Abstract
A central tenet of good diagnostic laboratory practice is protecting laboratory staff from contact with sample-borne pathogens and dangerous chemicals. Automated sample-processing systems can reduce or eliminate the risk of exposure to infectious samples while providing results on par with, or better than, those from manually processed samples. In addition, hands-free automated processing may enable analysts to focus on higher order activities while eliminating the risk of repetitive strain injuries associated with manual pipetting. Here, we describe a semi-automated tuberculosis interferon-γ release assay (IGRA) workflow that includes an automated high-throughput sample-processing system. The system automates cap removal, automates sample mixing and aspiration of blood from lithium heparin collection tubes, and aliquots blood samples into multiple blood assay tubes for downstream testing without manual intervention. We show that automated results are comparable to manual methods without risk of analyst exposure or repetitive strain injury.
Introduction
Safe practices are paramount to protect laboratory personnel, especially those exposed to infectious samples and hazardous materials. At the same time, modern diagnostic laboratories face increasingly high-throughput demands with the need for quick turnaround times. Manual workflows, including repetitive activities such as pipetting, may create bottlenecks in throughput and turnaround times, introduce risks of user variability and errors, and also subject laboratory analysts to strain injuries.
A more alarming threat to laboratory personnel is that of laboratory-acquired infection due to accidental exposure. 1 Pathogens that pose the greatest risk within the diagnostic laboratory include Brucella spp., Mycobacterium tuberculosis, Salmonella spp., Shigella spp., Rickettsia spp., and Neisseria meningitidis. 2 In the case of highly communicable tuberculosis (TB), laboratory protection is especially important because although the mortality rate has decreased, this disease remains one of the top 10 causes of death worldwide. 3
Although the traditional tuberculin skin test (TST) detects TB infection, it cannot distinguish between active disease and asymptomatic latent tuberculosis infection (LTBI). 4 In addition, incidences of false positives and false negatives may occur using TST methods. 4 Newer in vitro methods use heparinized whole blood to measure CD4 and CD8 T-cell activation, in the form of interferon-γ (IFNγ) secretion, in response to peptide antigens that simulate ESAT6 and CFP10 mycobacterial proteins. A highly specific and sensitive enzyme-linked immunosorbent assay (ELISA) qualitatively detects IFNγ as an indirect indication of TB infection, and those patients testing positive may be immediately referred for advanced diagnostic evaluation and treatment.
Here, we describe the feasibility of automating blood sample aliquoting from lithium heparin collection tubes into blood assay tubes as part of a semi-automated TB interferon-γ release assay (IGRA) workflow. We use a specialized liquid-handling platform to pipette whole-blood samples from the source collection tube into each of four target blood assay tubes. Both TB antigen tube 1 (TB1) and TB antigen tube 2 (TB2) contain peptide antigens from ESAT6 and CFP10 designed to elicit an immune response from CD4 T cells. The TB2 tube contains additional peptides to induce a response from CD8 T cells. The blood assay tubes in combination provide a broad picture of the immune response. The mitogen tube is an IFNγ-positive control for each tested sample, and the negative control or “nil” tube values adjust for background interference when subtracted from the IFNγ levels detected in the other three tubes. After the blood assay tubes are thoroughly mixed and incubated, we harvest plasma and test for the presence of IFNγ via ELISA methods. We report results in International Units per milliliter (IU/mL) calculated from a prepared standard curve using standards included in the kit.
In addition to automating sample aliquoting steps, the platform also automates cap removal from the vacuum-style blood collection tubes, and mixes and aliquots blood samples into blood assay tubes for downstream testing. Because these hands-free steps do not require manual intervention, analysts have fewer opportunities to come in accidental contact with infectious samples compared to fully manual methods. We show that results are comparable to those of manual methods, and also enable analysts to focus on higher order activities.
