Abstract
Rapid bacterial identification (ID) and antibiotic susceptibility testing (AST) are in great demand due to the rise of drug-resistant bacteria. Conventional culture-based AST methods suffer from a long turnaround time. By necessity, physicians often have to treat patients empirically with antibiotics, which has led to an inappropriate use of antibiotics, an elevated mortality rate and healthcare costs, and antibiotic resistance. Recent advances in miniaturization and automation provide promising solutions for rapid bacterial ID/AST profiling, which will potentially make a significant impact in the clinical management of infectious diseases and antibiotic stewardship in the coming years. In this review, we summarize and analyze representative emerging micro- and nanotechnologies, as well as automated systems for bacterial ID/AST, including both phenotypic (e.g., microfluidic-based bacterial culture, and digital imaging of single cells) and molecular (e.g., multiplex PCR, hybridization probes, nanoparticles, synthetic biology tools, mass spectrometry, and sequencing technologies) methods. We also discuss representative point-of-care (POC) systems that integrate sample processing, fluid handling, and detection for rapid bacterial ID/AST. Finally, we highlight major remaining challenges and discuss potential future endeavors toward improving clinical outcomes with rapid bacterial ID/AST technologies.
Introduction
The rise of antibiotic-resistant bacteria, including Gram-positive methicillin-resistant Staphylococcus aureus (MRSA) and vancomycin-resistant enterococci (VRE), as well as Gram-negative extended spectrum β-lactamase (ESBL)–producing Enterobacteriaceae and carbapenem-resistant Enterobacteriaceae (CRE), presents a major health threat in the United States and worldwide.1–4 According to the Centers for Disease Control and Prevention (CDC), more than 2 million people are infected annually with antibiotic-resistant infections, with >23,000 deaths in the United States alone. 1 Aggressive bacterial infections associated with antibiotic resistance are often managed within intensive care units (ICUs) with high associated costs, which impose significant healthcare, economic, and social burdens. The Alliance for the Prudent Use of Antibiotics (APUA) estimates that antibiotic-resistant infections cost the U.S. healthcare system more than $20 billion each year. 5
The development and use of rapid tests for the identification (ID) of resistant bacteria has been identified as one of the priorities to combat antibiotic resistance.1–4 Unfortunately, existing bacterial detection methods are limited in their inability to rapidly detect and identify pathogens that typically occur at low concentrations in biological samples. For instance, to rapidly detect <1 to 100 colony-forming units (CFU)/mL in blood, which are commonly found in adult bloodstream infections (BSIs), 6 remains a major global unmet need. Conventional bacterial cultures (18–24 h), which are used to grow bacteria, coupled with susceptibility testing (6–24 h), require several days to obtain a result. In many cases, a several-hour subculture step is further required to isolate and enrich bacterial strains prior to antibiotic susceptibility testing (AST) profiling, although several technologies for AST testing directly from positive blood cultures have recently been demonstrated.7–9 Therefore, the lack of rapid diagnostics in the current paradigm of clinical microbiology has resulted in either use of unnecessarily broad empiric antibiotics or a delay of several days in administering the appropriate antibiotics. Inappropriate initial antimicrobial therapy is associated with significantly increased mortality, particularly for BSIs.10,11 For instance, Kumar et al. reported that inappropriate initial antimicrobial therapy for septic shock occurs in about 20% of patients and is associated with a fivefold reduction in survival. 12 Likewise, several studies have demonstrated that reduced time for bacterial ID/AST using rapid tests is directly correlated with improved mortality and reduced healthcare cost.13–15 Therefore, rapid diagnostic tests would be exceedingly valuable in directing early therapy, improving clinical outcome, and enabling better antibiotic stewardship.15–25
Emerging automated rapid microbiology methods, especially those employing miniaturized microfluidic devices (or lab-on-a-chip systems) and nanotechnologies, offer unique opportunities to combat the crisis of antibiotic resistance.23,26–29 These microsystems often operate in small confined volumes so that bacterial growth or biochemical reactions can be accelerated. These devices typically utilize small amounts of sample and reagents and do not need expensive equipment. They are also amenable for high-throughput, highly parallel, and single-cell analysis. They can be readily automated, enabling sample preparation, fluid handling, analysis, and detection in an integrated fashion. Due to their small size, they can potentially be incorporated into low-cost portable devices for bacterial ID/AST at the point of care (POC). Collectively, with these appealing features, including superior assay time, cost, and amenability at the POC, these emerging micro- and automated systems can serve as alternative or complementary tools to conventional systems for bacterial ID/AST to provide timely guidance to infectious disease management.
In this review, we start with a brief overview of basic bacterial ID/AST test principles and summarize conventional systems to provide the context of the challenges that new rapid methods need to address. We then focus on miniaturized microsystems that utilize phenotypic and/or molecular mechanisms for rapid bacterial ID/AST. We also cover several of the most recent automated AST technologies that do not necessarily belong to “microsystems” but will likely make an immediate impact in the field. We touch on multiplexing, automation, and system integration, including combining both phenotypic and molecular tests, as well as POC systems. These example technologies are summarized in Table 1 . We also attempt to provide critical analyses of these technologies with regard to their pros and cons throughout the review and summarize the remaining challenges for future development.
Selected Examples of Emerging Microbiology Methods for Rapid AST.
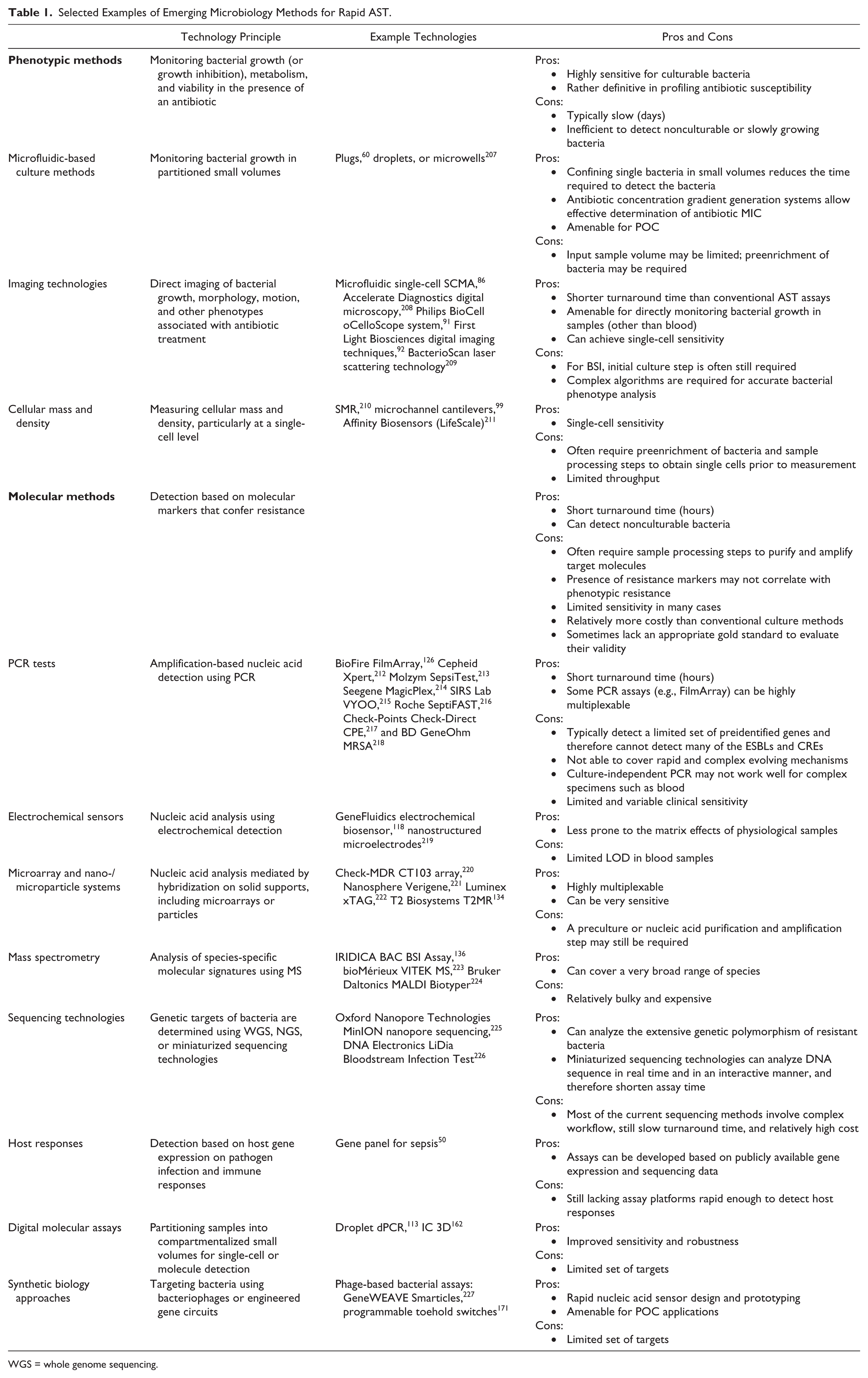
WGS = whole genome sequencing.
