Abstract
The detection of bacterial pathogens plays an important role in many biomedical applications, including clinical diagnostics, food and water safety, and biosecurity. Most current bacterial detection technologies, however, are unsuitable for use in resource-limited settings where the highest disease burdens often exist. Thus, there is an urgent need to develop portable, user-friendly biosensors capable of rapid detection of multiple pathogens in situ. We report a microfluidic chip for multiplexed detection of bacterial cells that uses antimicrobial peptides (AMPs) with species-specific targeting and binding capabilities. The AMPs are immobilized onto an electrical impedance microsensor array and serve as biorecognition elements for bacterial cell detection. Characterization of peptide immobilization on the sensors revealed robust surface binding via cysteine-gold interactions and vertical alignment relative to the sensor surface. Samples containing Streptococcus mutans and Pseudomonas aeruginosa were loaded in the chip, and both microorganisms were detected at minimum concentrations of 105 cfu/mL within 25 min. Measurements performed in a variety of solutions revealed that high-conductivity solutions produced the largest impedance values. By integrating a highly specific bacterial cell capture scheme with rapid electrical detection, this device demonstrates great potential as a next-generation, point-of-care diagnostic platform for the detection of disease-causing pathogenic agents.
Introduction
Bacterial pathogens are a major cause of human and veterinary morbidity and mortality worldwide.1–3 In particular, bacterial meningitis, pneumonia, tetanus, and diarrhea are leading causes of death in the developing world, especially among neonates and infants. 4 Recent studies indicate that neonatal infectious account for 1.5 to 2 million deaths each year in developing countries.5,6 The emergence of antibiotic-resistant pathogens further emphasizes the seriousness of this problem, particularly in the context of current issues related to biosecurty.7,8 In addition, microbial contamination of food and water inflict major losses in production and economic security. 9 Therefore, new point-of-care biosensors are needed to rapidly detect pathogenic bacteria and minimize infectious outbreaks.
Current technologies for detecting microbial pathogens, such as enzyme-linked immunosorbent assay (ELISA) 10 and polymerase chain reaction (PCR), 11 offer quantitative, high-sensitivity measurements. However, these methods are generally time-consuming, require expensive laboratory facilities and equipment, and rely on skilled technical expertise. Micro-electro-mechanical systems (MEMS)–based biosensors have recently been developed for bacterial detection to circumvent some of these problems.12–14 In general, these technologies rely on antibodies for molecular recognition because of their high specificity for antigen binding. Antibody-based MEMS biosensors, however, have limited stability in high-temperature environments and require highly specific, antibody-antigen pairs for each target. 15 Most environmental and clinical samples contain diverse cell populations, making it extremely challenging to target specific bacterial species. For example, recent results from the Human Microbiome Project identified more than 10,000 different microbial species residing in the human body. 16 In the mouth alone, more than 700 bacterial species have been identified, 17 making species-specific detection extremely challenging using current antibodies-based approaches. Antimicrobial peptides (AMPs) offer a promising, alternative approach for biorecognition due to their ease of synthesis, 18 high stability,19,20 and enhanced target selectivity.21,22 Bacterial biosensors employing AMPs have been reported.23,24 Mannoor et al. 24 have developed a microcapacitive sensor employing magainin I, a natural AMP found on the skin of African clawed frogs. This device employs a single sensor for high sensitivity, label-free detection of pathogenic bacteria, however, its specificity is limited because of the broad-spectrum binding activity of magainin I toward Gram-negative bacteria. Thus, the development of a point-of-care biosensor for rapid, high-specificity, multiplexed pathogen detection in environmental and clinical samples has yet to be realized. In this work, we present a microfluidic biosensor chip using synthetic AMP-coated microsensors for species-specific targeting and rapid detection of bacterial cells. An electrical impedance sensing scheme is employed for rapid, multiplex, label-free detection.25,26 Using this device, we simultaneously detected Streptococcus mutans and Pseudomonas aeruginosa, two common human pathogens,27–29 within polymicrobial samples. Our AMP-based, electrical bacterial biosensor will facilitate the future development of additional microsensor-based diagnostic technologies.
Materials and Methods
Bacterial Culture and AMP Preparation
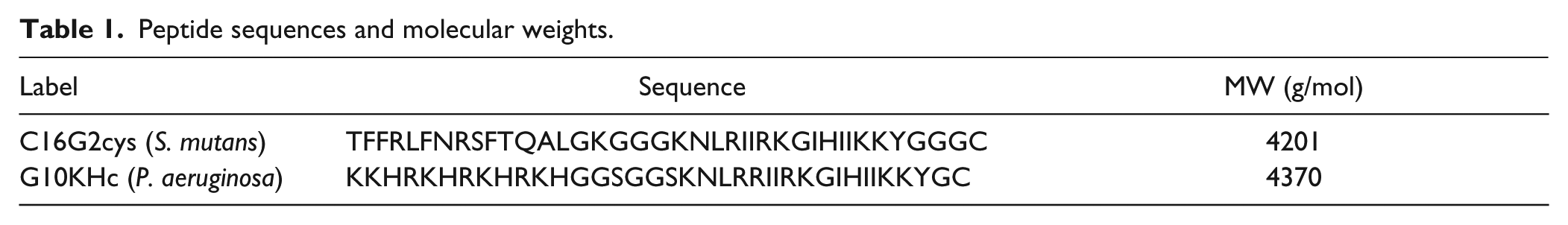
S. mutans was grown in brain-heart infusion or Todd-Hewitt medium (Difco, Franklin Lakes, NJ) at 37 °C under anaerobic conditions (80% N2, 10% CO2, and 10% H2). 30 P. aeruginosa was grown in Luria-Bertani (LB) broth (Fisher Scientific, Waltham, MA) at 37 °C under aerobic conditions. The AMPs C16G2cys (S. mutans) and G10KHc (P. aeruginosa; Table 1 ) were synthesized and purified as previously described.31,32 Peptide solutions were prepared in deionized (DI) water:methanol (1:1) at 1.0 µM, stored at −17 °C, and thawed to room temperature prior to experiments.