Materials and Methods
Materials
Samples and Assay Components
QuantiFERON-TB Gold Plus (QFT-Plus) Blood Collection Tubes (cat no. 622536) consisting of antigen-coated tubes corresponding to nil, TB antigen 1 (TB1), TB antigen 2 (TB2), and mitogen; and QuantiFERON-TB Gold Plus Reference Lab Pack ELISA Kit (cat no. 622832), including microplate strips coated with murine antihuman IFNγ monoclonal antibody, IFNγ standard, diluent, conjugate, wash buffer, enzyme substrate, and enzyme-stopping solution, were donated by QIAGEN, Inc. (Germantown, MD). Vacuette tubes (6 mL, lithium heparin 13×100 mm green cap–black ring, nonridged; cat no. 456088) were obtained from Greiner Bio-One (Monroe, NC). The blood samples were obtained via volunteer blood donation programs in which volunteers donated blood specimens with written informed consent under institutional review board (IRB)-approved protocols provided by Advarra (Columbia, MD). Occupational Health Consultants (Rockville, MD) supplied nonresponsive blood samples in accordance with IRB protocol Pro00008793, and LABWORKS LLC (Springfield, VA) provided low- and high-responsive blood samples per IRB protocol Pro00008997.
Instrumentation
The Aliquot STARlet assay-ready workstation from Hamilton Company (Reno, NV) fully automates sample aliquoting of blood and other biological fluids from up to 64 source tubes to up to 256 target tubes for downstream applications. The automated prototype instrument used in this study included one pipetting channel and one specialized tool, called a VacuTube channel. In contrast, the market-ready version has four pipetting channels and four VacuTube channels, as seen in Figure 1 . With the assay-ready workstation, source collection tubes containing whole-blood samples were automatically loaded and scanned with a barcode reader to enhance workflow efficiency and protect against errors with end-to-end sample tracking. Specialized VacuTube channels automatically decapped source collection tubes, and liquid-handling channels mixed and aspirated samples. After aspiration, each sample was precisely dispensed into target blood assay tubes. A variety of blood tube and cap styles are compatible with the Aliquot STARlet, and both tubes and caps were securely positioned within their respective carriers