Abstract
Advances in understanding many of the fundamental mechanisms of cancer progression have led to the development of molecular targeted therapies. While molecular targeted therapeutics continue to improve the outcome for cancer patients, tumor heterogeneity among patients, as well as intratumoral heterogeneity, limits the efficacy of these drugs to specific patient subtypes, as well as contributes to relapse. Thus, there is a need for a more personalized approach toward drug development and diagnosis that takes into account the diversity of cancer patients, as well as the complex milieu of tumor cells within a single patient. Three-dimensional (3D) culture systems paired with patient-derived xenografts or patient-derived organoids may provide a more clinically relevant system to address issues presented by personalized or precision medical approaches. In this review, we cover the current methods available for applying 3D culture systems toward personalized cancer research and drug development, as well as key challenges that must be addressed in order to fully realize the potential of 3D patient-derived culture systems for cancer drug development. Greater implementation of 3D patient-derived culture systems in the cancer research field should accelerate the development of truly personalized medical therapies for cancer patients.
Introduction
Globally, cancer-related fatalities are the leading cause of mortality. Although significant progress has been made in the treatment of cancer since the advent of chemotherapy, 7.6 million deaths (13% of all deaths) were attributed to cancer in 2008. 1 It is projected that this number will continue to rise, with an estimated 13.1 million deaths in 2030. 1 While the prognosis of certain cancers, such as leukemia and subsets of stomach and breast cancer, has greatly improved with the use of targeted therapies, inter- and intratumoral heterogeneity continue to contribute to cancer resistance and relapse in many cancers.
It is now widely accepted that cancer cells within the same tumor are not identical, but rather heterogeneous coexisting clonal variants that contribute to drug resistance and disease progression. 2 Additionally, every patient’s tumor is unique. Based on this understanding of cancer, increasing attention is being placed on developing patient-centric treatments. Personalized medicine is an approach for disease prevention and treatment that takes into consideration variations in genes, environment, and lifestyle in individuals. Indeed, patient-tailored mutation detection assays are already being used to identify the specific cancer subtypes to guide the use of targeted therapies, such as in the case of non-small cell lung cancer. However, the large number of molecular and epigenetic alterations characteristic of many cancers and our incomplete understanding of the genotype–phenotype relationship suggest that complementary strategies are needed, in addition to the genomic approach, for personalized medicine. Specifically, functional testing of patient cells may play a critical role in guiding personalized treatment strategies.
In line with the recent focus on personalized oncology, technologies that recapitulate the genetic and phenotypic heterogeneity of individual patient tumors for functional drug testing have gained traction. These include the replacement of traditional cancer cell lines with patient-derived xenograft (PDX) models by the U.S. National Cancer Institute in its drug screening program 3 and the development of new methods to grow patient tumor tissues directly as patient-derived organoids (PDOs) in vitro. 4 In this review, we describe the advantages and limitations of these patient-specific platforms and discuss how three-dimensional (3D) culture technologies can be leveraged to overcome some of these limitations ( Fig. 1 ). We believe the merging of 3D culture technologies with PDX and PDO models may provide more clinically relevant systems to truly bring personalized medicine into reality.
