Abstract
Recent advancements in science and engineering are revolutionizing our understanding of an individual’s disease, and with this knowledge we are gaining an increasingly sophisticated understanding of how discovery can be transformed to deliver personalized medicines. To reach this future state, we must reengineer our approach to enable the use of more relevant human cellular models earlier in the drug discovery process. Stem cells and primary human cells represent more disease-relevant models than immortalized cell lines; however, due to both availability and cost, their use is limited in lead generation activities. Miniaturization of cellular assays below microtiter plate volumes will enable the use of more relevant cells in screening, but this would require a change in how test molecules are introduced to the biology. With these shifting paradigms, Discovery Supply teams at GlaxoSmithKline (GSK) are modernizing our sample handling approaches. Various emerging technologies such as microarrays, nanowells, and microfluidic devices could bring fundamental changes in conventional sample handling support as we transition from microtiter plates to well-less platforms. The discussion here is exploratory in nature and reviews ongoing proof-of-concept experiments. Our ultimate goal is to industrialize the sample management platforms to support future miniaturized biological assay systems.
Introduction
Most large Pharma companies maintain libraries of millions of small molecules to support their lead generation and optimization efforts. To enable high-throughput and reliable screening support, there have been significant investments in large infrastructure for automated storage, retrieval, and dispensing of these small-molecule compound libraries. These facilities tend to be complex, occupy large square footage in laboratories, and are built to serve standard microtiter plate formats. Over time, compound handling groups have made tremendous efforts to increase the density and miniaturization of sample supply from 96- to 384- to 1536-well plates and reduce sample supply volumes from microliters to nanoliters. In addition to the advancements in compound handling and screening platforms, the Pharma industry has made tremendous efforts in the augmentation and modernization of their small-molecule libraries, adding new chemically diverse and more lead-like and drug-like compounds. Over the last two decades, due to the advancement in screening platforms and imaging techniques, the speed of screening millions of compounds has accelerated dramatically. Similarly, computational tools and techniques have also been developed to meet the growing demand for analytics of the large data sets generated from these high-throughput screens (HTSs).
With these continued technological advances, program success rates are low, as evidenced from data showing that 15% or less of molecules entering development phases successfully transition to the clinic.1–4 This is without including the failure of transitioning HTS hits to candidate molecules, which is largely undocumented in the literature. High attrition rates of drug candidates are accompanied by long cycle times and increasing costs required to bring drugs to the market. The current drug discovery model remains a largely linear and iterative process ( Fig. 1 ) that was designed in the 1980s to meet the paradigm of “one-size-fits-all” medicine, with many drugs lacking efficacy in 50% or more of the population. 5 This linear and iterative approach is a slow, resource-intensive process where less relevant cell systems such as generic cell lines are commonly used in hit identification efforts. Our goal is to enable the use of more human relevant cell systems at all phases of drug discovery, including the hit identification and lead optimization phases, with the expectation that hits identified with more relevant cell systems will translate more effectively to the clinic. One way to enable the use of these cell systems is to miniaturize the cellular assays used in early drug discovery, such that fewer cells per compound test would be needed. The key emphasis of this article is on miniaturized biology, focusing on the challenges and opportunities for handling small-molecule samples. The technologies for handling compounds and introducing them into ultra-miniaturized cellular assays are explored, along with an array-based technology for cellular assays.
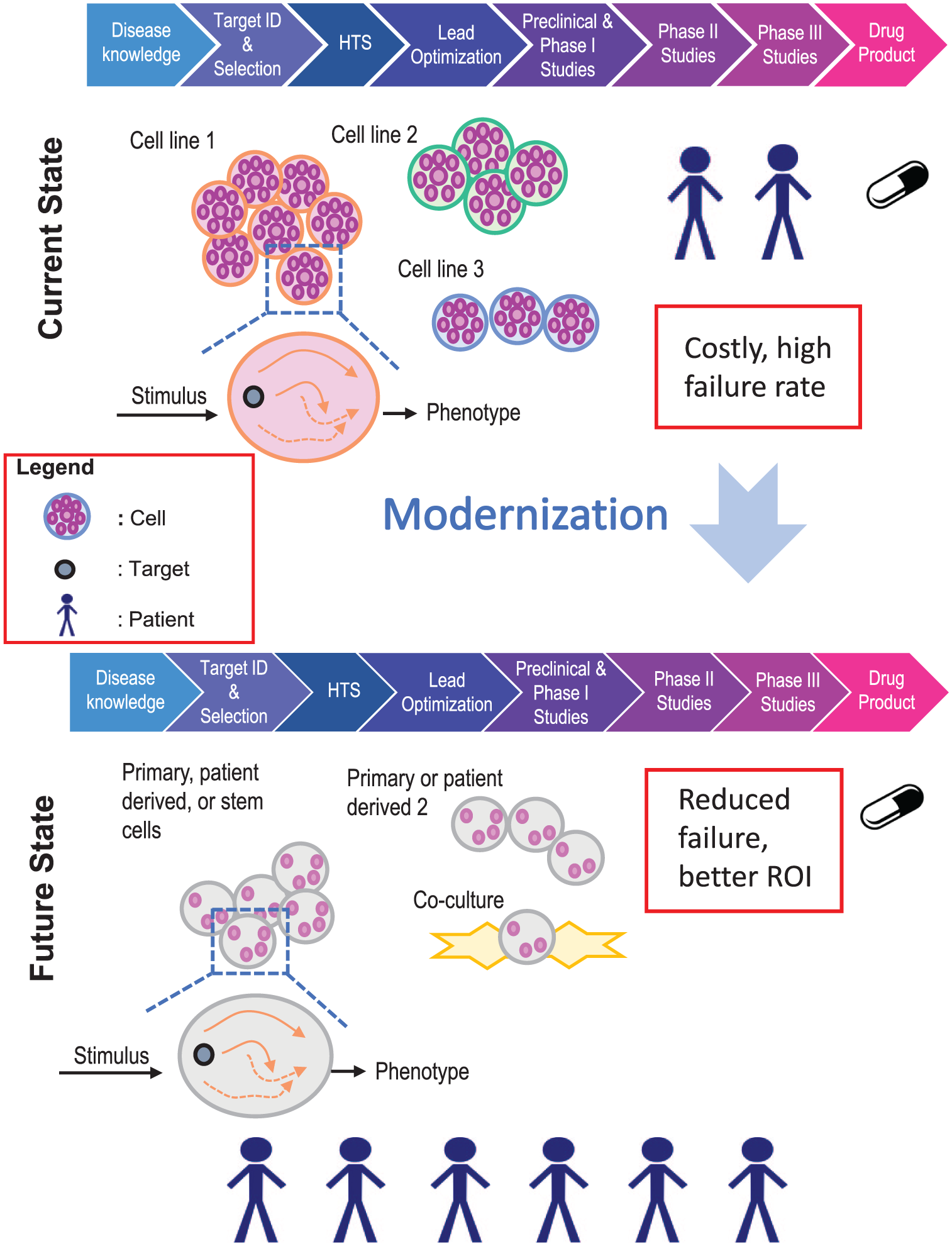
Schematic representation of the current drug discovery model and the desired future state, where disease-relevant cell models and patient samples drive drug discovery at the early stages of the process.