to eliminate unintended movement or biohazardous spills and enable inversion mixing. After Aliquot STARlet distributed samples to each target blood assay tube, the VacuTube channels automatically placed the appropriate caps back onto each source collection and target blood assay tube. We then manually transferred target blood assay tubes off-deck to complete the remaining steps of the assay procedure. Optionally, analysts may choose to send downloadable Aliquot STARlet run files to an integrated Laboratory Information Management System (LIMS).
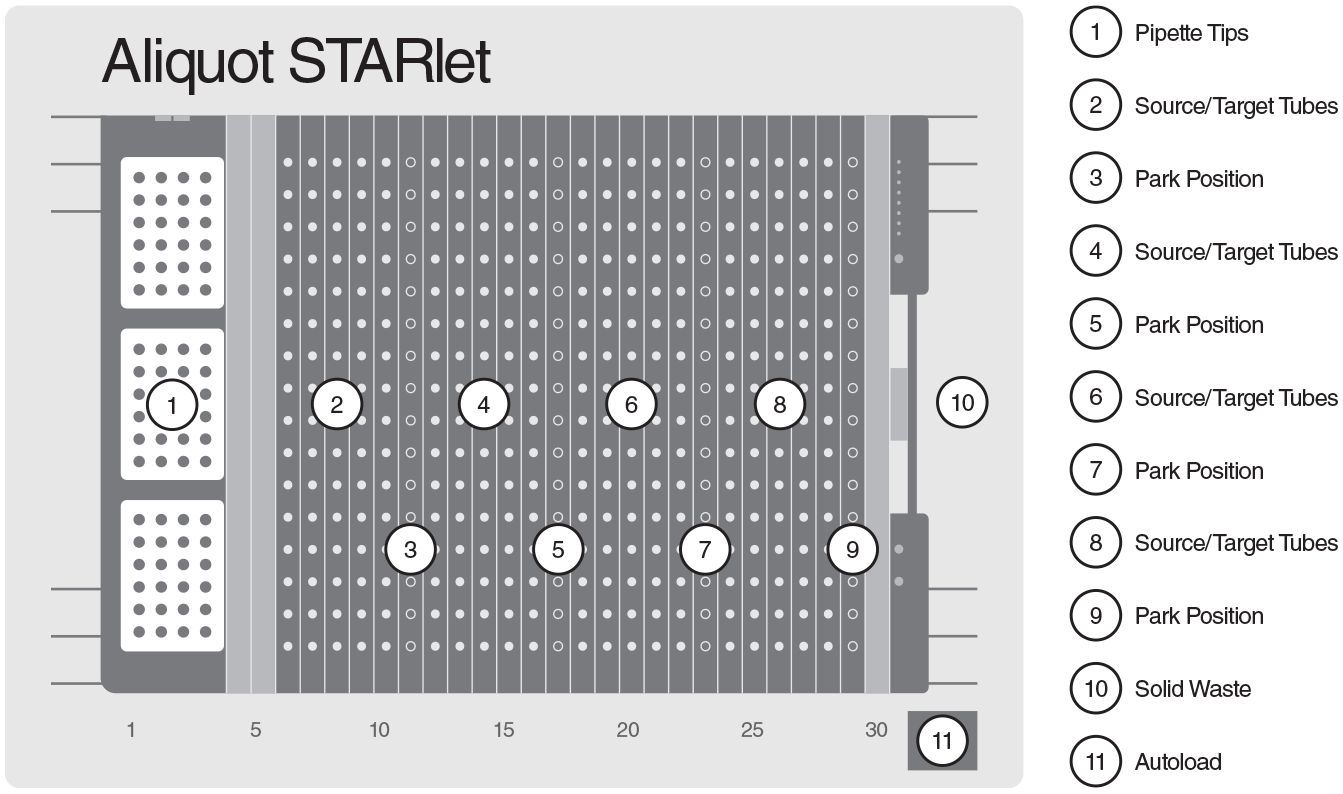
Aliquot STARlet deck layout, showing source and target tube positions as well as tip stations.
Additional instrumentation included the XS205 Dual Range analytical balance (Mettler Toledo, Columbus, OH), Sorvall Legend XT centrifuge (Thermo Fisher Scientific, Waltham, MA), HCS rotary shaker (QIAGEN), and ELx50 microplate washer and ELx800 absorbance reader (BioTek Instruments, Winooski, VT).
Methods
Dispensing Evaluation Sample Processing
We loaded two lithium heparin source collection tubes containing 6 mL of donated whole blood at room temperature along with target blood assay tubes into vial inserts. We then placed the vial inserts containing tubes into vial sample carriers, and loaded the carriers onto the Aliquot STARlet deck. Specialized VacuTube channels on the Aliquot STARlet removed source collection and target blood assay tube caps and transferred them to separate, specialized park positions. The Aliquot STARlet aspirated and dispensed the whole-blood samples three times using the liquid-handling channel with disposable CO-RE tips (Hamilton Company) to ensure complete mixing. The instrument then aspirated each whole-blood sample and dispensed 1 mL into each of four target blood assay tubes. The VacuTube channels replaced all tube caps onto their respective tubes, and we then transferred target blood assay tubes off-deck for gravimetric analysis. We also performed manual blood dispensing in parallel using additional donated blood in lithium heparin collection tubes and a manual pipette. We repeated automated and manual dispensing steps to reach a total of 40 replicates per method, after which we manually weighed each target blood assay tube and recorded the data.
