Abstract
Bloodstream infections are major contributing factors of morbidity and mortality among children. Precise and timely identification of causative agents can improve the clinical management and outcome of the infection, potentially saving lives. Electrochemical biosensors previously described by Gao et al. (2017) have the potential to deliver greater speed and discrimination. However, to date there are no data that determine whether the age of the host would cause bacteria to demonstrate different growth characteristics, or whether pediatric samples would behave differently using this electrochemical biosensor. The importance of this knowledge gap is clear: the preclinical testing phase of this line of research is limited by the relative lack of pediatric healthy blood volunteers to complete this work. Therefore, in this study we have applied this novel technology to diagnose bacteria spiked into pediatric blood and compared directly with adult blood samples. Only 180 µL of blood was utilized from both adult and pediatric volunteers and inoculated with Escherichia coli 67, and the signals generated at different time points were compared. We were able to demonstrate that the signals generated by adult and pediatric blood were not significantly different with this detection technology.
Introduction
Bloodstream infection (BSI)–associated sepsis remains a key challenge in medical practice.1 –4 This is particularly true in pediatrics. Accurate statistics on pediatric sepsis worldwide are lacking, but it has been estimated that infections are responsible for approximately 60% of deaths in children aged less than 5 years.5,6 The World Health Organization (WHO) has declared that the four main causes of death in children globally are related to infectious diseases: pneumonia, 1.9 million deaths/year; diarrhea, 1.6 million deaths/year; malaria, 1.1 million deaths/year; and measles, 550,000 deaths/year. 5 A recent study revealed a point prevalence of 8.2% for pediatric severe sepsis and septic shock in pediatric intensive care units worldwide, with a mortality of 25%. 7 Furthermore, treatment attributable to inappropriate antibiotic therapy has been demonstrated to lead to long-term stays in the hospital, increased organ failure, and increased mortality. 8 Therefore, rapid identification of common bacterial pathogens would likely facilitate appropriate species-specific therapy to be started in a timely manner, leading to better clinical outcomes. The current gold standard for the detection of bacteria in patients with BSI is blood culture, along with agar plate culture, which can take up to 1–3 days to identify the source of infection, and an additional 1–2 days may be needed for determining its antibiotic sensitivity.9,10 Therefore, there is a clear need for new diagnostic tests that can identify bacterial species rapidly and accurately in children with life-threatening infections.
Biosensors have been integrated in multiple biological applications, such as clinical diagnostics, environmental monitoring, and forensics.11 –13 Further, biosensors have gained wide popularity due to the potential of overcoming the difficulties encountered by conventional techniques, which can be laborious, complicated, and slow.11,13 –15 There are different types of biosensors, such as optical, thermal, bioluminescent, and electrochemical, characterized depending on the type of reactant. 16 Nevertheless, optical and electrochemical biosensors have become the most widely utilized.11,13,15 However, most optical biosensors still require amplification of the analyte in order to elevate the signal to a measurable level. Among these amplification methods, real-time PCR has ascended as the most widely accepted method and is now accepted as the benchmark for the detection of nucleic acids. 17 Real-time PCR has several limitations, with the most important actualities being the time requirement for the sample processing and amplification, false-positive results, and high instrumentation and labor costs. 11 Electrochemical biosensors furnish superior speed and discrimination at a low price.11 –13 Initially, electrochemical biosensors did not have sufficient sensitivity for applications beyond glucose monitoring and clinical chemistry analysis.11,18 However, recent engineering advancements, such as self-assembly monolayer coating for minimizing nonspecific binding and enzymatic amplification for signal enhancement, have dramatically increased the sensitivity of electrochemical biosensors. Moreover, the ability of electrochemical sensors to tolerate matrix effects of physiological samples has been demonstrated to allow direct detection of bacteria in raw physiological samples.9,19,20
In a previous study, we described an electrochemical biosensor for the rapid identification of bacterial pathogens in whole blood. 9 Additionally, we were able to demonstrate that the electrochemical detecting platform is portable and bedside testing was feasible. With the long-term goal of rapidly identifying bacteria in the small-volume samples that are necessary in children, in the present study we have tested, for the first time, this technology in further preclinical work to diagnose bacteria in pediatric blood. The importance of this knowledge gap is clear: the preclinical testing phase of this line of research is limited by the relative lack of pediatric healthy blood volunteers to complete this work. With the minimal volume requirement, the electrochemical biosensor opens the possibility to evaluate the feasibility of pediatric blood diagnostics. Therefore, in this study we have applied this novel technology to diagnose bacteria spiked with Escherichia coli 67 in pediatric blood and compared it directly with adult blood samples, and compared the signal generated at different time points. We hypothesized that there are no significant differences in the blood matrix of children and adults with this detection technology, and that preclinical testing in adult samples would be generalizable to studies aimed at pediatric BSI.
