Abstract
A persistent challenge in developing personalized treatments for hematologic cancers is the lack of patient specific, physiologically relevant disease models to test investigational drugs in clinical trials and to select therapies in a clinical setting. Biomicrofluidic systems and organ-on-a-chip technologies have the potential to change how researchers approach the fundamental study of hematologic cancers and select clinical treatment for individual patient. Here, we review microfluidics cell-based technology with application toward studying hematologic tumor microenvironments (TMEs) for the purpose of drug discovery and clinical treatment selection. We provide an overview of state-of-the-art microfluidic systems designed to address questions related to hematologic TMEs and drug development. Given the need to develop personalized treatment platforms involving this technology, we review pharmaceutical drugs and different modes of immunotherapy for hematologic cancers, followed by key considerations for developing a physiologically relevant microfluidic companion diagnostic tool for mimicking different hematologic TMEs for testing with different drugs in clinical trials. Opportunities lie ahead for engineers to revolutionize conventional drug discovery strategies of hematologic cancers, including integrating cell-based microfluidics technology with machine learning and automation techniques, which may stimulate pharma and regulatory bodies to promote research and applications of microfluidics technology for drug development.
Keywords
Introduction
Developing novel drugs and selecting effective therapies for the treatment of hematologic cancers in clinical settings both remain challenging endeavors in biomedical research. Recently, the Tufts Center for the Study of Drug Development (Tufts CSDD) declared the average estimated cost of taking a new compound from bench to bedside to be approximately US $2.6 billion. 1 Besides investigational new drugs, pharmaceutical companies are also including combination therapies and repurposed drugs in clinical trials. Immense efforts are being made to determine effective treatments for patients specifically diagnosed with hematologic cancers. However, over the last decade, the number of terminated clinical studies relative to the number of drugs that have received FDA approval have increased significantly ( Fig. 1 ). Clinically, while newly diagnosed patients respond to different drugs and enter into an initial period of remission, subsequent relapses and numerous adverse side effects often occur with increasing frequency.1,2 Determining an effective treatment at this stage is more challenging as it is a trial-and-error process that exposes patients to increasing toxicity of a certain regimen of drugs.2,3 Furthermore, the unresolved issue of drug resistance and the lack of sufficient biological information from patient-derived heterogeneous tumor samples upon which appropriate therapies can be selected for each individual are two major shortcomings of current clinical practice.4,5
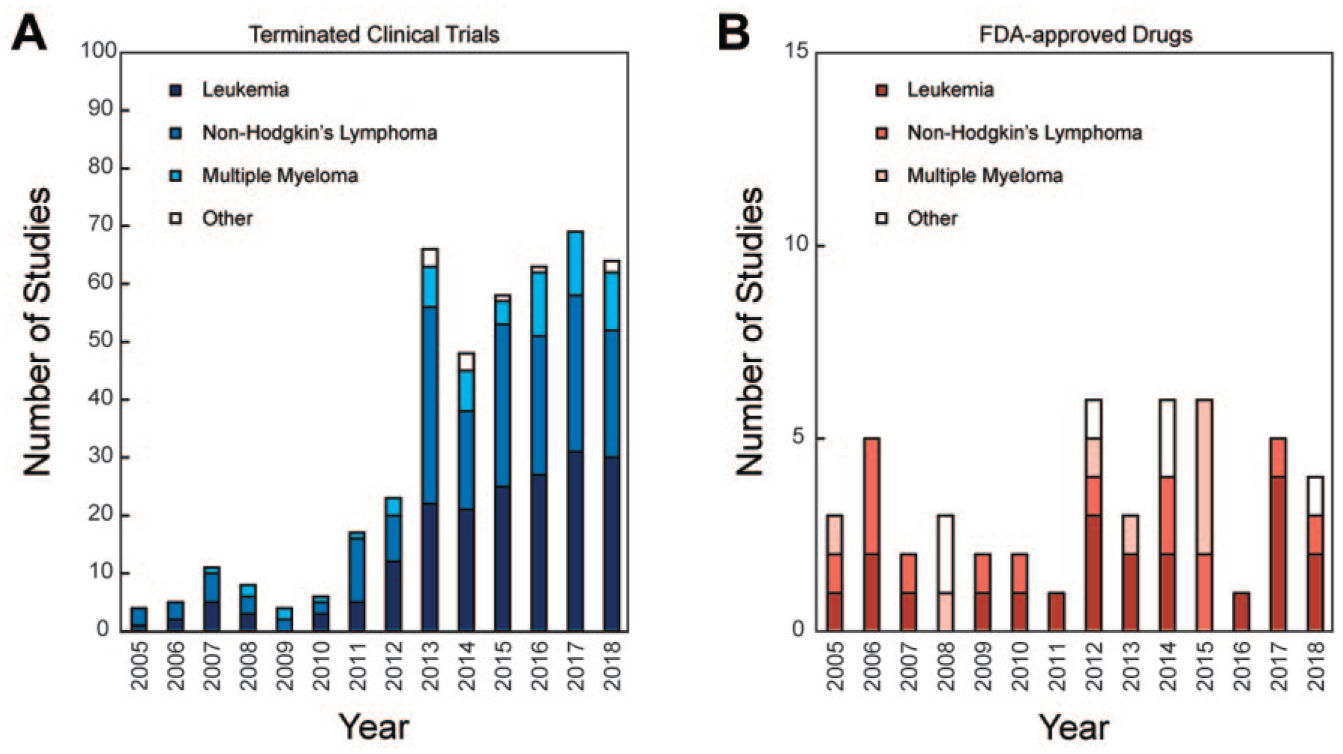
Comparison between (
Such unsatisfactory clinical translation in hematologic cancer drug development can largely be attributed to poor clinical efficacy, adverse side effects and toxicity issues, and a high rate of morbidity at different phases of clinical trials,6,7 which influence the decision-making process in the drug development pipeline. In clinical settings, a variety of techniques are used for hematologic cancer diagnosis, classification, prognostication, and therapeutic decision-making, including those based on immunohistochemistry (IHC), enzyme linked-immunosorbent assay (ELISA), electrophoretic mobility shift assays (EMSAs), fluorescent in situ hybridization (FISH), flow cytometry immunophenotyping, chromosome analysis, molecular testing, and microbiology testing. 8 These laboratory techniques all have various limitations: (1) most fail to recapitulate key pathophysiological features of human disease, including complex tumor heterogeneity, drug diffusion kinetics and dosage, and tumor–stromal interactions, and (2) most require more biological starting material than can be adequately obtained from patients. To meet these challenges, new models, new assay formats, and innovative screening technologies need to be developed, which would then contribute toward more successful clinical translation and improved patient care.
The emerging fields of “organ-on-a-chip” technology and cell-based microfluidic devices—or biomicrofluidic systems—have shown a significantly greater potential than traditional in vitro assays for mimicking the various elements and conditions of the tumor microenvironment (TME).2,3 Because these technologies allow the use of lower volumes of patient sample, the amount of experimental conditions can be increased while single-cell behavior and function can be observed.9–12 Also, biologists can operate passive pumping microfluidic devices easily and use standard tools found in most biology labs. 13 These technologies have the potential to produce improved tumor models, specifically for different hematologic cancers to test different FDA-approved drugs and to facilitate treatment selection for a patient. Furthermore, these models have the potential to be developed in parallel with anticancer drugs and could be used as companion diagnostic assays to guide the selection of patient cohorts for clinical trials. To further advance such biomicrofluidic models and to extend the understanding of the future potential of this technology in the research of hematologic malignancies, it is critical to review the most recent advances in the field, especially specific developments for hematologic cancers.
The purpose of this review is to highlight recent advances in microfluidics technology that specifically pertain to the study of hematologic cancers and their associated TMEs, and the application of these technologies for drug discovery and clinical diagnostics. The review covers state-of-the-art developments in biomicrofluidic platforms currently being developed or used for hematologic cancers, and also outlines chemotherapeutic drugs and different modes of immunotherapies that are currently under clinical investigation or have received FDA approval. This review also outlines key considerations for developing a physiologically relevant microfluidic device for testing different modes of cancer immunotherapy and chemotherapeutic drugs for different hematological cancers in a clinical setting and drug development pipeline. The rationale for bringing these different elements together in one review is to provide an outlook for future opportunities that can leverage advantages of microfluidics while considering the drug development landscape.
