Abstract
Pathogenic contamination and resistant bacterial infections remain critical concerns in both developed and developing countries. Rapid and sensitive detection of pathogens is still a key requirement for both environmental and clinical settings. This article introduces a simple, colorimetric, cost-effective, and high-throughput system based on a positively charged iron oxide/enzyme complex for the detection of both gram-positive and gram-negative bacteria in water between 103 and 108 cfu/mL. This study provides an effective strategy for the identification and purification of pathogen contamination in drinking water.
Introduction
Exposure to bacteria in drinking water is a serious public health concern. The World Health Organization estimates 502,000 diarrheal deaths induced by drinking contaminated water each year. 1 Any technology for rapid and ultrasensitive detection of pathogenic bacteria is highly desired for food and water safety. Advances in nanotechnology have allowed researchers to develop various strategies for rapid and sensitive bacteria detection in water. They have a similar sensitivity to the current technologies, like PCR, 2 plate counting, and enzyme-linked immunosorbent assay (ELISA). 3 For example, Rotello’s group reported a colorimetric method based on gold nanoparticles (Au NPs) and β-galactosidase.4–6 The activity of β-galactosidase was initially inhibited after precomplexation with Au NPs and recovered after being substituted by bacteria. The reactivated β-galactosidase allowed the quantitative detection of bacteria in both solution and strip paper with the detection limit of 100 cfu/mL. Based on this similar mechanism, other studies replaced Au NPs with ethylenediamine (EDA)-modified graphene oxide (GO), which could achieve the detection of Escherichia coli (E. coli) at concentrations as low as 100 cfu/mL and Staphylococcus aureus (S. aureus) at 200 cfu/mL. 7
Due to their unique magnetic properties, magnetic nanoparticles (MNPs) are also widely explored for bacteria separation and detection.8,9 For example, Nugen et al. prepared bacteriophage-tagged MNPs for the specific recognition and separation of E. coli. 10 Lee et al. developed magneto-DNA probes capable of rapid and specific profiling of bacterial 16S rRNAs using a miniaturized nuclear magnetic resonance (NMR) device. 11 In general, MNPs act as either concentrating affinity probes to enrich bacteria or magnetic resonance contrast agents. However, existing methods required either an additional step with another agent for the bacteria quantification or fancy NMR equipment for the magnetic resonance detection.
This article aims to develop a strategy that combines the features of β-galactosidase and MNP. This strategy can achieve the simultaneous separation and quantification of bacteria without the need for additional agent or equipment. Briefly, a β-galactosidase/MNP complex is developed to enrich and remove the bacteria ( Fig. 1 ). The binding of bacteria on the complex proportionally detaches the β-galactosidase from MNPs, and the freed β-galactosidase can be quantified with colorimetric assay. Due to the high sensitivity of β-galactosidase, we demonstrate the detection of gram-negative E. coli and gram-positive S. aureus with the limit of 1000 cfu/mL in 2 h. In addition, the bacteria were removed from the water samples by applying magnetic field because of the complexation with MNPs.