Methods
Electrochemical Sensors
The multiplex electrochemical sensor chip consisted of 16 gold sensor electrodes with reference and auxiliary electrodes surrounding a circular working electrode (see
Fig. 2
). Gold electrodes for electrochemical sensing were fabricated by sputtering with a shadow mask on plastic substrates. Each biosensor was coated with a self-assembled monolayer consisting of hexanedithiol (HDT), a casein blocker, a dried DNA detector probe, and a thiolated capture probe (
Figs. 1A
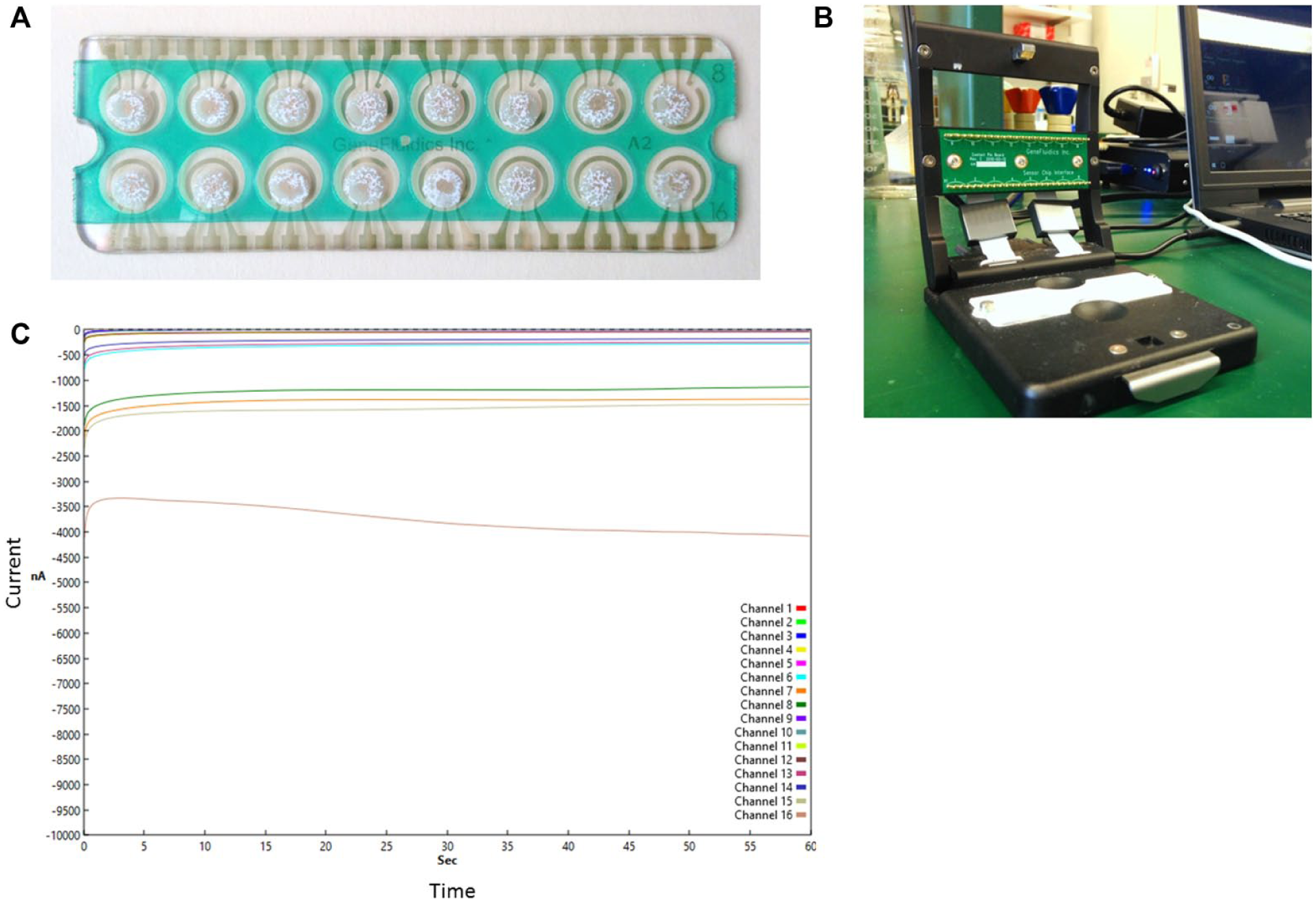
Configurations of the electrochemical chips (
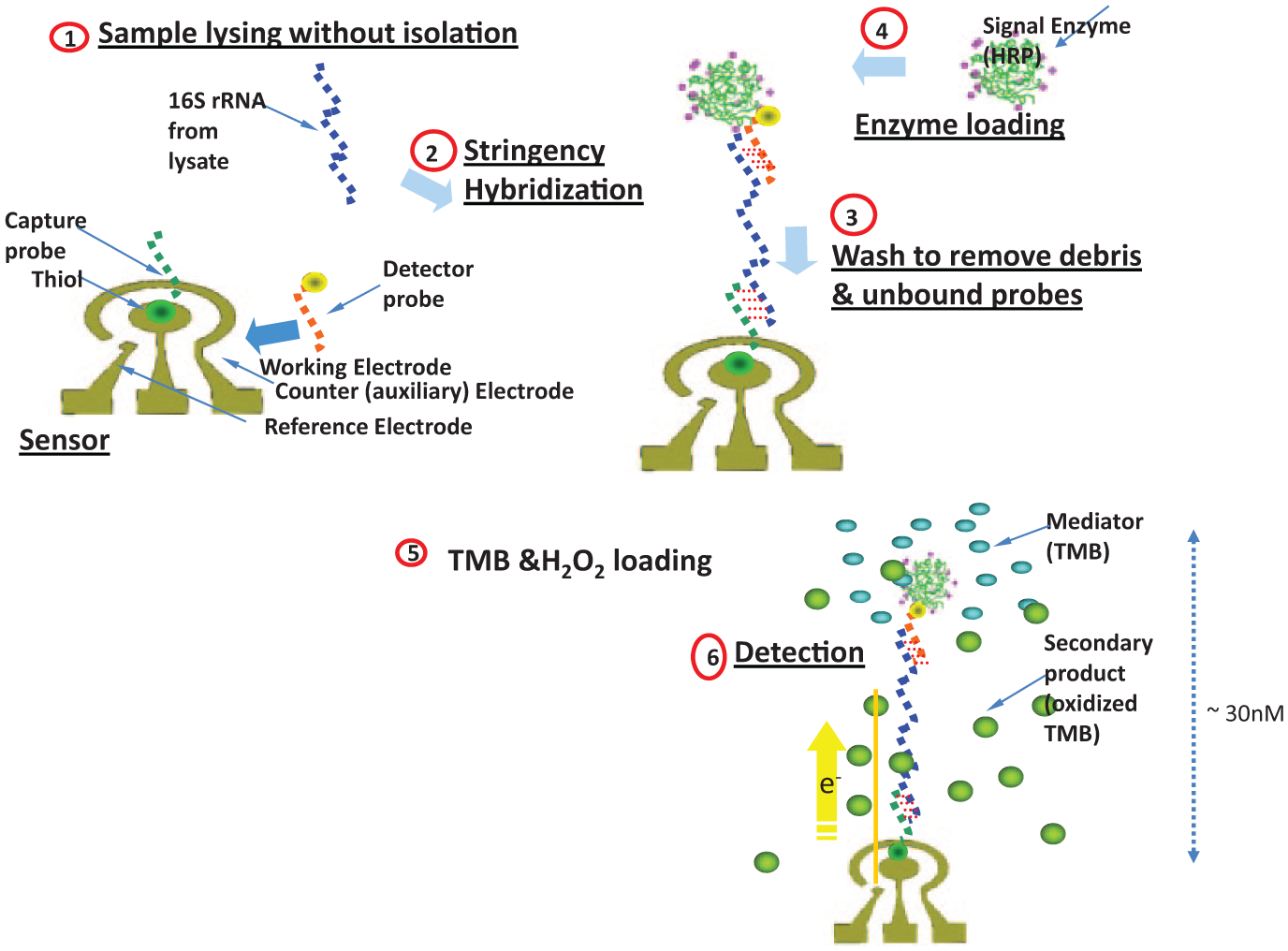
Illustration of binding of bacterial 16s rRNA to the probe and generation of the signal.
E. coli probes
Two pairs of oligonucleotide probe ( Table 1 ), including electrochemical capture and detector probes, were utilized. Probes were synthesized by Integrated DNA Technologies (Coralville, IA). The capture and detector probes were modified with thiol and fluorescein at the 5′ end and 3′ end, respectively ( Table 1 ).