Overview of the Tumor Microenvironment in Hematologic Malignancies
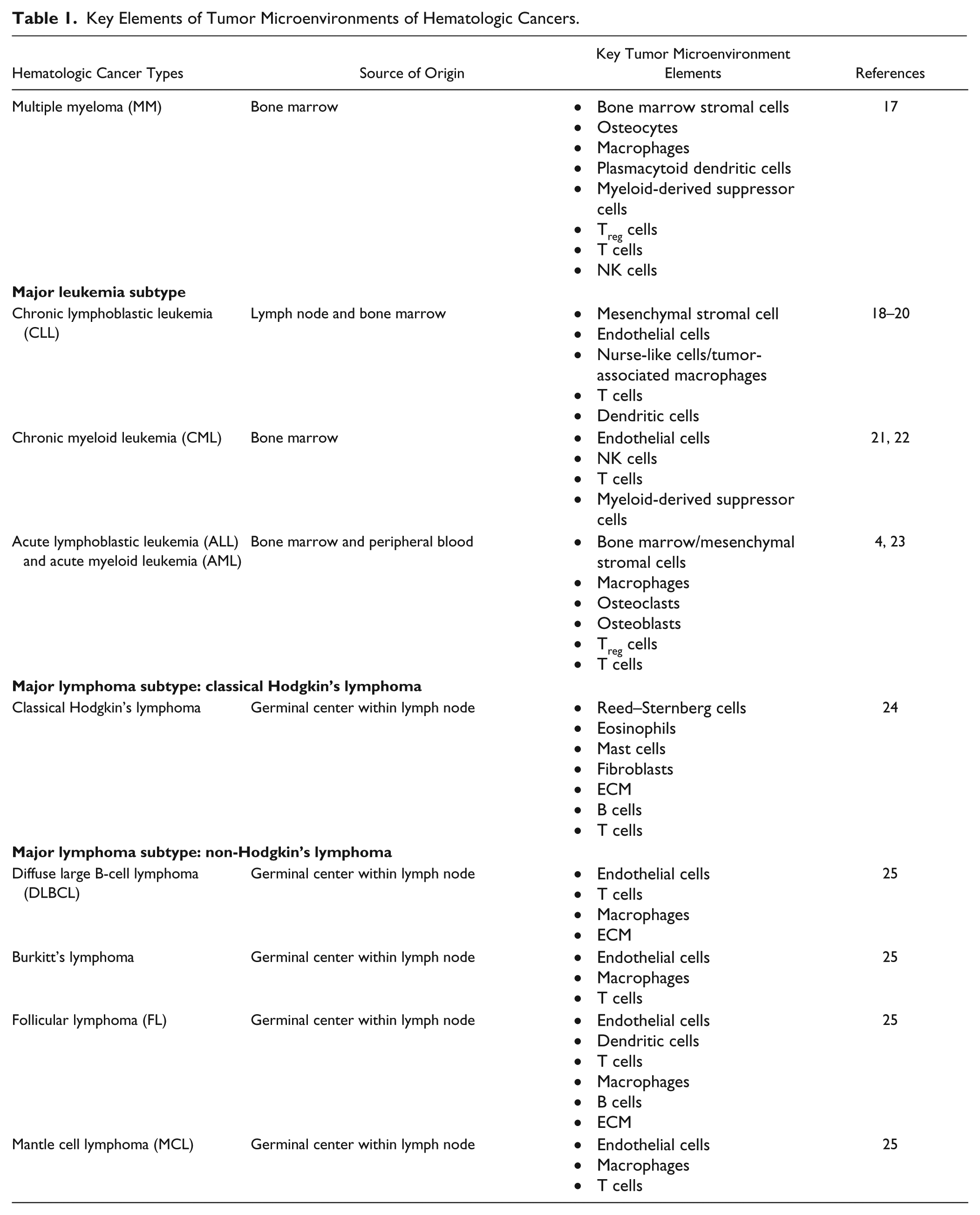
Hematologic cancers, also known as liquid tumors, originate in the blood, bone marrow, and lymph nodes and are characterized by their presence in the lymphatic and circulatory systems ( Fig. 2 ). 14 Three major categories of hematologic tumors are leukemia, lymphoma, and multiple myeloma (MM); leukemia and lymphoma cancers are incredibly diverse by nature and can further be divided into subcategories based on the clinical manifestations and site of origin ( Table 1 ). Based on the percentage of leukemia cells in blood or bone marrow and based on the predominant lineage of malignant cells, leukemias are categorized as acute or chronic and myeloid or lymphoid, respectively. 15 Similarly, based on the presence of Reed–Sternberg cells and other clinical manifestations lymphomas are categorized as classical Hodgkin’s lymphoma or non-Hodgkin’s lymphoma.15,16 Table 1 includes sources of origin and key elements of TMEs of the major subcategories of hematologic cancers. Figure 3 shows a diagram of hematopoietic cell lineages and the origin of different cancers. A detailed description of the biology of these cancers is beyond the scope of the current review. Interested readers are directed to the following reviews for more detailed discussions on these topics.15,16
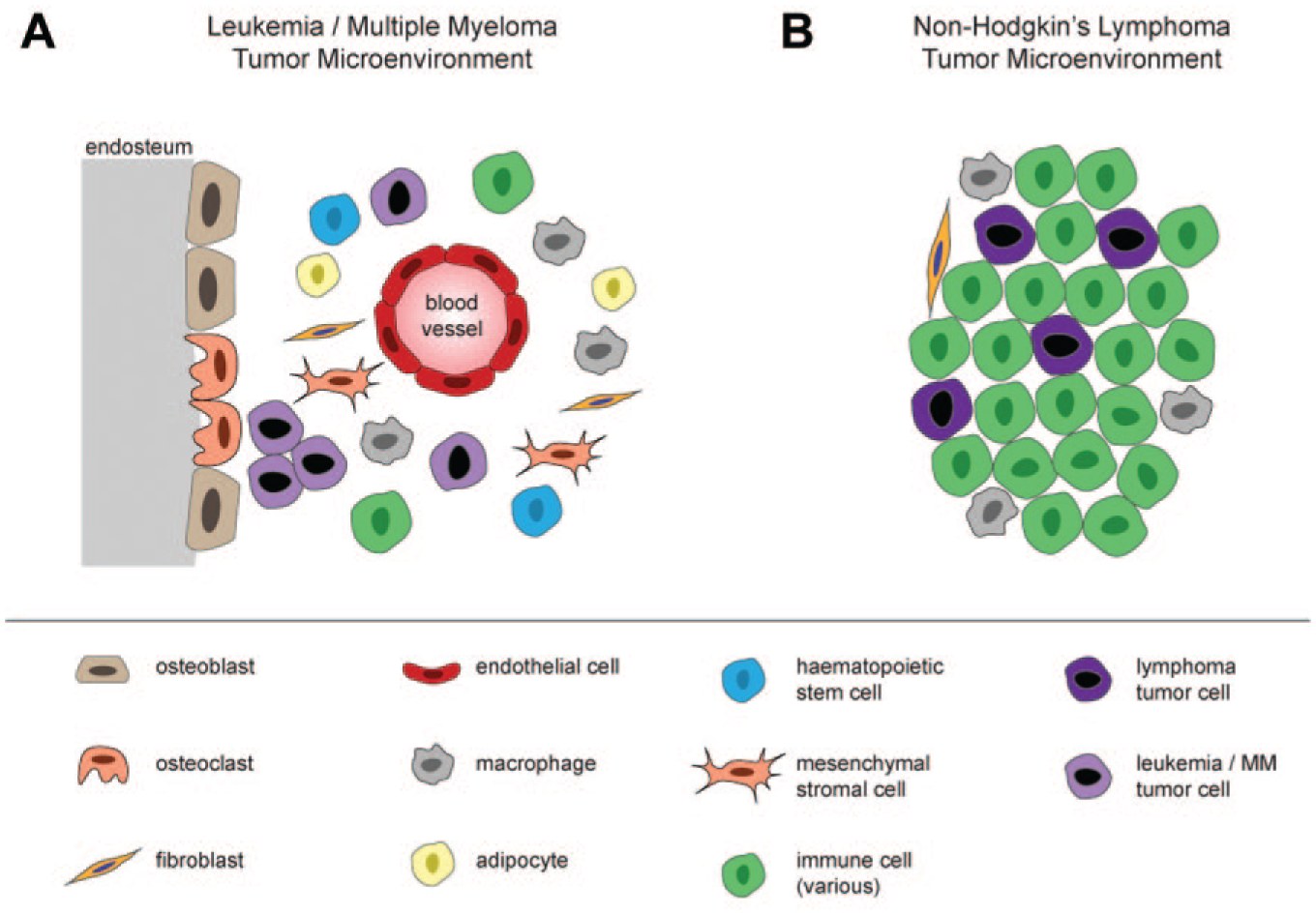
Hematologic TMEs consist of heterogenous mixtures of cells and biological components, with major differences between two major classes: (
Key Elements of Tumor Microenvironments of Hematologic Cancers.