Bacterial ID/AST Test Principles
Antibiotic susceptibility can be identified using phenotypic assays by monitoring bacterial growth (or growth inhibition), metabolism, and viability in the presence of an antibiotic, or using molecular assays to identify biomarkers that confer resistance.20,30,31 Phenotypic methods are by far the most commonly employed and include broth microdilution, antibiotic gradient methods (e.g., E-test), and disc diffusion assays. 32 These methods can categorize bacterial strains into resistant or susceptible groups and provide minimum inhibitory concentration (MIC) to describe the efficacy of the antibiotics against a particular infection. Numerous commercial automated systems (e.g., BD Phoenix [Franklin Lakes, NJ] and bioMérieux VITEK [Shanghai, China]) are now being widely utilized in clinical microbiology laboratories. Phenotypic growth-based technologies rely on the measurement of biochemical or physical parameters that reflect the growth of microorganisms, including CO2 production (by fluorometric or pH sensors), turbidity, color,33,34 fluorescence or impedance changes on microbial consumption of biochemical and carbohydrate substrates in the media, bacterial autofluorescence, 35 heat production 36 or consumption, and culture container pressure changes due to microbial respiration. In addition, bacteria can be detected by viability staining and detection of metabolic activity (e.g., SYTO9/propidium iodide and resazurin) using flow cytometry, solid-phase fluorescence imaging (typically for bacteria collected on a membrane after sample filtration), electrochemistry, 37 and ATP bioluminescence assays. 38 Phenotypic assays for bacterial ID/AST can be highly sensitive (1 CFU per sample) for culturable bacteria and arguably more definitive than molecular methods in profiling antibiotic susceptibility as they directly characterize bacterial growth in the presence of antibiotics. However, because these in vitro phenotypic assays rely on the growth of bacteria in the presence or absence of antibiotics, they are slow (days) and are not capable of or efficient in detecting nonculturable or slowly growing bacteria, and can therefore lead to false negatives. In addition, they do not take into consideration any variables of the host response.
Molecular tests for bacterial ID/AST rely on the ID of biomarkers indicative of bacterial genera, species, and strains or antibiotic resistance, including genetic materials, proteins, enzymes, and metabolites. 39 For instance, conserved 16S and 23S ribosomal DNA (rDNA) or RNA (rRNA) regions have been used as pan markers for broad-spectrum bacterial detection.40,41 Repetitive DNA called VNTR (variable number tandem repeat) and 16S-23S intergenic spacers that represent sources of genetic polymorphisms and variability can also be used for bacterial genotyping. Precursor rRNA (pre-rRNA) is an intermediate stage in the formation of mature rRNA and has been used as a marker for bacterial metabolism, viability, and growth rate. 42 Antibiotic resistance genes (e.g., mecA gene for MRSA; vanA/B genes for VRE; TEM, SHV, OXA, and CTX-M gene families for ESBLs;43,44 and KPC, NDM, OXA-48, VIM, and IMP genes for CREs 45 ), as well as antibiotic-responsive mRNA transcripts, 46 have been widely used as markers for rapid AST. Compared with DNA targets, bacterial RNA markers are more abundant in the cell, and can potentially be indicative of viability to differentiate live and dead bacteria, but are more prone to degradation. Furthermore, host responses, including gene expression, on pathogen infection and immune responses47–49 have also been exploited as alternative approaches for rapid bacterial detection. For instance, recent analyses of the publicly available gene expression and sequencing data have led to the discovery of a robust set of genes for distinguishing patients with sepsis from patients with sterile inflammation. 50
A large number of platforms have been developed for detecting genetic markers, including (1) amplification-based assays, such as PCR, reverse transcription–PCR (RT-PCR), ligase chain reaction (LCR), transcription-mediated amplification (TMA), 51 recombinase polymerase amplification (RPA), rolling circle amplification (RCA), 52 nucleic acid sequence-based amplification (NASBA), and loop-mediated isothermal amplification (LAMP); 53 (2) non-amplification-based assays, such as fluorescence in situ hybridization (FISH), molecular beacons, 54 locked nucleic acid (LNA), 55 and peptide nucleic acid (PNA) probes;56,57 (3) electrochemical methods; (4) microarray-, microbead-, and nanoparticle-based hybridization assays; (5) mass spectrometry (MS); and (6) sequencing technologies. In addition to the widely used genetic markers, protein-, enzyme-, and metabolite-based molecular signatures can also be used for bacterial ID/AST using techniques such as immunoassays, MS, and Raman and infrared spectroscopy.17,21
The major advantage of molecular-based approaches is that they could significantly reduce turnaround times and rapidly provide antibiotic resistance information. However, a major limitation of molecular tests is that the presence of resistance markers may not always correlate with phenotypic resistance, especially for ESBLs and CREs. 58 In addition, they are unable to detect uncharacterized or unknown resistance mechanisms and can miss new resistance markers as bacteria quickly evolve. Furthermore, molecular-based approaches also have technological hurdles, such as upstream sample processing, that make them arguably more difficult to automate than the well-established phenotypic-based culture methods. Finally, it is important to note that phenotypic and molecular assays are often utilized in combination with emerging rapid bacterial ID/AST assays, 59 and in fact, the majority of the above-mentioned molecular methods still require a preculture step due to the low numbers of target pathogens that exist in complex biological samples.
Emerging Microtechnologies and Automated Systems for Phenotypic-Based Bacterial ID/AST
A number of microsystems have recently been demonstrated to measure phenotypic characteristics of bacteria, including growth, viability, morphology, and metabolism. In particular, confining bacteria in small, discrete volumes, especially at the single-cell level, in microfluidic devices can potentially accelerate biochemical reactions and bacterial growth, making the bacterial marker concentration in the isolated environment reach the detectable level much quicker. These systems therefore represent appealing alternatives to the conventional phenotypic assays due to their reduced sample-to-answer time, simplicity, portability, and single-cell analysis capability.
Microfluidic Device-Based Culture Methods for Bacterial ID/AST