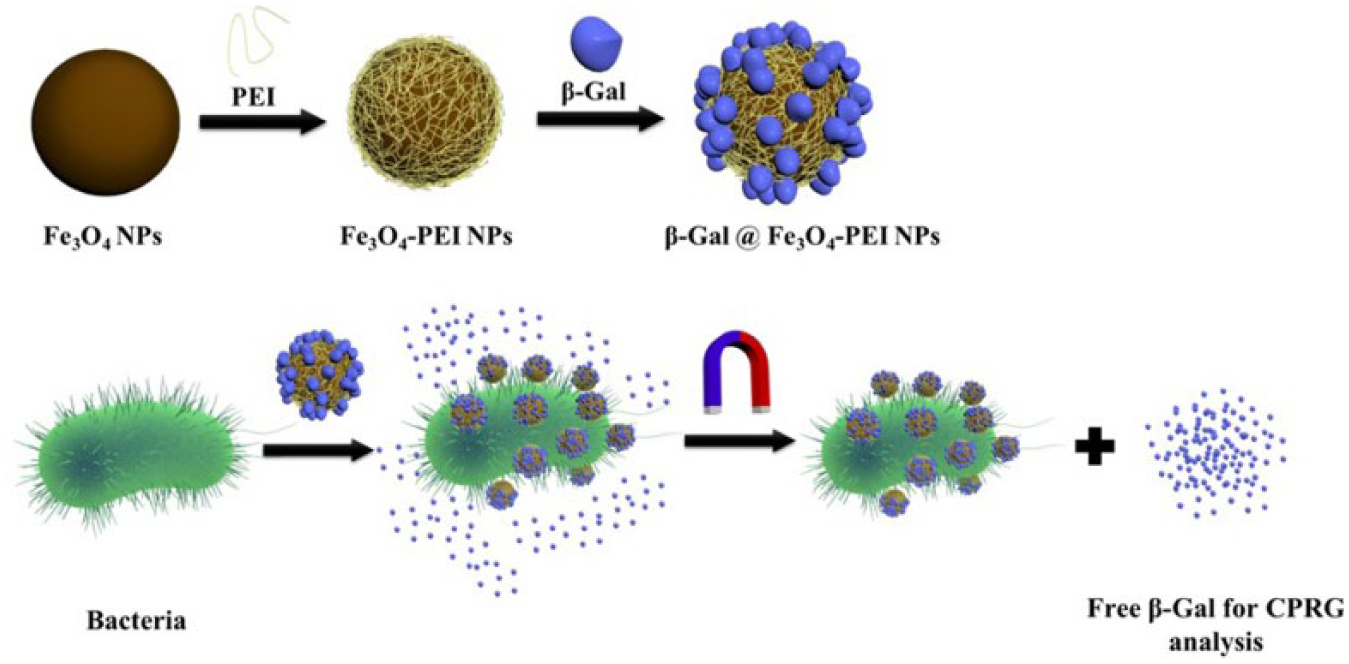
Schematic illustration of the synthesis (
Materials and Methods
All materials except those mentioned specifically were obtained from Sigma-Aldrich (Singapore). Luria-Bertani (LB) broth and LB agar were bought from Becton, Dickinson and Company (Franklin Lakes, NJ). Chloroform (CHCl3) was purchased from TEDIA (Fairfield, OH). E. coli ATCC 25922 and S. aureus ATCC 6538 were from ATCC (Manassas, VA).
Instruments for the Particle Characterization
The hydrodynamic diameter and zeta potential were examined with the ZS Particle Sizer (Malvern Instruments, Malvern, UK). FTIR spectra were acquired with a Fourier-transform infrared spectroscopy–attenuated total reflection (FTIR-ATR) spectrometer. Particle morphology was examined using a transmission electron microscope (TEM, JEOL 2010) operated at 200 kV.
Bacterial Culturing
E. coli and S. aureus were cultured overnight by transferring a colony from the agar plates to culture tubes containing sterilized LB broth medium. Then small aliquots were taken from the tubes and transferred to 10 mL of fresh LB medium to further culture for 4 h. Then obtained bacteria were washed and diluted with 5 mM phosphate buffer (pH 7.4) 5 for experiments.
Plate Counting
Bacterial solution (~100 µL) was uniformly sprayed on the surface of LB agar and cultured in a 37 °C incubator. After 24 h of culture, colonies were counted to derive the initial concentration. All the samples were measured in triplicate.
Synthesis and Modification of Fe3O4 MNPs
Ten-nanometer Fe3O4 MNPs were synthesized by the reduction of Fe(acac)3 in oleyl amine solvent at 300 °C for 2 h. 12 Modification of Fe3O4 MNPs with branched polyethylenimine (PEI, 25 kDa) was achieved by adding 1 mL of Fe3O4 MNP solution (1 mg/mL in chloroform) into 1 mL of CHCl3 containing 3 mg of PEI under stirring. Solution was then dried under a nitrogen flow to obtain the Fe3O4/PEI film, which was dispersed in 0.7 mL of sterile deionized water to get a final concentration of 1 mg/mL. The Fe3O4/PEI complex was separated from free PEI through high-speed centrifugation and redispersed in 5 mM phosphate buffer (pH 7.4). The Fe concentration of the Fe3O4/PEI complex was quantified through inductively coupled plasma mass spectrometry (ICP-MS, Elan DRC-e, PerkinElmer, Waltham, MA).