Mixing Evaluation Sample Processing
We loaded a lithium heparin source collection tube containing donated whole blood at room temperature and four target blood assay TB1 tubes into a vial insert that was firmly affixed to a vial sample carrier. Then, we loaded the vial sample carrier onto the Aliquot STARlet deck. VacuTube channels on the Aliquot STARlet removed and parked source collection and target blood assay tube caps, as previously described. The Aliquot STARlet aspirated and dispensed the whole blood three times using the pipetting channel with CO-RE tips to ensure complete mixing, then it aspirated the blood and dispensed it into four target blood assay TB1 tubes. All caps were automatically replaced by the VacuTube channel, as previously described, and we repeated mixing and aliquoting five additional times in the same manner. We repeated the process, including replicates, using a second blood sample from the same donor for a total of 24 target blood assay TB1 tubes. After completion, we transferred the target blood assay tubes off-deck, where we manually shook them 10 times using an up-and-down motion and placed them into a 37 °C ± 1 °C incubator for 16–24 h. We also manually processed donor blood with a total of 24 target blood assay tube replicates using a handheld pipette according to the package insert procedure.
Signal Comparison Sample Processing
We processed samples in the same automated manner as those in the mixing evaluation study, and aliquoted them into the nil, TB1, TB2 and mitogen target blood assay tubes. Likewise, we performed manual processing using the same donor’s blood sample. The process was repeated throughout time using whole-blood samples from a total of 23 donors.
Assay Procedure
After incubation, we centrifuged manually and automatically prepared target blood assay tubes at 2000–3000 relative centrifugal force (RCF) (g) for 15 min until the gel plug separated cells from plasma. After removing tubes from the centrifuge, we used a manual pipette to harvest plasma from each target blood assay tube into a cluster tube. Before starting the ELISA, we prepared the working-strength conjugate and four dilutions of IFNγ standard. To start the QFT-Plus ELISA, we added 50 µL of working-strength conjugate to the ELISA plate well, followed by 50 µL of harvested plasma from each manually and automatically prepared sample. Then, we dispensed 50 µL of each diluted IFNγ standard into the appropriate wells.
We covered and mixed the plate for 1 min at 500–1000 rpm, incubated it at room temperature for 120 ± 5 min, then washed the plate wells at least six times with 400 µL of working-strength wash buffer. The washing step included a 5 s soak period between each wash cycle. After removing residual wash buffer from the plate, we added 100 µL of enzyme substrate solution to each well, covered and mixed the plate for 1 min at 500–1000 rpm, and incubated it at room temperature for 30 min. After incubation, we added 50 µL of enzyme-stopping solution to each well and shook the plate on a plate shaker to mix the well contents. Within 5 min of stopping the reaction, we transferred the plate to a microplate reader for absorbance reading at 450 nm with a reference wavelength between 620 and 650 nm.
Results and Discussion
Dispensing Evaluation
We compared manual and automated dispensing performance using a total of 40 whole-blood samples that were aliquoted from the source collection tube to the four target blood assay tubes. For gravimetric assessment of dispensing, we determined each donor’s blood density and used it to calculate dispensed volume. When comparing plotted results per Figure 2 , blood assay tubes dispensed using manual methods exhibit a wider degree of variation than those dispensed using the automated method. In addition, per Table 1 , the automated dispensing method resulted in a mean dispensing volume closer to the target 1000 µL, with threefold higher precision and accuracy compared to the manual method.