Ismagilov and coworkers reported a plug-based microfluidics culturing method for rapid detection and drug susceptibility screening ( Fig. 1 ). 60 This method separates individual bacteria using stochastic confinement into nanoliter volume droplet plugs where bacterial growth and variability are measured using resazurin as an indicator. They demonstrated that confining single bacteria in nanoliter plugs enables eliminating the preincubation step and reducing the time required to detect the bacteria (a bacterium can be detected in a 1 nL plug in 2 h). They further demonstrated that a combination of stochastic confinement with a microfluidic hybrid method could screen many antibiotics in a single experiment to identify MIC in approximately 7 h. This method could also be used to distinguish between sensitive and resistant strains of S. aureus in complex biological matrices, including human blood plasma. In another study from Ismagilov’s group, they demonstrated that the plug-based microfluidic system is capable of integrating single-bacterium encapsulation and culture enrichment, then splitting the plugs into arrays of identical daughter plugs, and finally characterizing and analyzing each array using independent techniques, such as cellulase assays, cultivation, cryopreservation, Gram staining, and FISH. 61 In a separate study, Kelley and coworkers reported an electrochemical approach to obtain a rapid AST profile within 1 h. 37 In their assay, bacteria in urine samples are captured and then cultured in miniaturized wells with antibiotics. Resistance profiles are determined based on the electrochemical reduction of a redox-active molecule resazurin. This approach can detect bacteria at concentrations of 1 CFU/μL or above and exhibits similar MIC profiles for the antibiotics they tested when compared with conventional culture-based methods. Different versions of microfluidic-based culturing methods have been demonstrated for rapid bacteria ID and/or AST,26,62–78 including those for polymicrobial cultures. 79 In addition, microfluidic-based systems that combine bacterial culture and antibiotic concentration gradient generation represent a particularly powerful approach to quantitatively investigate the antibacterial effect of antibiotics to obtain MIC.21,80–83 Furthermore, AST assays have been demonstrated using paper-based portable culture devices that might find utility in low-resource environments.84,85
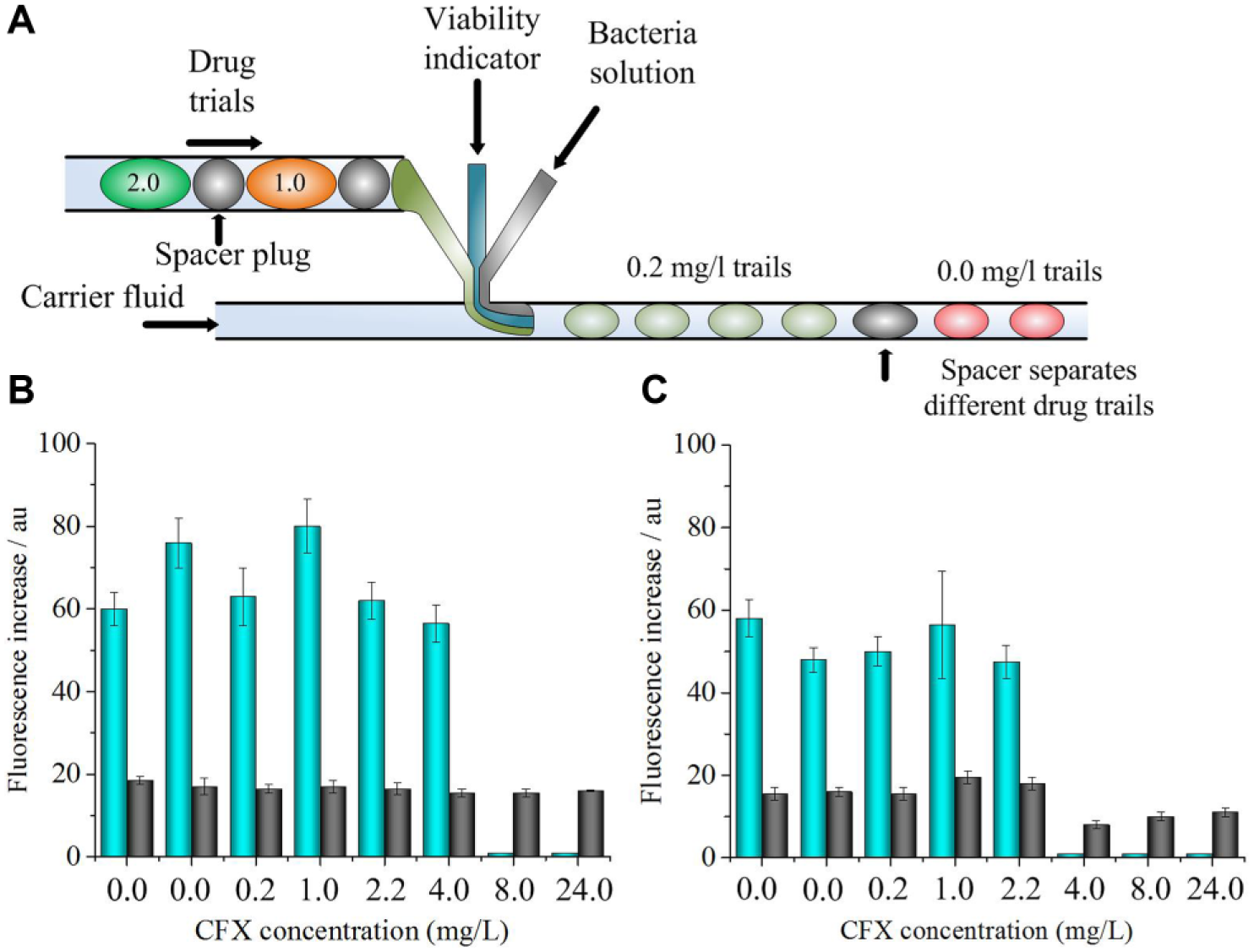
(
Imaging Technologies for Bacterial ID/AST
Direct imaging of bacterial growth, morphology, motion, and other phenotypes associated with antibiotic treatment, particularly at a single-cell level, represents an emerging tool for rapid bacterial ID/AST. For instance, Kwon and colleagues developed a rapid AST method based on the single-cell morphological analysis (SCMA) ( Fig. 2 ). 86 SCMA profiles antibiotic susceptibility by automatically analyzing and categorizing morphological changes of single bacterial cells in the presence of antibiotics. Specifically, they utilized a microfluidic agarose chip 87 to immobilize bacteria upon gel solidification so that bacteria can be readily imaged. The microfluidic agarose chip is also composed of channels containing antibiotics and nutrients that can diffuse into the agarose through openings between the channels and wells ( Fig. 2 ). They further integrated the microfluidic chips with a 96-well platform for high-throughput analysis. Time-lapse bright-field imaging of single cells was then performed. Automated image processing and data interpretation were used to profile the response of bacteria to antimicrobial agents based on different morphological patterns, including dividing, filamentary formation, and swelling. Using this method, they tested four different standard strains from the Clinical Laboratory Standard Institute (CLSI) and 189 clinical samples, including ESBLs, MRSA, and VRE. The SCMA method demonstrated 91.5% categorical agreement and 6.51% minor, 2.56% major, and 1.49% very major discrepancies when they compared their results with the standard broth microdilution test. The SCMA method can obtain AST results in only 3–4 h, although the current system still requires a preculture step.
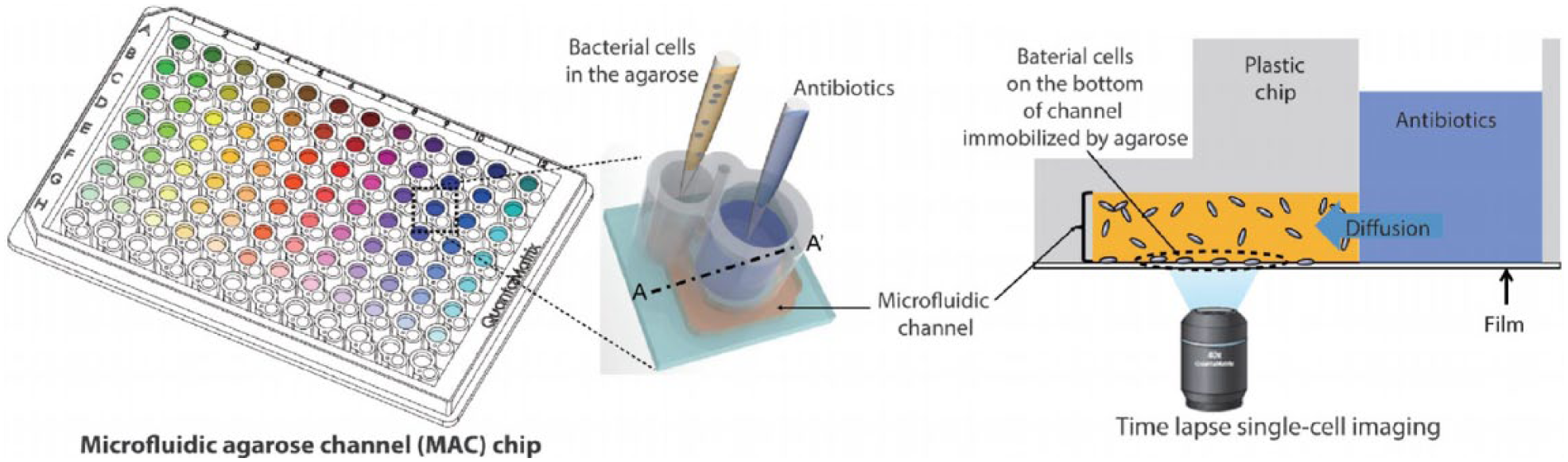
Schematic of an SCMA platform. The microwells are integrated on a 96-well plate. Bacteria are immobilized in agarose cultures. Antibiotics and nutrients are diffused to the agarose culture. The bacteria morphology under antibiotics could be monitored by an imaging system under the plate. 86 (This figure is modified from reference 86 with permission.)
In another example, Accelerate Diagnostics (Tucson, AZ)88–90 is developing an automated digital microscopy system for rapid bacterial ID/AST, for which they have recently received Food and Drug Administration (FDA) marketing authorization. In their system, bacterial inoculum from positive blood culture is pipetted into the independent fluidic channel of a custom disposable multichannel fluidic cassette where bacteria are sequentially immobilized onto the transparent flowcell channel using an electrokinetic concentration. For bacteria ID, immobilized cells were identified using in situ hybridization of fluorescently labeled oligonucleotide probes that can be universal for broad-range bacteria or specific for different species (up to 19 targets covering pathogens responsible for 85%–90% of BSI cases). For AST, time-lapse imaging and analysis of individual bacteria in response to antibiotics over time are performed. Polymicrobial AST and MIC determination can be obtained by interpretation of cell morphology, division rates, and growth patterns, and mass changes. According to Accelerate Diagnostics, bacterial ID and AST using the Accelerate system following positive blood culture can be obtained in 1 h with 95% overall agreement compared with the VITEK 2 system and 5 h with 91% agreement against broth microdilution, respectively. In addition, Philips BioCell (Amsterdam, Netherlands) has developed the oCelloScope system, which scans growing bacteria using digital time-lapse microscopy. 91 First Light Biosciences, Inc. is also developing digital imaging techniques for rapid bacterial ID/AST. 92 BacterioScan (Spectral Platforms, St. Louis, MO) exploits laser scattering technology to rapidly differentiate growth versus no growth of bacteria in clinical samples, which allows them to determine both the presence or absence of pathogenic microorganisms in a test sample and the concentration of microorganisms for antimicrobial susceptibility profiling. Several other technologies for bacterial ID/AST that are based on optical imaging and algorithm analysis of bacterial phenotypes in response to antimicrobial agents or other stresses are currently being developed.93–96
Measuring Cellular Mass and Density for Bacterial ID/AST
Manalis and colleagues demonstrated a suspended microchannel resonator (SMR) integrated with picoliter-scale microfluidic control to measure buoyant mass and growth rates of individual bacterial cells. 97 Using this technique, they can monitor cellular density changes during osmotic shock recovery, a phenomenon that allows them to differentiate between antibiotic-resistant and antibiotic-susceptible bacteria. 98 Etayash et al. applied a microchannel cantilever with its internal surface functionalized with bacteria binding molecules (e.g., antimicrobial antibodies or peptides) to capture target bacteria. By measuring the cantilever deflection, nanomechanical infrared spectra, and resonance frequency shift for mass adsorption, they were able to selectively identify single bacteria with a sensitivity of one cell per microliter and measure their responses to antibiotics. 99 Longo et al. also demonstrated that the fluctuations of highly sensitive atomic force microscope cantilevers could be applied to detect low concentrations of bacteria, analyze their metabolism, and quantitatively monitor their response to antimicrobial agents within minutes. 100 LifeScale (Santa Barbara, CA) is currently developing an automated instrument that can rapidly determine microbial growth and response to stressors by simultaneously measuring both concentration and microbe mass for rapid bacterial ID/AST. 101 In general, bacterial ID/AST assays based on cellular mass and density measurement require preenrichment of bacteria and often sample processing steps to obtain single cells prior to measurement.