Synthesis of Fe3O4/PEI/Enzyme Complex
Fe3O4/PEI MNPs were coupled to β-galactosidase via the electrostatic interaction. In brief, 1500 µL of Fe3O4/PEI solution with a concentration of 1 mg/mL was mixed with 50 µL of enzyme (1 mg/mL) in 5 mM phosphate buffer and incubated for 15 min at room temperature. The free enzyme in supernatant was removed by magnetic separation and subsequently quantified using a chlorophenolred-β-
Bacterial Enrichment and Quantification with Fe3O4/PEI/Enzyme Complex in Known Solutions
The Fe3O4/PEI/enzyme complex was incubated with known solutions containing various concentrations of bacteria ranging from 102 to 107 cfu/mL for 1 h. During the incubation, gentle shaking (80 rpm) was applied to accelerate the replacement process. Supernatant containing free enzyme molecules was later separated from the MNP complex under the magnet, and free enzyme molecules were quantified by CPRG assay, recording the absorbance at 574 nm after reaction with 500 µM CPRG.
Bacterial Enrichment and Quantification with Fe3O4/PEI/Enzyme Complex in Unknown Solutions
The study was performed in a single-blinded manner, with the preparation and analysis kept separate. Solutions containing various amounts of bacteria were performed by investigator 1, who was kept separate from the data acquisition workflow. Investigator 2 collected the raw data and performed analysis. Thereafter, the identity of the various groups was revealed by investigator 1 and matched with the experimental data. The sample process was the same as that with known solutions.
Results
Synthesis and Characterization of Fe3O4/PEI NPs