Peptide sequences and molecular weights.
Fabrication of the Microfluidic and Biosensor Chips
Poly(dimethylsiloxane) (PDMS) molds were fabricated on silicon wafers via photolithography (Karl Suss, Garching, Germany) and DRIE (Unaxis, Schwyz, Switzerland). A PDMS mixture (Sylgard 184) was poured onto the mold, cured for 2 h at 80 °C, and cut into individual chips. A polyethylene glycol surface coating was applied to the PDMS chips as previously described 33 to minimize cell attachment on the channel walls. The fabrication of the microsensor array was performed via photolithography to pattern AZ4620 photoresist as a shadow mask for metal evaporation. Cr and Au were evaporated (CHA Mark 40) onto glass slides (Fisher Scientific), and lift off was performed via sonication in acetone. Devices were assembled by bonding the PDMS and glass chips together, which were subsequently rinsed in isopropanol and DI water and dried using compressed N2. Device assembly was performed under a microscope to ensure proper alignment of the microchannels with the microsensor array. The bonding between PDMS and glass is reversible such that PDMS can be repeatedly removed and rebounded to glass. Although the bonding is not permanent, leakage was not observed during experimentation.
Immobilization of Peptides on the Microsensor Array
AMPs were immobilized onto the microsensors by flowing 1.0 mM peptide solutions inside a microfluidic chip for subsequent incubation. This chip ( Fig. 2A ) is composed of four individual channels that flow over the capture electrode of each sensor. Peptide solutions were manually dispensed into the inlets of the device using a pipette and pumped through the channel using a syringe. After a 40 min incubation period, the peptide solutions were flushed out of the channels, followed by rinsing in DI water. The PDMS chip was detached from the microsensor array, which was rinsed in DI water and dried using compressed N2.
Characterization of Immobilized Peptides Using AFM
Surface topologies of AMP-coated sensors were captured in air using a Digital Instruments (Santa Barbara, CA) MultiMode Scanning Probe Microscope (SPM) with a Nanoscope 3A controller operating in tapping mode. Specimens were mounted to steel discs, which were magnetically attached to the stage. Silicon probes (Veeco Probes, Camarillo, CA) were used with a typical tapping frequency of 240 to 280 kHz and a nominal scanning rate of 0.8 to 1.0 Hz. Images were analyzed and processed using Digital Instruments Nanoscope R IIIa software.
Optical and Fluorescent Imaging
Device assembly was performed under a Leica microscope (DM4000M) equipped with a CoolSNAPHQ Monochrome CCD camera (Photometrics, Tucson, AZ). Peptide immobilization and cell binding were visualized using fluorescence microscopy. Images were captured using RS Image software. Fluorescent images were processed (imaging stacking, contrast enhancement) using ImageJ software.
Electrical Impedance Measurements
Impedance measurements were carried out using a 4294A precision impedance analyzer (Agilent, Palo Alto, CA), which has an impedance accuracy of ±0.08%. Kynar wiring (30 AWG) was used to connect the impedance analyzer to the biosensor chips. Spring-loaded connector pins (Digi-Key, Thief River Falls, MN) were soldered at one end of the wires, facilitating contact to the electrodes. A polycarbonate chip holder was fabricated using a MAXNC 15 computer numerical controlled mill. After securing the chip in the holder, samples were dispensed into the inlet using a syringe and incubated for 20 min to allow the bacterial cells to bind to the immobilized peptides. Unbound bacteria were removed by flushing with phosphate-buffered saline (PBS; Fisher Scientific). After three rinsing cycles, PBS (pH 7.2, 38 mS/cm), low-conductivity buffer (LCB), or DI water (0.1 µS/cm) was pumped into the device for impedance measurements. The LCB solution was 10 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES), 0.1 mM CaCl2, 59 mM D-glucose, and 236 mM sucrose in DI water, having a pH and conductivity of 7.35 and 0.28 mS/cm, respectively. A 100 mV excitation voltage was applied for all measurements, and impedance was scanned from 100 Hz to 100 MHz in a logarithmic scale with 200 data points for each scan. Each measurement lasted 5 s. Experiments were performed using new chips, which were discarded after each measurement.