Bead Rotation and Motion-Based Bacterial ID/AST Techniques
Kopelman’s laboratory has developed asynchronous magnetic bead rotation (AMBR) biosensor systems102,103 for monitoring the growth and drug susceptibility of individual bacteria. Magnetic beads assume a specific rotational spin when brought into a revolving magnetic field, which can be influenced by the binding of bacteria. Using this phenomenon, they demonstrated that the AMBR biosensor can be used to sensitively monitor individual bacterial cell growth dynamics, including cell elongation, generation time, lag time, and division, as well as their sensitivity to antibiotics in a rapid fashion. They further demonstrated that the AMBR biosensors could operate in microfluidic droplets, which potentially enables highly parallel and long-duration experiments. 104 When Escherichia coli was exposed to various concentrations of gentamicin in droplets, a 52% change in the sensor rotational period was observed within 15 min, thus enabling rapid AST measurements. They further utilized their AMBR biosensor as a viscometry method to rapidly measure bacterial growth and drug sensitivity by monitoring changes in the suspension’s viscosity as bacteria proliferated in droplets. 104 For another example, Chuang and colleagues developed a rapid bead-based AST method using optical diffusometry. 105 In their study, the diffusivity (or Brownian motion) of bacterium–particle complexes was used as a sensitive indicator for bacterium–particle binding, as well as antibiotic sensitivity. By analyzing the temporal diffusivity change of particles attached to bacteria, an AST assessment of the response of single Pseudomonas aeruginosa to gentamicin was obtained within 2 h.
Emerging Microtechnologies and Automated Systems for Molecular-Based Bacterial ID/AST
Molecular tests for bacterial ID/AST utilize molecular markers that are indicative of the presence of bacteria and antibiotic resistance. The vast majority of molecular tests in this area use nucleic acid markers, which we focus on in the next several sections, with the detection platforms including PCR, electrochemical methods, microarrays, micro- and nanoparticles, MS, and sequencing technologies. We also introduce synthetic biology-based approaches, as well as emerging platforms for non–nucleic acid markers, such as β-lactamases.
Molecular Bacterial ID/AST Testing Using Nucleic Acid Markers
PCR-Based Assays
Numerous PCR-based tests have been developed for bacterial ID/AST applications, although many of them still utilize samples from culture-enriched samples. 106 These assays typically detect a small set of preidentified nucleic acid targets, such as 16S or 23S rDNA (or rRNA), for broad-range bacteria, 107 species- or genus-specific targets, or resistance genes. For instance, BioFire (now a bioMérieux company) has recently introduced the FilmArray platform, a closed and fully automated system that combines DNA extraction from clinical samples, nested multiplex PCR, post-PCR melt curve analysis, and data interpretation. The FilmArray blood culture identification (BCID) panel can analyze a set of 24 Gram-positive, Gram-negative, and yeast pathogens and 3 antibiotic resistance genes (mecA, vanA/B, and KPC) associated with BSI. In a recent clinical evaluation, Altun et al. demonstrated that the FilmArray BCID panel identified microorganisms in 153/167 (91.6%) samples with monomicrobial growth. 108 When polymicrobial growth was analyzed, the FilmArray could detect all target microorganisms in 17/24 (71%) samples. Their study showed that the FilmArray is a rapid (65 min) ID method with overall robust performance in direct ID of bacteria and yeasts from positive blood culture bottles.
Several culture-independent PCR assays15,16,109,110 (e.g., Cepheid Xpert, Molzym SepsiTest, Seegene MagicPlex, SIRS Lab VYOO, Roche SeptiFAST, Check-Points Check-Direct CPE, and BD GeneOhm MRSA) have been developed for the detection of bacteria and/or antibiotic resistance directly from raw samples. In general, these assays have a short sample-to-answer turnaround time (1–8 h). Some of them have been adopted in clinical settings for less complex clinical samples (e.g., BD GeneOhm MRSA for nasal swab samples) or unculturable pathogens. However, most have not been widely used, particularly with whole blood clinical samples, because of their limited and variable clinical sensitivity (30%–90%),111,112 as well as a large discrepancy with conventional culture methods that makes interpretation difficult. Indeed, conventional PCR is typically not sufficiently sensitive and robust to detect low-abundance targets. Some of these drawbacks can be potentially addressed by the recent digital PCR (dPCR) systems by which extracted nucleic acids are partitioned into many individual reactions and quantified digitally (1 or 0). The dPCR format permits absolute quantitation of target DNA/RNA with improved precision and reproducibility without the need for a standard. For instance, Ismagilov’s group has recently employed dPCR to measure DNA replication of the target pathogen and demonstrated that their digital AST (dAST) can determine the susceptibility of clinical isolates from urinary tract infections (UTIs) after only 15 min of exposure to clinically relevant antibiotics. 113 Another general issue associated with PCR is contamination from nonpathogenic bacterial species (e.g., staphylococcal) introduced during the testing process, as well as from background bacterial nucleic acid materials that exist in PCR reagents (e.g., Taq polymerase) that are manufactured using bacterial sources. 114 Careful screening of vendors and using methods that remove or suppress contaminations are often required to achieve robust PCR performance, especially for detecting low-abundance targets. 115 Finally, PCR assays are typically designed to detect a limited set of preidentified genes, which are not able to cover rapid and complex evolving mechanisms associated with infectious bacteria. In particular, it remains a challenge for conventional PCR to detect many of the ESBLs and CREs that are highly variable and often differ from each other by single-nucleotide polymorphisms (SNPs).43–45
Electrochemical Methods
Electrochemical sensors have also been widely used for nucleic acid analysis. For instance, Liao et al. demonstrated a rapid (approximately 3.5 h) AST assay from clinical urine samples by direct culture of urine samples in the presence of antibiotics, followed by analyzing 16S rRNA levels using an electrochemical sensor.116–118 Clinical validation using patient urine samples demonstrated that this test was 94% accurate in 368 pathogen-antibiotic tests compared with standard microbiological methods. Together with GeneFluidics, Inc., the same team has been developing a multiplex electrochemical biosensor system for rapid pathogen ID in blood samples ( Fig. 3 ). 119 Their portable, multichannel potentiostat is integrated with a disposable, 16-sensor chip. The chip is fabricated by gold deposition on a plastic substrate, on which target bacterial rRNA can be detected amperometrically following sandwich binding by the capture probe and the detector probe. This electrochemical sensor is potentially less prone to the matrix effects of physiological samples and does not require nucleic acid amplification. They evaluated the system using spiking bacterial clinical isolates in whole blood and positive blood culture bottles. The reported system achieved a limit of detection (LOD) of 290 CFU/mL in culture media, which may be limited for directly detecting bacteria in blood specimens but could be useful for postculture samples. Furthermore, Kelley’s group has made a series of innovative advances in the use of electrochemical sensors for pathogen detection, including integrated electrical bacterial lysis,120,121 nanostructured microelectrodes to improve sensitivity, 122 solution circuit chip for multiplexed detection, 123 and PNA clamps for point mutation detection. 124
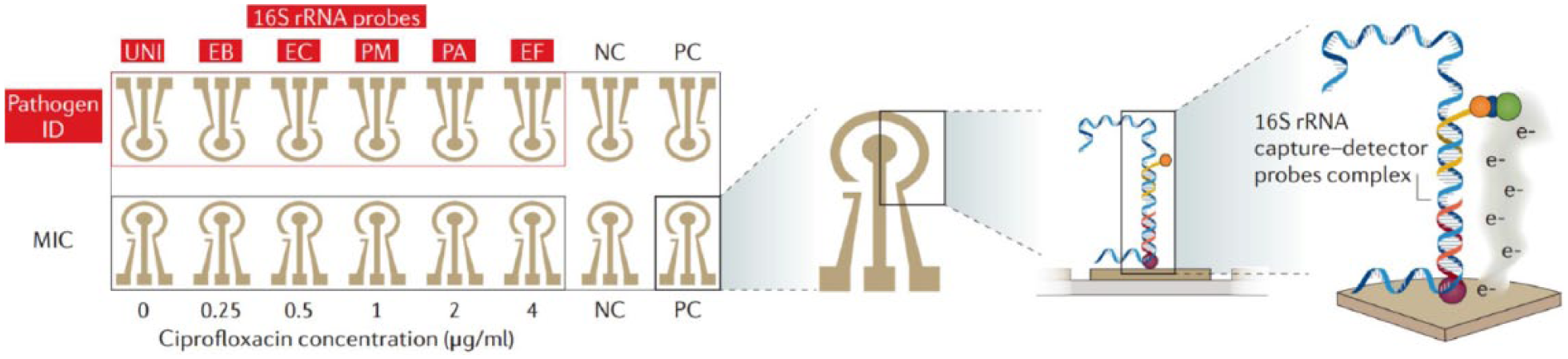
An electrochemical biosensor array consists of 16 sensors with DNA probes for the detection of different bacterial species. Sensors are integrated into a potentiostat. Every sensor is composed of a working electrode, a peripheral reference electrode, and an auxiliary electrode. The hybridization of probes and targets can be facilitated by electrokinetic heating and mixing.116,119 (This figure is modified from reference 116 with permission.)