Fe3O4 NPs with a radius of 10 nm were synthesized using the traditional high-temperature deposition method (
Fig. 2a
). The as-synthesized NPs were hydrophobic and made hydrophilic through complexation with cationic branched PEI. The free PEI postsurface modification was removed through high-speed centrifugation. We optimized the mass ratio between PEI and MNPs during the synthesis of the Fe3O4@PEI complex. As shown in
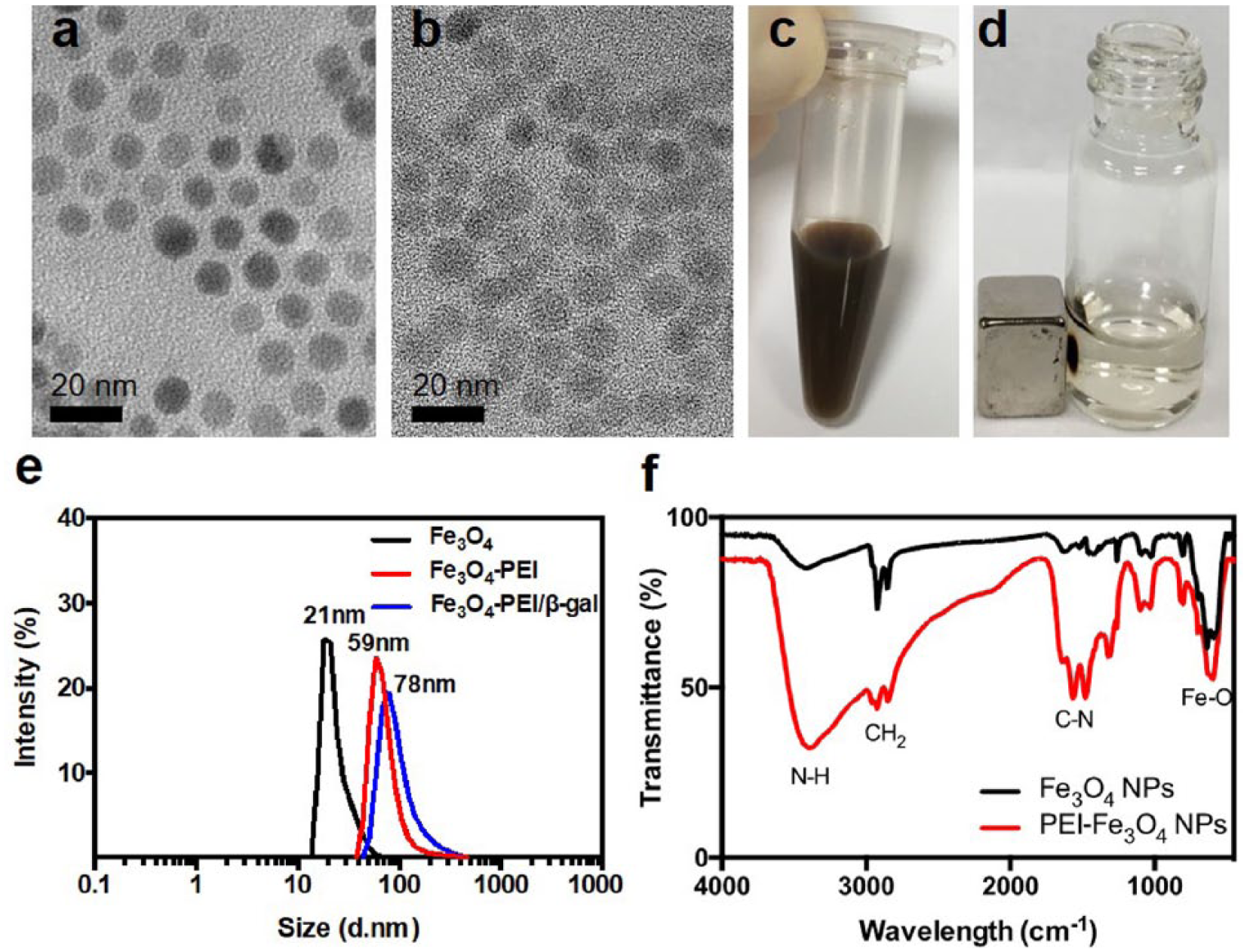
Characterization of Fe3O4/PEI NPs. TEM images of (
The modified particles dispersed well in aqueous solution and could respond to the magnet attraction ( Fig. 2c , d ). There was no significant change of particle size after the PEI modification under TEM ( Fig. 2b ); however, the hydrodynamic diameter changed from 21 to 59 nm ( Fig. 2e ). The FTIR result reveals that there are strong absorptions from N-H stretch vibration (3300–3500 cm–1) after the modification, which confirms the successful modification of NPs with PEI ( Fig. 2f ).
Modification of Fe3O4/PEI NPs with β-Galactosidase
The coating of cationic PEI provided a positive surface charge (i.e., +22 mV) for Fe3O4/PEI NPs in phosphate buffer, which allowed the complexation with β-galactosidase via electrostatic interactions. Fe3O4/PEI/β-galactosidase was magnetically separated with free β-galactosidase. After the complexation, the zeta potential dropped from +20 to +10 mV. The prepared Fe3O4 NPs and Fe3O4 NP/PEI and Fe3O4 NP/PEI/β-galactosidase complexes had a hydrodynamic diameter of about 21, 59, and 78 nm, respectively ( Fig. 2e ). The PEI coating increased the hydrodynamic diameter of Fe3O4 NPs from 21 to 59 nm. The further increase from 59 to 78 nm was due to the binding of the enzymes.