New Approaches in Sample Compound Handling
Sample management infrastructure at GlaxoSmithKline (GSK) integrates automation, IT and robotic, to exploit small-molecule and biological collections to support the current screening paradigm. Millions of small-molecule samples are stored in a plate-based format in 100% DMSO, with the capability to dispense compounds with noncontact, tipless technology down to volumes as low as 2.5 nL. Target assay plates include 1536-well microtiter plates and analytic support matrices, for example, Maldi. 6 Individual compounds are managed at the container level using 2D barcodes by an integrated software platform that interconnects systems, from the registration of incoming compounds to the preparation of solutions for assay-ready plate delivery.
New paradigms for sampling handling will be needed to support new ultra-miniaturized platforms. Within this context, we identified several technologies and explored them for their potential use to deliver sample handling solutions in support of miniaturized biology. Considering the current landscape of technology and developing science, we chose to pursue proof-of-concept experiments using an array-based approach. While technologies such as microfluidics7–10 could ultimately be useful in handling and dispensing compounds in a high-throughput environment in the future, the compound presentation represents a significant challenge and would require major changes to sample management infrastructure. The array technology provides an approach that allows the addition of compounds directly onto the array with both existing liquid handling capability and instruments brought in for evaluation.
Numerous commercially available array printers were identified, with wide-ranging technologies such as noncontact acoustic, contact, and noncontact piezo-based pin tools and injection-based dispensers. Based on instrument capability and compatibility with experimental plans, we selected two instruments to test for compound dispensing from the existing plate-based libraries onto an array-based assay system. These two instruments were the acoustic Echo 555 dispenser by Labcyte (San Jose, CA) and solid contact pin 2470 Arrayer from Aushon Biosystems (Billerica, MA), a company owned by Quanterix.
The Echo is an adaptive focused acoustic dispenser capable of dispensing increments of 2.5 nL droplets onto any substrate type. Its technology utilizes focused acoustic energy that precisely transfers solutions from a source to an assay substrate.11,12 It is noncontact, fast, and accurate, and can handle multiple solution types from DMSO to aqueous-based surfactants. GSK Discovery Supply’s HTS liquid collection is dispensed into assay-ready plates using Echo units. We sought to explore whether the Echo could be utilized in this ultra-miniaturized space, as a cost-effective and simple approach to maintain data integrity for both dispensing compounds and tracking locations, as data integration tools were already in place. The Echo was utilized for compound sample dispensing at its low limit volume dispense of 2.5 nL.
The 2470 Arrayer is a fully enclosed automated printer used for the fabrication of microarrays of biological materials (typically protein and antibody arrays). Also of note is that this instrument has been used to generate microarrays of covalently bound small molecules for use in protein binding screens. 13 The 2470 Arrayer uses a proprietary solid pin ( Fig. 2A ) printing system that is capable of precise deposition volumes ranging from 150 to 11,225 pL, depending on pin size diameters, which range from 85 to 350 µM. It comes with environmental control enabling the liquid transfer area to maintain constant humidity up to 80%. The pins of the instrument were designed to pick and dispense liquid from the source to a destination. The liquid remains held by surface tension on the pin, before moving the pin to a solid substrate, where the pins touch that surface to release some liquid.
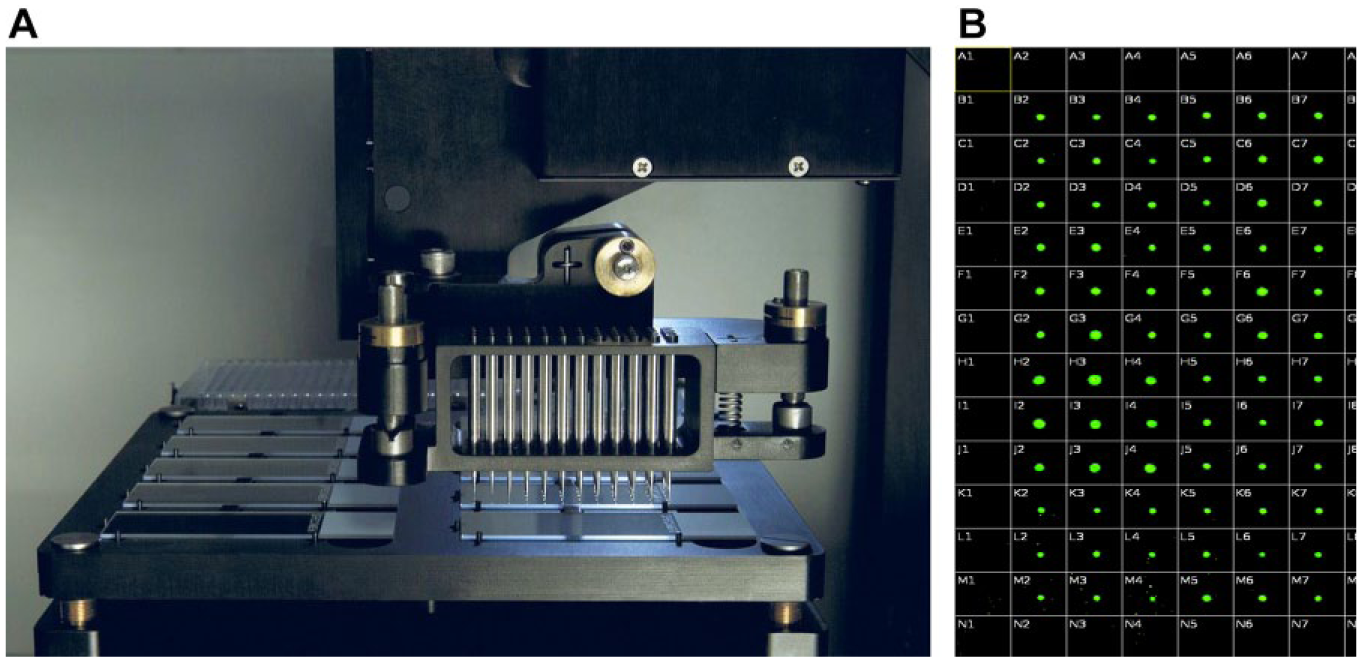
The 2470 Arrayer from Aushon Biosystems. (
These two liquid handlers were evaluated for compound printing, dispense times, evaporation rates, and compatibility with miniaturized cellular arrays. Sample reconstitution was also explored as a way to preprint and deliver compounds on slides in a high-throughput environment.
Materials and Methods
Instrumentation
The dispensing instruments were Labcyte’s Echo 555 and the 2470 Arrayer from Aushon Biosystems. Fluorescent microscopy was conducted using the GE InCell 2200.
Materials
The microtiter source plates used included the Greiner clear, flat-bottom, 384-well plate (cat. 781201); Nunc 384 Well PP Plate (cat. 267459); and Greiner clear, V-bottom, 384-well plate (cat. 781280).
Aquarray slides (1 mm) were purchased under agreement from the Karlsruhe Institute of Technology KIT/Aquarray (Karlsruhe, Germany).
The petri dish used for slide incubation was from Thermo Fisher Scientific (cat. R37112). U-2 OS cells were obtained from ATCC (cat. HTB-96). Stem cell derived cardiomyocytes were purchased from Cellular Dynamics (Madison, WI), and grown as per the vendor’s instructions, in media supplied with the cells.