Microarray and Nano-/Micro-Particle-Based Nucleic Acid Assays
A challenge in detecting Gram-negative ESBLs and CREs is that there are numerous distinct mechanisms of β-lactamase variants.43–45,125 PCR-based approaches, as described above, typically only detect a handful of targets, 18 with a few exceptions, including two-step nested PCR (e.g., the FilmArray system) and ligation-mediated real-time PCR. 126 By comparison, several nucleic acid detection platforms, including microarrays, nanoparticles, and microparticles, are particularly amenable for highly multiplexing and SNP analysis in a single assay (although a preculture or nucleic acid amplification step may still be required).18,21
In a recent study, for instance, Cuzon et al. evaluated the Check-MDR CT103 array (Check-Points, Wageningen, Netherlands) for the rapid detection of ESBLs, including TEM, SHV, and CTX-M; plasmid-mediated cephalosporinases (CMY-2-like, DHA, FOX, ACC-1, ACT/MIR, and CMY-1-like/MOX); and CREs (KPC, OXA-48, VIM, IMP, and NDM). 127 The Check-MDR CT system can simultaneously detect up to 100 specific resistance markers with single-nucleotide specificity. Briefly, following whole cell DNA extraction, a multiplex ligation detection reaction (LDR) was used to produce DNA molecules that are subsequently PCR amplified. The PCR products were next hybridized to a low-density DNA microarray system. Images were acquired using an array tube reader and interpreted with the software that automatically translates the data into the presence or absence of a specific target gene. A total of 187 Gram-negative bacilli isolates possessing different bla genes were tested in this study. 127 Specificities and sensitivities of 100% were recorded for most bla genes. For another example, Great Basin Scientific (Salt Lake City, UT) has developed a system where they combine isothermal helicase-dependent amplification and a DNA array on a silicon chip, which multiplexes up to 64 distinct targets in a single assay. In a recent clinical study for C. difficile detection using this system, 130 patient samples were tested and a clinical sensitivity of 97% and a specificity of 100% were achieved. 128
The Verigene system (Nanosphere, now part of Luminex, Austin, TX) currently offers automated, multiplex capabilities that detect both Gram-positive and Gram-negative pathogens, as well as a panel of drug resistance markers (mecA for meticillin; vanA and vanB for vancomycin; and CTX-M for the detection of ESBLs, IMP, KPC, NDM, OXA, and VIM for carbapenemases) from positive blood cultures. 129 The Verigene tests run on the Verigene Processor and Reader platforms, which extract and purify nucleic acids, followed by hybridization to specific oligonucleotide-labeled gold nanoparticles on a microarray. In a recent clinical study where 173 positive cultures were tested, Ward et al. reported that the Verigene assay can accurately identify target organisms that are featured on the Verigene panel (with occasional false-positive results [6/173]), and 27.95 h earlier than conventional methods. 129 Luminex also has barcode bead-based technology (Luminex xTAG) for highly multiplexed analysis of nucleic acid markers.
Magnetic nanoparticles (MNPs) have also been used for the rapid detection of pathogens. Weissleder and Lee and coworkers have reported a magneto-DNA nanoparticle system that is capable of rapid and specific profiling of pathogens in clinical samples. 130 In their procedure, nucleic acids were first extracted and PCR amplified. An amplified single-strand DNA product was then captured by beads conjugated with capture probes, before hybridizing with MNPs to form a magnetic sandwich complex. Samples were subsequently analyzed using a miniaturized nuclear magnetic resonance (NMR) device system. The use of a magnetic detection strategy allows near-background-free sensing, which could potentially simplify and speed up the assay. This approach permits both universal and specific detection of various clinically relevant bacterial species. The authors claimed they could achieve sensitivity down to single bacteria in clinical specimens within 2 h. 130 A similar approach has been applied to detect M. tuberculosis and their drug resistance strains from mechanically processed sputum samples. 131 The same group has also reported a microfluidic chip-based micro-Hall (μHall) platform for measuring single, magnetically tagged bacteria directly in clinical specimens. 132 In this approach, target bacteria are first labeled with MNPs using cycloaddition chemistry with a density of 104–106 MNPs per cell, which makes bacteria superparamagnetic. The sample then flows through a μHall sensor array microfluidic device, where hydrodynamic focusing is applied to confine bacteria in close proximity to the sensor surface for single-cell detection. The authors applied the μHall chip for enumerating Gram-positive bacteria and demonstrated a LOD of ∼10 bacteria with an assay time 50 times faster than that of conventional assays. T2 Biosystems (Lexington, MA) recently received FDA approval for their T2 magnetic resonance (T2MR) Candida test, which also employs MNPs ( Fig. 4 ). In their workflow, the T2Dx instrument automatically performs all steps after sample loading, including blood cell lysis and Candida cell concentration, Candida cell lysis, PCR amplification, DNA target hybridization to capture supermagnetic nanoparticles, and measurement of T2MR induced by agglomeration of supermagnetic particles. In recent clinical trials,133,134 T2MR demonstrated an overall specificity per assay of 99.4% with a mean time to negative result of 4.2 ± 0.9 h, and the overall sensitivity was 91.1% with a mean time of 4.4 ± 1.0 h for detection and species ID. The LOD was 1–5 CFU/mL depending on the Candida species. This technology represents a great advance in system automation that allows direct analysis of whole blood specimens to detect pathogens within hours of sample collection.
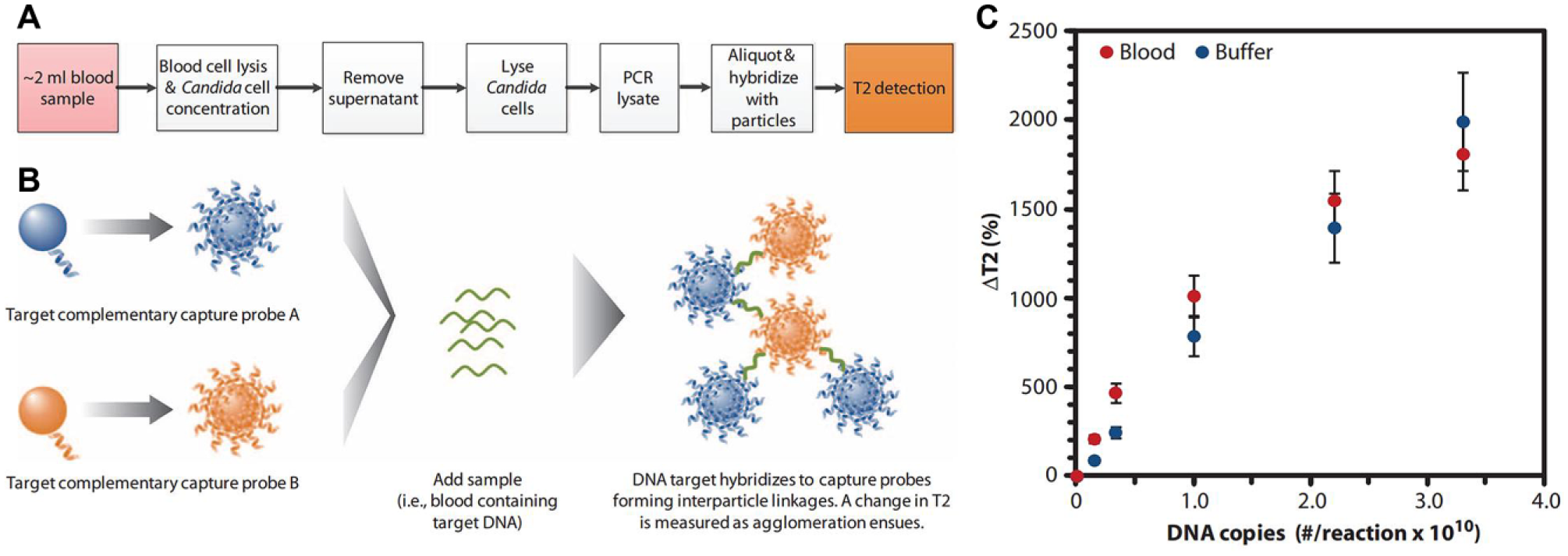
Detection of target pathogens from whole blood using MNP biosensors. (
Mass Spectrometry Methods
As we discussed above, conventional molecular methods such as PCR for the detection of microbial nucleic acids from a clinical specimen are limited in sensitivity and in the breadth of coverage. This remains an unmet need for technologies that are capable of identifying diverse pathogens directly from uncultured specimens, especially blood samples.22,25,135 Analysis of amplified microbial nucleic acids using MS may help to address this issue. For instance, the IRIDICA BAC BSI Assay (Ibis Biosciences, an Abbott Company, Carlsbad, CA) can identify hundreds of diverse organisms based on species-specific genetic signatures using electrospray ionization–MS (ESI-MS). Briefly, their automated system includes extracting DNA from larger volumes of whole blood (5 mL) and amplifying conserved bacterial and fungal genes (covering >95% of the eubacteria and Candida species associated with human infection), as well as antibiotic resistance markers (mecA, vanA, vanB, and blaKPC), using a mismatch- and background-tolerant PCR chemistry. An automated desalting and DNA debulking process is then performed to prepare amplicons for downstream ESI-MS. With their onboard analysis program, this method is capable of discriminating amplicon sequence variants on the basis of multilocus base composition signatures from different species. The IRIDICA assay can detect more than 780 bacterial and candidal species. The mean LOD for the assay is 39 CFU/mL, with a range of 0.25–128 CFU/mL, depending on the target species. 136 The method can provide organism IDs directly from uncultured blood in less than 8 h. Interestingly, in a recent study, 137 the IRIDICA BAC BSI Assay produced twice as many positive detections as culture across 285 clinical blood specimens from sepsis patients. This suggests that emerging molecular assays such as the IRIDICA BAC BSI Assay could identify clinically relevant pathogens that are difficult to grow in conventional culture. On the other hand, this discrepancy between conventional gold standard culture methods makes data interpretation difficult. Furthermore, the IRIDICA BAC BSI system is relatively bulky and expensive, and its market penetration is yet to be determined.