To calculate the loading capacity of enzymes on the complexes, we first generated a standard curve between CPRG concentrations and maximum reaction velocities (Vmax) (
Sensitivity of Fe3O4/PEI/β-Galactosidase Complexes in Bacterial Enrichment and Quantification
Fe3O4/PEI/β-galactosidase complexes were first used to confirm the already-known concentrations of gram-negative E. coli in a series of solutions. Briefly, solutions containing a range of bacterial concentrations were incubated with Fe3O4/PEI/β-galactosidase complexes. Enzymes on the complexes were proportionally replaced by E. coli due to electronic interaction between the bacterial membrane and Fe3O4/PEI/β-galactosidase NPs. The higher the concentration of bacteria was, the more enzymes that were freed. The enzymes detached from the complexes were quantified by the CPRG assay. Specifically, the supernatant containing detached β-galactosidase was mixed with CPRG and the dynamic change of absorbance at 574 nm was monitored (
Fig. 3a
and
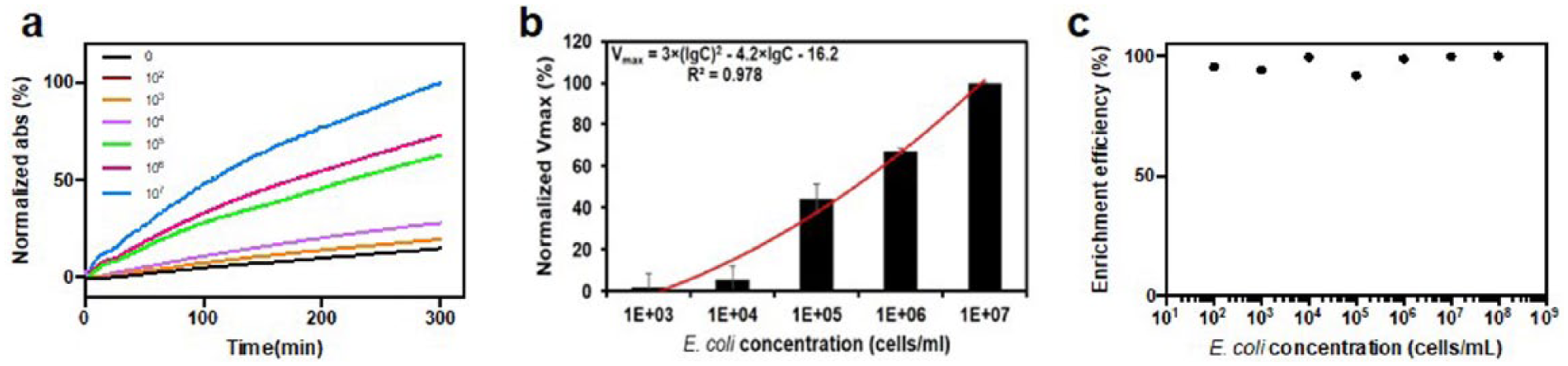
Fe3O4/PEI/β-galactosidase complexes for the enrichment and analysis of E. coli in samples with known concentrations. (
We further looked into the quantification of gram-positive S. aureus using a similar protocol to understand the sensitivity of this system (
Fig. 4a
and
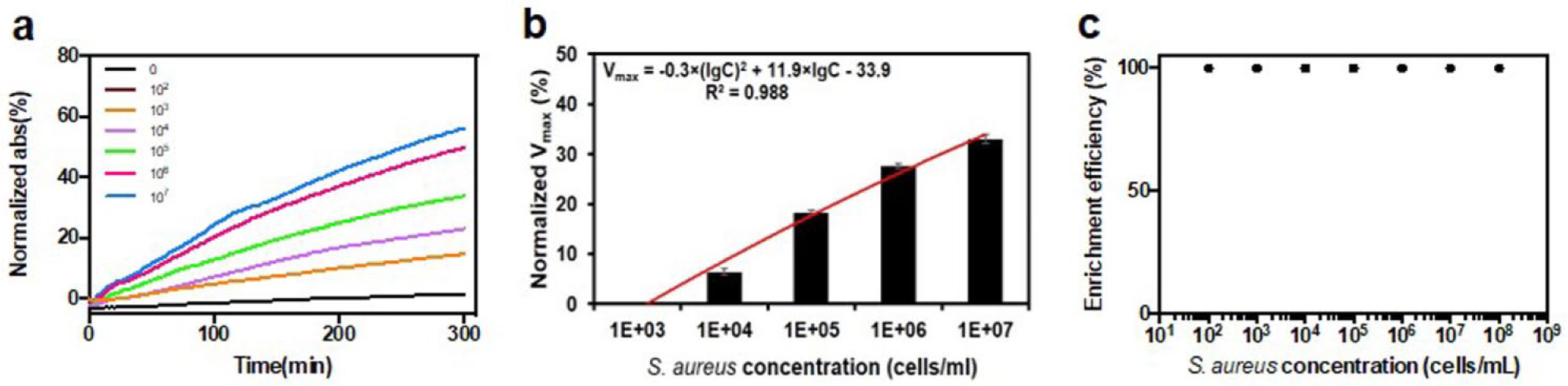
Fe3O4/PEI/β-galactosidase complexes for the enrichment and analysis of S. aureus in samples with known concentrations. (
To better understand the different performances of Fe3O4/PEI/β-galactosidase NPs between gram-positive and gram-negative bacteria, we studied the surface charge of two kinds of bacteria ( Fig. 5 ). EC-exp and SA-exp represent the surface charges of E. coli and S. aureus, respectively, at the exponential phase, while EC-sta and SA-sta represent the charges of E. coli and S. aureus, respectively, at the stationary phase. The surface charge of E. coli was around 10 mV lower than that of S. aureus for both exponential and stationary phases.