The following was purchased from Gibco: Dulbecco’s Modified Eagle’s Medium Nutritional Mixture F-12 (Ham) without phenol red (DMEM/F12 [1:1]; cat. 11039), GlutaMax (cat. 35050), and 1× Hank’s balance salt solution (HBSS) with calcium chloride and magnesium chloride (cat. 14025-126).
Fetal bovine serum (cat. 26140095) and Hoechst 3334 (cat. H3570) were obtained from Life Technologies.
Cytochalasin D (cat. C8273), high-performance liquid chromatography (HPLC)-grade acetonitrile (cat. 43851-4L), HPLC-grade DMSO (cat. 34869-4L), and HPLC-grade water (cat. 270733-4L) were purchased from Sigma-Aldrich (St. Louis, MO). Formaldehyde solution (3.7%; cat. 15682) was obtained from Electron Microscopy Sciences.
For the ActinRed 555 ReadyProbes reagent, fluorescent microscopy images were captured with the GE InCell 2200 instrument, and bright field images were captured with an iPhone from an OLYMPUS IX50 Phase Contrast Inverted Microscope.
Formic acid (cat. M1116701000) was from EMD Millipore.
Cell Staining and Imaging Protocol
Cells were harvested and resuspended in media at 2 × 105 cells/mL and applied to the surface of the glass array slide with a pipette tip. After several minutes, excess liquid was removed either by tilting the slide or by aspirating excess liquid. The slide was placed in a covered petri dish containing a wet pad to maintain high humidity and placed in a tissue culture incubator. After 4 h, compounds were added with the indicated instruments. The slide was then returned to the incubator overnight, and the following day the slide was rinsed and then fixed with 3.7% formaldehyde in HBSS buffer, permeabilized with 0.1% Triton in HBSS, and then stained with Hoechst 33342 (3 µM in HBSS buffer) and phalloidin reagent at 1 drop/5 mL HBSS. After 1 h, the slide was rinsed three times with HBSS buffer and imaged on the GE InCell 2200 instrument or with a bright field microscope and iPhone. Numeric data from fluorescent micrographs were derived from the images using the Harmony image analysis software within the Columbus system (PerkinElmer, Waltham, MA).
LC/MS Sample Preparation and Protocol
Three hundred fifty-two compounds (20 µL,10 mM DMSO solution) were dispensed in duplicate into two Greiner clear, V-bottom, 384-well plates. Plate 1 was analyzed immediately by Waters Acquity (Manchester, UK) liquid chromatography/mass spectrometry (LC/MS). Plate 2 underwent solvent evaporation using a Genevac HT-24 (4 h, full vacuum at 1050 rPM, chamber temperature 40 °C throughout). Following solvent evaporation, samples were reconstituted using 20 µL of DMEM cell growth medium and incubated at 37 °C for 24 h, mimicking current assay protocols. Plate 2 was then sent to LC/MS analysis.
Analyses were performed using a Waters Acquity ultra-performance liquid chromatography (UPLC) instrument with a Waters Acquity BEH C18 column 1.7 μm (50 × 2.1 mm i.d.) held at 50 °C and running a 2 min linear gradient (0.1%–100% B in 1.5 min, held for 0.4 min, 100%–3% B in 0.1 min). Reagent A was water containing 0.1% (v/v) formic acid. Reagent B was acetonitrile containing 0.1% (v/v) formic acid. UV diode array detection was a summed signal from 210 to 350 nm. The evaporative light scattering detector (ELSD) gas pressure was 40 psi with a drift tube temperature of 50 °C and nebulizer temperature of 12 °C. Mass detection was by a Waters QDa mass spectrometer, alternating scans between positive and negative electrospray ionization (scan range 100–250 AMU, scan time 0.27 s with an interscan delay of 0.10 s). Reconstitution was measured by comparing plate 1’s compound peak areas with those of plate 2 areas measured after resolubilization.
Results and Discussion
Proof-of-Concept Compound Handling Experiments
The Labcyte Echo and Aushon Biosystems 2470 Arrayer were used in proof-of-concept experiments focusing on the addition of compounds to cells that had been arrayed onto a glass surface. The goal was to test an array-based platform for adherent cell assays with a high-content imaging readout. These initial experiments were aimed at assessing the biology and development of a miniaturized high-content imaging readout, and testing the two liquid handlers for their ability to add compounds to the cells. These experiments represent early proof-of-concept ideas toward the ultimate assessment of the feasibility of running cellular small-molecule screens in an ultra-miniaturized platform.
Labcyte Echo Evaluation
The first Echo experiment explored the maximum droplet density in terms of spatial arrangement and was also aimed at recording the dispense time and droplet evaporation time. Using the Echo to dispense 2.5 nL, we tested various droplet-to-droplet pitches ranging from 500 to 1000 µm on a glass slide. Experiments showed that the Echo successfully achieved a reproducible density of 1820 spots, with a pitch of 750 µm center to center ( Fig. 3 ). The dispense time was 4 min 42 s and DMSO spots began evaporating after 35 min of resting on an open benchtop at room temperature.
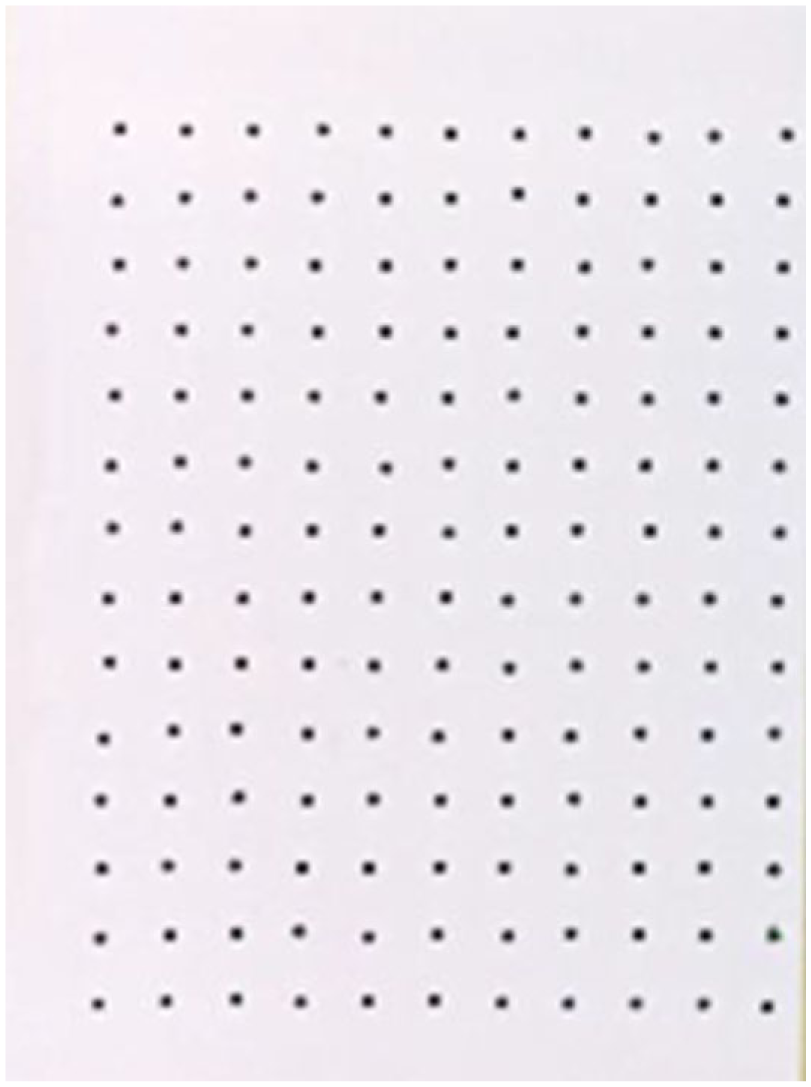

Image showing a 2.5 nL dispense of black dye at 1.5 mm pitch spacing with the Echo 555 acoustic dispenser.