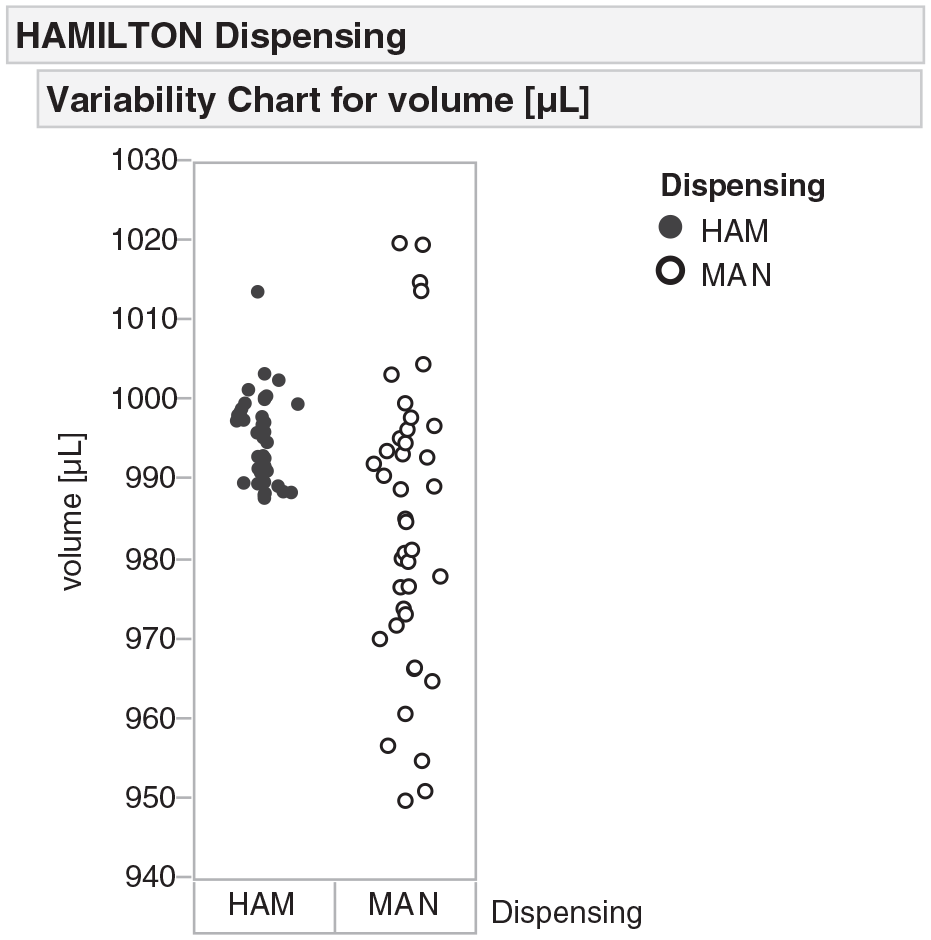
Gravimetric analysis of whole-blood sample dispensing. Comparison of 10 source tubes dispensed into 40 destination tubes either manually (MAN) or by the automated instrument (HAM). A wide degree of variation occurs using the manual methods, while samples dispensed using automated method are more consistent.
Gravimetric-Based Comparison of Manual and Automated Dispensing Methods.
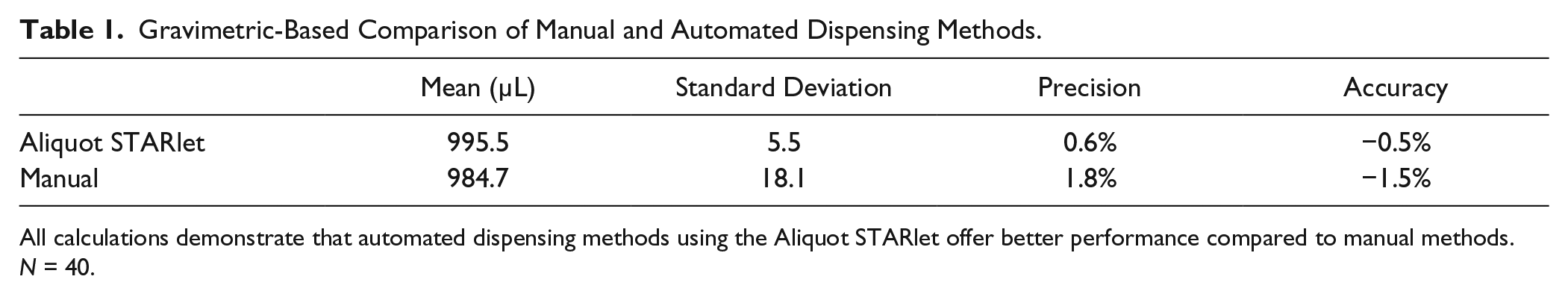
All calculations demonstrate that automated dispensing methods using the Aliquot STARlet offer better performance compared to manual methods. N = 40.