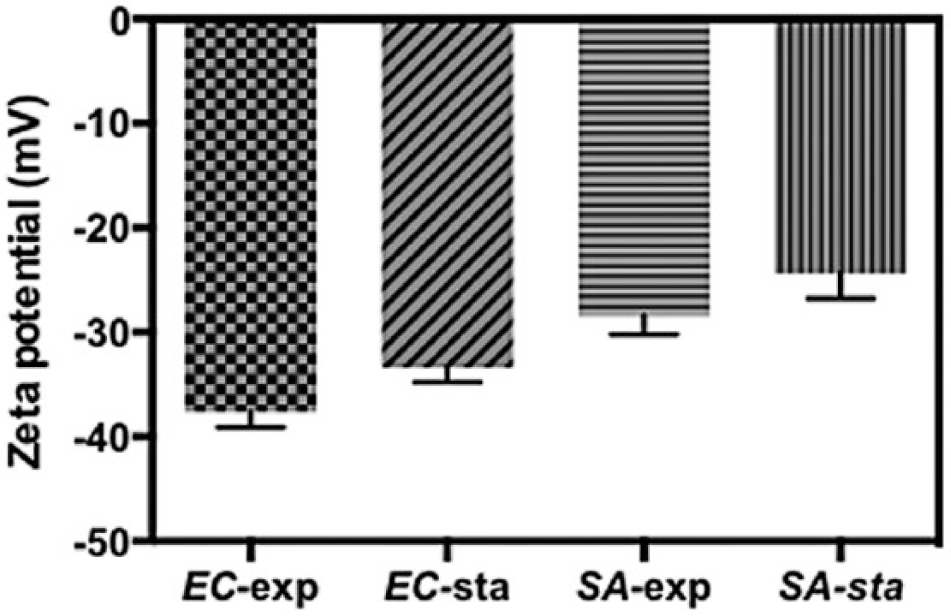
Zeta potentials of E. coli and S. aureus (exponential phase and stationary phase).
Fe3O4/PEI/β-Galactosidase NPs for the Enrichment and Quantification of E. coli and S. aureus in Unknown Solutions
The standard curves derived above allowed us to predict the bacterial concentration in an unknown sample ( Fig. 6 ). To evaluate the efficacy of this system, a blind assay was conducted. Briefly, E. coli and S. aureus were cultured for 7–9 h randomly with different numbers of inoculation colonies. This step was carried out by one person, samples were labeled only by numbers, and the corresponding sample names were recorded at the same time. Then the measurements were blindly conducted by another person ( Table 1 ).