The next Echo experiment investigated aqueous dispenses and evaporation rates using the same conditions as above. Although dispense precision and accuracy were on target, evaporation was evident almost instantly, with spots having already evaporated by the time the dispensing was completed. These initial experiments gave indications of the rapid evaporation of these small-volume samples and underlined the need to consider how drying of samples might affect their activity in future experiments.
Aushon Biosystems 2470 Arrayer Evaluation
The first Aushon experiment tested the transfer of Bodipy dye in 100% DMSO from a Nunc 384 Well PP Plate to an Aquarray slide at a drop volume of 350 pL using 110 µm pins in an 80% humidified environment. A 16-pin printing head with 110 µm pins (4 × 4 well orientation) was used to transfer liquid from a Nunc 384 Well PP source plate into 12 × 12 grids/field on a 1 mm spot slide, and images were taken on the InCell 2200 imager ( Fig. 3B ). The printer demonstrated precise spotting capability; however, the total transfer time for 384 compounds took more than 4 min to complete, resulting in evaporation, even with a high-humidity setting. This is due to the fact that the design of the instrument did not allow for free humidified air circulation throughout the inner chambers of the instrument. In addition, repeat transfers were needed with only 16 pins in the head, extending the dispense time and exacerbating the evaporation issues. The manufacturer offers a head with 48 maximum pins; a customized pin head with a greater number of pins would be desirable to help reduce dispense time and therefore evaporation.
Miniaturized Cellular Detection Technologies
There are many emerging platforms where miniaturized cellular readouts have been implemented. Some systems have been designed to detect secreted antibodies or cytokines from single cells, and can be applied more broadly to clonal cell line selection of antibody-producing cell lines. 14 In the functional genomics area, transcript detection at the single-cell level has become more and more mainstream, with the application of analytics to realize new capabilities in this area.15,16 While single-cell analyses can be done, none of these platforms were designed for high-throughput, small-molecule screening applications. Our goal for cellular assay miniaturization proof-of-concept experiments in arrays is to explore miniaturized cellular assay platforms where fewer cells could be tested for each compound tested, with the ultimate desire to scale the experiments up to thousands or millions of compounds for any given assay. We chose to combine an open array format with high-content imaging, so that the open array could be accessible for compound delivery, as a testing ground for miniaturization beyond the 1536-well format.
Array-Based Cellular Assays
For biochemical screening, an HTS of 2 million compounds in 1536-well plate format can be performed in less than 1 week. Challenges are faced when biology becomes complex, where cells require maintenance and differentiation, and compound incubation times extend into days. The logistics of such experiments become insurmountable, and the labor, time, and money needed for the growth, expansion, and maintenance of complex cell systems becomes prohibitive. In many cases, media exchanges and long incubation times restrict these assays to 384-well plates, because of the technical challenges encountered in the adaptation of complex assays into 1536-well plates. With the goal of utilizing disease-relevant cell systems in early drug discovery, we have begun to explore the use of arrays for miniaturized cellular assays, in a small-molecule screening environment.
Development of High-Content Image-Based Readout
As a test system, we initiated experiments with an array-based platform, where alternating hydrophobic and hydrophilic patterning on the surface creates an array onto which cells can be grown. The published accounts of this device have shown successful results with cellular image-based assays. 17 This array device was developed at the Karlsruhe Institute of Technology by Dr. P. Levkin and colleagues, and was purchased through their commercial entity, Aquarray. We aimed to do proof-of-concept experiments in a drug discovery setting to understand the device’s applicability to small-molecule screening.
Shown in Figure 4 are images taken during the process of loading cells onto the array. First, an aqueous cell suspension was loaded onto the surface of the device, as shown in Figure 4A . Then the excess liquid was removed by tilting the slide and allowing excess liquid to run off, leaving behind liquid trapped in the hydrophilic areas of the pattern on the slide ( Fig. 4B ). On the 1 mm array shown, the volume trapped in each area of the array was approximately 80 nL, based on the vendor’s prior characterization of this device. Examination under a microscope using bright field showed that cells were contained within the aqueous liquid that was trapped on the array ( Fig. 4C ), increasing magnifications using 4×, 10×, and 20× objectives.
