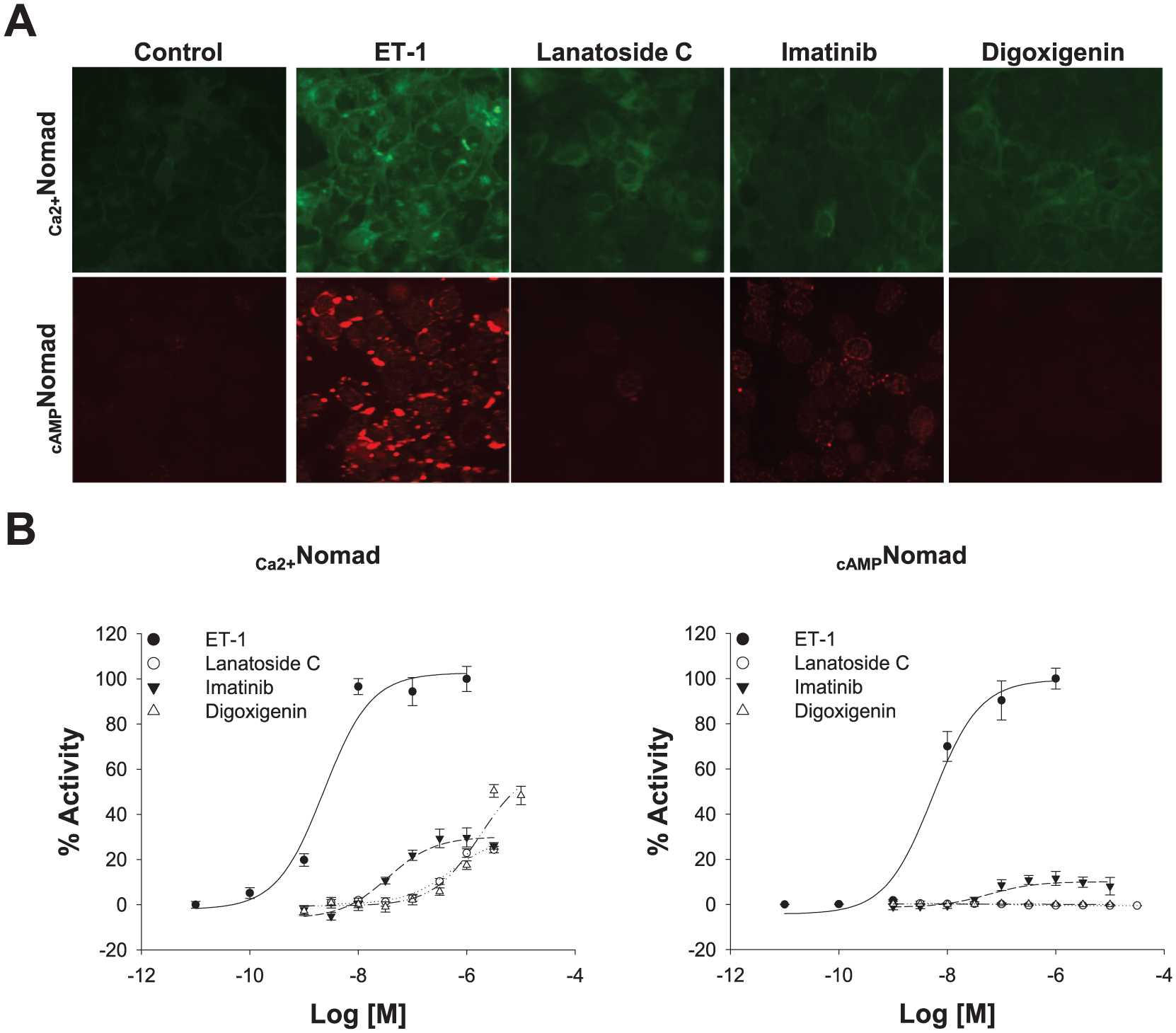
(
Discussion
The principal objective of this work was to develop a novel tool for multiplexing high-throughput cell-based GPCR assays. Our results confirm that the Nomad Biosensors allow for the detection of changes in second messengers at physiologically relevant concentrations. The Nomad Biosensors described here have been tested with different agonists and a specific antagonist of the GPCR of interest, ETBR. We have further demonstrated that Nomad Biosensors are specific for the receptor under study and that the technology is adaptable to study different types of GPCRs.
Actually, the reliability of the Nomad Biosensors for conducting screening campaigns was validated using a large library of chemical compounds with Z′ values for the assay of >0.7, which confirms the high reproducibility and robustness of this system.
Conceptually, Nomad Biosensors are FP-based reporters engineered with a single FP that is coupled to a specific signal-sensing element that transduces the signal upon ligand binding to its specific GPCR. Importantly, this design allows numerous advantages compared with other fluorescence-based sensors. For instance, the screening assays based on Forster resonance energy transfer (FRET) biosensors are often complicated to set up. The single FP-based Nomad Biosensors allow for minimal perturbation, a multiplicity of measurable parameters, and suitable timescales, offering the signals required for an automated detection. Furthermore, luminescence indicators for measuring cAMP, like the Glo sensor, 22 which have the advantage of being very sensitive, lack potential for use in multiplexed applications. On the contrary, the Nomad Biosensors can be combined within the same cells to study different second-messenger signals, allowing easy analysis of putative biased compounds.
Other cAMP sensors suitable for screening campaigns that have been recently improved are the cADDis biosensors, 23 which offer robust assay performance with Z′ > 0.75. However, this technology requires the tedious task of performing baculovirus transduction, which increases the steps and costs of the assay. The main goal of the Nomad Technology was to make the screening assays easier, less expensive, and more reliable since the assays that are currently available require additional reagents, cell lysis, complex liquid handling steps, and importantly, GPCR labeling. With Nomad Technology, the receptor under study is label-free, reducing its desensitization and the potential signaling problems that the presence of a tag could confer to the receptor.
Currently, GPCRs are among the most important therapeutic targets for the pharmaceutical industry. Indeed, global GPCR drug screening is a rising market. The discovery of the ability of some compounds to induce different signaling cascades mediated by second messengers, a phenomenon known as functional selectivity, has become a breakthrough in this field.24,25 Nomad Biosensor-based screening not only optimizes GPCR-targeted drug design but also is amenable to the testing of large numbers of compounds, emerging as a useful strategy to identify novel GPCR ligands.
In conclusion, Nomad Technology provides accurate quantitative results and represents a powerful first approach in the search for novel targets of GPCRs to provide new insights into the general mechanisms of the signal transduction of GPCRs in living cells.
Footnotes
Acknowledgements
The authors would like to thank Dr. Laura Escobar from Achucarro Basque Center for Neuroscience for her technical assistance with flow citometry. We thank Dr. Elisabete Hernández and Dr. Isbaal Ramos for helpful discussions about this work.
Supplementary material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Basque government (Global Lehian 48AFG 201700013 and Gaitek IG201500197/IG-2014/00297).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