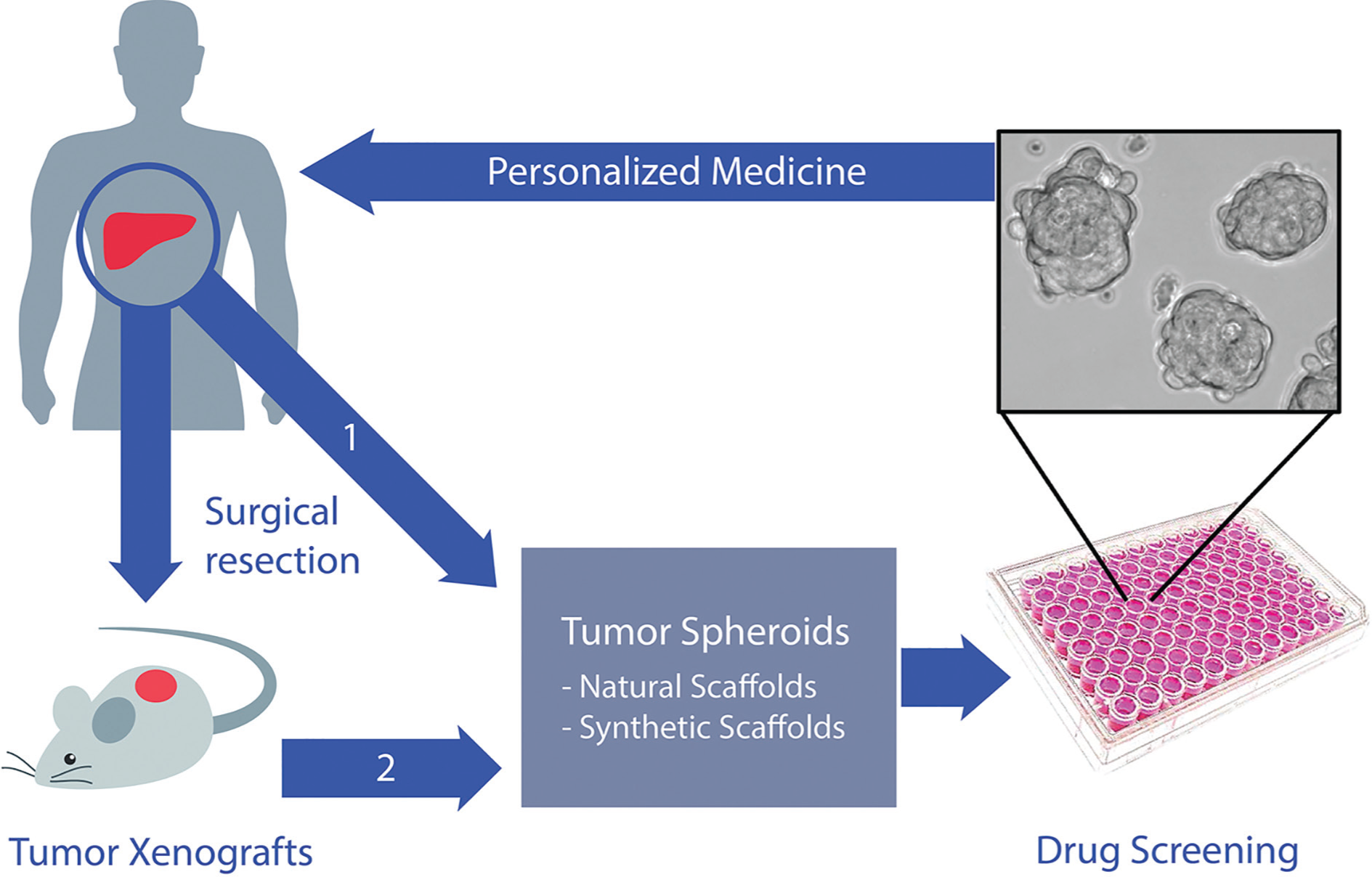
Personalized cancer therapy using 3D spheroid cultures. Tumors derived from patients can be either (1) grown directly in 3D culture or (2) passaged in xenografts in immunocompromised mice to expand the tumor for 3D culture. Tumor spheroids can be subsequently used for high-throughput drug screening or drug combinations to identify promising drug candidates for personalized therapy.
Patient-Specific Tumor Models
Patient-Derived Xenograft Models
As described by Mardis in a recent review, the publicly available multicentered comprehensive next-generation DNA and RNA sequencing efforts of The Cancer Genome Atlas (TCGA) demonstrate that a vast number of mutations in cancer patients drive tumor progression, rather than a few common mutations. 5 PDX models, generally created by direct implantation of freshly resected patient tumors into immunocompromised mice in the absence of any in vitro culture, have been shown to well preserve this tumor heterogeneity inherent in patients. 6 Additionally, PDX models have been reported to retain key characteristics of the parental tumor, including histology, gene expression, and copy number variants.7,8 However, the use of PDX models for patient-tailored therapy has its limitations. A major shortcoming is the high variability in engraftment frequency and growth rate among cells derived from different patients. This could be directly related to the absence of appropriate supporting matrices, growth factors, and/or tumor immunity at the site of implantation. Another major challenge is the lack of patient-specific stromal and immune components. Tumor–stroma mismatch may lead to clinically irrelevant drug efficacy derived from PDX models, particularly when the stromal contribution to treatment response is critical. Lastly, the high cost and relatively long duration involved in generating PDX models make the use of these models for personalized drug testing particularly challenging, especially for patients with limited survival time. These disadvantages have led to the recent development of PDO cultures for rapid drug screening as an alternative platform for personalized medicine.9–12
Organoid Cultures
Organoid cultures are 3D complex multicellular constructs that can self-assemble to recapitulate specific developmental programs. The landmark study for growing in vitro intestinal organoids was first reported by Sato and colleagues, who successfully defined culture conditions for growing murine intestinal organoids without mesenchyme but with the use of key factors (including R-spondin-1, EGF, Noggin, and Matrigel). 12 In a follow-up study, they showed that human intestinal organoids require additional factors, such as nicotinamide, prostaglandin E2, an Alk inhibitor, and a p38 inhibitor for long-term expansion. 10 These organoids can undergo long-term expansion, demonstrate the ability for self-renewal, and are genetically stable with the capacity for differentiation upon withdrawal of key growth factors. More importantly, the establishment of PDOs as a disease model for use in patient stratification has already been shown for cystic fibrosis. 13 To date, organoid cultures of prostate, colorectal, and pancreatic cancers have been established.14–16 As an example, Gao et al. reported the establishment of prostate cancer PDOs that recapitulate the genomic landscape of the disease, including TMPRSS2-ERG fusion, SPOP mutation, SPINK1 overexpression, and CHD1 loss. 15 van de Wetering et al. showed that colorectal PDOs recapitulate the clonal heterogeneity of the original patient tumors. 16 These studies suggest that in vitro PDO cultures are improved, if not complementary platforms to PDX models, for personalized drug testing. However, while organoid cultures are a significant advance toward the goal of personalized drug testing, similar to PDX models, tumor microenvironmental control is lacking in these cultures. Specifically, PDOs are purely epithelial cultures, devoid of any stromal cells. This is a major limitation of the PDO model, as tumors are not merely masses of aberrantly proliferating tumor cells. Rather, tumor tissues are a complex mix of tumor cells, different subsets of immune cells, fibroblasts, and endothelial cells embedded within a dynamic 3D extracellular matrix (ECM) that collectively contribute to tumor progression and therapeutic sensitivity. 17 The importance of the surrounding tumor microenvironment in supporting the tumor phenotype has indeed, for a few decades, been advocated and reflected in 3D tumor modeling, which is described in the next section.