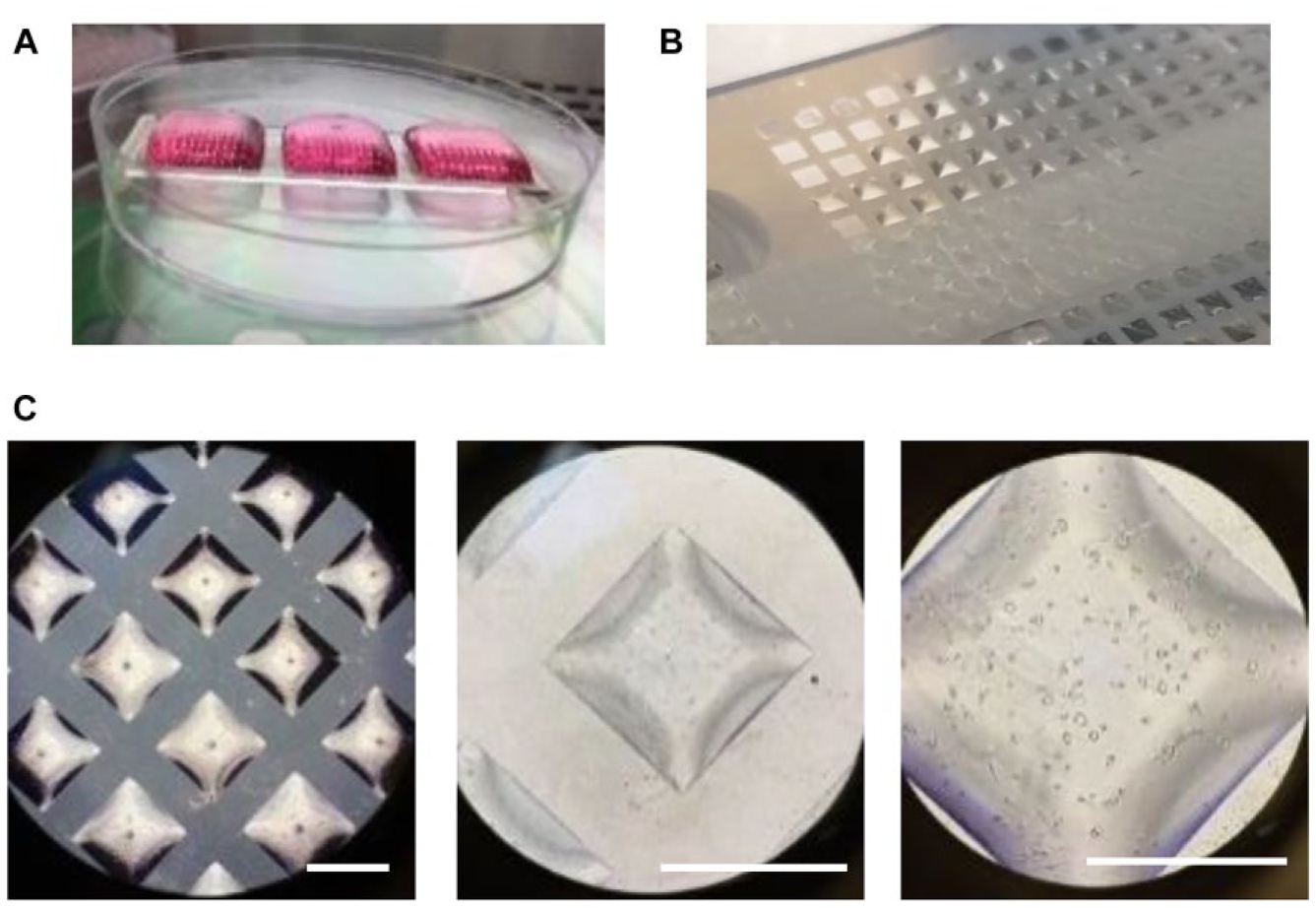
Seeding cells onto the Aquarray platform. (
Once it was ascertained that cells could be loaded onto the array successfully, fixing and staining of cells on the array were done to develop a high-content imaging readout. Shown in Figure 5 are images taken at low magnification ( Fig. 5A ) and then at a higher magnification with a 40× objective ( Fig. 5B ) after cells had been fixed and stained with Hoechst and phalloidin to stain nuclear DNA and actin, respectively. The results indicated that cell growth and morphology on the device were similar to the results observed in 384-well microtiter plates (not shown). Initial cell fixing, and staining results also showed that protocols could be successfully performed on the array in 1 mm, 500 µm, and 350 µm sided squares, using methods similar to the standard staining protocols used for 384-well plate imaging.
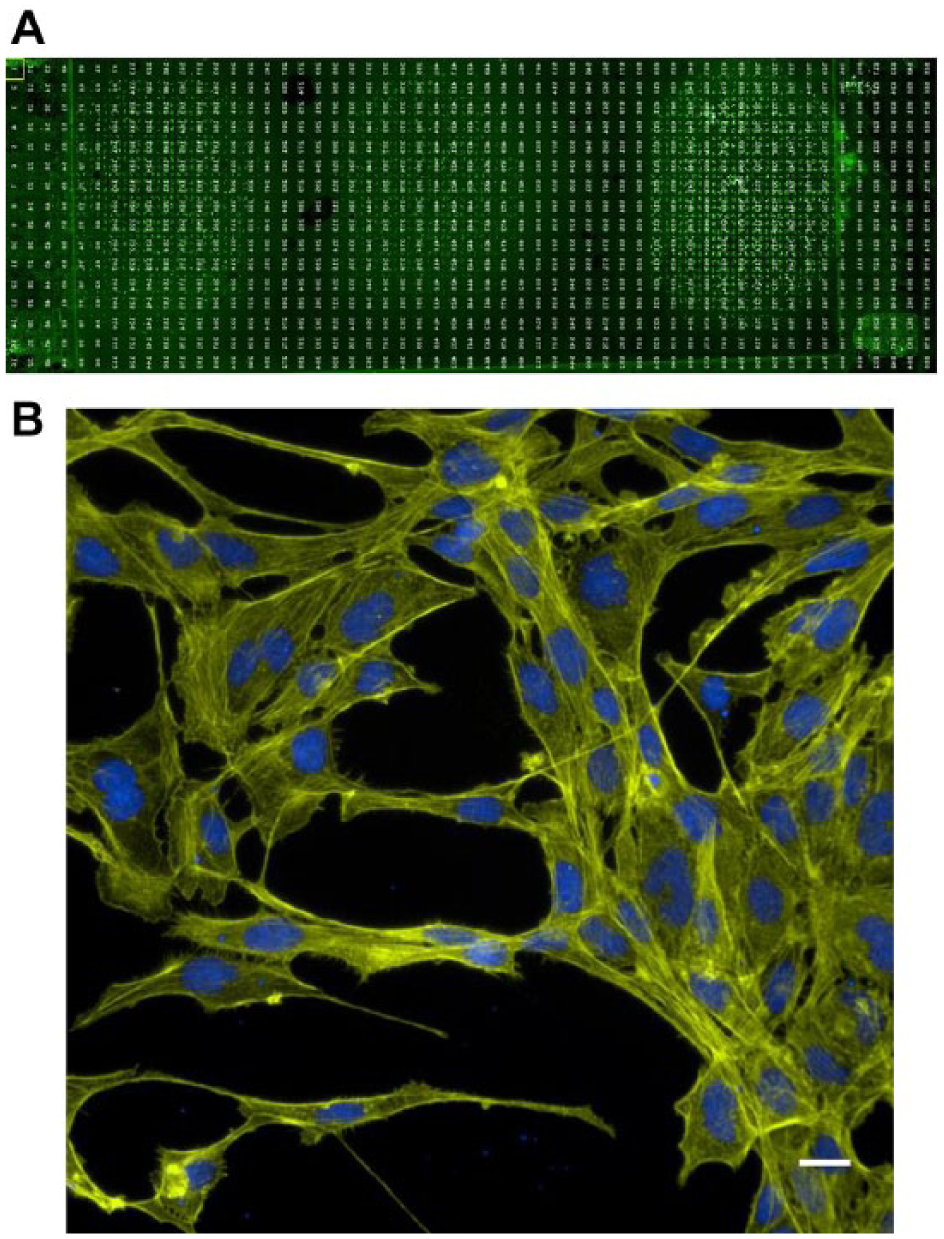
High-content imaging on the Aquarray platform. (
Addition of Compounds to the Array with the Echo 555
To understand the potential for use of this platform in small-molecule drug screening, the Echo 555 instrument was first tested for the addition of compounds to cells. Using the method described above, we allowed the cells to attach onto the surface of the Aquarray slide. Since the lowest-volume dispensing capability of the Echo 555 instrument is 2.5 nL, cell experiments were planned with an intermediate dilution scheme. Compound stock solutions (1 mM) in 100% DMSO were first diluted down to 32% DMSO with cell media to create a 384-well source plate, such that the addition of 2.5 nL of this source to cell array spots of approximately 80 nL would result in a final DMSO concentration of 1% and a final compound concentration of 10 µM. This protocol mimics the current 384-well and 1536-well cellular assay screening protocols, where compounds are screened at 10 µM concentrations in 1% DMSO. The 10 µM concentration represents the highest concentration in the dose–response curve.