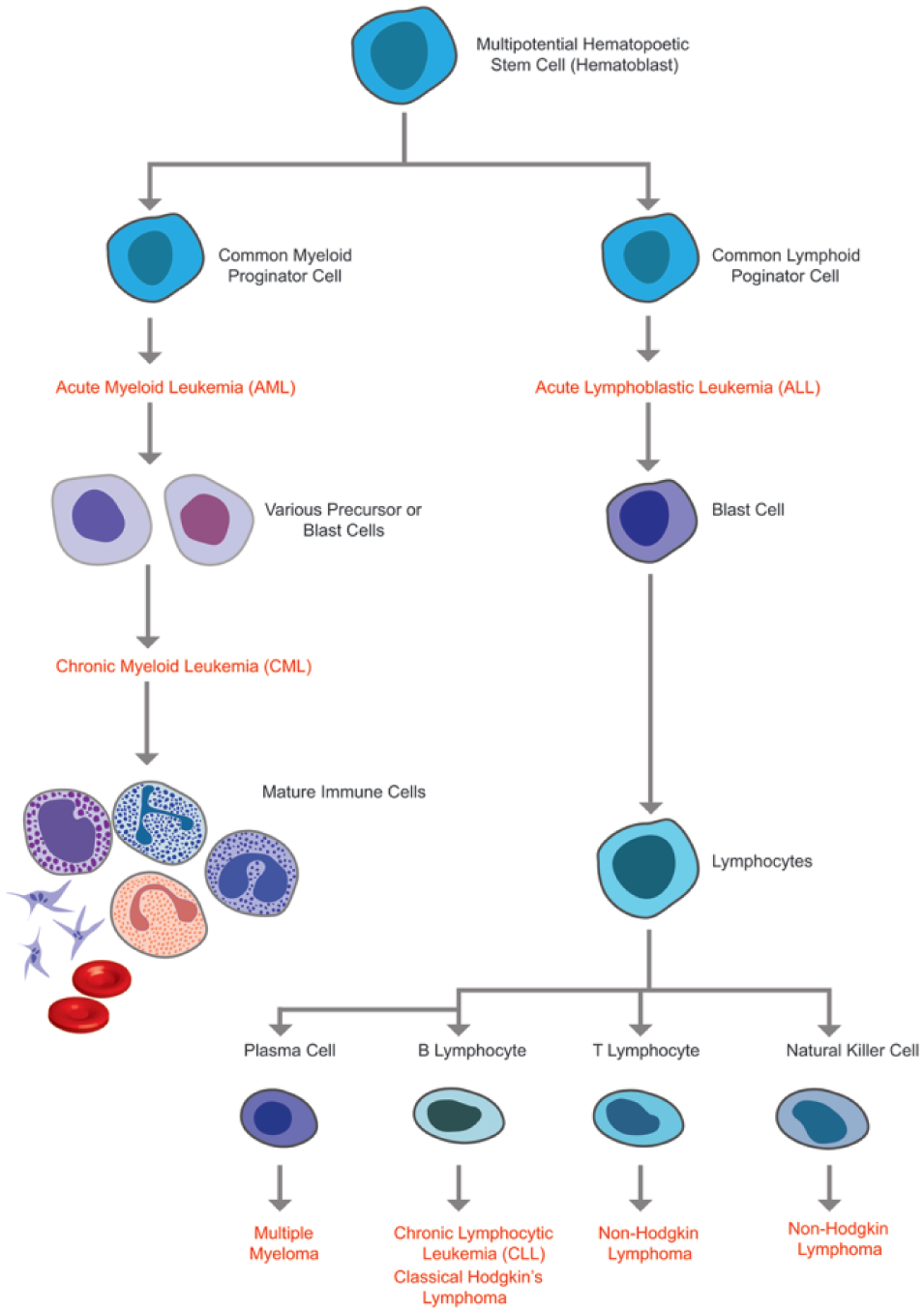
Hematopoiesis and development of different hematologic cancers within normal hematopoietic cell formation.
A large body of evidence supports the view that all hematologic cancers exhibit profound heterogeneity, considerable complexity and dynamics, and the existence of a distinct pool of cancer-initiating cells, which are also referred to as cancer stem cells. 14 Cancer stem cells do not divide rapidly, exhibit resistance to conventional chemotherapies, and possess the ability of self-renewal after therapies. 26 Among different hematologic cancer stem cells, leukemia stem cells have been extensively studied.25,26 A number of therapeutic approaches to target leukemia stem cells are currently under investigation with a few nearing the clinic. 27
More than a century ago, the first microscopic examinations of one subtype of hematologic cancer revealed that the liquid tumor comprised not only the malignant cells but also a variety of nonmalignant cells and extracellular matrix (ECM), that is, the TME. 25 The interaction of liquid tumor cells with the ECM plays a crucial role in tumor cell behavior, such as growth, proliferation, inflammation, immune suppression, angiogenesis, drug resistance, and metastasis. 28 Tumor cells secrete ECM-degrading enzymes resulting in the invasion of malignant cells, and develop adhesion via specific cell surface receptors with ECM components, which causes drug resistance. It has also become increasingly evident that reprogrammed stromal cells of the liquid TME exhibit inflammatory responses, which also enable cancer pathogenesis, progression, and drug resistance. The reprogrammed stromal cells mainly include mesenchymal stem cells, endothelial cells, fibroblasts, monocyte-derived nurse-like cells, and Reed–Sternberg cells (specifically found in Hodgkin’s lymphoma).4,5 Also, bone cells that reside in the bone matrix, such as osteoblasts (OBs), osteoclasts, adipocytes, and osteocytes, not only play a significant role in tumor cell progression and chemotherapy resistance but also cause bone lesions. 29 Notably, the role of the TME significantly varies between hematologic cancer subtypes. Also, the TME exhibits a variety of extrinsic survival signals, which often trigger resistance to targeted and chemotherapeutic drugs.
It has been experimentally demonstrated that both innate and adaptive immune systems play a significant role in eradicating liquid tumor cells through the activation of T-cell and humoral-mediated immunity. However, strong evidence demonstrates that in many cases antitumor T cells and natural killer (NK) cells not only are suppressed by the TME, but also promote tumor progression by immunoselection. 30 Furthermore, plasmacytoid dendritic cells (pDCs) and macrophages induce tumor cell proliferation and survival by direct interactions. 31 Such an immunosuppressive milieu causes failure to chemotherapy, targeted therapy, and radiotherapy. Hence, novel immunotherapies to enhance antitumor immunity are currently under investigation. 31
The presence of angiogenesis in liquid tumors has been supported by many studies that include the finding of increased bone marrow microvessel density and increased levels of plasma pro-angiogenic cytokines and growth factors (e.g., vascular endothelial growth factor [VEGF]) that promote neovascularization.32,33 Few proteasome inhibitors (PIs) and immunomodulatory drugs (IMiDs) have shown successful inhibition of angiogenic activity in MM to date, and other drugs are currently under clinical trials. 34 Formation of neoangiogenic networks is associated with the creation of hypoxic regions in the bone marrow microenvironment, which induce metabolic changes, survival, migration, metastases, and chemoresistance.35,36 Although migratory behavior is consequential in leukemia and lymphoma cells, 37 recent studies reported the presence of metastatic myeloma cells in hypoxic bone marrow regions. 38 Myeloma metastasis is mediated by a hypoxia-induced epithelial mesenchymal transition (EMT)-like phenotype, 38 which has implications on the prospective development of hypoxia- and metastasis-targeted drug and diagnosis strategies to obtain better clinical outcomes for patients.
Herein, we have presented key features and a series of major events (e.g., liquid tumor initiation, angiogenesis, hypoxia) of a complex hematologic TME that typically influence tumor progression and drug resistance. Although many reviews of TMEs are available,34,39 the intent of this summary is to provide readers a succinct review on the TME as a preliminary step in developing biomicrofluidic systems.
Challenges in Drug Development and Treatment Selection
Current Therapeutic Agents in Hematologic Cancer Treatment
Recent advances in molecular biology and genetic engineering have demonstrated outstanding success in the treatment of hematologic cancers with the advent of numerous chemotherapies, targeted therapies, and immunotherapies. Nevertheless, each treatment category has its strengths and limitations.