Sequence of Synthetic DNA Oligonucleotides Used to Form a Hybridization Complex with Target E. coli 16s rRNA.
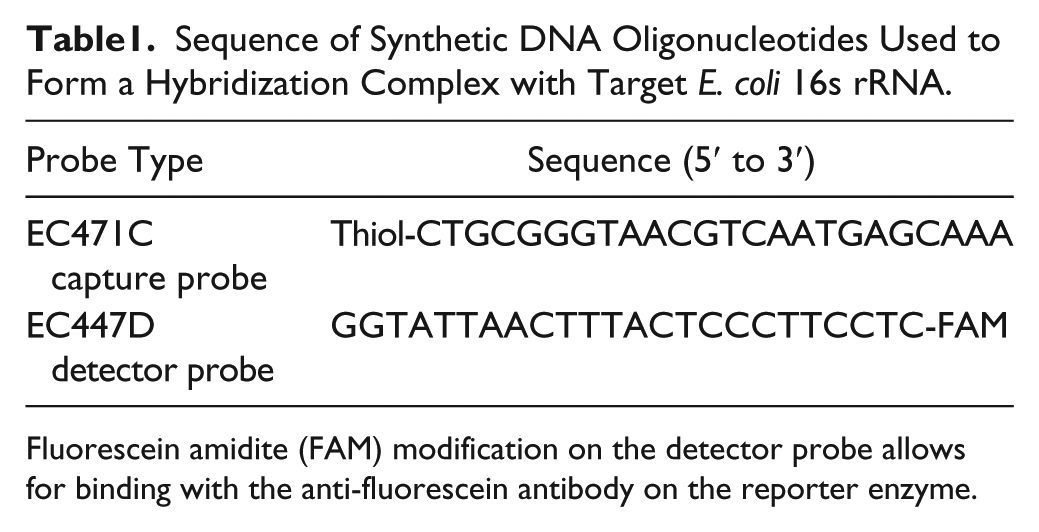
Fluorescein amidite (FAM) modification on the detector probe allows for binding with the anti-fluorescein antibody on the reporter enzyme.
Bacterial Cultures
E. coli 67, a clinical isolate listed in the Centers for Disease Control and Prevention (CDC) carbapenemase detection panel, was used for spiking blood samples. E. coli 67 was grown in 1 mL of Mueller–Hinton (MH) broth (Difco, MD) at 37 °C for 60–90 min to grow the bacterial strain to the exponential growth phase as measured by optical density at 600 nm. The culture was then serially diluted to generate concentrations of 10 (E1), 100 (E2), 1000 (E3), and 10,000 (E4) colony-forming units (cfu)/mL E. coli. These four concentrations were used to spike both adult and pediatric blood. Additionally, 100 µL of each dilution was plated on two replicates of MH agar plates to enumerate the exact number of bacteria spiked into the blood. MH plates were incubated at 37 °C overnight, and the number of cfu was counted the next day to calculate the number of bacteria spiked into the blood samples.
Blood Collection and Spiking
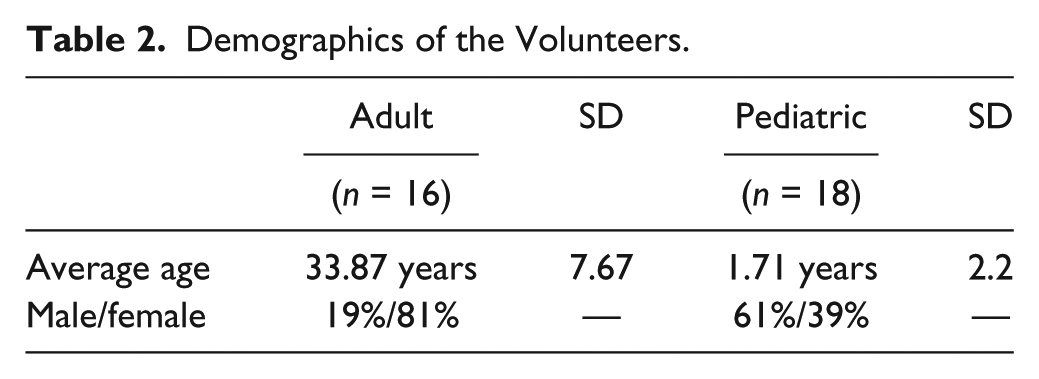
The study was reviewed and approved by the Institutional Review Board of the Penn State College of Medicine. Healthy adult and pediatric blood specimens were collected from volunteers after informed consent was obtained. Sixteen adult blood samples and 18 pediatric blood samples were collected ( Table 2 ). All samples were analyzed within 4 h of collection. Volunteers who had antibiotic treatment within 48 h before blood collection were not eligible for the study. One milliliter of blood was collected into vacutainer tubes (BD, Franklin Lakes, NJ) containing sodium polyanetholesulfonate, and 180 µL of the collected blood was spiked with 20 µL of bacterial culture. Then, 200 µL of MH was added to the blood and centrifuged at 21,000g for 10 min. Supernatant was removed and pellet was resuspended in 200 µL of MH broth and incubated at 37 °C for 4 h. Readings were taken at 120, 180, and 240 min. At each time point, 60 µL of spiked blood was used for the assay. At the end of each incubation, 18 µL of 1 M NaOH was added to the 60 µL of sample, incubated 4 min at room temperature to lyse cells, and neutralized with 12 µL of 1 M HCl. Sensor chips were then loaded with 6 µL of lysate. Positive and negative controls were loaded with 6 µL of MH broth on indicated sensors on the chip.
Demographics of the Volunteers.
Amperometric detection of 16S ribosomal RNA
Electrochemical biosensing was performed using a portable, multichannel potentiostat ( Fig. 1B ). The multiplex electrochemical sensor used a sandwich binding scheme with a capture probe and a detector probe for detecting the target 16S ribosomal RNA (rRNA) ( Fig. 2 ). The target 16S rRNA was immobilized by the capture probe and was detected by the detector probe bound with HRP. The HRP catalyzes the redox reaction to generate an amperometric signal with the addition of 3,3′,5,5′-tetramethylbenzidine (TMB)-H2O2 solution (K-Blue Low Activity TMB Substrate; Neogen, Lexington, KY) and H2O2 substrates ( Fig. 2 ). Briefly, 6 µL of cell lysate was loaded to the each sensor and incubated at 37 °C for 30 min. Chips were placed inside a petri dish to prevent drying. Then, chips were washed thoroughly and dried and loaded with 6 µL of HRP enzyme (150 U/mL) and incubated at room temperature for 15 min. After incubating at room temperature, chips were washed and dried and 40 µL of TMB-H2O2 solution was quickly added. Thirty seconds after adding TMB-H2O2 solution, chips were read using a potentiostat. An example of the current outputs of 16 electrochemical sensors simultaneously is shown in Figure 1C . The first sensor in the chips is the negative control, which is similar to other sensors, and it is loaded with MH broth.