Design of the Biosensor Chip
Our device consists of a gold microsensor array on a glass substrate and a PDMS microfluidic chip ( Fig. 1 ). Each sensor is composed of three electrodes: an excitation electrode (EE), a ground electrode, and a capture electrode (CE). The CE is coated with AMPs and serves as the binding sites for the target bacterial cells. All electrodes are 200 µm in width, and the CE has dimensions of 200 µm × 200 µm. The vertical spacing between the CE and the EE/CE is 50 µm, and the horizontal spacing between each sensor is 500 µm. Because of its compact design, additional sensors can be added to the array, enabling for the detection of additional cell types while still maintaining portability.
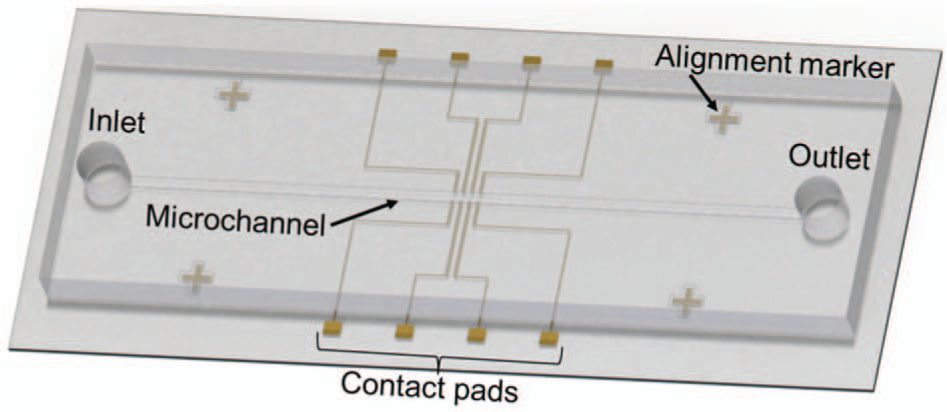
Schematic of the biosensor chip for cell detection measurements. The chip consists of a microsensor array and a PDMS microchannel.
Two PDMS chips are employed in this work: chip A is used for peptide immobilization, and chip B is used for impedance measurements. Chip A ( Fig. 2A ) consists of four channels corresponding to each sensor in the array. Each channel has a separate inlet and outlet having, diameters of 2 mm and 1 mm, respectively. The channels are designed to flow across the CE without contacting the electrode leads to minimize nonspecific cell binding. Channels are 200 µm wide and expand to 250 µm as they cross over the CEs to ensure optimal coverage of the peptide solution, accommodating for possible misalignment between the PDMS chip and sensor substrate. Chip B ( Fig. 2B ) consists of a 400 µm wide channel with an inlet and outlet having diameters of 2 mm and 1 mm, respectively. The channel height is 50 µm, which allows the cells to be in close proximity to the CEs, thereby enhancing AMP-cell recognition and binding. A channel height <50 µm can impede the fluid flow and potentially clog the channel. Cross-shaped alignment marks are positioned at the corners of the PDMS and sensor substrates to facilitate device assembly.
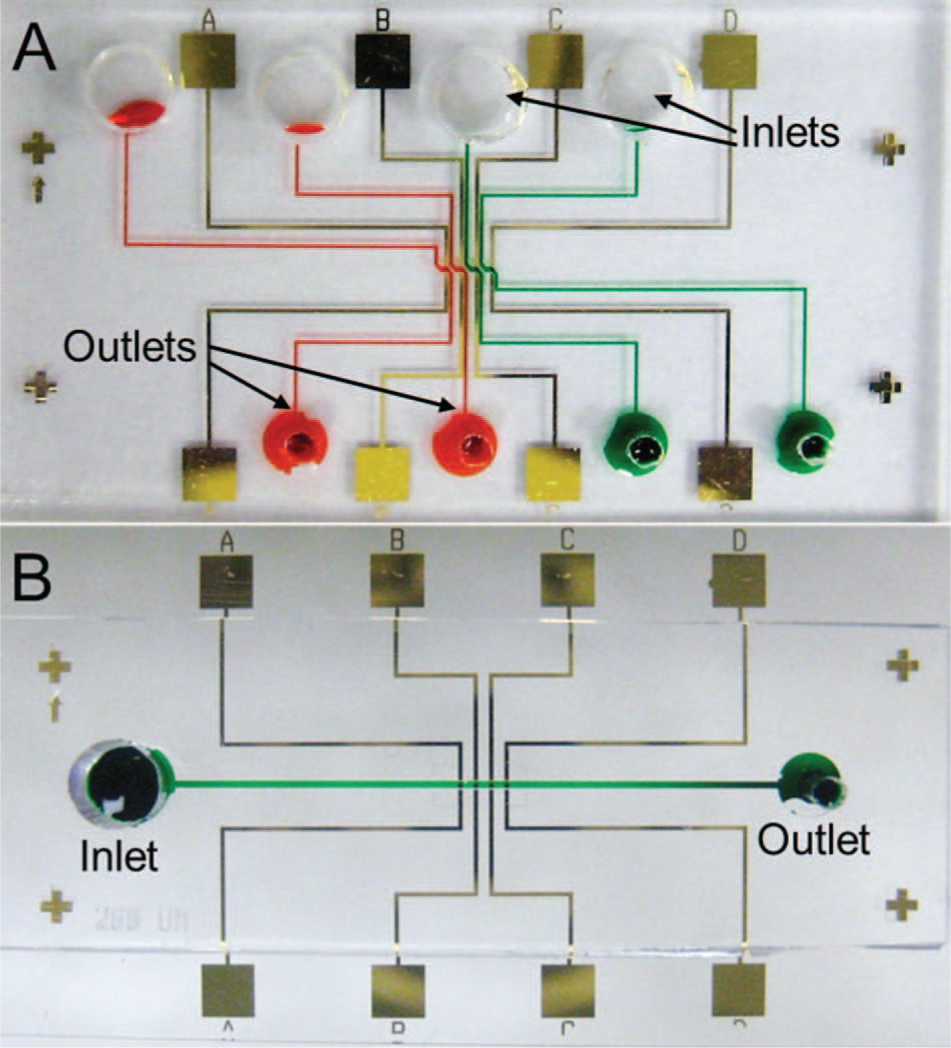
Photographs of the PDMS chips used for (A) peptide immobilization and (B) impedance measurements. Channels are filled with colored dye for improved visualization.