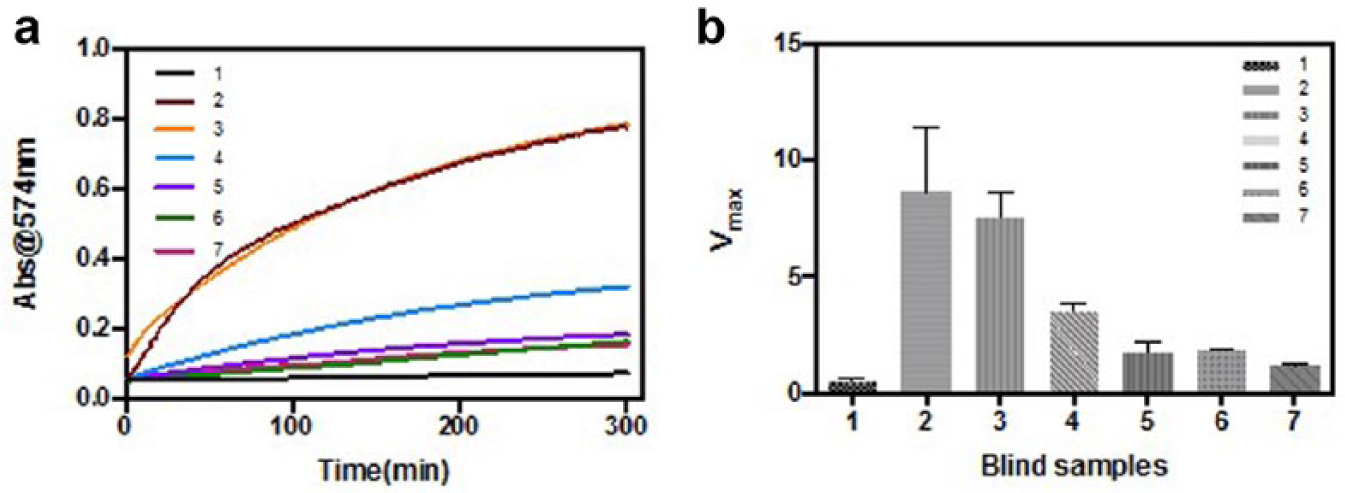
Fe3O4/PEI/β-galactosidase complexes for the enrichment and analysis of bacteria in samples with unknown concentrations. (
Bacteria Samples Used in the Blind Assay and the Quantification from Both Plate Counting and the Fe3O4/PEI/β-Galactosidase Complex Assay.
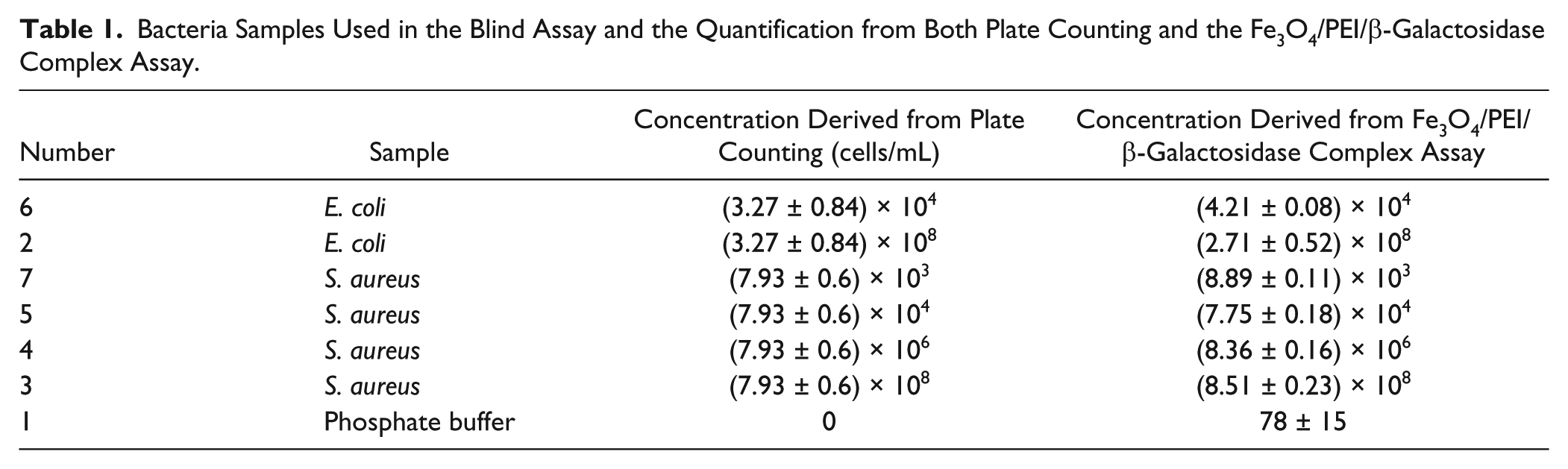
As shown in Figure 6a , the control group had the lowest absorbance at 574 nm, while samples 2–3 showed the highest absorbance. The derived Vmax confirmed this observation ( Fig. 6b ). Samples 6–7 ended up with similar absorbance and Vmax to the control. Vmax values were brought back to the standard curves in Figures 3b and 4b for the calculation of bacterial concentrations. Simultaneously, the concentrations were examined with the standard plate counting. As shown in Table 1 , results from two kinds of assays matched well for all samples except sample 2 (E. coli at the highest concentration, 107 cfu/mL).