Limit of Blank Assay
For the limit of blank (LOB) assay, 180 µL of blood was collected (adult n = 8, pediatric n = 9) and spiked with 20 µL of MH broth with no bacteria. Then, samples were centrifuged at 21,000g for 10 min. Pellet was resuspended in 200 µL of MH broth and incubated at 37 °C for 4 h. Readings were taken at 120, 180, and 240 min. At each time point, 90 µL of spiked blood was used for the assay. The reaction was stopped with the addition of 27 µL of 1 M NaOH and incubated for 4 min at room temperature to lyse the bacterial cells, and 18 µL of 1 M HCl was added to neutralize the pH. Six microliters of lysate was loaded to each sensor (sensors 2–15) to measure the signal generated from blank blood samples. For positive and negative control sensors, 6 µL of MH broth was loaded.
Statistical Analysis
Power Analysis
Given the preliminary experience under E4 concentration, the equivalence limits were selected as −356.0 and 676.0. Thus, an equivalence test of means using two one-sided tests on data from a parallel-group design with sample sizes of 17 in the adult group and 17 in the pediatric group can achieve 80% power at a 5.0% significance level when the true difference between the means is 0.0, and the standard deviation (SD) is assumed to be 400. Summary statistics are provided with mean and SD for each continuous variable (e.g., age and current) and percentages for each categorical variable (e.g., gender). The normality check was conducted for continuous variables based on Shapiro–Wilk tests. The group comparisons of signals (current [nA]) at each time point were based on two-sample t tests. Mixed-effect models were adopted to investigate the difference between adult and pediatric groups in the mean level of signal and the growth characteristics over time under each concentration level (10, 100, 100, and 1000 cfu/mL) ( Table 3 ). In particular, the fixed effects include the time (120, 180, and 240 min), which was treated as a categorical variable; the group indicator (1 = adult; 0 = pediatric); and their interaction (if not significant, then excluded from analysis). Also, only a random intercept was considered in the model to accommodate the potential correlation among repeated measures of signals ( Table 3 ). The parameter estimation was based on the restricted maximum likelihood, and the analysis was conducted by utilizing SAS PROC MIXED. All the hypothesis tests are two-sided with a significance level of 0.05.
Statistical Analysis.
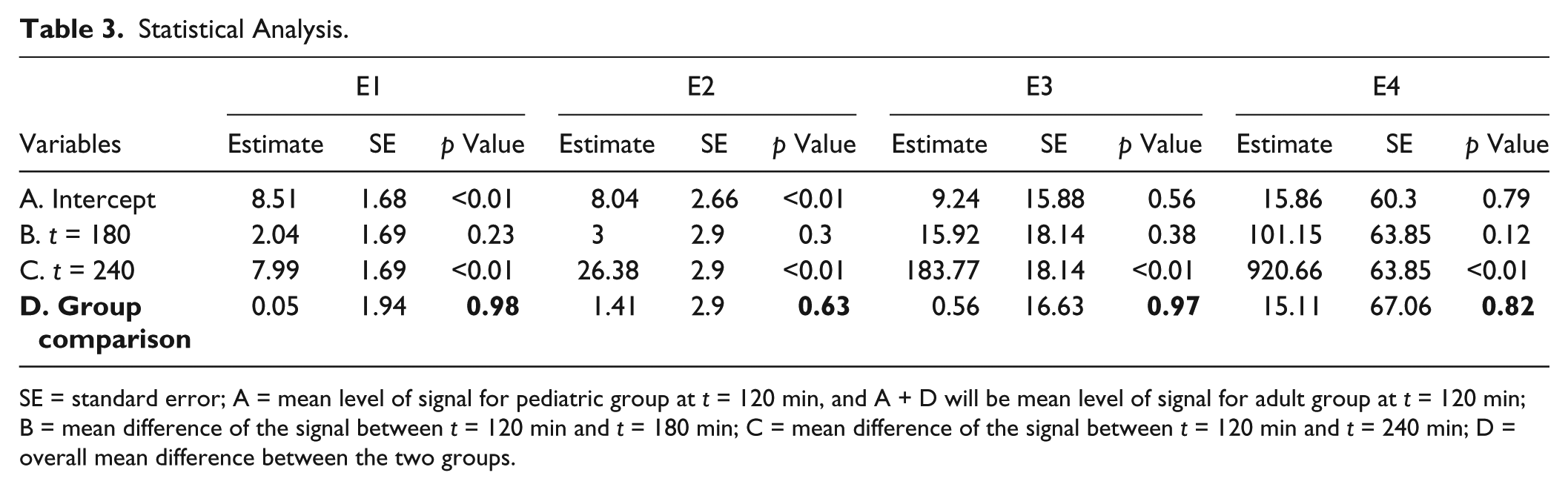
SE = standard error; A = mean level of signal for pediatric group at t = 120 min, and A + D will be mean level of signal for adult group at t = 120 min; B = mean difference of the signal between t = 120 min and t = 180 min; C = mean difference of the signal between t = 120 min and t = 240 min; D = overall mean difference between the two groups.
Results
Limit of Blank
The Clinical and Laboratory Standards Institute defines LOB as the highest signal expected to be generated when replicates of a blank sample (without analyte of interest) are tested. LOB is calculated by measuring the replicates of a blank sample and calculating the mean result and the SD. This aids in determining the background signal. We investigated the variation of LOB signal between adult and pediatric samples, and we were able to demonstrate that the LOB signal is not significantly different between the two groups (p = 0.95) ( Fig. 3A ). Additionally, we compared LOB signals at different time points for both adult and pediatric samples, and we demonstrated that the LOB signal is comparable between the adult and pediatric populations over time, although the LOB signal had a marginal upward trend in both groups (p > 0.05) ( Fig. 3B ).