Figure 6 shows the results from a dose–response experiment that was performed on an Aquarray slide device. Images in Figure 6A show the expected cell morphology resulting from the treatment of cells with cytochalasin D, compared with the DMSO-only control. The graphs in Figure 6B show the results from a cytochalasin D threefold serial dilution dose–response curve experiment, with an image analysis output based on the actin staining intensity.
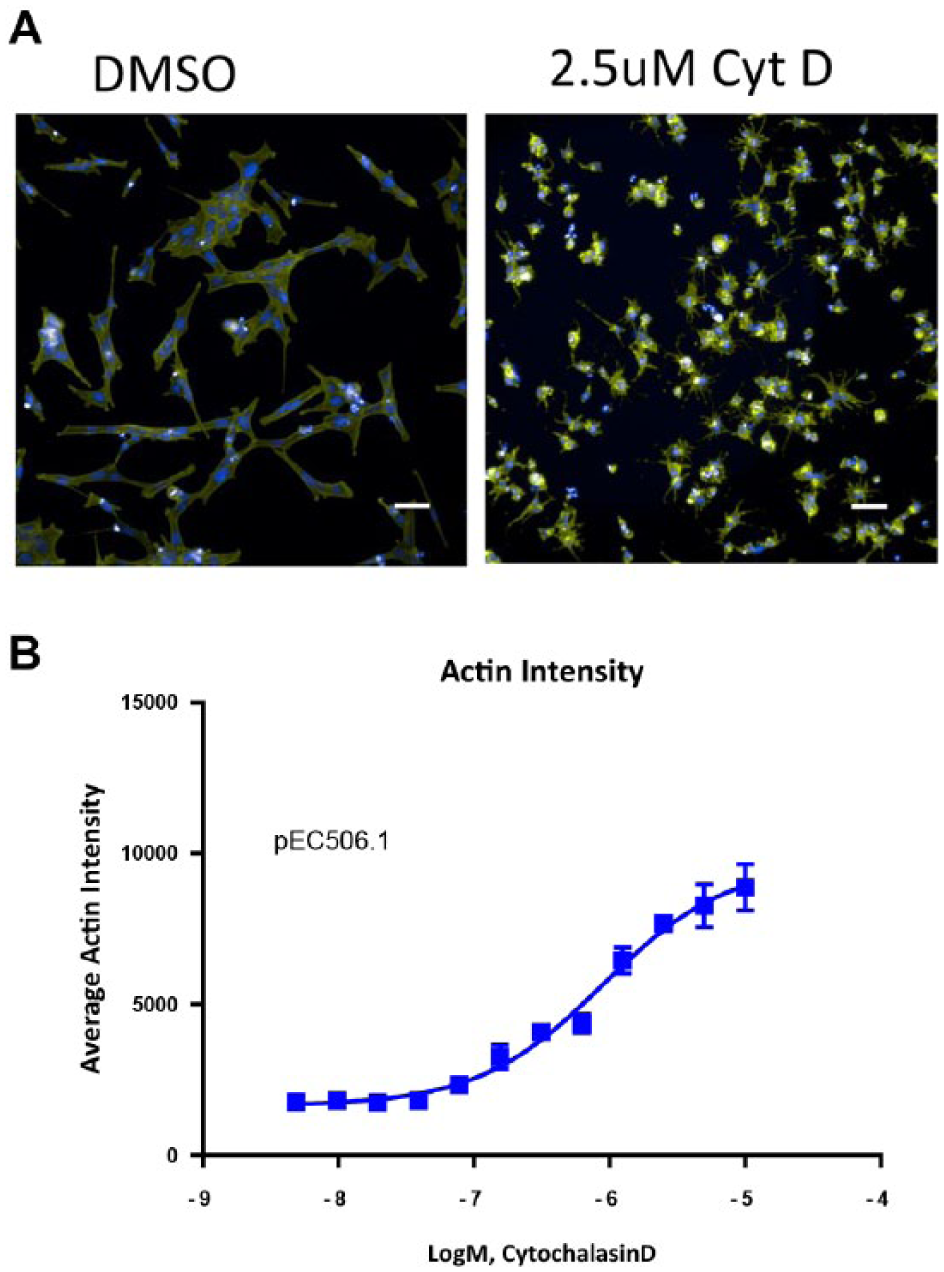
Compound treatment on the Aquarray platform. (
The next experiments were focused on running more compounds in a single-concentration, high-throughput screening mode. Approximately 700 random compounds were added to the array, at 1 compound per array spot using the Echo 555, to give a 10 µM final screening concentration. In addition to the random compounds, cytochalasin D was added in multiple locations as a control. After compounds were added to the cells on the array, the slide was incubated in a tissue culture incubator overnight at 37 °C. The arrays were then processed as described to stain and image the slide. As shown in Figure 7 , results indicated that compounds were successfully added, based on the comparison of the DMSO-only wells to the wells where the actin-disrupting cytochalasin D was added. The scatterplot in Figure 7A shows a clear separation of the control wells from the sample and DMSO wells. The data calculated in Figure 7B show that good statistics were observed, with a signal-to-background average of 2.2 and a Z′ of 0.4. These data represent excellent cellular assay statistics, in an assay volume of approximately 80 nL, where minimal assay optimization was performed.