3D Tumor Engineering
Cancer cells have historically been grown as monolayers on two-dimensional (2D) flat surfaces. One of the earliest studies demonstrating the need for culturing cells in 3D was that by Petersen and colleagues, who reported that while normal and malignant epithelial cells appeared phenotypically similar under 2D culture conditions, they behaved very differently when grown as 3D cultures. 18 Organized, growth-arrested acini structures, similar to normal breast architecture, formed when nontransformed breast epithelial cells were cultured in 3D basement membrane matrix (Matrigel). However, the same matrix supported the formation of disorganized cellular masses that did not respond to the growth inhibitory cues from the matrix when cells derived from breast carcinoma were cultured in it. This landmark study underscored the limitations of using 2D culture systems in providing insights into cancer cell behavior and led to the inception of 3D tumor modeling.
Tumor Spheroids
The tumor spheroid is the most commonly employed 3D cancer model, a cellular aggregate that forms when nonadherent conditions are used to culture transformed cells. 19 By supporting a 3D cellular architecture and in vivo–like cell–cell contacts, spheroids recapitulate the in vivo tumor architecture more closely than 2D monolayer cultures. Indeed, spheroids have been reported to more closely reflect the cell morphology, 20 growth kinetics, 20 signaling pathways, 20 and drug response 21 of solid tumors. Additionally, the spheroid model has been actively pursued for drug testing applications, as the 3D structure of spheroids is thought to mimic that of avascular tumors—a necrotic core surrounded by a viable rim as a result of diffusion limitations. 22
Several techniques have been developed for the generation of 3D spheroids. These methods are fundamentally based on the principle that for spheroids to form, the adhesive forces between cells should be larger than that between cells and the culture surface. 23 Some of these methods include the use of spinner flasks and rotary cell culture systems.22,23 While the use of such suspension cultures enables the mass production of spheroids and long-term culture, control over the size and composition of the resulting spheroids is poor. Another suspension culture approach is the liquid overlay method, where cells are grown on nonadherent surfaces, such as agar or poly-HEMA.24,25 While this technique is simple in execution, the control over spheroid uniformity is also limited, which makes consistency in results difficult to achieve. Greater control over spheroid formation can be exerted by generating spheroids using the hanging drop method, where cells spontaneously form aggregates at the bottom of an inverted droplet of cell suspension. Newer hanging drop platforms compatible with high-throughput screening instruments have been developed to overcome the challenge of handling these droplets, traditionally formed on the underside of culture plate lids. 26 Spheroids can also be fabricated by using microwell arrays, where nonadhesive hydrogels are micromolded to form deep recesses with concave bottoms to force cells to form aggregates.27,28 Lastly, microfluidic approaches have also been undertaken to generate spheroids with tunable size by controlling the cross-sectional area of the microchannels. 29 While useful as a model of avascular tumors and micrometastases, tumor spheroid models are simplistic, as they are solely dependent on the ability of cells to self-assemble and produce their own ECM. As such, spatiotemporal control over tumor–ECM and tumor–stromal cell interactions is limited. 30 To address these challenges, tools in the tissue engineering field—such as the use of 3D scaffolds and biological moieties to guide tissue formation—have been leveraged for the development of more sophisticated and advanced 3D tumor models.