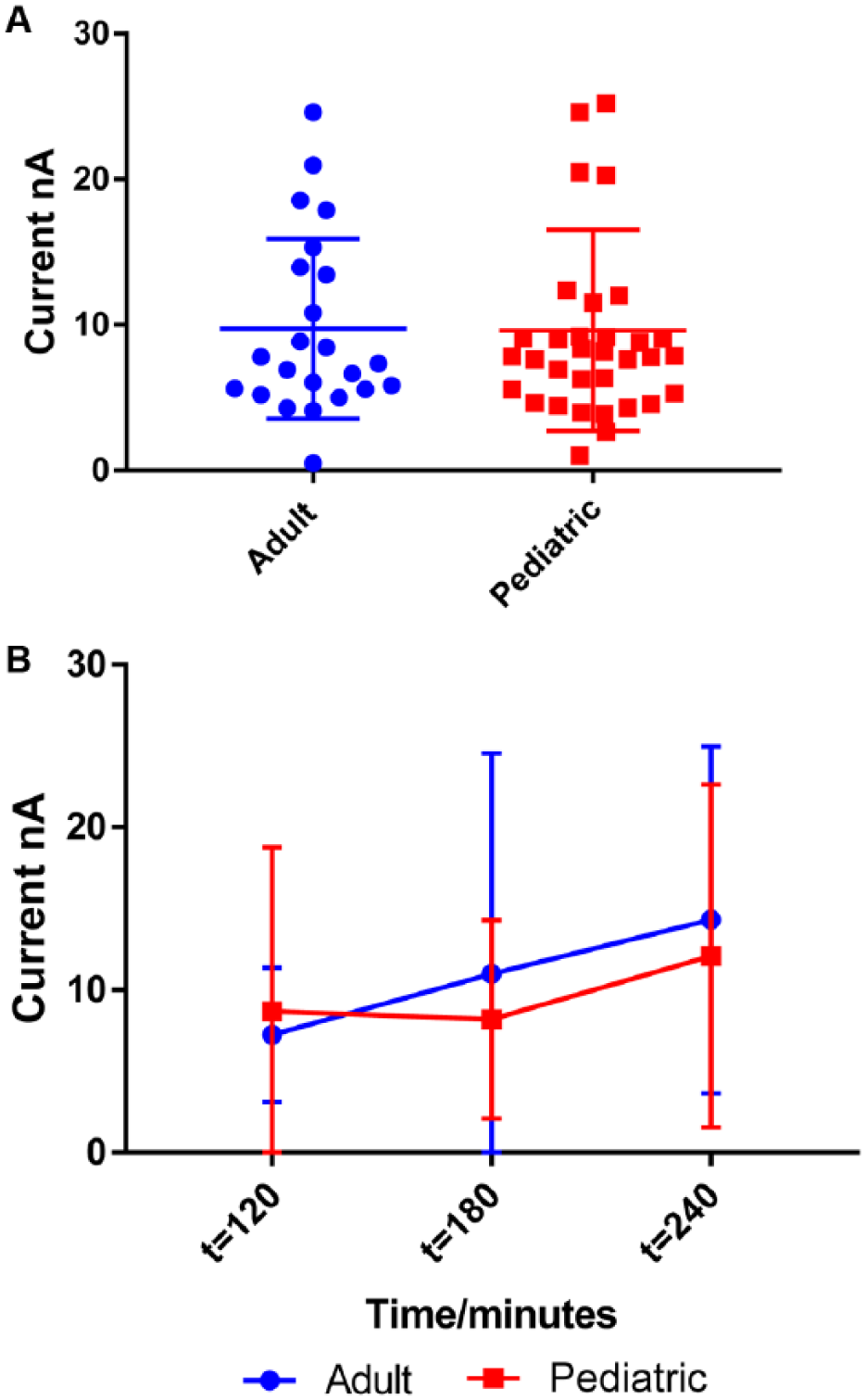
Comparison of adult and pediatric LOBs. (
Amperometric Performance of E. coli 67–Spiked Adult and Pediatric Blood Samples
A clinical isolate of E. coli 67 was acquired from the CDC and utilized for the spiking of blood samples. Four serial dilutions of E. coli 67 were used for spiking and reading at 120, 180, and 240 min. Each day, one adult and one pediatric blood sample were used for the analysis, with the same bacterial culture and chips from the same batch ( Table 2 ). All four concentrations were loaded in the same chips.
E. coli 67 E1 Detection
Four sensors from electrochemical chips were committed for the E1 concentration throughout the study (sensors 2, 3, 9, and 10). One chip was dedicated to each group, and altogether, three chips were used for each blood sample at three time points. Each dot in
Figure 3A
represents a signal generated from a single sensor. Each day, bacteria were plated on MH plates and the amount of bacteria in the culture was enumerated (cfu/mL). The average number of bacteria in E. coli E1 was 28.82 cfu/mL. In this experiment, we were able to establish that signals generated from both adult and pediatric samples for E. coli E1 were similar (