Standard chemotherapeutic agents could be divided into two broad categories: cell cycle-specific drugs and cell cycle-nonspecific drugs, 40 which influence tumor cells and TMEs following both intrinsic and extrinsic mechanisms of action. This therapy has been proven a curative mode of treatment for certain hematological malignancies, such as childhood acute lymphoblastic leukemia (ALL) and subgroups of Hodgkin’s and non-Hodgkin’s lymphoma. 41 Nonetheless, in other hematologic malignancies, responses are generally partial, brief, and unpredictable. Furthermore, different research groups proved the nonspecific targets of chemotherapeutic cytotoxins for malignant cells, thus explaining their narrow therapeutic index and significant toxicities. 42
Similar to chemotherapies, targeted therapies influence tumor cells and TMEs following both intrinsic and extrinsic mechanisms of action that enhance antitumor immunity, inhibit biological activities of tumor-causing elements, disrupt cell adhesion and migration, and induce apoptosis.40,43–48 However, unlike the mechanism of action of standard chemotherapies, targeted therapies follow a mechanism that specifically acts on a well-defined target or biological pathway to destroy malignant cells. As a result, these therapies are less toxic, more effective, and show a higher therapeutic index.41,49 Major subcategories of targeted therapies include PIs, kinase inhibitors, IMiDs, histone deacetylase inhibitors (HDACis), and monoclonal antibodies.27,41,50
Another emerging therapy in hematologic cancer treatment is immunotherapies that include chimeric antigen receptor T (CAR-T) cells, bispecific antibodies, tumor vaccines, and immune checkpoint inhibitors.41,51 Unlike the other two major categories of treatments, immune-based therapies specifically target immune cells, boost up immunity to kill malignant cells in advanced-stage cancer patients, eliminate large tumor masses, and elicit immunological memory that can lead to prolonged protection from cancer relapse.52–54 Interested readers can refer to an in-depth review published by Vasekar and colleagues 55 for a detailed discussion on the mechanisms of action of immune checkpoint inhibitors, CAR-T cells, and other modes of immunotherapy.
Table 2 outlines the mechanisms of action of each subcategory of the three major therapies and lists the corresponding FDA-approved drugs. For more in-depth explanation of these therapies, interested readers can refer to reviews published by Hamilton and colleagues 41 and Im and Pavletic. 68
List of Major Therapies and Corresponding FDA-Approved Drugs (as of October 18, 2018).
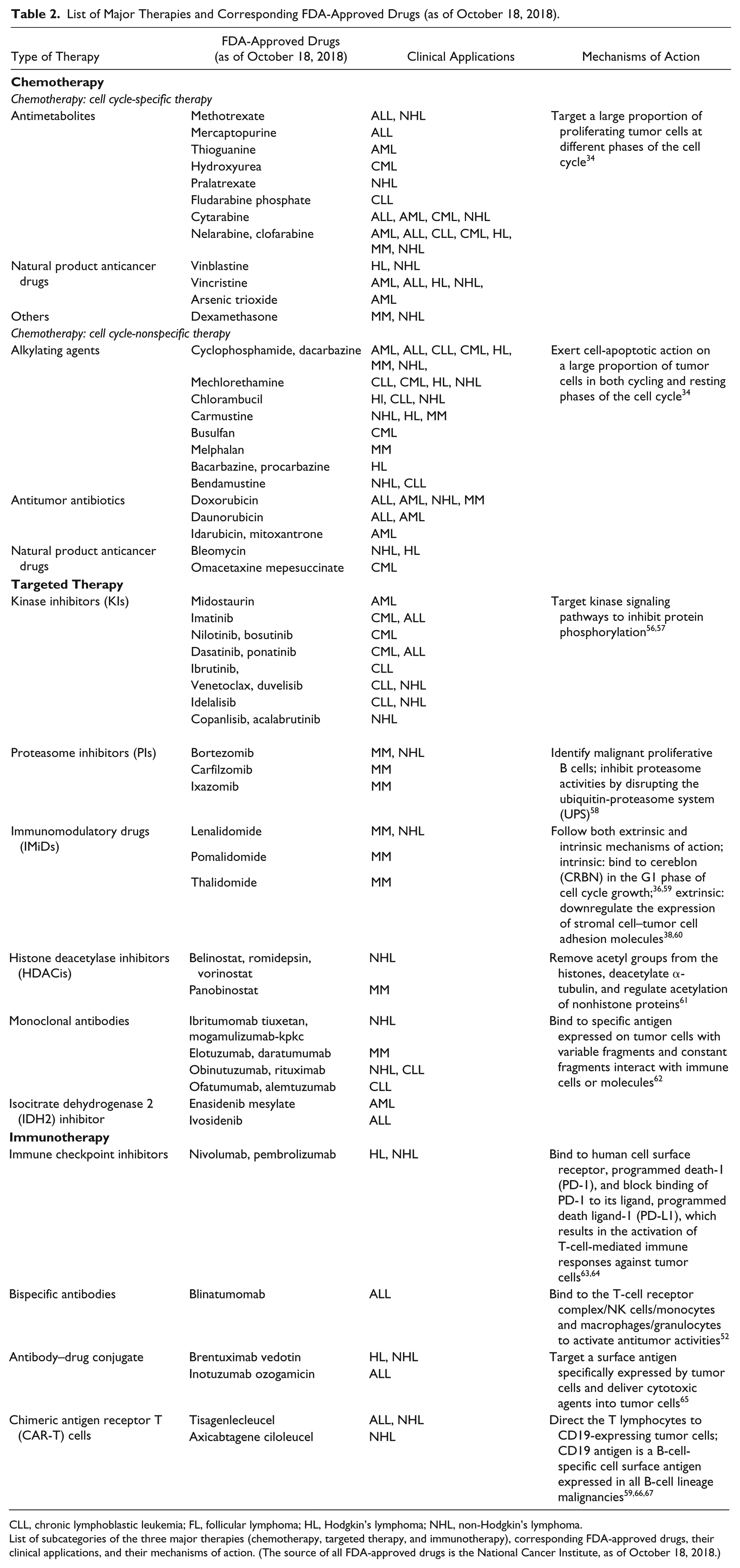
CLL, chronic lymphoblastic leukemia; FL, follicular lymphoma; HL, Hodgkin’s lymphoma; NHL, non-Hodgkin’s lymphoma.
List of subcategories of the three major therapies (chemotherapy, targeted therapy, and immunotherapy), corresponding FDA-approved drugs, their clinical applications, and their mechanisms of action. (The source of all FDA-approved drugs is the National Cancer Institute, as of October 18, 2018.)
Need for Personalized Medicine
Although a number of therapy options have been discovered for different hematologic cancers to date, effective therapy selection remains challenging in clinical settings. Optimal therapy selection for a patient includes an assessment of the patient’s age, health, and treatment preference; clinical manifestations of the disease; and the treatment situation (e.g., first- vs second-line treatment, response vs nonresponse to last treatment).69–71 Assessment of these parameters is combined with genetic analyses including karyotype tests, FISH, PCR, and next-generation sequencing, 72 as well as cytology and histology analyses 73 for diagnosis, classification, and risk stratification. 72 Risk stratification is further combined with empirical methods to select therapies for a patient, a trial-and-error process that often exposes patients to adverse side effects. 74 Accumulating evidence suggests most hematologic cancer patients respond to initial treatments, but relapse and fail to respond to further therapies, resulting in drug resistance.25,57,75,76 Although there are several stand-alone and combination therapeutic options available to use in the latter case,77,78 rapidly identifying an effective therapy is critical for a patient with a short median survival rate.79–81 Hence, to select therapies for individual hematologic cancer patients in a clinical setting, it is instructive to address the following issues: (1) reducing the time to identify effective therapies for patients, (2) identifying the sensitivity and resistivity of patients to specific therapeutic drugs, (3) identifying patients for whom a drug could induce adverse side effects, and (4) personalizing treatment selection to improve treatment decision-making and clinical outcomes.
To improve clinical outcomes and to reduce the morbidity rate of hematologic cancer patients in the clinic, biopharmaceutical companies are increasingly focusing on the development of new anticancer drugs and repurposed drugs. However, a vast majority of them cannot meet the expected clinical efficacy and exhibit adverse side effects and toxicity in patients, and as a result fail to reach the bedside.82,83–85 One of the major underlying reasons of this is the lack of a patient-specific diagnostic tool to predict the sensitivity or resistivity of a new therapeutic agent. 7 In clinical trials, a patient cohort is exposed to a new therapeutic agent, which may or may not be safe and effective, leading to severe side effects or morbidity.7,86 Figure 1A shows that the number of terminated clinical studies for hematologic cancers has increased from 2005 to 2018, and drugs are more likely to fail in clinical trials today than in 2005. The major reasons for the failure of these studies include drug-induced adverse side effects sometimes resulting in mortality. Figure 1B also shows that between 2005 and 2018, only a few drugs (relative to the large number of terminated clinical studies) received FDA approval to be used clinically. Hence, more reliable, accurate, and personalized diagnostic models are needed for obtaining information that is essential for the safe and effective use of a therapeutic agent on a specific patient in the early phase of a clinical trial.6,86,87
Companion Diagnostics
Such a patient-specific diagnostic model is often referred to as a CDx and has the potential to predict therapy for a specific patient in the clinic and select the patient cohort for a particular therapeutic agent in clinical trials. 88 According to the USFDA (2016), a CDx is defined as an in vitro device that provides information that is essential for the safe and effective use of a corresponding drug or biological product. There are four areas where this assay could be used: (1) to identify patients who are most likely to benefit from a drug, (2) to identify patients likely to be at increased risk of serious adverse reactions as a result of treatment with a drug, (3) to monitor the drug response for the purpose of adjusting treatment (e.g., schedule, dose, and discontinuation) to achieve improved safety or effectiveness in clinical settings, and (4) to identify patient populations for which a new drug will be the most effective in the drug development pipeline.87,88 While a CDx can improve patient outcomes in the clinic and reduce the failure of a new drug compound in the development pipeline, manipulating limited patient samples and choosing corresponding readouts remain challenging, 6 and it is clear that we need additional approaches to develop a robust CDx platform for hematologic cancer research.