Scaffold-Based 3D Tumor Models
By harnessing tissue engineering techniques, it is envisioned that complex 3D tumor models can be engineered with control over matrix composition, density, stiffness, and spatial positioning of different cell types. 31 This approach may generate more in vivo–like tumor models that support mechanistic studies of the role of the tumor microenvironment in tumor progression and acquisition of drug resistance. Biologically derived matrices, such as collagen and Matrigel, are highly popular substrates that have been used for 3D cancer cell culture. Encapsulated tumor cells form clusters similar to traditional spheroids, but unlike traditional spheroids, cells experience not only cell–cell but also cell–ECM interactions that can influence their phenotype.31,32 Collagen type I gels are often used to provide a 3D microenvironment for cancer cell growth because it contains the tripeptide Arg-Gly-Asp (RGD), a short amino acid sequence found in several ECM components, such as fibronectin, which binds to integrin receptors on the cell surface. 33 Collagen gel cultures have been employed to study tumor drug sensitivity, 34 replicate the prevascularized stages of in vivo solid tumor progression, 33 and construct an in vitro organotypic model of normal and malignant breast to investigate the role of cell–cell interactions in breast cancer progression. 35 Matrigel is a commercially available product consisting of basement membrane proteins such as collagen type IV, perlecan, and laminin, extracted from the culture of Engelbreth–Holm–Swarm tumor cells. 36 This extract forms a 3D gel at 37 °C via protein aggregation. Matrigel has been used widely in cancer research, 37 as well as in the generation of PDO cultures.14–16 However, the use of Matrigel for drug screening purposes is limited, as it is largely ill-defined in composition; its compositional complexity makes it difficult to understand cell signaling and elucidate the role of individual microenvironmental components on tumor behavior. 38 Additionally, batch-to-batch variability may affect the reproducibility of experimental results. 38
To confer greater control over matrix properties than biologically derived scaffolds and to maintain reproducibility of results, synthetic scaffolds have also been developed and investigated for use in 3D tumor modeling. Synthetic polymers such as polylactide and poly(ethylene glycol) (PEG) are widely used in the tissue engineering field, as they enable great flexibility in the design of biochemical and biophysical properties, 39 thereby permitting the systematic evaluation of the effects of tumor microenvironmental components on tumor progression. However, many of these polymers inherently have limited bioactivity and lack the ability to support critical cell–matrix interactions. 38 To overcome this limitation, synthetic scaffolds can be modified with desired biological motifs to mimic the in vivo ECM.40,41 As an example, Fong et al. demonstrated the use of semisynthetic hyaluronan hydrogels functionalized with integrin-binding RGD motifs and matrix metalloprotease-sensitive substrates as a platform to support the coculture of osteoblastic cells with prostate cancer PDX cells to recreate the prostate cancer–osteoblast unit in bone metastasis in vitro. 42 In the presence of the functionalized matrix, osteoblastic cells were observed to organize around the prostate cancer spheroids, closely mimicking the architecture of prostate cancer metastases in bone. Additionally, in vivo–like tumor–stromal signaling mediated by the fibroblast growth factor axis was also recapitulated in vitro. Given that the complex in vivo microenvironment makes it difficult to employ PDX animal models to investigate tumor–stromal interactions, this engineered model was one of the first to demonstrate the feasibility of leveraging 3D technologies to create in vitro PDX models to enable controlled mechanistic studies to be carried out (faster, higher throughput, and more cost-effective than their in vivo counterpart) in parallel with in vivo studies. Specifically, 3D PDX in vitro models enable the role of various microenvironmental components to be teased apart, a challenging feat with in vivo PDX models. In another example, Gill et al. modified PEG hydrogels with bioactive peptides to study the role of the matrix on epithelial morphogenesis and epithelial-to-mesenchymal transition in lung cancer. 43 Interestingly, metastatic lung cancer cells were able to form lumenized epithelial spheres comparable to that seen with Matrigel. On this note, Gjorevski et al. very recently demonstrated the use of PEG hydrogels to define the key ECM moieties that are necessary for intestinal organoid formation and showed that different mechanical environments and ECM components are necessary for different stages of the process. 44 Serving as a defined substitute for Matrigel, it is to be expected that similar progress will soon be made in elucidating the key ECM components that govern PDO formation for various cancers.15,16