Mixing Evaluation
Because whole blood is a mixture of solids in a liquid (blood cells in plasma), proper mixing is necessary to ensure a homogeneous sample component distribution prior to aliquoting for downstream testing. Therefore, using antigen response data from TB nonresponsive, low-responsive, and responsive whole-blood samples, a comparison was made between the mixing performance of the automated method and manual method. Per each of the three donor samples in lithium heparin source collection tubes, we used manual and automated methods to aliquot replicate whole-blood samples into target blood assay tubes. Then we performed the QFT-Plus test, as previously described, for a total of 48 assayed samples. To remove sources of variability between manually and automatically prepared samples, we processed them in parallel using a single ELISA plate. We plotted antigen response results in IU/mL from each target blood assay tube per Figure 3 , showing similar replicate data for each whole-blood sample subject. The difference in mean response from the collective 48 assayed samples was −0.045, and a P-value was 0.889, confirming that automated mixing methods are equivalent to or better than manual mixing methods. The automated methods also reduced the unnecessary risk of exposing laboratory personnel to hazardous blood samples.
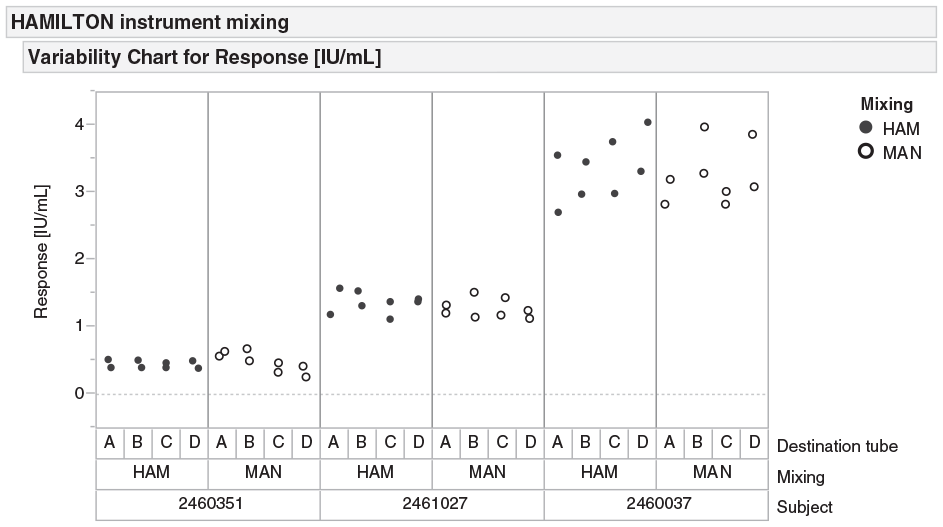
Antigen response in whole-blood samples of three different subjects. When comparing results, values produced via the automated system (HAM) are on par with or greater than those produced using the manual method (MAN).