Fig. 4
) (p >0.05). Further, as expected, we observed that signals increased with time for both groups, and the signals remained equivalent between the two groups over time (
Figs. 4
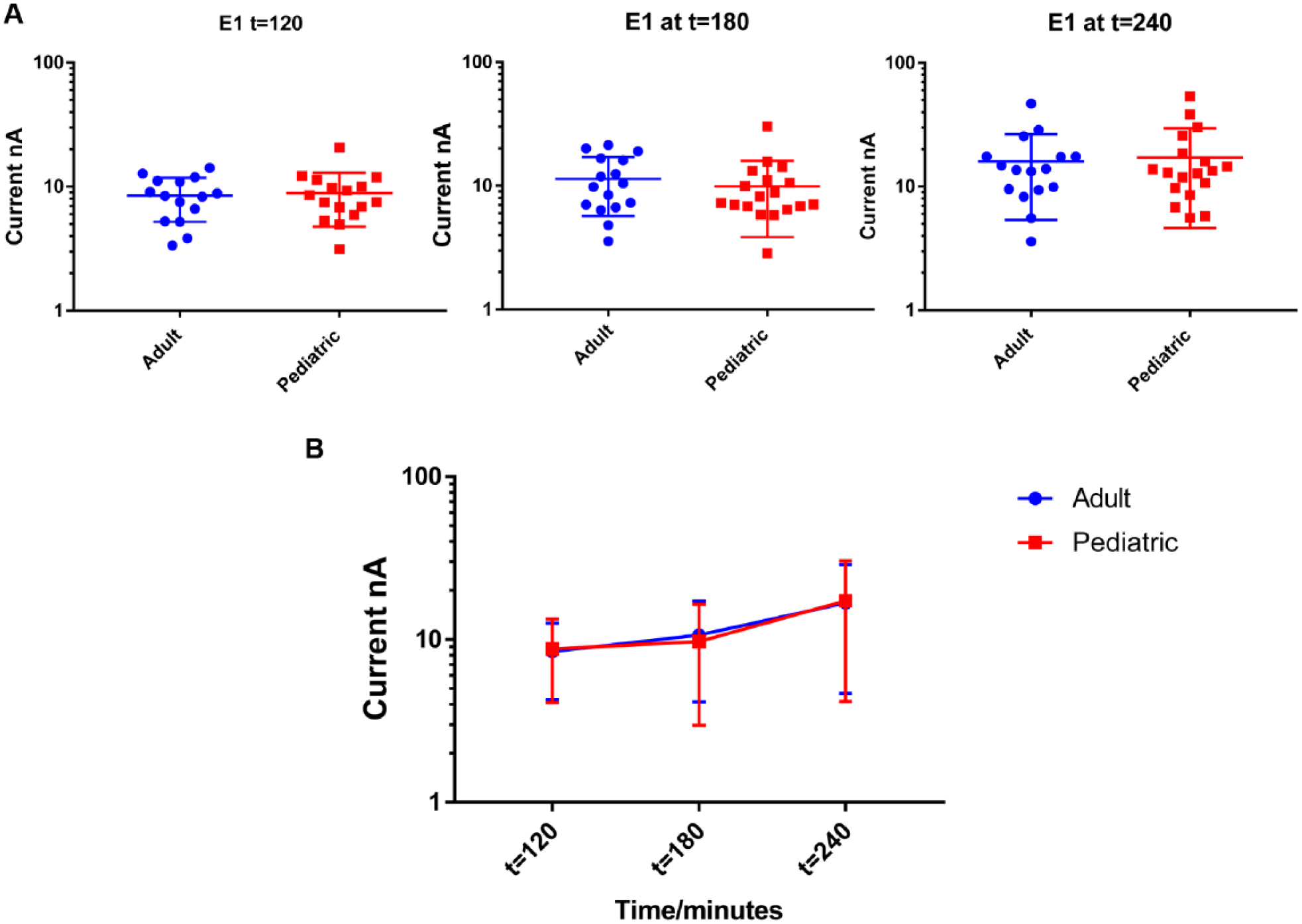
Comparison of signal from the E. coli E1 concentration at t = 120, t = 180, and t = 240. (
E. coli 67 E2 Detection
Sensor 4, 5, 11, and 12 in the electrochemical chips were used for the E2 concentration and three chips were utilized for one adult and one pediatric blood sample each day at all three time points. The average number of bacteria spiked was 288.22 cfu/mL.
Figure 5A
shows the signal from each sensor for both adult and pediatric blood samples at different time points. At this concentration, we were also able to demonstrate that signals generated from both groups are very similar (p > 0.05). Additionally, we observed that signals from both adult and pediatric blood samples increased in a similar fashion (
Figs. 5
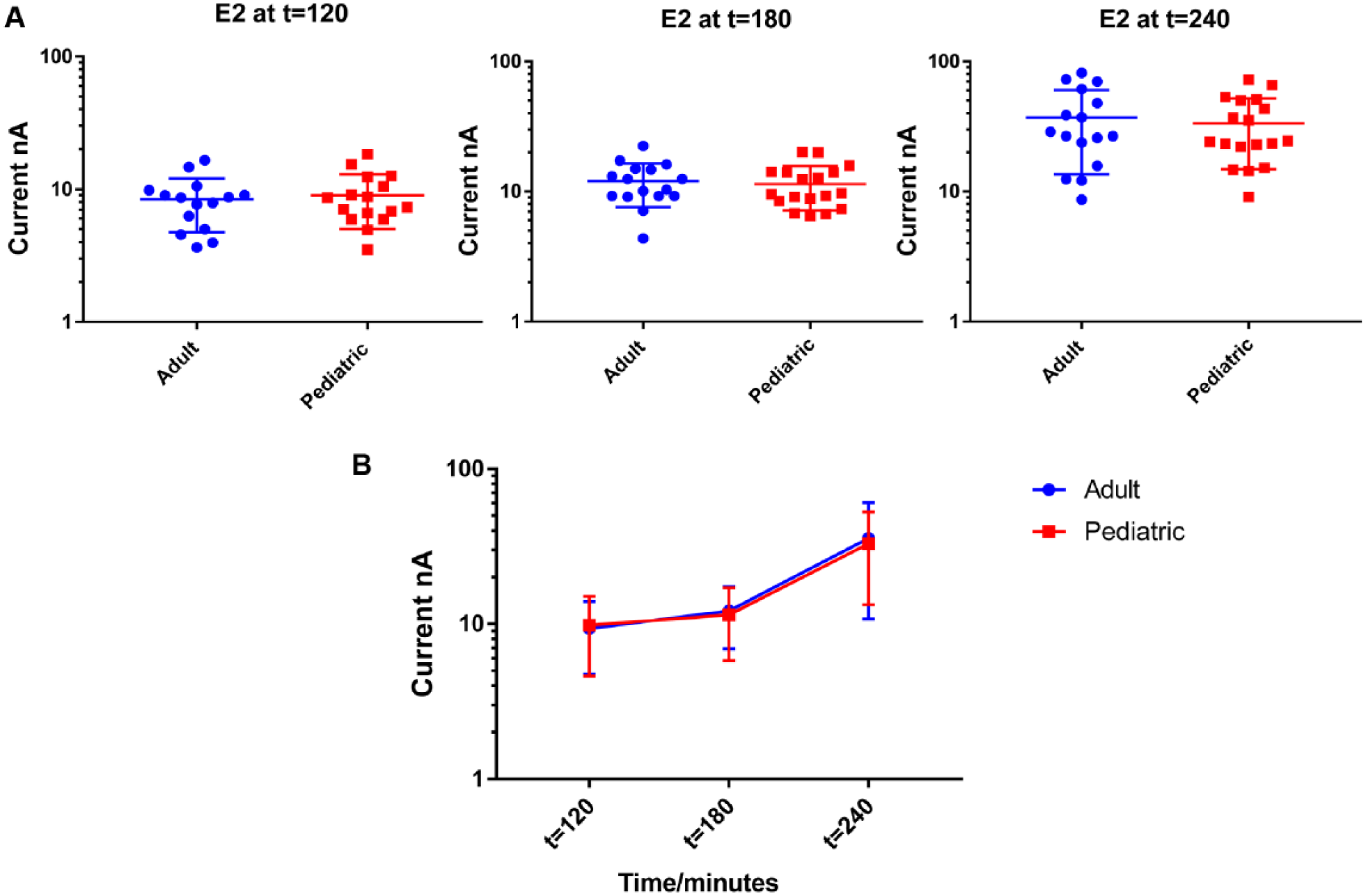
Comparison of signal from the E. coli E2 concentration at t = 120, t = 180, and t = 240. (A) Signal from four sensors (sensors 4, 5, 11, and 12) for 16 adult and 18 pediatric blood samples. (B) Average signal and SD of E. coli E2.