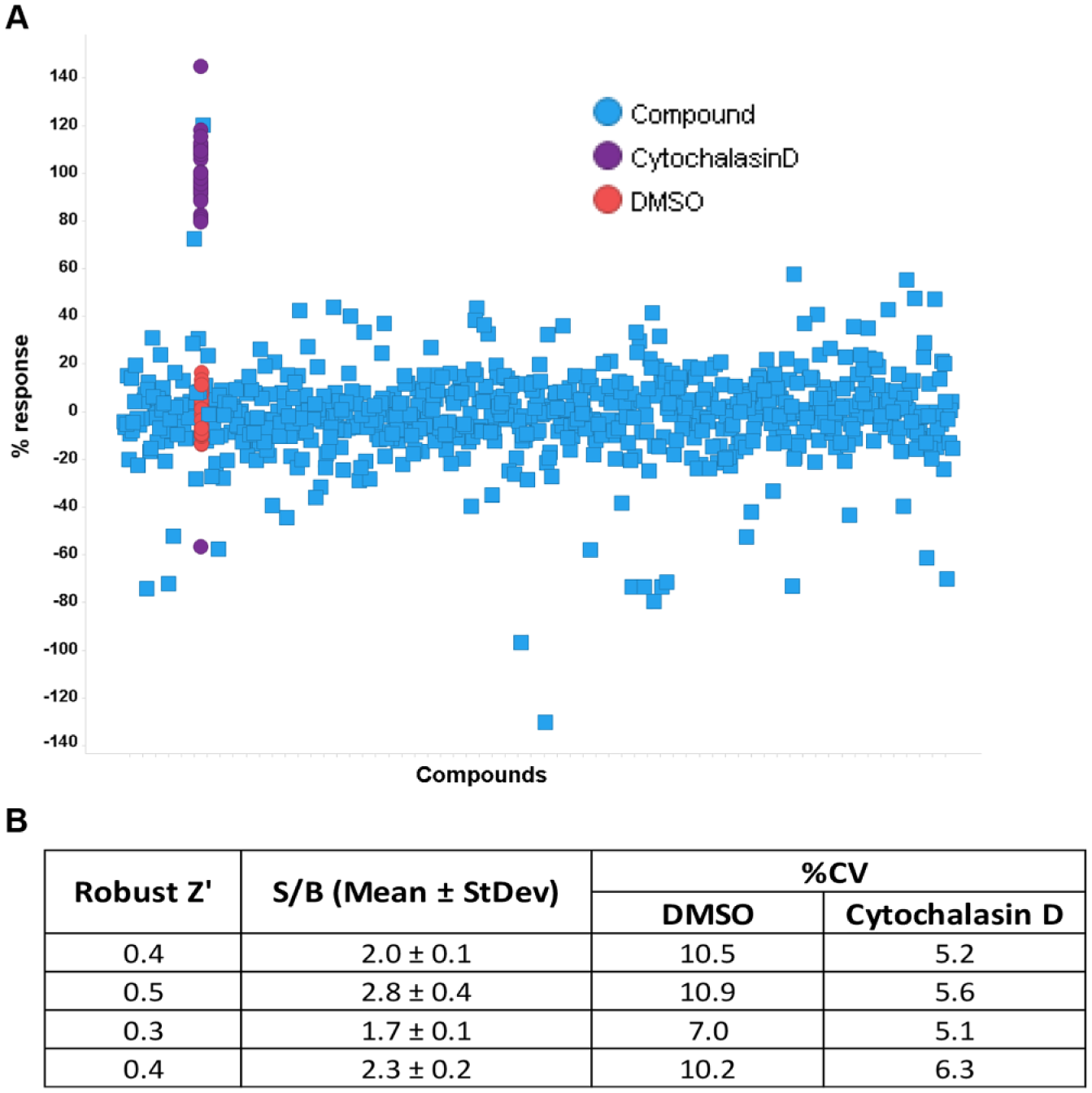
Seven hundred-compound screen on the array. Seven hundred random compounds from the GSK compound collection, plus cytochalasin D as a control compound, were added to cells on the array. Statistics were generated based on the comparison of cytochalasin D to DMSO only. (
With these preliminary results, our observations indicated that in terms of both cell morphology and the cell responses observed, biology miniaturization down to below 100 nL was feasible. Since our goal will ultimately be to enable the use of cell samples that are either limited in supply or expensive, a more complex cell type was grown and tested on the system. Shown in Figure 8 is an image of stem cell-derived cardiomyocytes that were grown on the slide, stained, and imaged. The cell viability and cell morphology were as expected; cells showed signs of beating after several days of incubation on the array (data not shown). Future experiments will be aimed at developing disease models for small-molecule screening on this array system, and continuing the proof-of-concept experiments to develop and test methods for scaling this process so that it can be used in an industrial setting.
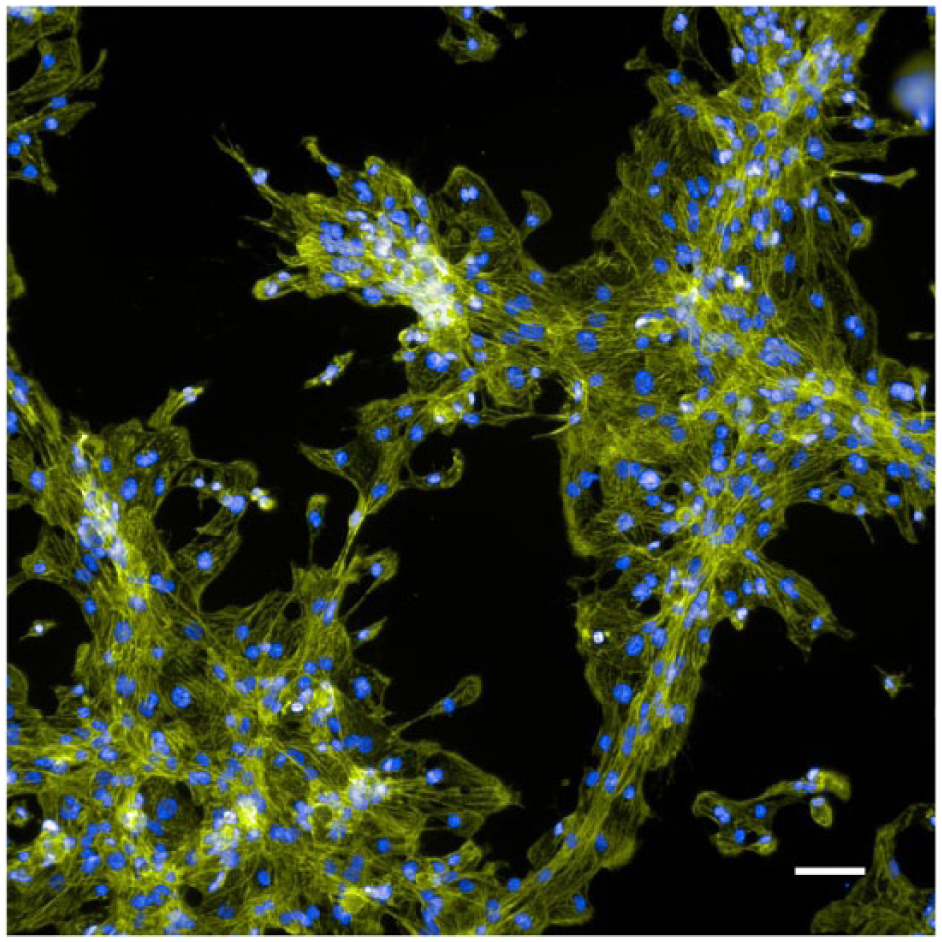
Stem cell-derived cardiomyocytes showed viability and expected cell morphology on the array platform. Cells were grown for 3 days in a 100 mm dish before being transferred to the array device, where they were incubated for an additional 4 days. Cells were fixed and stained with Hoechst (blue) and phalloidin (green) to stain nuclear DNA and actin, respectively. Image was taken with a 20× objective on the InCell 2200 instrument. Scale bar is 50 µm.
Compound Reconstitution Experiments
The miniaturized biology results showed that sample evaporation is a major issue that needs to be solved to move forward with ultra-miniaturization of cellular assays. GSK has undertaken various experiments to investigate the reconstitution of dried compounds of diverse chemical properties. Initial testing with 2-hydroxypropyl-β-cyclodextrin excipient18,19 with a small number of compounds did not show improved compound resolubilization of dried-down compounds (data not shown). Further experiments will be needed for more conclusive results with various other excipients. DMSO–glycerol solvent mixtures are also being explored to slow sample evaporation. 20 Finally, GSK conducted a pilot experiment purposefully drying down compounds and reconstituting them with cell media.
To start initial experiments, ~1500 compounds were selected from the GSK small-molecule library to be analyzed using LC/MS. The selection was done so that half of the set was designed to cover diverse molecular properties that could affect drying down and half of the set was selected to be representative of the GSK small-molecule chemical space. From that list, a total of 352 compounds were selected for this pilot experiment.