Signal Comparison
We made a final comparison by measuring the stimulation response level after manual and automated blood aliquoting. Using Aliquot STARlet or a manual pipette, we aliquoted three groups of samples (TB nonresponsive, low-responsive, and responsive) in parallel. After stimulation and plasma harvesting, we determined IFNγ concentration in plasma by the QFT-Plus ELISA according to the instructions for use. These data are shown in
Figure 4
. In five TB responsive samples (
Fig. 4A
), automated methods yielded comparable signal recovery, expected variability and fluctuation in data points, and no discrepancy in data values between automated and manual methods. Six TB low-responsive samples (
Fig. 4B
) yielded results similar to the responsive sample results. Finally, automated methods yielded expected results in TB nonresponsive samples (
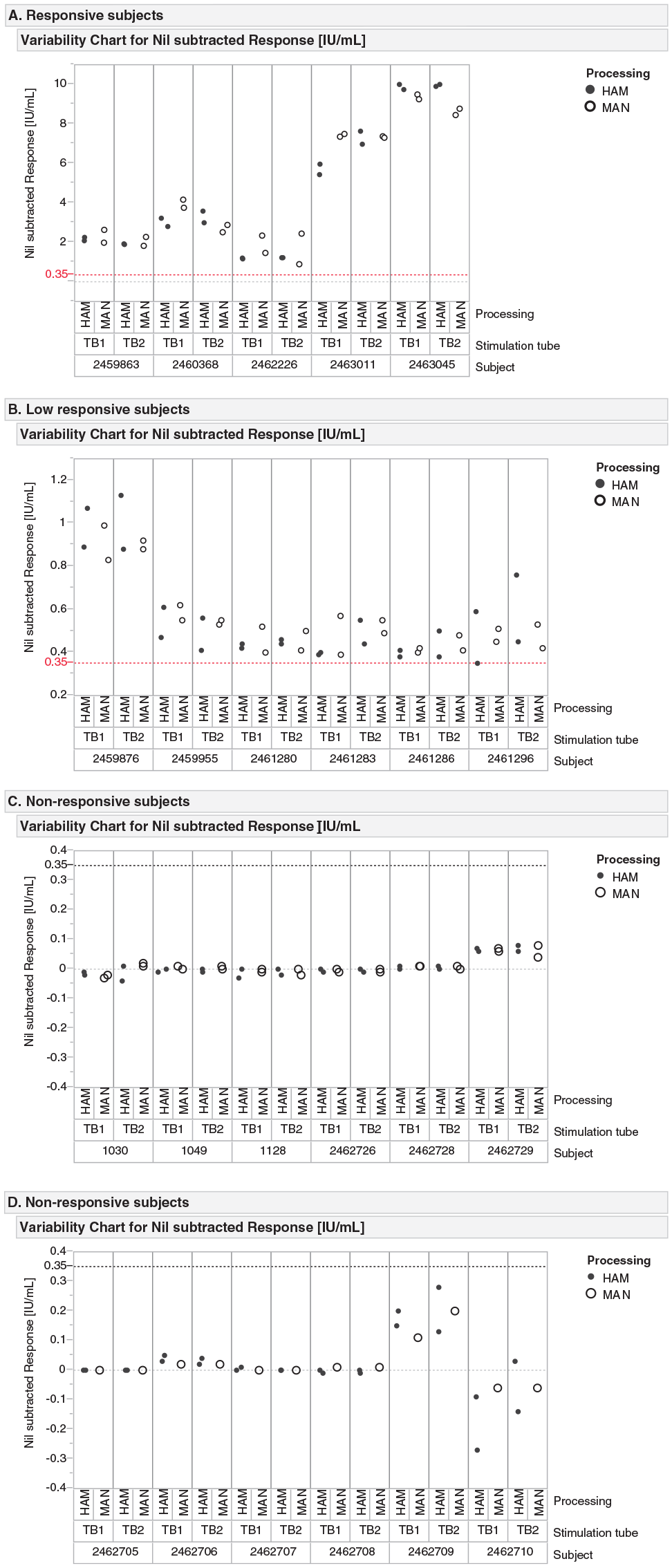
Nil subtracted antigen response from (
Footnotes
Acknowledgements
We previously presented these data in “Specialized Automated Workflows to Mitigate Risk of Infectious Sample Exposure to Testing Personnel,” at the SLAS 2019 International Conference & Exhibition, February 2–6, 2019, poster ID 539036.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Neil Grossman, Peter Haviernik, John Wolff, and Chih-lung Fu are full-time employees at QIAGEN, Inc., in their respective positions. QIAGEN manufactures and sells assay chemistries such as the type discussed in this article. Kevin Miller, Brandon Bare, and Eric Sindelar are full-time employees at Hamilton Company in their respective positions. Hamilton Company manufactures and sells automated liquid-handling platforms such as the one discussed in this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