Sequencing Technologies
Bacterial genome sequencing represents another great tool that can address the extensive genetic polymorphism of resistant bacteria.138–140 A number of sequence-based methods, including especially next-generation sequencing (NGS), are now available to identify most bacterial species and resistance genes. In particular, great progress has been made on the technical feasibility of antimicrobial resistance prediction with whole bacterial genome sequencing. For example, Zhao et al. 141 sequenced the genomes of cultured Campylobacter coli and Campylobacter jejuni strains and compared the predicted resistance based on the detection of 18 resistance genes and 2 gene mutations with the phenotypic resistance to 9 antibiotics. The overall correlation between phenotypic and genotypic resistance is 99.2% with 1025 phenotypic results for 114 strains. Note that these sequencing techniques are often coupled with upstream PCR amplifications. For instance, the SepsiTest (Molzym, Bremen, Germany) incorporates automated nucleic acid extraction, broad-range PCR amplification, and downstream sequencing analysis for species ID. Note that most of the current sequencing methods involve complex workflow (e.g., library preparation) and quality control and suffer from interfering contamination, lack of a gold standard, still slow turnaround time, and relatively high cost. Some of the recent advances in the use of miniaturized sequencing systems and single-cell sequencing technologies, as exemplified below, can potentially enable sequencing as routine and practical microbial diagnostics.
MinION nanopore sequencing (Oxford Nanopore Technologies, Oxford, UK) has recently been applied for rapid bacterial ID/AST142–147 ( Fig. 5 ). The MinION is a miniaturized and portable device that measures electrical impedance as DNA passes through arrayed nanopores. It generates DNA sequence data in real time and in an interactive manner, which has great potential to significantly shorten the sample-to-result time. A recent study reported that MinION nanopore sequencing can identify bacterial species and strain information within 1 h of sequencing time, initial drug resistance profiles within 2 h, and a complete resistance profile within 12 h. 146 Note, however, that DNA extraction and library preparation can still take up to 5 h prior to sequencing. 146 For another example, DNA Electronics, Inc. (Carlsbad, CA) developed a label-free nucleic acid analysis technology using a semiconductor chip. In their system, nucleotides that are incorporated during DNA amplification or sequencing release hydrogen ions that can be detected as an electrical signal. 148 Their platform integrates sample preparation steps (e.g., bacterial enrichment, cell lysis, and DNA purification), on-chip amplification and genotyping to identify the bacterial species and strains, and sequencing to identify any antimicrobial resistance genes. The company claims that their LiDia Bloodstream Infection Test takes approximately 3 h to generate clinically actionable information, directly from an uncultured blood specimen. Furthermore, several droplet- or microwell-based single-cell sequencing technologies have been demonstrated, which can be useful to address the heterogeneity issue of a mixed microbial population.149–154 In particular, combining bacterial culture enrichment in small-volume compartments with downstream genetic analysis, including PCR and sequencing, represents a great approach to obtain both phenotypic and molecular information.61,72,155
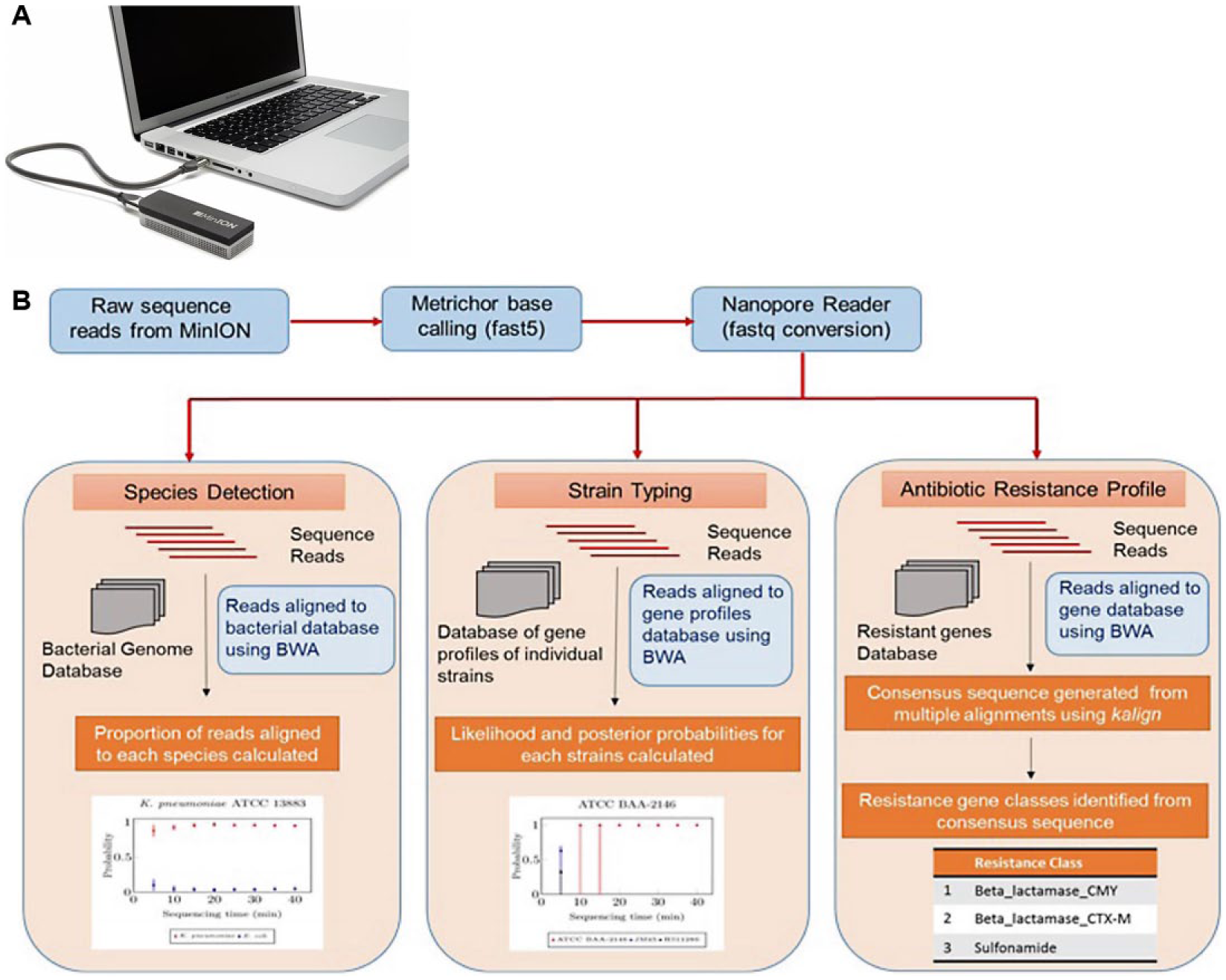
(
Molecular Bacterial ID/AST Testing Using Enzyme, Protein, or Metabolite Markers
Apart from nucleic acid markers, protein-, enzyme-, antigen-, and metabolite-based molecular signatures can also be used for bacterial ID/AST as analyzed by techniques such as MS, Raman and infrared spectroscopy, and immunoassays. For instance, Ingber’s group reported a broad-spectrum sepsis diagnostic using microbead-modified mannose binding lectin linked to the Fc portion of human IgG1 that detects pathogen-associated molecular patterns (PAMPs) in blood. 156 Automated matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) MS (e.g., bioMérieux VITEK MS and Bruker Daltonics MALDI Biotyper [Billerica, MA]) has recently been introduced to clinical microbiology labs for rapid microorganism ID based on distinct protein and peptide mass spectrum, compared with a reference database. MS can also be used for AST profiling by measuring antibiotic resistance markers such as β-lactamases that degrade antibiotics, and antibiotic degradation due to resistance enzymes. 31 β-Lactamases can also be detected by chemiluminescent or fluorescent substrates.157,158 For instance, the RAPIDEC CARBA NP test detects carbapenemase-producing bacteria based on the detection of hydrolysis of the β-lactam ring of imipenem, which leads to the color of a pH indicator changing.157,158 In addition, Rao’s laboratory has developed a series of fluorogenic sensor compounds for β-lactamases and carbapenemases.159,160 These chemical sensors can be integrated with droplet microfluidics for enumerating bacteria in samples. 161
We recently developed a technology called Integrated Comprehensive Droplet Digital Detection (IC 3D) that holds the potential to rapidly (1–3 h) and selectively detect bacteria directly from a large volume (milliliters) of unprocessed blood in a one-step, culture-free reaction. 162 The IC 3D system integrates real-time, bacterium-detecting fluorescence chemistries, droplet microfluidics, and a high-throughput particle counter system ( Fig. 6 ). In our first proof-of-principle study, fluorogenic DNAzyme sensors, 163 isolated by in vitro selection to specifically react with protein markers produced by target bacteria, are mixed with whole blood samples within a microfluidic channel, which is then encapsulated into tens of millions of individual picoliter droplets. DNAzyme sensors fluoresce instantaneously in the droplets that contain target bacterium, which can be counted by a high-throughput particle-counting system that can robustly and accurately detect single fluorescent particles from milliliter volumes within several minutes. Using E. coli as a target, we demonstrated that the IC 3D can selectively detect both stock isolates of E. coli and clinical isolates in spiked whole blood at single-cell sensitivity within 1–3 h. Moreover, the IC 3D can provide absolute quantification of target bacteria within a broad range of low concentrations with LOD in the single-digit regime. We are currently applying the IC 3D technology to target a broader panel of pathogens and to rapidly profile antibiotic resistance directly from blood samples.