Recent advances in biomicrofluidic and microfabrication technologies have demonstrated the capability of manipulating minute patient-derived samples in microfluidic cell-based platforms to test different anticancer drugs with quantitative readouts in hematologic cancer research, which shows the promising application of this platform to developing a personalized diagnostic model to select therapy in the clinic. In addition, this platform has the ability to mimic the pathophysiology of a hematological cancer TME, giving relevant information about physiological responses and drug responses in vitro. This is achieved due to greater spatial and temporal control over stimuli and microenvironmental factors. Moreover, the in vitro microscale geometries use small reaction volumes, making it easier to fill this space with optimum quantities of cells and to use relevant drug concentrations. This in turn preserves critical cellular functionality in vitro. 11 By leveraging these aspects of biomicrofluidics and microfabrication techniques, optimal geometries can be generated to perform multiple drug assays, reach higher throughput, and achieve desired sensitivities. 89 Furthermore, this platform could be used to identify the efficacy and compound-associated toxicity for a specific patient to guide patient cohort selection in clinical trials. Hence, such a biomicrofluidic system has the potential to be used as a biomicrofluidic companion diagnostic tool or “biomicrofluidic CDx” assay. Currently, no biomicrofluidic system has yet received regulatory approval to be used as a personalized diagnostic tool in the clinic, nor in the drug development pipeline for hematologic cancer treatment, but efforts are underway. Therein, it is imperative to review state-of-the-art development of biomicrofluidic systems that specifically pertain to the study of hematologic TMEs, and the application of these technologies for drug discovery and clinical diagnostics. The following section outlines different biomicrofluidic systems developed to study different hematologic cancer microenvironments and to test various anticancer drugs on patient-derived samples.
Biomicrofluidic Systems for Hematologic Cancer Research
To date, a major application of biomicrofluidic systems in hematologic cancer research is testing different kinds of patient-derived liquid tumor cells, either in monoculture or in coculture with different stromal cells, under exposure to different chemotherapeutic agents in two-dimensional (2D) and three-dimensional (3D) platforms. The 2D or 3D geometries were specially designed to capture the complex cellular dynamics and interactions between nonadherent liquid tumor cells and other adherent cells, and to study tumor cell microenvironment-mediated drug resistance. Other applications include augmenting the viability of primary liquid tumor cells in vitro and studying different TME processes (e.g., angiogenesis) by controlling different microenvironmental factors spatially and temporally in tailored geometries. This section covers recent advances in the development of drug testing assays and TME models using different biomicrofluidic techniques for three major types of hematologic cancer: MM, leukemia, and lymphoma.
Multiple Myeloma
Young and colleagues developed a poly(dimethylsiloxane) (PDMS)-based compartmentalized microscale coculture platform to study cell–cell interactions between small populations of nonadherent MM cells and adherent bone marrow stromal cells (BMSCs), and to quantify subcellular transcription factor localizations at a single-cell level. 90 This device was also used to test chemotherapeutic drugs to validate its potential as a drug testing assay. Pak and colleagues further applied this device as an ex vivo microfluidic-cis-coculture (MicroC 3 ) chemotherapeutic drug testing platform by coculturing patient-derived CD138+ MM cells with the same patient’s CD138− tumor-companion mononuclear cells and assessing patient tumor cell response to bortezomib treatment. This pilot study showed excellent correlation between patient clinical responses and MicroC 3 readout ( Fig. 4B ), demonstrating potential as a predictive diagnostic assay for MM. 74
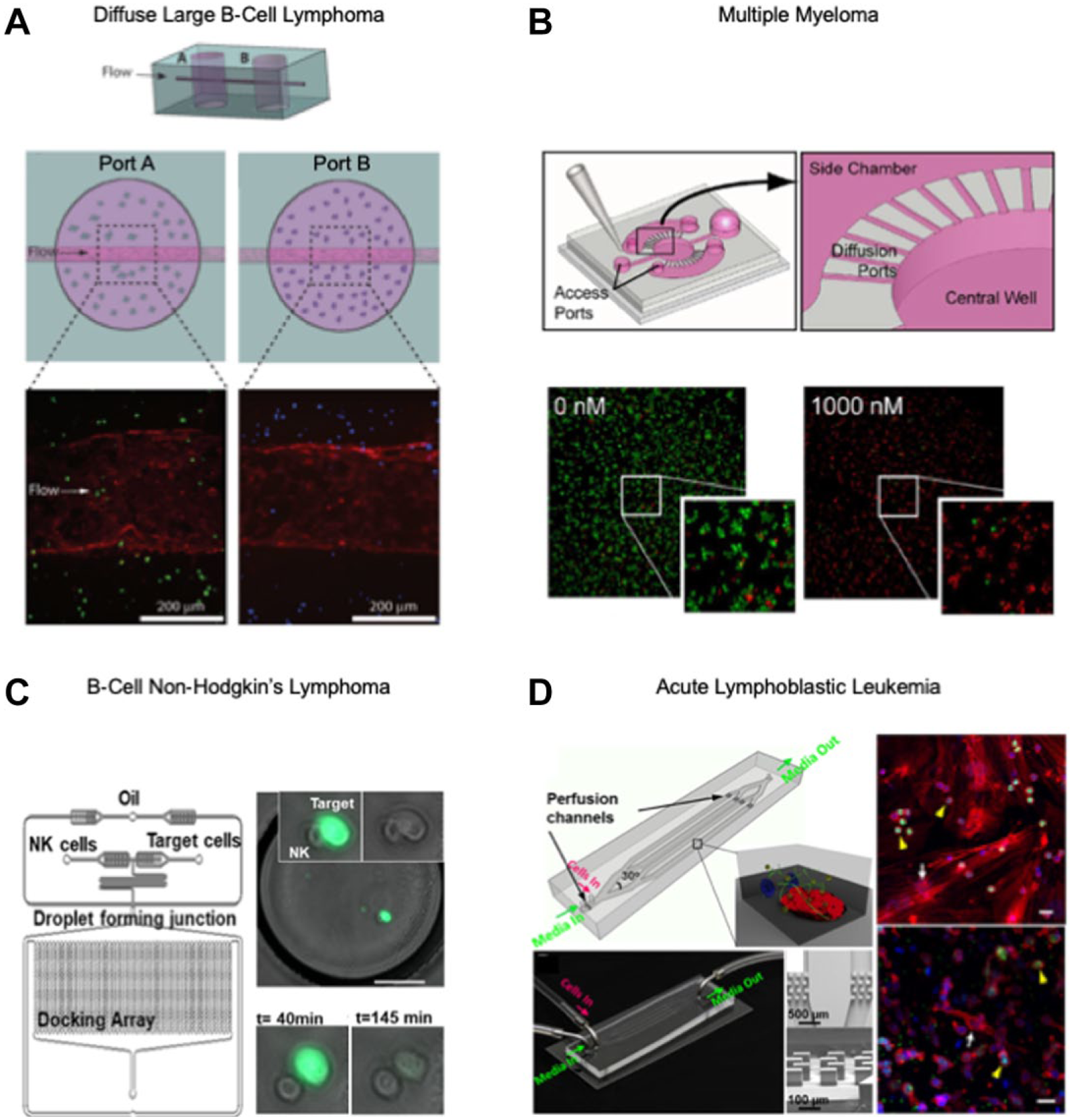
Recent biomicrofluidic systems for studying hematologic cancers. (
Culturing and long-term preservation of patient-derived primary MM cells in vitro have remained challenging in the laboratory and have hindered assay development. Zhang and colleagues developed a 3D ossified tissue culture model in a hexagonal microfluidic device supplemented with patient plasma media to recapitulate interactions between OBs, bone marrow mononuclear cells, and primary MM cells, and demonstrated the viability of primary MM cells after 3 weeks of culture. 94 However, they observed reduced viability and osteogenic activity of OBs in the presence of primary MM cells and patient plasma, which compromised the viability of MM cells over 3 weeks. To address this issue, they used a perfusion-based well plate device, optimized the perfusion flow rate and patient-derived plasma composition in the culture medium, and replenished the OBs during the culture to prolong MM cell viability beyond several weeks. 95 Nevertheless, while solid ossified tissues mimic bone marrow physiological conditions, these are technically challenging to reproduce and adapt. De la Puente and colleagues developed a 3D bone marrow culture model derived from the bone marrow supernatant of MM patients; recapitulated the interaction among MM cells, stromal cells, and endothelial cells; and induced an oxygen gradient, drug gradient, and drug resistance in the model, showing considerable advancement in spatiotemporal complexity and the number of cell types involved. 96
Leukemia
Aljitawi and colleagues reported an in vitro 3D human bone marrow culture model that incorporated the mesenchymal stem cell microenvironment in coculture with leukemia cell lines and was tested with doxorubicin. 97 They observed a continuous increase of drug resistance in the 3D culture model compared with the 2D culture models and significant differences in drug response between different leukemia cell lines. These findings imply that the drug resistance and variable drug response are dependent on the in vitro coculture system used. Furthermore, the probable cause of drug resistance in a leukemia microenvironment was confirmed by Bruce and colleagues. 93 They developed a 3D microfluidic ALL cell culture device encapsulating primary human BMSCs, OBs, and leukemic cells in a collagen gel and exposed them to physiological flow and the chemotherapeutic agent cytarabine. They compared the drug sensitivity in 2D-static, 3D-static, and 3D-flow cultures and reported a gradual decrease in sensitivity in 3D models, which alluded to the protective role of the leukemia TME in tumor cell survival ( Fig. 4D ).