Overall, the majority of compounds did not come back into solution. Figure 9A shows a comparison of quantification of samples before and after dry-down (blue and orange, respectively). Roughly 14% of the total samples were recovered at greater than 80%, shown in the bar graph in Figure 9B . For 48% of the samples, less than 1% of the original material was recovered.
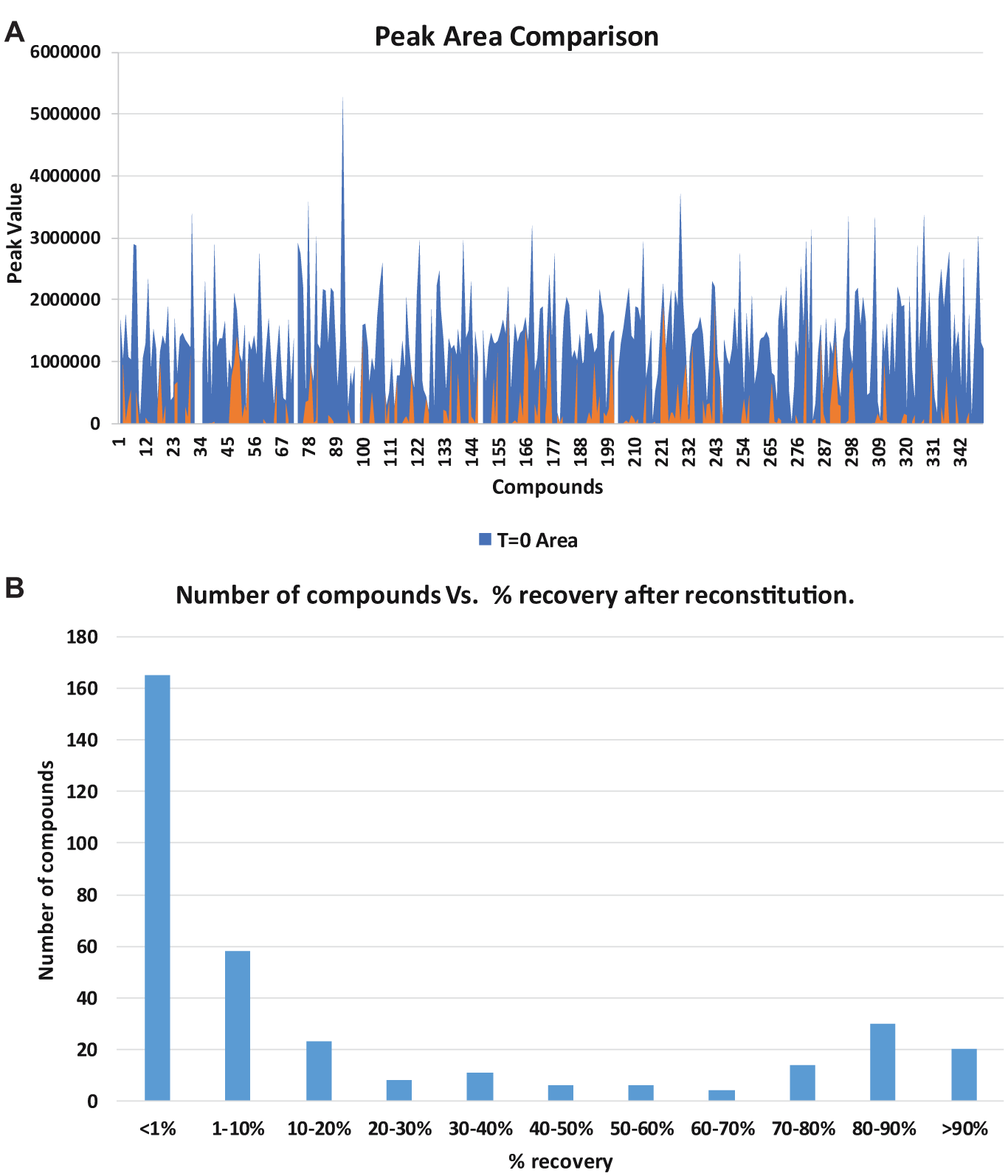
Compound reconstitution experiment. (
Some of the main challenges we faced included sample evaporation and the lack of liquid dispensing equipment that could flexibly dispense desired small volumes into non-SBS plate formats. Since our current drug screening infrastructure has been built around microtiter plates and larger-volume assays, new equipment will be needed to scale these assays into a drug screening process for routine assays. For example, we did not have liquid handlers on site that we could use to dispense cells onto the Aquarray device; the volumes needed were smaller than our current cell dispensers could achieve. Because of this, we were not able to test the Aquarray assay in a compound-first mode, with cells added on top of preprinted compounds. Attempts at dispensing cells with the Echo 555 and the 2470 Arrayer were not completely satisfactory, so we are currently pursuing evaluations of other instrumentation and methods. In addition, much more experimentation will be required to ascertain the robustness and reproducibly of a wide range of cell types and disease biology to understand how broadly this miniaturized platform can be applied.
So, while evaporation is a big issue during dispensing compounds and cells, one advantage of testing miniaturized biology with an open array system was that we could access the surface with the compound dispensers and successfully dispense compounds onto the array. With this system, we were able to show that compounds could elicit cellular responses in the 80 nL volumes tested, and that robust Z′ assay statistics could be obtained. We intend to continue to explore this platform for more complex cellular systems, and to develop higher-throughput processes for the introduction of higher numbers of compounds into the assays.
For compounds, one of the biggest challenges is that very small volumes of compounds evaporate quickly. There are at least two ways one could solve this issue; one would be to dispense compounds in high-humidity environments to prevent evaporation, and the other would be to develop methods to consistently redissolve dried compounds once they have been completely dried. Methods for the latter could include the addition of excipients to help prevent complete drying or changing the properties of the dried compound such that it reenters the aqueous phase more readily once buffers or media have been added.
If high-throughput screening capability materializes for these ultra-miniaturized platforms, then screening small molecules in high throughput with primary human samples and stem cell-derived systems could be enabled. With a 10- to 100-fold reduction in the cell numbers required for assays, fewer cells would be required for screening, compared with our current screening paradigms. The implementation of an array-based miniaturized screening platform will require a change in infrastructure in order to move away from SBS standardized microtiter plate formats. Other peripheral devices designed to hold, incubate, or transfer slides or liquids will be needed to build a process that will be amenable for routine compound screening.
Another main area of future work will be focused on assessing the behavior of small-molecule compounds in these small volumes. As mentioned, compound drying and reconstitution needs to be investigated. In addition, more compounds will need to be tested, since it is not yet clear what issues will be encountered with a wide range of different compound properties.
Finally, the high-content imaging readout captures only some cellular responses, and there is a desire to develop methods for multiplexing the high-content imaging readouts with secreted proteome and RNA transcript readouts. These areas represent aspirational space and have not yet been investigated.
Footnotes
Acknowledgements
The following GSK colleagues are gratefully acknowledged for their contributions:
Stan Martens for valuable discussions and input regarding this perspective
Pat Brady for providing computational expertise in the selection of small-molecule compounds Stephen Vitale, Stephen Besley, and Jamal Johnson for running the compound reconstitution experiment, and Chris Watson for early work on the Aquarray platform
The following technology partners are gratefully acknowledged for their support and assistance:
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