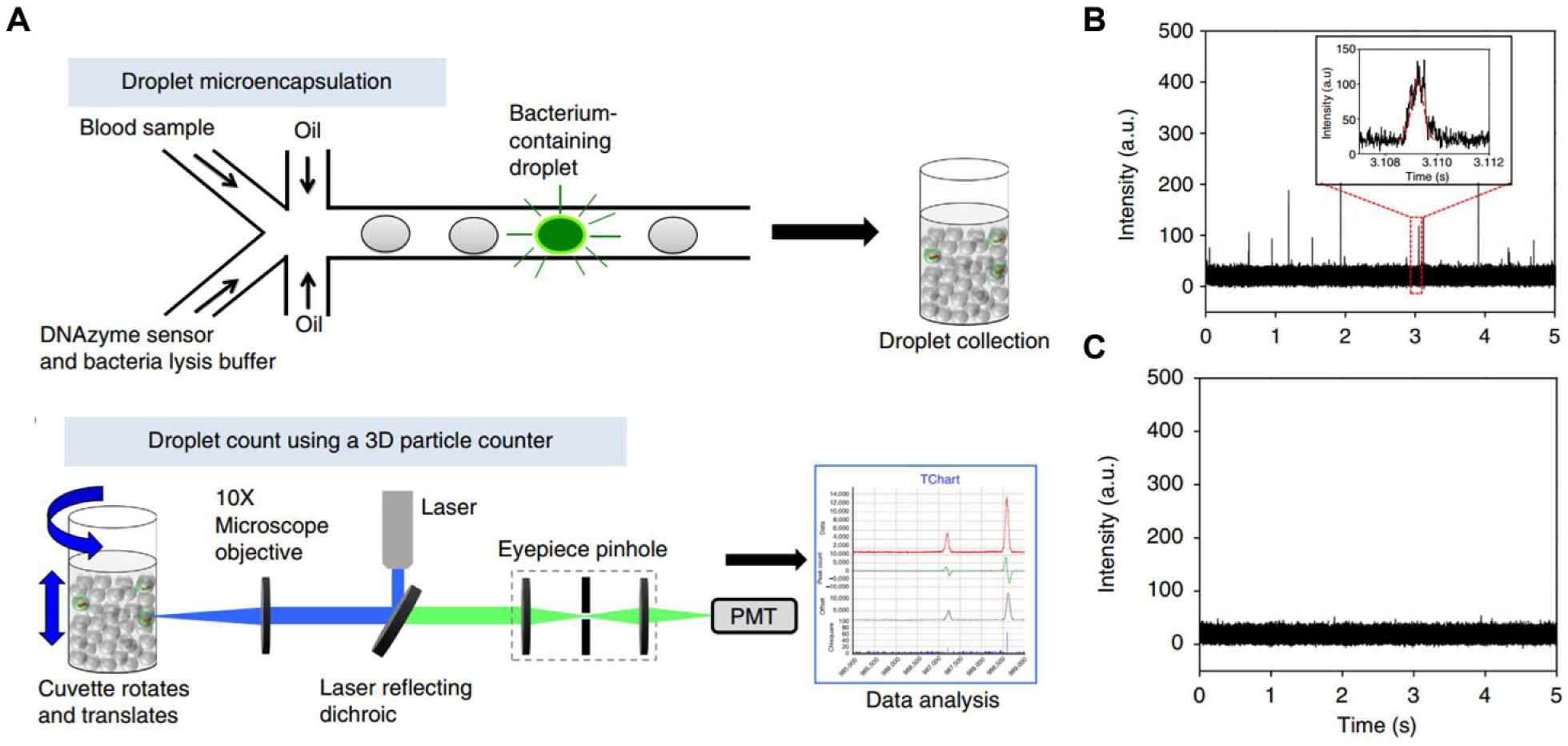
(
Molecular Bacterial ID/AST Testing Using Synthetic Biology Approaches
Synthetic biology approaches using bacteriophages or engineered gene circuits represent another emerging field that can aid the development of rapid bacterial ID/AST tests.164–169 For instance, due to their inherent selectivity to bacteria, ease of use, and cost-effective and straightforward production, phages have been extensively exploited for bacterial ID in the past few decades. Phage-based bacterial assays typically exploit events, including phage binding, amplification, reporter delivery, or lysis. The FDA has approved several phage-based tests, including those for Mycobacterium tuberculosis and S. aureus and their respective resistant strains. For example, MicroPhage’s KeyPath blood culture test for MRSA/MSSA (methicillin-susceptible S. aureus) utilizes phages to identify S. aureus. If the target bacteria are present, phages will be amplified and assayed by downstream phage-specific antibodies. To differentiate MRSA and MSSA, cefoxitin (CFX) is added in the assay where MRSA (but not MSSA) can grow and amplify phages to produce a positive readout. A recent study demonstrated that this KeyPath test produced 91.8% sensitivity and 98.3% specificity for the detection of S. aureus. 170 GeneWEAVE (recently acquired by Roche, Basel, Switzerland) has been developing gene-carrying particles called “Smarticles” that can bind and deliver genes to specific pathogens to produce light for detection. Synthetic biology methods offer shorter design-to-production cycles, as they can be rationally designed, rapidly tested, and deployed as POC diagnostics to tackle emerging pathogens. For instance, Yin and Collins’s team recently reported programmable toehold switches for RNA detection, 171 which can be integrated into a simple, inexpensive paper-based, cell-free system for POC applications. 172 Integrating this paper-based diagnostic with emerging genome-editing tools (e.g., CRISPR) offers further versatility for rapid nucleic acid sensor design and prototyping.173,174
Microtechnologies for Sample Processing, System Integration, and Automation in Bacterial ID/AST
The workflow in a conventional clinical microbiology laboratory is often manual and laborious and requires skilled personnel. Great advancements have been made in recent years toward clinical microbiology automation, with a part of or the entire process of specimen inoculation, processing, bacterial culture, detection, and analysis now being automated for different purposes (although the complexity of microbiology prevents a simple “one-size-fits-all” system). 18 For instance, platforms such as BD Phoenix, bioMérieux VITEK, and Siemens MicroScan have replaced manual methods for inoculation, reading, and analysis for bacterial ID. Total laboratory automation (TLA) has also been available, including BD Kiestra TLA, bioMérieux full microbiology laboratory automation (FMLA), and Copan WASPLab. Microtechnologies and microfluidics have much to offer for automation and system integration by miniaturizing processes such as pathogen capture, separation, and enrichment; cell lysis; nucleic acid extraction and amplification; and detection. Numerous partial or fully integrated microfluidic diagnostic devices for infectious diseases have been reported,175–178 including Cepheid’s GeneXpert system. 179 As this subject has been extensively reviewed elsewhere,23,180–187 here we only introduce a few recent examples in the areas of bacterial ID/AST.