Khamenehfar and colleagues developed a patient-specific dielectrophoretic microfluidic device to study, with single-cell resolution, multidrug resistance (MDR), heterogenous drug responses, and drug failure in relapsed adult acute myeloid leukemia (AML) patients, treating cancer cells with different efflux transporter (ABCB1, ABCC1, and ABCG2) inhibitors. 82 They found the presence of MDR-positive leukemic blast cells in patients who had achieved complete remission, suggesting that these cells were responsible for subsequent relapse.
Houshmand and colleagues developed a 3D microfluidic assay to test the drug treatment of azacitidine and cytarabine on AML cell lines cocultured with bone marrow mesenchymal stem cells. 98 They further quantified B-cell lymphoma 2 (BCL2) expression in leukemia cells using quantitative reverse transcription PCR (qRT-PCR) to compare the reduced rate of apoptosis in 2D and 3D models. The decreasing rate of apoptosis in the 3D model indicated drug resistance.
Zheng and colleagues reported a 3D microfluidic biomimetic model to study leukemia cell-induced bone marrow angiogenesis. They incorporated endothelial cells, leukemia cell lines, and bone marrow stromal fibroblasts to promote and visualize angiogenesis. Quantitative measurements of secreted angiogenic factors in monoculture and coculture indicated a synergistic relationship among three types of cells in angiogenic induction. 99
Lymphoma
To study the complex interactions among different cell types of the diffuse large B-cell lymphoma (DLBCL) TME, Mannino and colleagues created a “lymphoma-on-chip” model by encapsulating DLBCL tumor cells and immune cells in an unconventional hydrogel-based microfluidic model that was traversed by a perfused, vascularized, fully round endothelialized microchannel. 91 This model successfully recapitulated the key complexities and interactions of the in vivo DLBCL TME in vitro ( Fig. 4A ).
Sarkar and colleagues developed a droplet generation and docking array platform integrating genetic and microfluidic analyses to demonstrate the dynamic interactions of NK cells with B-cell non-Hodgkin’s lymphoma (b-NHL) and to evaluate the NK cell-based cytotoxicity on lymphoma cells at a single-cell level ( Fig. 4C ). 92 Tian and colleagues engineered an integrin-specific ligand functionalized 3D “lymphoma organoid” encapsulating either malignant B or T lymphoma cells with follicular dendritic cells in a polyethylene glycol-MAL scaffold. 100 This model demonstrated enhanced proliferation, clustering, and drug resistance to chemotherapeutics and a new HDACi, panobinostat.
To develop a biomicrofluidic CDx that could be used both in a clinical setting and for the drug development process, it is instructive to develop a drug testing assay prototype that may involve other cellular components, such as bone cells, lymph cells, and immune cells, as well as the cytokines and growth factors contained in the bone marrow supernatant, and different novel therapeutic agents ( Table 2 ). To achieve desirable cellular complexity and physiological response, it is imperative to identify the appropriate geometry of the biomicrofluidic device and choose appropriate materials and microfabrication techniques. 101 It is indispensable to factor in a number of other key considerations in the development of a biomicrofluidic CDx, which are outlined in the following section.
Key Considerations for Advancing Biomicrofluidic CDx Platforms
As mentioned, the development of a viable biomicrofluidic CDx platform, and in general any biomicrofluidic system, requires careful consideration of many key design and operational factors. In addition, for CDx platforms in particular, it is crucial to also consider FDA regulatory issues. The USFDA (2016) in vitro diagnostics regulation stipulates three key concepts recommended for CDx development: (1) analytical validity, (2) clinical validity, and (3) clinical utility. 102 To develop a biomicrofluidic CDx, these key concepts need to be coupled with fundamental concepts of microfluidics and microfabrication techniques. In this section, we propose an application of these three fundamental concepts to the development of a biomicrofluidic CDx for therapy selection and drug development for hematologic cancer, and outline associated key considerations. For the purpose of this review article, we only focused on the engineering design and validation phase of the development process.
Analytical Validation
Analytical validation of a CDx refers to establishing the acceptability of the device with regards to its sensitivity, specificity, accuracy, and precision. 102 To satisfy these standards in a biomicrofluidic platform, biological and engineering aspects have to be taken into account. Biological aspects may involve a comprehensive literature review of a liquid TME of interest. Based on the review, key functional features could be identified to mimic the critical functions of the original physiological system in a multiscale biomicrofluidic device, which will be used as a prototype assay. These key functional features may include biomarker of interest, drug metabolism, cellular physiology, systemic paracrine effects, biomarkers, secreted soluble factors, cytokines, and chemokines. Such findings may govern the engineering aspects of the device, which include the complexity of the geometry, functionality, and material choice. To ensure that a biomicrofluidic system provides accurate and unbiased data, and to optimize the geometry of the device, it is critical to carefully simulate the fluid velocity, shear stress, surface tension, and transport of the biomolecule of interest within the system using computational tools (e.g., COMSOL Multiphysics or other commercially available software), and further validate the model with experimental results. Moore and colleagues applied a combination of experimental and computational approaches to measure and predict the concentration pattern of various cytokines (IL-6, VEGF, and TNF-α) predominantly found in the MM TME, 103 using the previously developed MicroC 3 device geometry.74,90
Clinical Validation
Clinical validation of a CDx refers to establishing the ability of the device to identify, measure, and predict the concept of interest to an acceptable degree. 102 To achieve clinical validation, the predictive potential of the device, for example, to measure biomarker activity, has to be assessed at clinical phases I and II using representative primary hematologic cancer cells derived from patients or obtained from a biobank. Clinically equivalent dosing of drugs can be used in the device that can predict in vivo pharmacokinetic properties of a single drug or biologic, or a combination of drugs. This drug-response data can further be evaluated using automated image-based screening platforms and image analysis. A comparison of obtained drug-response data between patients and patient-derived cells would enable the evaluation of the predictive potential of the device. It is also instructive to perform statistical analyses using retrospective dose-response data obtained from cell lines to determine the minimum patient-derived cell population size for each test. Moore and colleagues investigated the effect of using Pittsburgh Heterogeneity Indices (PHIs) on reducing MM tumor cell population size in a microfluidic device, and further found the minimum population size in the device without altering the heterogeneity indices. 104 Predictive potential involves critical pharmacokinetic parameters of drugs, including bioavailability, clearance, and efficacy. Based on the obtained predictive potential, the clinical cutoff of a drug must be selected. Several studies have shown the potential of biomicrofluidic platforms to model such critical pharmacokinetics parameters.105–108 To achieve clinical validation, several reproducible tests are performed across several laboratories. Finally, the device is used to demonstrate the ability to predict the endpoints (i.e., sensitivity, resistivity, selectivity, positive predictive value [PPV], and negative predictive value [NPV]) and clinical safety and efficacy of the drug. 109
Clinical Utility
The clinical utility of a CDx relates to confirming the ability of the device to improve health outcomes, or to provide useful information about diagnosis, treatment, management, or prevention of a disease. 102 To achieve clinical utility, improved patient outcomes, which include benefits, harm reduction, and patient-reported outcomes, have to be assessed using a newly developed biomicrofludic CDx with a hematologic cancer drug. 102
In Table 3, we present a set of key considerations to take into account when testing investigational new drugs, repurposed drugs, combination therapies, and biologics. We anticipate that no single model or assay will accurately recapitulate the full complexity of the hematologic cancer conditions that it intends to represent. However, we believe that careful consideration of the limitations of existing devices and the defining key factors outlined in Table 3 will lead to a more critical evaluation of the predictive outcome of a biomicrofluidic CDx in clinical settings and the drug development pipeline. Also, we anticipate that broad communication of newly proposed ideas and technologies across diverse scientific disciplines will both stimulate and guide further innovative solutions. In Figure 5 , we propose a stepwise framework for the development of a biomicrofluidic CDx model for hematologic cancer, which could be applied in therapy selection and the drug development pipeline.