Results and Discussion
Characterization of Peptide Immobilization
Immobilization of the AMPs on the sensors was carried out via self-assembled monolayer (SAM) formation. The quality of SAM formation is influenced by several factors, including the nature of the chemical interaction between the substrate and adsorbate, as well as the type and strengths of intermolecular interactions between the adsorbates that are necessary to hold the assembly together.
34
In this work, the AMPs were synthesized with a cysteine on their C-terminus for robust thiol-gold binding.
35
Fluorescent-tagged AMPs were incubated at various time periods and observed using fluorescent microscopy to investigate AMP incubation timing. After a 40-min incubation, the sensors exhibited uniform peptide coverage across their surfaces, as shown in
AFM scans were performed to characterize the binding of AMPs to the gold sensors. As shown in Figure 3 , AMPs can be clearly observed on the gold surface. These three-dimensional scans reveal that the peptides are vertically aligned, validating that binding occurs at the C-terminal cysteine residue. Because the C16G2cys and G10KHc AMPs do not contain additional cysteines, peptide misalignment is unlikely. This upright orientation of the AMPs is crucial for effective cell recognition and binding. Topographic data also suggest that the AMPs are arranged vertically. Peptide aggregates measure ~16 nm in height, matching closely with measurements of similar-sized peptides. 36 The AMPs also demonstrate robust surface binding, being able to withstand repeated rinsing in DI water/methanol solution. From these results, it can be determined that the unique design of the AMPs enables for optimal surface alignment, surface uniformity, and binding strength.
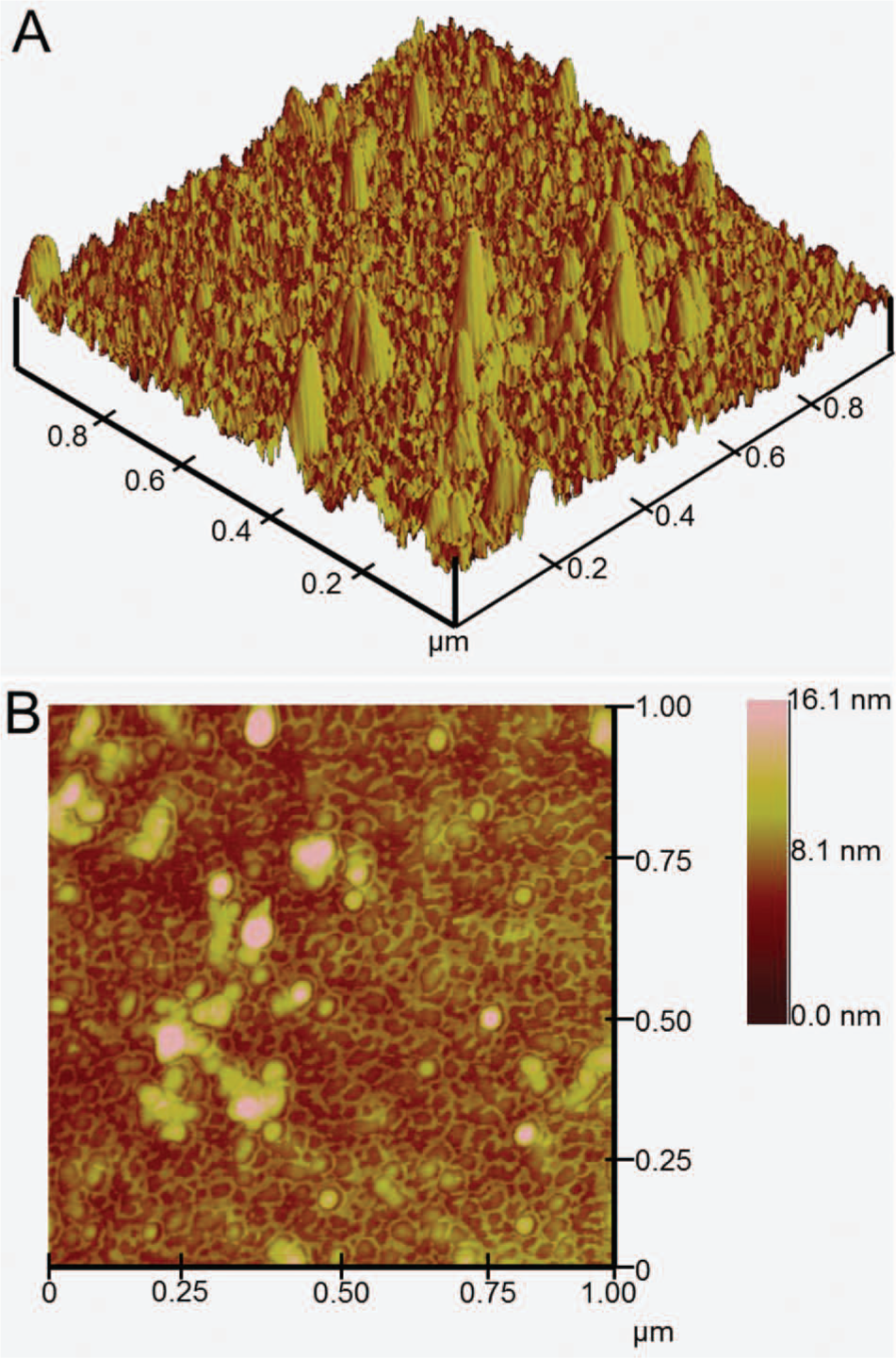
Atomic force microscopy images of antimicrobial peptides immobilized on gold. The scan size and z-scale are 1.0 µm x 1.0 µm and 16.1 µm, respectively.