Sample processing to enrich, purify, and amplify target bacterial cells or biomarkers from a raw specimen is essential in developing robust diagnostics and POC tests, as pathogens typically exist in low numbers in complex biological samples. Much of the effort has therefore been put into developing microfluidic-based systems for sample preparation upstream of bacterial detection.184,188–197 For instance, Hung and Han’s team reported an inertial microfluidics to rapidly isolate bacteria from whole blood in a label-free fashion with the efficient recovery of even low-abundance bacteria (10–50 bacteria/mL) ( Fig. 7 ). 198 The isolated bacteria were then concentrated via centrifugation and lysed as input for quantitative RNA detection using the NanoString technology. This strategy can be used for both species ID and AST by analyzing rRNA and mRNA profiles on antibiotic treatment, respectively. This RNA assay still requires 8 h and a >105 CFU/mL input bacterial concentration for direct susceptibility testing. Ohlsson et al. recently demonstrated an integrated system for rapid sepsis diagnosis with acoustic bacterial separation, enrichment, and subsequent microchip-based PCR detection. 199 In their system, a blood sample is first processed in an acoustophoretic chip to remove red blood cells. Bacteria were then enriched from remaining plasma by acoustic trapping and eventually released to chips for PCR detection and ID. The entire process can be completed in less than 2 h and can achieve a LOD of 1000 bacteria/mL. Weissleder and Lee’s team recently developed a polarization anisotropy diagnostic (PAD) system that integrates a disposable cartridge for sample preparation and multiwell detection, as well as assays to perform bacterial nucleic acid amplification and detection using a library of sequence-specific probes to assess bacterial burden, pathogen types, antibiotic resistance, and virulence. 200 PAD measurement, which is based on probe fluorescence anisotropy change upon binding to target bacterial nucleic acid, is controlled through a custom-designed application in a smartphone. By applying PAD to detect clinically relevant healthcare-associated infection pathogens, the authors demonstrated that the system can achieve an accuracy comparable to that of bacterial culture, but with a much shorter turnaround time (~2 h) and can operate on site. Indeed, the integration of phenotypic and molecular testing with a mobile phone and digital healthcare tools can be particularly effective for infectious disease surveillance, screening, and diagnosis at the POC.201–204
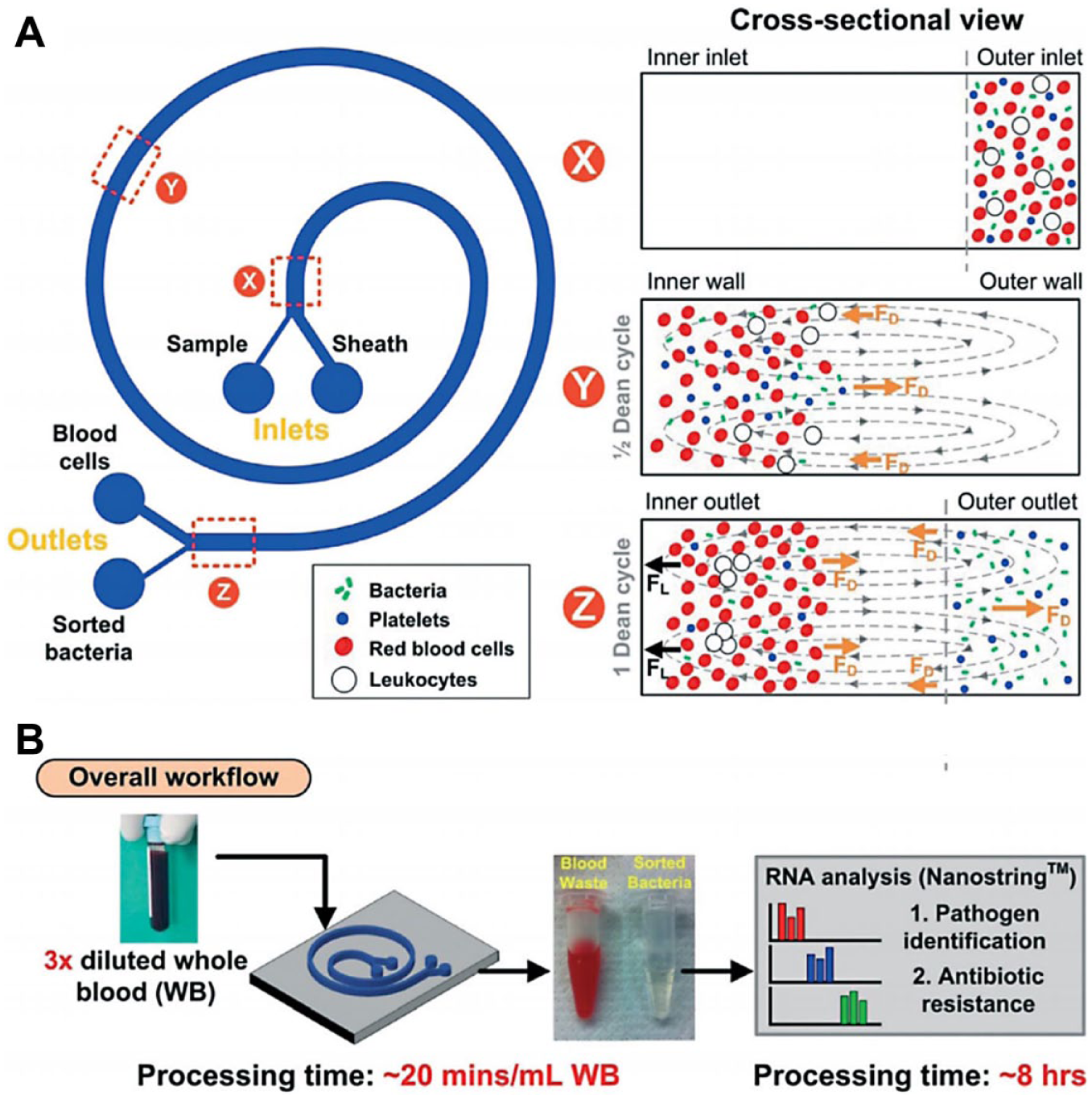
(
Summary
Emerging microtechnologies and automated systems are transforming clinical microbiology by providing faster and more comprehensive and accurate results. However, a number of challenges remain before they can be adopted into routine clinical practice. Recent molecular assays can reduce assay time to hours but are often not sensitive enough to detect bacteria at low concentrations, especially for BSIs, and therefore still requiring a lengthy culture enrichment. How to avoid the initial culture step and potentially sample processing altogether without compromising detection robustness and sensitivity remains an answered question. Another challenge is the lack of an appropriate gold standard to evaluate the validity of these new tests. Bacterial cultures have been considered the gold standard, but they are limited for their inability for nonculturable pathogens. Large discrepancies have already been observed between recent molecular tests and traditional culture methods in clinical microbiology. In addition, molecular assays for AST do not always correlate to phenotypic resistance profiles, especially for ESBLs and CREs. Therefore, at least in the short term, these new rapid molecular tests will likely serve as a “rule-in” rather than “rule-out” function when it comes to resistant pathogens and be used in conjunction with, rather than replacing, culture-based methods. Indeed, tests integrating both phenotypic and genetic analysis will be particularly effective to provide more rapid and definitive actionable information for the physician. Furthermore, system automation and integration with required quality, reliability, and consistency will continue to be key hurdles for microfluidic technologies for bacterial ID/AST applications. As biological matrices are typically complex and highly variable, it is often inevitable to perform upstream sample preparation processes to achieve robust assay performance. Therefore, integration of microfluidics-based sample preparation steps, assay chemistries, and detection and analysis instruments into a fully automated, user-friendly, “sample-to-result” system represents a key future direction for both pathogen ID and AST, especially in a POC setting.23,183
Moving forward, the clinical value of these new technologies needs to be demonstrated. So far, there have been only a handful of studies in evaluating rapid microbiology methods in improving patient outcome, reducing healthcare costs, or improving antibiotic use, which produced overall encouraging yet mixed results.11,13,14,25,136,205,206 Most of the rapid testing methods still require a 4–8 h sample-to-answer turnaround time. Therefore, how and whether they can fit into current clinical workflow, especially for BSI management at the ICU, is yet to be determined. Future randomized and controlled trials of these new diagnostics and implementation strategies will need to be carried out. The outcome of these endeavors would likely influence reimbursement policies in an evidence- or value-based reimbursement system. In particular, many of the new rapid tests (typically $100–$250/test) cost significantly more than the conventional culture methods and the current paradigm of empirical treatment with broad-spectrum antibiotics without a diagnosis (i.e., $0 for the tests). The cost-effectiveness of these new microbiology tests, along with their clinical value, needs to be addressed in the future.
Footnotes
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: W.Z. is the founder of Velox Biosystems, a company that develops rapid diagnostics for infectious diseases, cancer, and other conditions. X.Y. is an employee of bioMérieux.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is supported by the National Institutes of Health (NIH)/National Institute of Allergy and Infectious Diseases (NIAID) (1 R01 AI117061–01).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