Key Considerations for Developing Biomicrofluidic CDx Platforms for Hematologic Cancer.
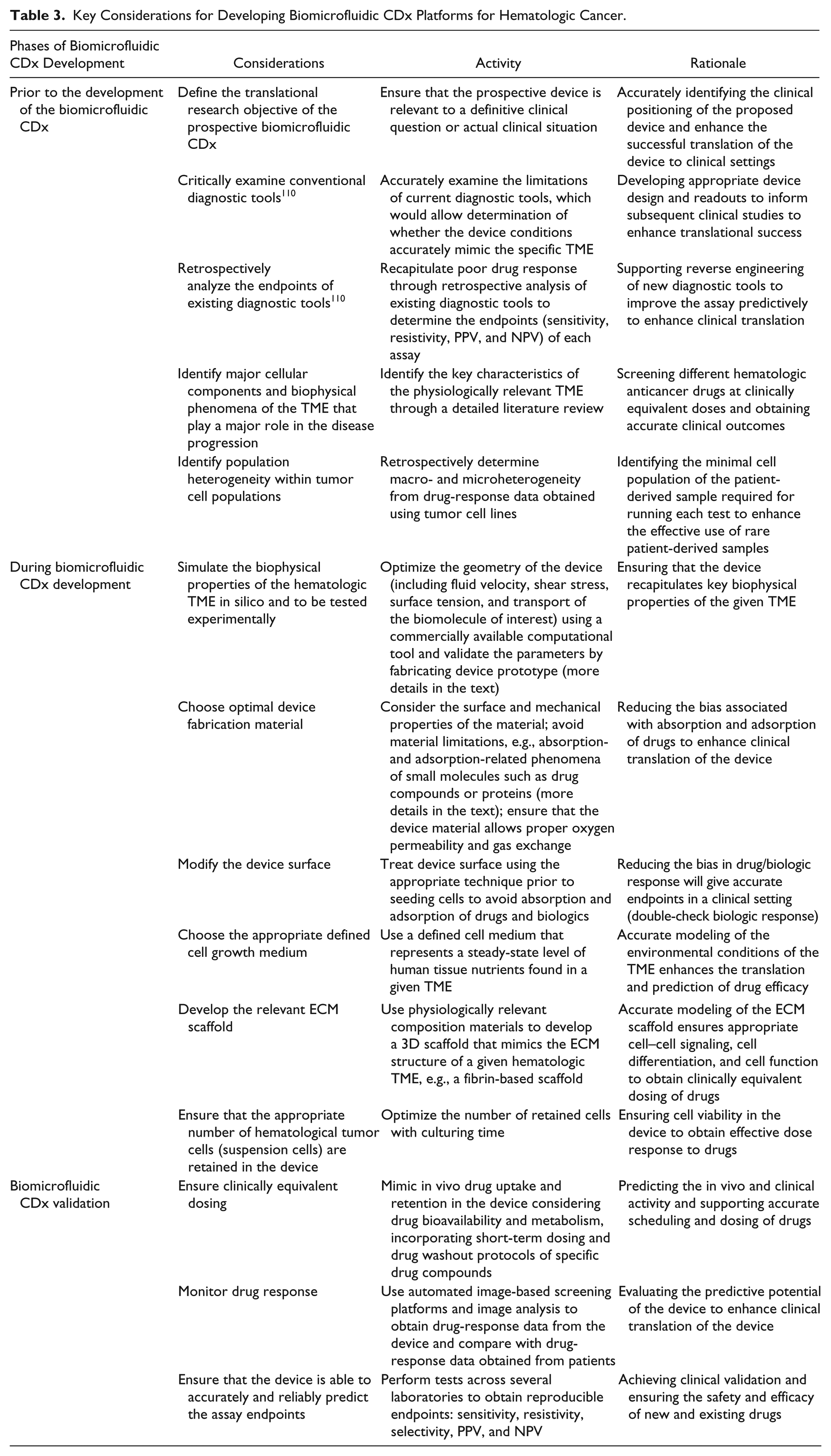
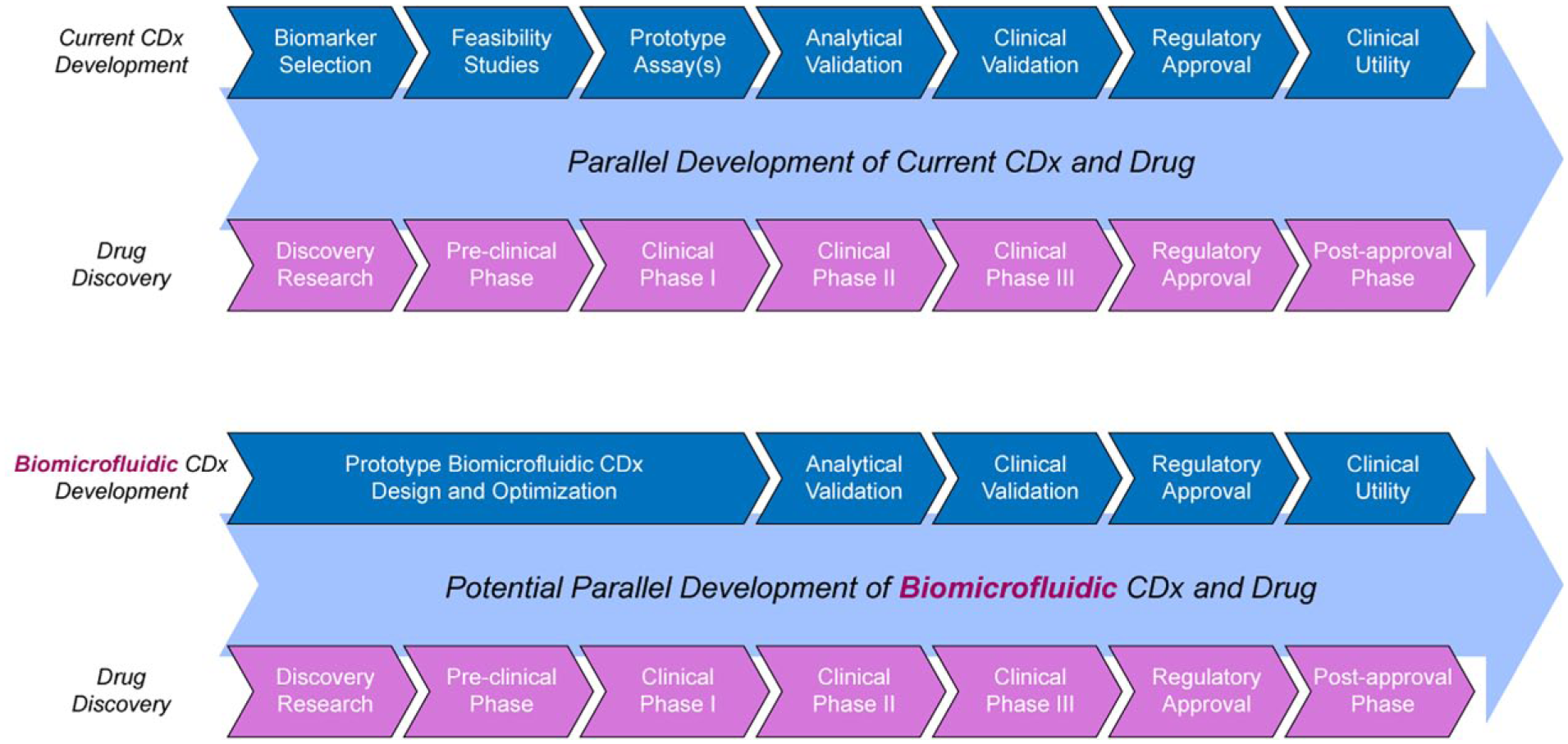
Proposed framework of a biomicrofluidic CDx development model with parallel development of an investigational new drug and conventional framework of a CDx development. The bottom stream represents the positioning of the proposed biomicrofluidic CDx in the drug-diagnostic co-development pipeline in comparison with the conventional framework of a CDx development.