High-Selectivity Cell Targeting and Binding
The ability to capture target analytes is highly dependent on the biorecognition elements employed. In the AMP approach, peptide affinity for their respective bacterial cells is governed by hydrophobicity, electrostatic/cationic interactions, and secondary folding structure.37,38 The AMPs used in this work were designed to selectively target S. mutans30,31 and P. aeruginosa 32 within polymicrobial samples, making this detection scheme highly specific and technologically advanced compared with antibody-based targeting schemes. The development of these peptides was achieved through an efficient screening process among rationally designed peptide libraries. Potential candidates with highest binding affinity to target cells were chosen for amino acid sequence modifications and were subsequently rescreened for binding optimization. Further, AMPs are more robust compared with antibodies because of their smaller molecular sizes (<50 amino acids), making them highly stable and more suitable for applications requiring an extended shelf life.
Binding assays against G10KHc- and C16G2cys-coated microsensors were performed using fluorescently labeled P. aeruginosa (red) and S. mutans (green). A mixture of S. mutans and P. aeruginosa was dispensed into the microchannel and incubated for 20 min ( Fig. 4A ). The solution was flushed out and rinsed with PBS three times to remove unbound cells. As shown in Figure 4B , the remaining florescent signals represent cells bound to the sensors. Sensors S1 and S2 exhibit strong preferential binding of P. aeruginosa ( Fig. 4C ) with negligible binding of S. mutans (green). Similarly, sensors S3 and S4 exhibit preferential binding of S. mutans ( Fig. 4D ) with negligible binding to P. aeruginosa. Immobilized bacterial cells are localized within the sensor regions ( Fig. 4B , outlined in white), demonstrating minimal cross- and nonspecific binding. These results demonstrate the high specificity and strong binding properties of AMPs, making them well suited as biorecognition elements for cell detection.
