Abstract
Bacterial infection is a leading cause of morbidity and mortality (from infants to the elderly) and accounts for more than $20 billion in healthcare costs in the United States each year. The pathogens responsible for many of the common infectious diseases, such as urinary tract infection (UTI) and ventilator-associated infections (VAIs), have proven to be highly adept in acquiring mechanisms of antimicrobial resistance. The use of broad-spectrum antibiotics by healthcare providers and the infiltration of antibiotics in the environment have accelerated the selection and growth of resistant pathogens. To further exacerbate the problem, the need for new antibiotics has far outpaced the development of new classes of antibiotics by the pharmaceutical industry (only two new classes of antibiotics have reached the market in the last 20 years), in large part due to prohibitive cost and historically poor return on investment to develop new antibiotics. Consequently, clinicians have limited treatment options, particularly in the neediest patients. To tackle this major global health issue, we are developing novel technological approaches for rapid definitive clinical microbiological analysis. These technologies will improve the clinical management of bacterial infections and reduce the improper use of antibiotics in current practice, hopefully limiting the spread of drug-resistant organisms.
Guidelines from the Infectious Diseases Society of America (IDSA) and the American Thoracic Society (ATS) recommend that early, appropriate antibiotics for bacterial infections should be prescribed in adequate doses. 1 However, discerning which antibiotics are “appropriate” is not always easy, and technical hurdles currently limit the ability of providers to make this determination swiftly. As a result, most patients with infections are initially treated with broad-spectrum antibacterial agents for 2–3 days until laboratory testing identifies the causative bug responsible for the infection. Ideally, clinicians and clinical labs would have the capability to make this identification much sooner. In this case, instead of using a sledgehammer approach to antibiotic selection, providers could craft a personalized drug regimen that targets specific causative organisms without overwhelming the body’s normal microbiota.
Bacterial culture, the cornerstone of care for infected patients, lacks sensitivity and more importantly is the most time-consuming step in the clinical microbiology workflow. For instance, clinical specimens such as urine or blood are cultured overnight on agar plates or blood tubes for growth, concentration, identification, and isolation in the clinical microbiology laboratory. Rapid microbiological analysis techniques that identify antimicrobial susceptibility phenotypically (which is independent of the evolving genetic basis of antibiotic resistance) represent the most significant unmet needs in clinical microbiological analysis. In this study, our approach rapidly determines the presence of bacteria and classifies them according to their shape and size for rapid phenotypic AST. 2 To demonstrate the clinical application potential, we designed our study using clinical urine samples, including negative samples.
The ability of the single-cell microbiological analysis system to rule out or confirm the presence of bacteria at a clinically relevant concentration, which is demonstrated in this study, could improve patient care and clinical lab workflow. Furthermore, the classification scheme, which includes Staphylococcus-like, Enterococcus-like, Pseudomonas-like, Klebsiella-like, and Escherichia coli-like bacteria, is tailored to identify several of the most common and deadly human pathogens. Rapid classification of the predominant pathogen in a sample can accelerate the selection of appropriate antibiotics for AST as well as molecular probes for pathogen identification. Importantly, using the single-cell AST system, pathogens can be classified based on morphology and genetic features. Moreover, antimicrobial resistance profiles of the bacteria can be determined in as few as 30 min, offering the potential for major improvements in clinical care.
Looking forward, the rapid single-cell microbiological analysis system will have a profound impact beyond pathogen detection.3–5 For instance, the effectiveness of the single-cell analysis system will enable frequent monitoring of the human microbiome, even among individuals without an infection. Increasingly, it is recognized that a wide range of diseases (from diabetes to cancer to inflammatory bowel disease) are impacted by the state of the microbiome. While it is obviously essential to determine if pathogens are present within clinical samples, it is also important to assess whether the body’s commensal microbes (the microbiome) are intact or have been displaced.6,7 However, there is currently no efficient way to make this assessment in routine clinical care. Current practice patterns thus tend to ignore the microbiota entirely unless patients have infections or are considered at unusually high risk for infection. Our study offers a first step toward a precision approach to this problem leveraging advancements in informatics, microbiology, and engineering ( Fig. 1 ). These advancements will eventually empower providers not only to react swiftly to the appearance of a specific pathogen, but also to proactively monitor the “health” of the microbiome in real time.