E. coli 67 E3 Detection
Further, we analyzed the E3 concentration from both groups at all three time points and demonstrated that signals from both groups were equivalent ( Fig. 6 ) (p > 0.05). For the E3 concentration, sensors 6, 13, and 14 were dedicated during the study. Also, we were able to demonstrate that the signals from both groups increased over time and were equivalent ( Figs. 6 and 8 ).
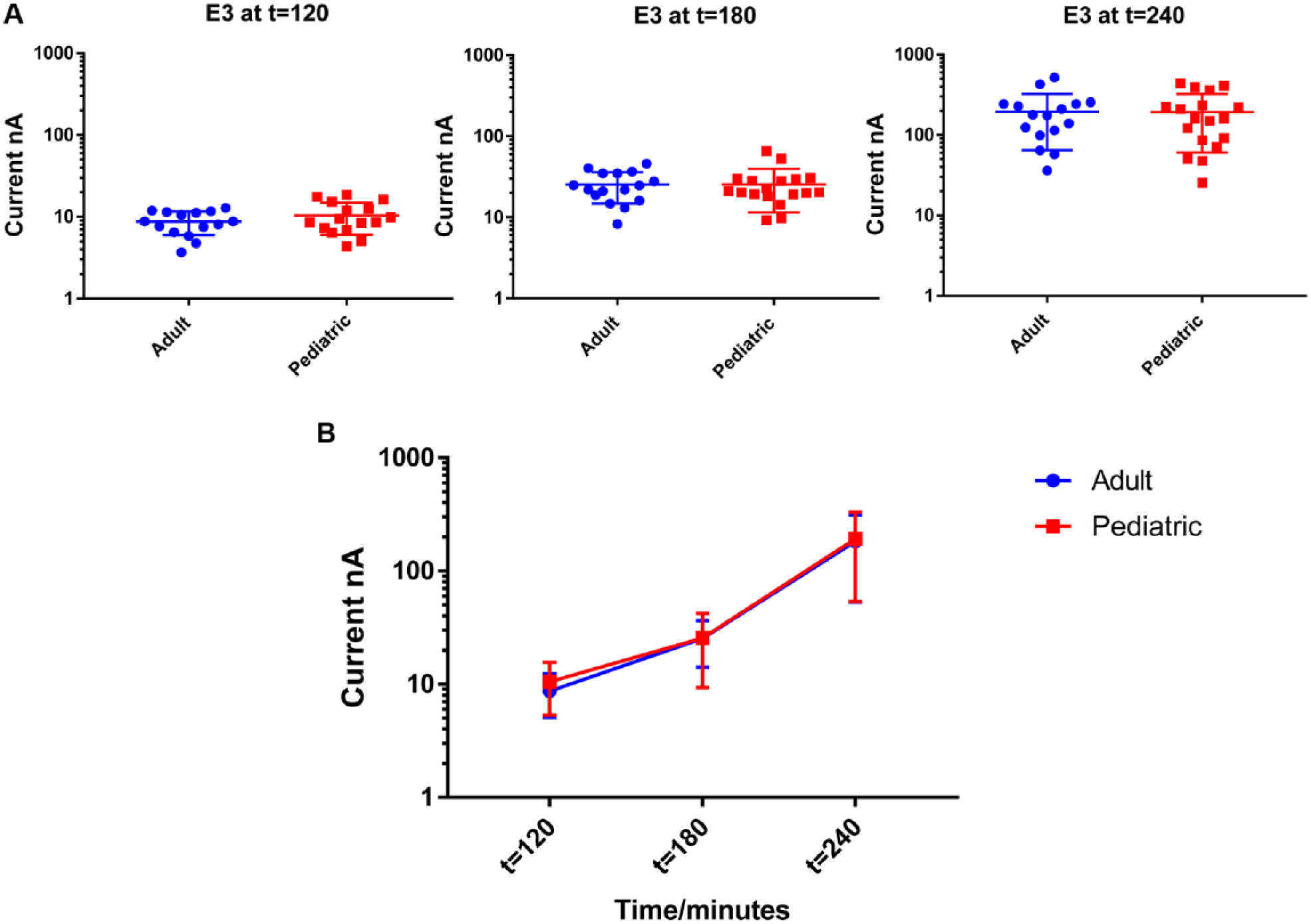
Comparison of signals from the E. coli E3 concentration at t = 120, t =180, and t =240. (
E. coli 67 E4 Detection
Finally, we examined the E4 concentration from both groups at 120, 180, and 240 min, and the signals indicated that both groups are comparable at all time points (
Fig. 7
) (p >0.05). Similarly, we were able to demonstrate that signals from both populations increased over time in parallel (
Figs. 7
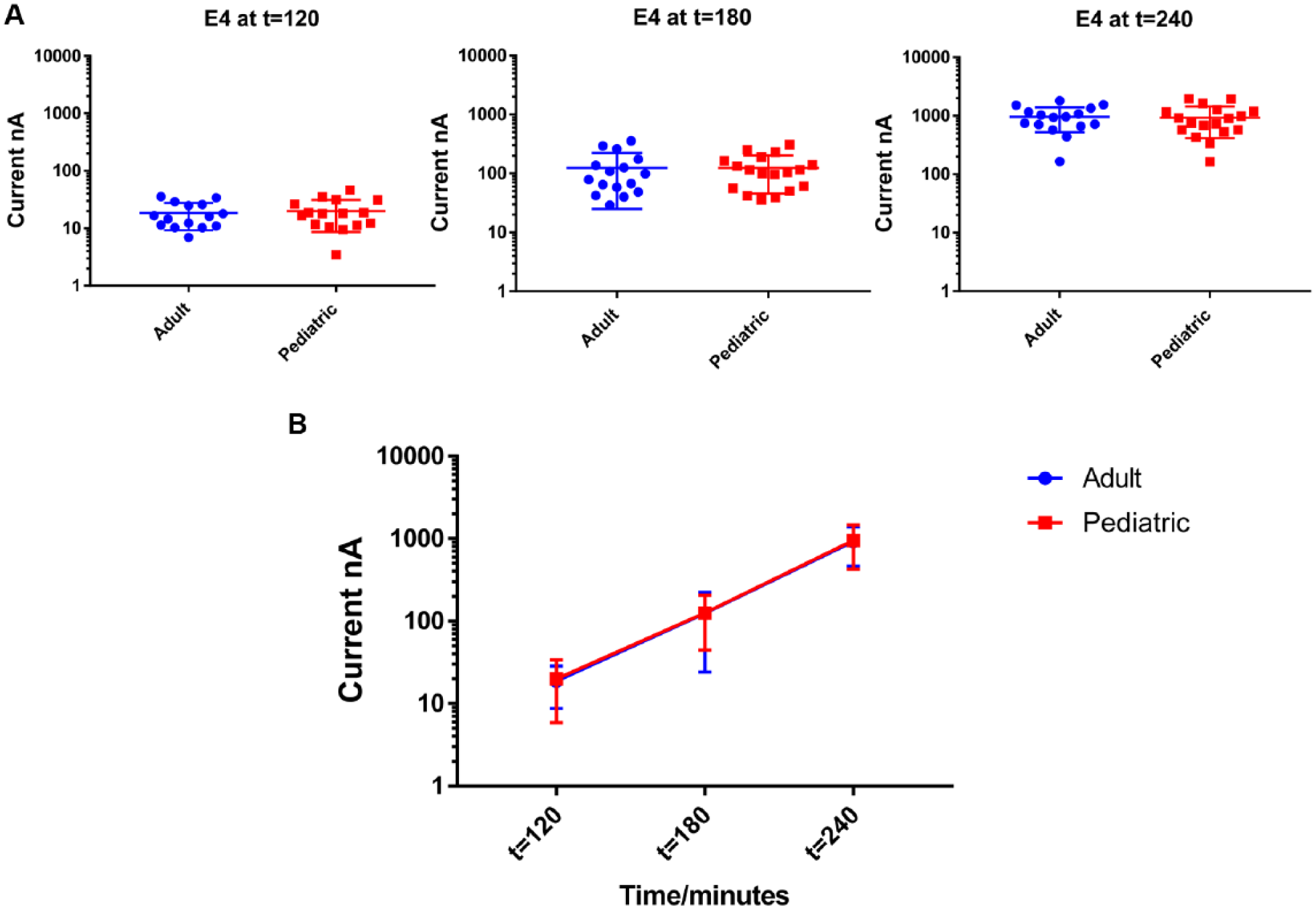
Comparison of signal from E. coli E4 concentrations at t = 120, t =180, and t =240. (
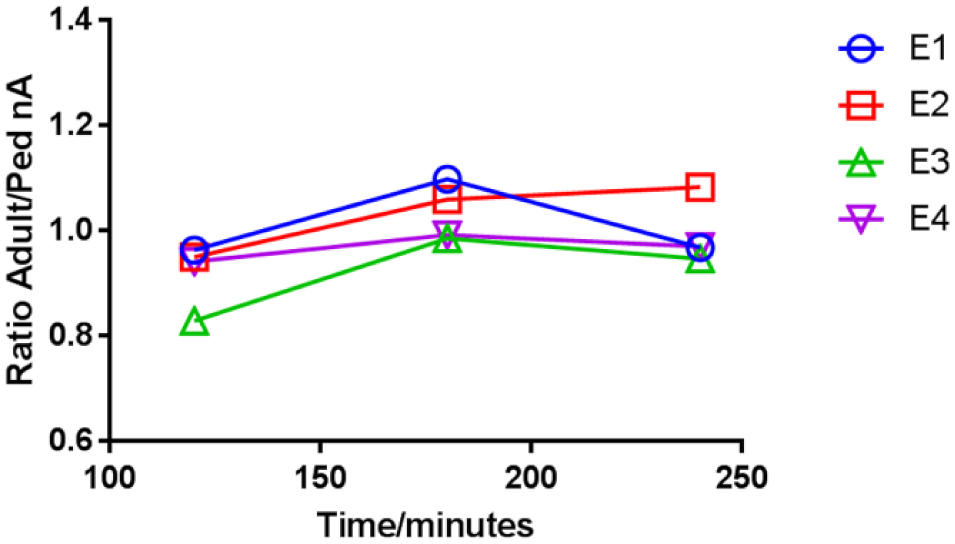
Ratio of the signal generated by adult and pediatric blood samples for each E. coli concentration (E1–E4) over time.
Discussion
In the present study, our overall goal was to evaluate the aptness of an electrochemical biosensor for the detection of BSI in pediatric blood. In a previous proof-of-concept study, the efficiency of this method was confirmed with human blood. 9 However, due to the unknown differences between the matrix of children and adults, we compared blood from both age groups simultaneously in order to evaluate the appropriateness of this method for the detection of BSI in pediatric blood. Data on comparison between adult and pediatric blood with detection methods are sparse. However, it is crucial to acknowledge the possible variations between adult and pediatric blood matrices and probable differences of bacterial growth and illuminate the effectiveness of the electrochemical biosensor for the detection of BSI in pediatric blood ( Fig. 3A ). An understanding of any potential differences is necessary to continue the preclinical testing of these types of devices. Using the LOB assay, we demonstrated that the matrix effects of adult and pediatric blood are comparable for the electrochemical sensor. Whole blood consists of various complex matrices, and most matrix components can skew the signal of the biosensor. For example, serum proteins have the ability to interact nonspecifically with analytes of interest and reduce the sensitivity. 19 However, with the method described in this present study, we were able to strongly substantiate the pertinence of this technology for the detection of BSI in pediatric blood. Further, we demonstrated that during the time of the assay, the LOB signal slightly increases; however, adult and pediatric blood behaved similarly ( Fig. 3B ).