Discussion
Fe3O4/PEI/β-galactosidase complexes were constructed with oleic acid-coated Fe3O4 NPs, 25 kDa branched PEI, and β-galactosidase. Fe3O4 NPs synthesized through thermal decomposition with oleic acid coating are well known for their uniform size and superior magnetic properties.
17
PEI is also widely known for its capability to bring the hydrophobic Fe3O4 NPs to aqueous solution.18,19 We compared the effectiveness of branched PEI with molecular weights of 0.8, 25, and 50–100 kDa (
The anionic enzyme (i.e., β-galactosidase) was then easily complexed with the Fe3O4/PEI NPs in solution. Similar to other systems reported,
5
positively charged Fe3O4/PEI NPs bound with β-galactosidase at the negatively charged residues around its active site through electrostatic interactions, resulting in the inhibition of enzymatic activity (
We examined the detection of two kinds of bacteria: gram-negative E. coli and gram-positive S. aureus. First, we generated a standard curve with the known concentrations of both bacteria ( Figs. 3b and 4b ). However, we realize that the sensitivity of this methodology is lower than the reported results from the Au NP/β-galactosidase and EDA-GO/β-galactosidase complexes, both of which allowed the detection of E. coli at concentrations as low as 100 cfu/mL. In comparison, Fe3O4/PEI/β-galactosidase complexes can provide a trustable readout when the bacterial concentration is above 1000 cfu/mL. This should be due to the strong absorption of Fe3O4 NPs at 574 nm, which presents a strong background and requires a higher concentration of CPRG metabolites. In addition, we noticed that there was aggregation of NPs when the bacterial concentration of E. coli was above 108 cfu/mL. This suggests the disruption of the cationic PEI coating on the Fe3O4 NPs. In other words, the binding between cationic Fe3O4/PEI and anionic bacterial membrane destabilizes the PEI coating. With a higher bacteria versus Fe3O4/PEI ratio, the destruction becomes more severe. This is confirmed by the less severe aggregation of NPs in the S. aureus solution with the same 108 cfu/mL concentration. S. aureus is less negatively charged than E. coli. ( Fig. 5 ), which means that more S. aureus is needed to achieve the same destabilization of the PEI coating.
Finally, we explored use of the derived standard curve to quantify the bacteria in unknown solutions ( Fig. 6 and Table 1 ). While the result obtained from the Fe3O4/PEI/β-galactosidase complexes is comparable to that from plate counting, we note that this method did not allow us to identify the type of bacteria. In other words, it only allows us to enrich and quantify a bacterium with the known species.
In summary, this article introduces an Fe3O4/PEI/β-galactosidase complex for the enrichment and quantification of both gram-negative and gram-positive bacteria. We can recurrently realize the quantification for bacteria between 103 and 108 cfu/mL. Despite the sensitivity needing further improvement, the method is universal for all kinds of bacteria, and the bacteria enriched can be used for further analysis. In the future, an antibody or aptamer could be modified onto the surface of NPs to further identify bacterial types in unknown samples.
Supplemental Material
Supplemental material for Detection of Bacteria in Water with β-Galactosidase-Coated Magnetic Nanoparticles
Supplemental material for Detection of Bacteria in Water with β-Galactosidase-Coated Magnetic Nanoparticles by Mingyue Cui, Hao Chang, Yang Zhong, Min Wang, Tianze Wu, Xiao Hu, Zhichuan J. Xu, and Chenjie Xu in SLAS Technology
Footnotes
Acknowledgements
We appreciate the support from Nanyang Environment & Water Research Institute, Nanyang Technological University.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