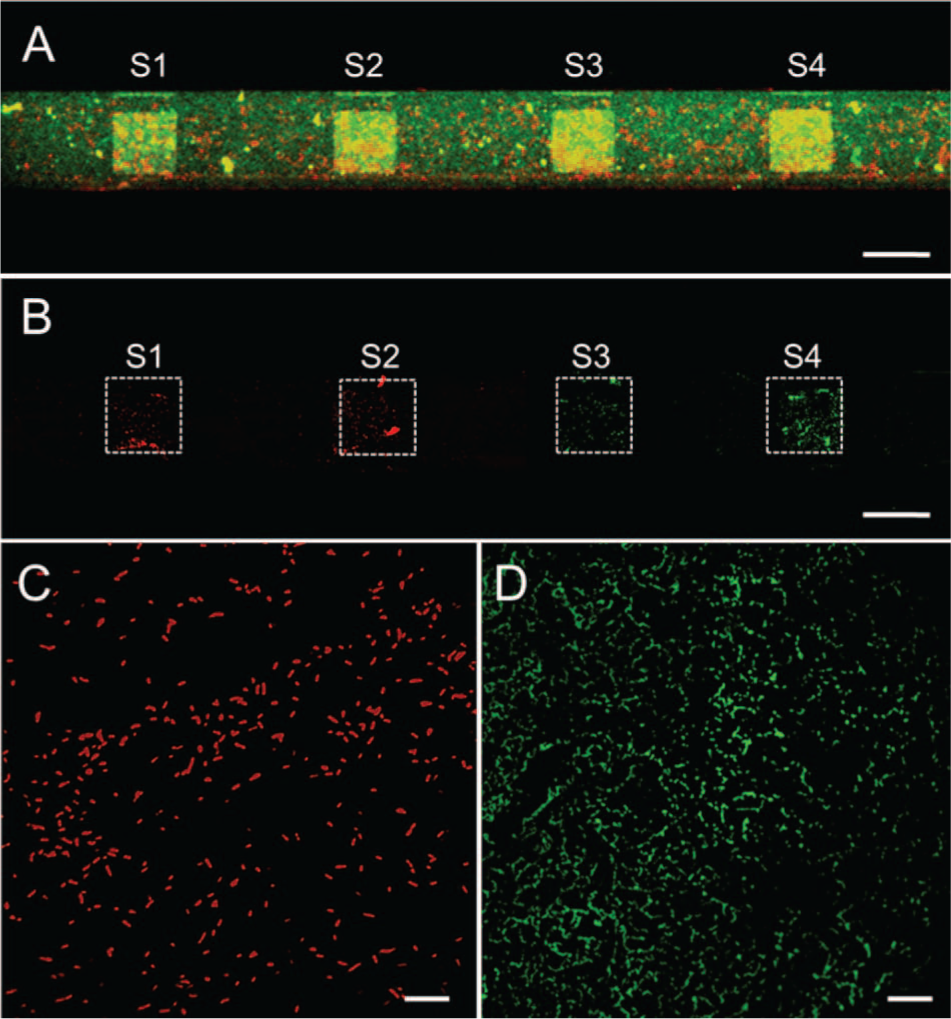
Fluorescent images showing the selectivity of antimicrobial peptides (AMPs). Sensors S1 and S2 are coated with G10KHc (P. aeruginosa AMP), and sensors S3 and S4 are coated with C16G2cys (S. mutans AMP). (
Characterization of the Impedance Sensor
Impedance measurements of NaCl solutions with increasing molar concentrations from 0.001 M to 1.0 M were performed to characterize the response of our platform. As shown in Figure 5 , different NaCl concentration solutions generated distinctive impedance spectra. This response is characteristic of electrical impedance spectroscopy measurements and can be described by an equivalent circuit analysis for cell impedance measurements.39,40 In the lower frequency region (flower, 102 Hz to 3 kHz), the impedance drops with frequency, primarily because of the electric double layer (EDL) at the electrode surfaces. At the middle frequency region (fmiddle, 3 kHz to 2 MHz), the impedance response is constant and dominated by the electrolytic solution impedance. This response is reflected in our data, which show a clear correlation between the magnitude of the impedance (at fmiddle) and the solution concentration; that is, the shift in impedance is on a similar order of magnitude as the solution concentration. For example, at 0.001 M NaCl, the largest impedance is observed (~200 kΩ at fmiddle) because of its low ionic concentration and electrical conductivity. Solutions of higher NaCl concentrations produce successively lower impedance responses owing to their higher conductivities, which facilitates the flow of current between the electrodes. At the upper frequency region (fupper, 2 MHz to 100 MHz), the solution conductivity has negligible impact on the impedance response. Rather, the impedance decreases with frequency and is primarily dominated by capacitive effects of the electrodes.
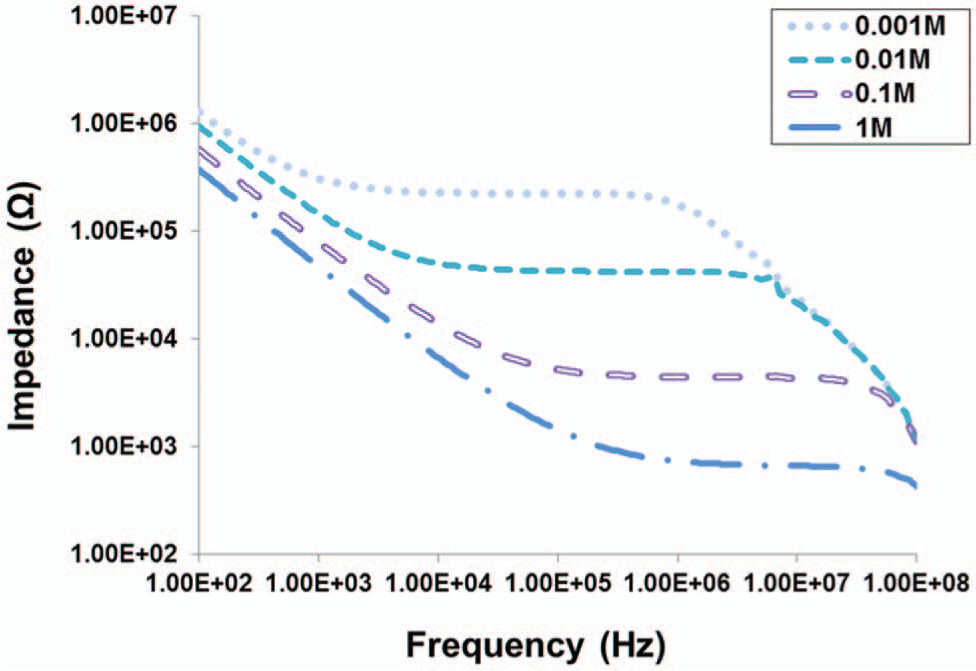
Impedance spectra of NaCl solutions with concentrations ranging from 0.001 M to 1 M. Measurements were performed immediately after loading the solutions using peptide-coated sensors.