Challenges and Opportunities for Biomicrofluidics for Hematologic Cancers
The biomicrofluidic systems described in this article have not yet reached the stage to be able to test drugs for patients diagnosed with hematological cancer in clinical settings. Significant challenges and opportunities lie ahead for microfluidic engineers to take biomicrofluidic diagnostic systems from bench to bedside. Some of these opportunities are more technical, such as developing a biomicrofluidic CDx to screen investigational new drugs, repurposed drugs, combination therapies, and immunotherapies, which would possibly reduce the huge discrepancy between “terminated” clinical trials and FDA-approved drugs for hematological cancers ( Fig. 1 ). From the pragmatic standpoint, challenges relate specifically to how an end user adopts the system in the clinic considering four key attributes of the system: user operability, compatibility, endpoint measurements, and product readiness. 111 The following discussion aims to synthesize both the future challenges and opportunities faced by engineers, clinicians, patients, and other stakeholders and, importantly, encourage further discussion and debate on the clinical relevance and application of a biomicrofluidic CDx to improve the clinical success rates of hematological cancer treatment. 3
Challenges
Although microfluidic 2D or 3D coculture models have shown immense potential in mimicking the physiologically relevant liquid TME on-chip, most of these devices involve culture of only two or three cell types, which limits the opportunity to explore in vivo drug-induced toxicity on the liquid TME accurately and potentially hinders researchers from gaining important subtle information about pharmacokinetics and pharmacodynamics (PKPD) characteristics of drug compounds. Typically, cell-based microfluidic assays have employed immortalized or transformed liquid tumor cell lines, which are known to possess substantial phenotypic drift. 112 Hence, culturing cells over extended periods and many passages may cause heterogeneity within a cell population, with variability within the same cell line between passages, batches, and even laboratories.113,114 Another major drawback of conventional cell culture technique is that cells are usually grown in standard incubators under high-oxygen partial pressure (~20%), which does not reflect the physiological in vivo steady-state conditions of human organ tissues. Such culture conditions do not accurately represent the unique liquid TME and also impact reactive oxygen species (ROS) production, mitochondrial function, and cell differentiation.111,115 A major practical limitation of 3D coculture is handling temperature-sensitive gels and managing the high cost of reagents. Because most liquid tumor cells are nonadherent by nature, it often becomes challenging to culture these suspension cells in devices, to execute immunostaining procedures and sample washing steps, to choose optimal adherent and nonadherent cell ratios, and to perform high-content analyses. 110
An important engineering consideration is the selection of appropriate device materials for fabricating biomicrofluidic systems. PDMS and glass are attractive materials for the development of these assays due to the fact that the fabrication is simple and rapid, and the end product is transparent and represents the optimal substrate to perform imaging with standard microscopic techniques.116–118 However, PDMS has been reported to show absorption and adsorption of small hydrophobic molecules such as drugs, which have been shown to be significantly higher compared with adsorption to other materials.119,120 Recently, Moore and colleagues showed that the half maximal inhibitory concentration (IC50) of different chemotherapeutic agents for MM tumor cells (bortezomib and carfilzomib) was significantly higher in PDMS microfluidic devices compared with thermoplastic devices, and that this bias was attributable to both higher levels of drug absorption into and adsorption on PDMS. 121 However, thermoplastic devices also have the issue of leaching potentially bioactive chemicals such as plasticizers, and both PDMS and thermoplastic materials have been reported to demonstrate potential adsorption of protein-coated drugs, which may also result in interpretation bias.122,123 Therefore, rationale material selection is required when fabricating cell-based microfluidic assays, and alternative new materials may be an interesting new avenue to investigate for applications in biomicrofluidic systems.
Perhaps one of the most challenging aspects of high-content screening of a 3D cell-based microfluidic assay is image acquisition and analysis, which requires advanced microscopy and image informatics solutions. 124 Conventional assay validation techniques, which are based on standard luminescence or fluorescence microscopy, use average population-based analysis and mask the population heterogeneity in response to culture conditions or drug exposure. Such assays also do not recognize or quantify subpopulations of cells carrying specific markers that could be recognized by single-cell analysis.
Translation of biomicrofluidic CDx into clinical settings is a major challenge. During the design phase of a biomicrofluidic CDx, user operability, compatibility, and endpoint measurements need to be considered to ensure adoption by clinicians. Key design considerations include automation, ease of handling, compatibility with existing equipment, and the ability to generate multiple data points. 111 Moreover, biomicrofluidic systems need to be developed to be compatible with standard clinical laboratory techniques. Even after a biomicrofluidic system reaches maturity, clinicians may continue to provide crucial feedback about usability, compatibility, and assay ability. In the case of hematological cancers, biomicrofluidic engineers may need to collaborate with a hematologist to understand the specialized clinical infrastructure and technology requirements that are specific to hematological cancers.
The parallel development of a drug and a biomicrofluidic CDx necessitates the integration of different areas of expertise, which naturally means that collaboration between researchers, clinicians, and pharma is critical. For instance, pharmaceutical companies may outsource traditional drug development to contract research organizations (CROs), while microfluidic companies may develop the biomicrofluidic CDx technology. However, these collaborations can lead to conflicts arising from different business strategies, financial goals, and logistical issues. 83 Furthermore, currently few CROs have the expertise necessary to conduct clinical trials needed for the approval of conventional CDx tools, resulting in delays in the clinical translation of the device due to regulatory requirements. Hence, co-development of a biomicrofluidic CDx and a pharmaceutical drug requires specialized expertise and facilities, and similar or greater regulatory challenges than those arising from developing the drug and platform independently can be expected.
Opportunities
With the advent of different modes of immunotherapy such as immune checkpoint inhibitors, CAR-T cells, monoclonal antibodies, and vaccines, the potential for personalized immunotherapies is emerging rapidly. Although different modes of immunotherapy exhibit immense potential in hematologic cancer research, many of them have thus far failed to show the desired efficacy in the clinic and clinical trials. Perhaps these issues could be addressed by developing a biomicrofluidic CDx incorporating different immune cell types to test patient-derived samples with different immunotherapeutic agents. However, such platforms require modeling of immune cell network properties, which involves a series of tasks, such as extracting parameters from extended imaging datasets, analyzing parameters using mathematical tools, and deriving a quantitative and predictive description of immune cell behavior and motility.125,126 These platforms need to incorporate certain key features of the immune system that play a vital role in immunotherapy in vivo, such as different immune cell types of the innate or adaptive immune system and cell–cell interactions at both short range (e.g., during development and antigen presentation) and long range (the homing and recirculation of T cells). 125 With more in-depth investigation of the immune system, such a biomicrofluidic CDx could be tailored to select different modes of immunotherapy for hematologic cancer in clinical settings and to identify responsive patient cohorts in clinical trials.
Although current biomicrofluidic systems rely heavily on fluorescence microscopy to acquire endpoint readouts, much information from a complex image cannot be acquired from fluorescently labeled microscopic images. Deep learning could be a solution in such cases. 127 For example, convolutional neural networks (CNNs) may be used for segmenting and subtyping individual fluorescently labeled cells using large numbers of fluorescent images and unlabeled images from phase contrast microscopy.115,128,129 In drug development, deep learning could be used to predict the activity of compounds, predict protein–ligand interactions, and predict compounds given a set of reactants or reaction steps given a final product. 115
Finally, academic–industrial partnerships are essential in the effort to develop and translate a cell-based microfluidic device like a biomicrofluidics CDx for clinical use and drug development. However, a gap exists in the development of such consortia in the area of diagnostic tool development, which results in lack of sufficient funding to conduct such large-scale research and development. Such academic microfluidic diagnostic systems and models have typically not been developed for the purpose of screening or guiding clinical decision-making, and thus remain to be validated for robustness, reproducibility, and clinical predictivity. 110 In the near future we may see this field develop more robust and reliable technologies, stimulating translational funding schemes and precompetitive research consortia to expand their support for the development of new diagnostic tools and screening technologies that can provide improved clinical outcomes.
Summary
Biomicrofluidic systems and organ-on-a-chip technology not only have shown remarkable promise in recapitulating major biological phenomena such as cell–cell interactions, drug resistance, and angiogenesis of complex and dynamic liquid TMEs, but also have achieved reliable drug-response data in vitro. One of the notable abilities of this platform in hematologic cancer research is testing patient-specific samples with different FDA-approved drugs and correlating the obtained responses with corresponding clinical responses obtained from patients subjected to those drug-containing therapies, enabling the identification of patients who are responsive or nonresponsive to a specific drug. As evidenced by the current advancements and abilities of these systems to test different anticancer drugs on patient-derived samples in coculture or multicell culture platforms in hematologic cancer research, it offers great promise as a potential tool for developing a personalized companion diagnostic device. This tool could be used for the purpose of therapy selection in clinical settings to reduce drug-induced toxicity in patients and to select responsive patient cohorts in different phases of clinical trials, which will make clinical trials more informative and efficient. Despite many current challenges, concerted efforts among engineers, biologists, oncologists, clinicians, and regulatory bodies may be able to bring the concept of a reliable and efficient personalized biomicrofluidic treatment platform to fruition, which will lead the field to major scientific breakthroughs in the near future.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors acknowledge financial support from the Cancer Research Society and the Ontario Ministry of Research and Innovation for an Early Researcher Award to EY.