Subsequently, we compared the signal from E. coli 67–spiked samples at different time points and were able to demonstrate that the growth of E. coli 67 in blood was comparable between both groups for all four concentrations ( Fig. 8 , Table 3 ). Thus, we can confirm that the matrix effects of adult and pediatric blood toward the growth of bacteria were very similar, indicating that this technology is suitable for the detection of BSI in pediatric blood. We were able to demonstrate that signals generated from both blood groups were similar at multiple bacterial concentrations and time series readings. Therefore, based on these data, the generalizability of the preclinical work performed in adult blood to children is confirmed. Furthermore, we determined that 180 µL of blood was sufficient to diagnose BSI, reducing the need for the much larger volumes that are needed in standard blood culture methods. Therefore, the impact of this novel testing method for the smallest of children cannot be overstated.
Future plans include testing the clinical utility of the biosensor for the diagnosis of different bacterial organisms. In addition, current work is being performed to begin validating this work in true clinical samples of BSI from children and adults. Further, methodology is planned to expand to identification of Gram-positive bacteria, such as Staphylococcus aureus and Streptococcus pneumoniae, as they account for the majority of BSIs in pediatric patients. The present technology, once it is clinically validated, has the potential to represent a vast improvement to time of BSI diagnosis, from the standard of 1–3 days to only 3 h, leading to earlier use of appropriate antibiotics, with the hope of saving lives.
Footnotes
Acknowledgements
Research reported in this publication was supported by the National Institute of Allergy and Infectious Diseases of the National Institutes of Health under Award Number R01AI117059, SB1AI088756, and the Eunice Kennedy Shriver National Institute of Child Health and Human Development of the National Institutes of Health under Award Number R44HD084033.
Authors’ Note
The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