Cell Impedance Measurements
Impedance measurements were performed to detect increasing concentrations of S. mutans and P. aeruginosa in PBS using the AMP biosensor. Figure 6A shows the impedance spectra following a 20 min incubation of AMP-coated sensors with 104 cfu/mL to 107 cfu/mL of S. mutans. Measurements were also performed on non–AMP-coated sensors as a control, which exhibits negligible cell binding. Bacterial cell concentration directly correlates with the impedance response due to the cells’ inherent electrical properties (e.g., membrane capacitance, cytoplasmic resistance). 41 The difference in impedance is most prominent at flower and diminishes at higher frequencies. Therefore, we focus on the impedance values at 102 Hz in our data analyses. Impedance measurements at 102 Hz of bound S. mutans and P. aeruginosa are plotted in Figure 6B . The impedances of P. aeruginosa are comparable to those of S. mutans, which also correlate with the cell concentration. P. aeruginosa produce slightly higher (4%–10%) impedance values compared with S. mutans at all concentrations tested. It is likely that other bacterial cells captured with other AMPs will produce similar impedance values at these cell concentrations. By comparing measurements from S. mutans and P. aeruginosa with the non-AMP, nonbacterial control, our biosensor exhibits a lower limit of detection of 105 cfu/mL for both bacteria, which is comparable to previously reported antibody- 14 and AMP-based 23 bacterial biosensors. Further optimization of the biosensor chip will improve detection sensitivity. For example, modifying the sensor geometry may allow for a lower concentration of cells to be detected. Also, the sensitivity of the sensor could be enhanced by employing conductive nanoparticles to facilitate signal transduction.
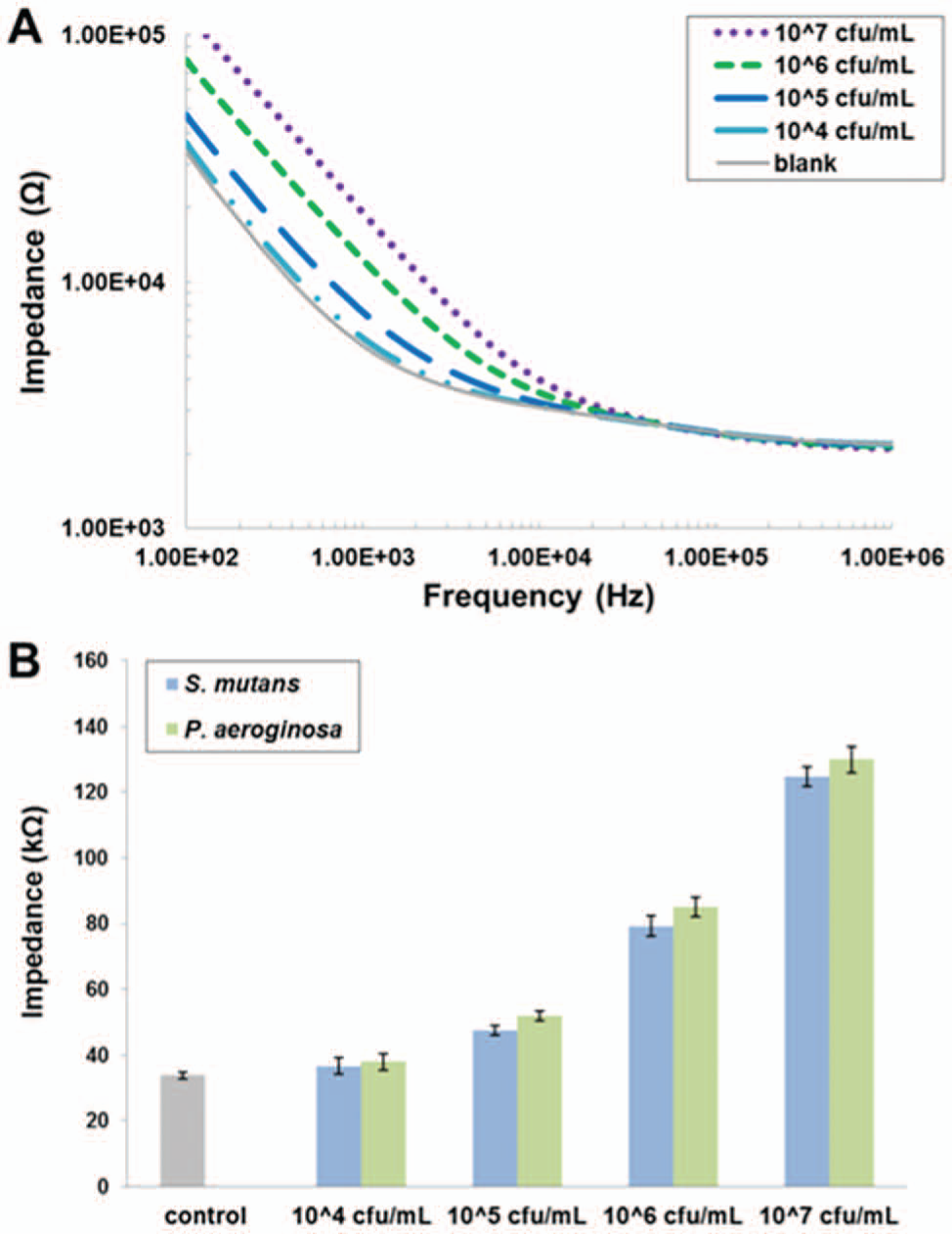
(
Impedance measurements were performed using S. mutans in three different solutions to evaluate the effect of solution conductivity on the response of the AMP biosensor. The impedance spectra of S. mutans at 107 cfu/mL in PBS (38 mS/cm), LCB (0.28 mS/cm), and DI water (0.1 µS/cm) are plotted in Figure 7 . Each data set includes a control measurement performed on sensors without AMP coating. As in Figure 5 , PBS, with the highest conductivity exhibits the lowest impedance, followed by LCB and DI water. We also see that the shift in impedance due to cell binding is most prominent at flower. S. mutans in PBS resulted in the largest increase in impedance compared with the control (270% at 102 Hz), whereas the impedance increases for LCB and DI water were 100% and 120%, respectively. Further, the impedance shift spans a wider bandwidth in LCB and DI water compared with PBS. This can be explained using the equivalent circuit analysis for cell impedance measurements described above. At flower, system impedance is dominated by the EDL at the electrode surfaces. PBS contains a relatively large concentration of ions, which enables ample current flow between the electrodes, thereby minimizing the impedance effects of the bound cells. In contrast, LCB and DI water have much lower ionic concentrations, and bound cells have a more substantial effect on the overall impedance response relative to the respective control. Based on these results, we conclude that measurements in higher conductivity solutions (PBS) are preferable in order to enhance impedance response, but measurements in low-conductivity media may be preferred if impedance responses over a broader frequency bandwidth are desired.