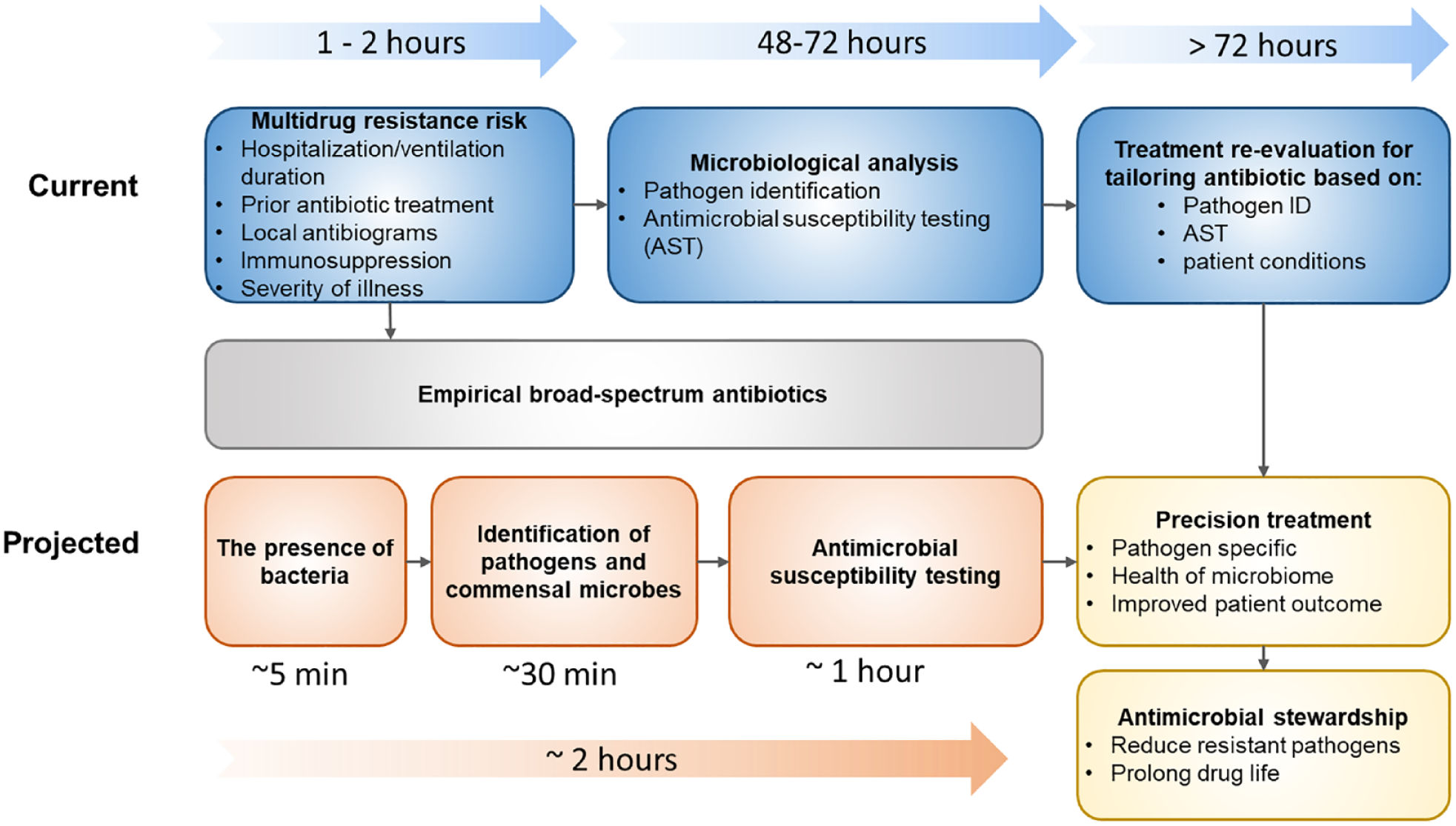
Rapid microbiological analysis for precision management of infections and dysbiosis.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is supported by the U.S. Defense Threat Reduction Agency (DTRA) (HDTRA114-AMD1), NIH NIAID (R01AI117032), and the Penn State Research Fund.