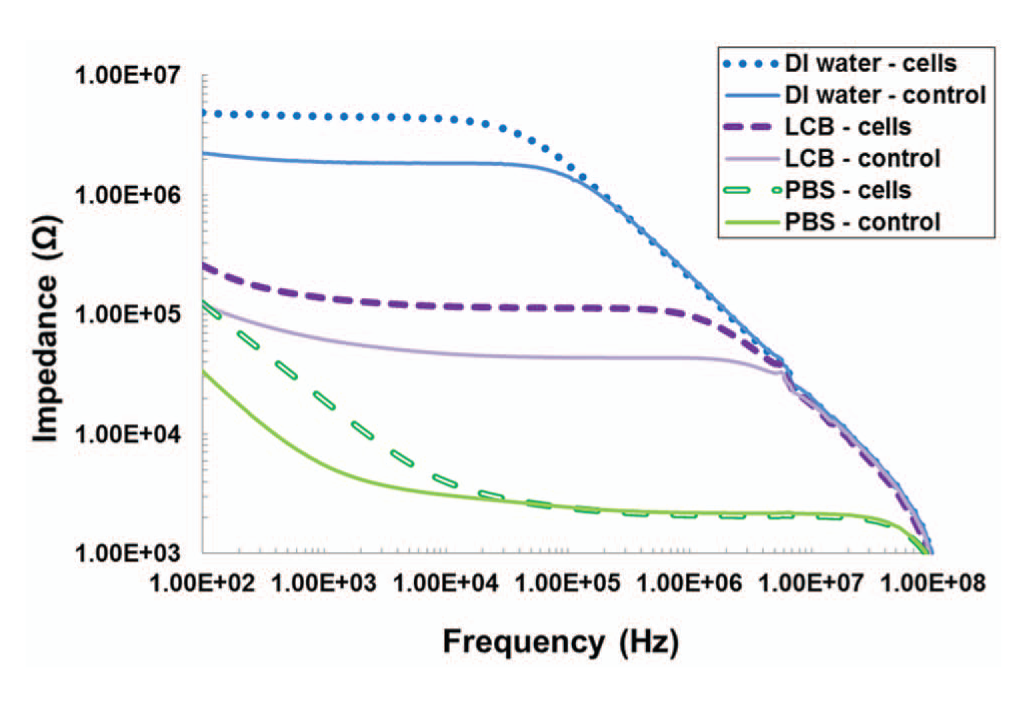
Impedance spectra of S. mutans at 107 cfu/mL in phosphate-buffered saline, low-conductivity buffer, and deionized water. Controls are measured from non–antimicrobial peptide–coated sensors with no bound cells.
Conclusions
We have presented a microfluidic biosensor for rapid and multiplexed detection of bacterial pathogens. This platform uses synthetic AMPs, which are designed with highly specific targeting and binding capabilities. AMPs were immobilized onto a microsensor array via thiol-gold binding, which resulted in vertical binding orientation and robust surface attachment. Assays characterizing peptide-bacterial cell binding were performed using fluorescently labeled S. mutans and P. aeruginosa cells. Based on these experiments, AMP-coated sensors demonstrated strong preferential binding to their corresponding targeted cells with negligible cross-binding. An electrical impedance sensing scheme is used for cell detection, in which S. mutans and P. aeruginosa cells at various concentrations can be distinguished based on impedance measurements, with a lower limit of detection of 105 cfu/mL. Each measurement requires only 5 s, and the entire detection process is completed within 25 min. Implementation of additional sensors will allow the detection of other pathogenic bacteria using their corresponding AMPs, enhancing the versatility and point-of-care diagnostic capabilities of this biosensor. With further development, we feel that this platform will become an enabling technology for further advancing microfluidic-based diagnostic tools to aid in the prevention of future outbreaks of infectious diseases.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the following agencies: NASA National Space Biomedical Research Institute (grant NCC 9-58-317) and UCLA T32 Dentist-Scientist & Oral Health Scientist Training Program (grant DE 007296).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