Beyond the Matrix
The examples above illustrate the tools that are available to design the tumor matrix in 3D, bearing in mind that not all ECMs are the same. Cells are dynamic entities that respond to their microenvironment, whether biochemical or physical cues. Using alginate hydrogels with tunable stiffness, Shin et al. recently reported that different myeloid leukemia subtypes have different proliferative responses to changes in matrix stiffness, and that matrix stiffness can modulate chemosensitivity. 45 This study highlights the importance of incorporating 3D mechanical cues in anticancer drug screening. With these tools, besides cancer cells, it is possible to modify the tumor microenvironment to house fibroblasts,46,47 vascular cells,48,49 and immune cells, 50 critical components in the tumor microenvironment that influence tumor heterogeneity and drug response. In one example, Yip et al. observed that the incorporation of stromal fibroblasts in coculture with liver carcinoma cells within collagen gels enhanced chemoresistance to an anticancer drug. 51 By coculturing T lymphocytes with lung carcinoma cells in 3D using a porous polystyrene scaffold, Alonso-Nocelo et al. observed that T lymphocytes enhanced the malignant phenotype, highlighting the importance and feasibility of engineering tumor-immune cell interactions in the tumor microenvironment in vitro. 50 Furthermore, it is also possible to incorporate perfusion to study the effect of mechanical forces exerted by interstitial flow in the tumor in 3D constructs. Such an approach was illustrated by Santoro et al.; Ewing sarcoma cells cultured in an electrospun fiber scaffold within a flow perfusion bioreactor exhibited a different cancer phenotype and drug sensitivity than the statically cultured counterpart. 52 Lastly, with the advent of 3D printing technologies in tissue engineering that enable the direct assembly of cells and ECMs to fabricate complex designs and structures, 53 we are one step closer to engineering a tumor unit representative of the patient’s tumor by combining relevant cell types within an appropriate matrix with high spatial control. Zhou et al. reported a method of 3D printing for HeLa cells with gelatin, alginate, or fibrinogen hydrogels to engineer a 3D in vitro model of cervical cancer. Cancer cells grown in the 3D printed construct were found to exhibit higher matrix metalloproteinase expression (related to cancer metastasis) and chemoresistance. 54 In another interesting study, Phamduy et al. demonstrated the feasibility of printing cancer cells onto live and intact microvascular tissue to study cancer cell dynamics and interaction with vasculature. 55 While these emerging studies suggest the potential of 3D printing technologies as tools for developing the next generation of complex 3D tumor models, further studies are needed to truly demonstrate the need for such high spatial control in modeling cancer.
In sum, we have an armamentarium of 3D culture technologies that can be leveraged to engineer the tumor microenvironment in vitro. As described, a major limitation of PDX and PDO models is the presence of inappropriate tumor stroma or lack thereof; the incorporation of PDX or PDO cells into 3D systems may circumvent this problem. The development of 3D PDX in vitro models may greatly increase the throughput and decrease the time needed for drug evaluation, overcoming the key limitations that currently hinder the use of these otherwise useful in vivo models for personalized medicine.
Future Directions: Combining 3D-Cultured Patient-Derived Cells with High-Content Screening
High-content screening (HCS) is frequently used at the preclinical stage of drug discovery due to its relatively low cost and larger and faster data output compared with animal studies. HCS also has the benefit of allowing for multiparametric phenotypic outputs that can be used for more comprehensive drug screens. 56 These HCS studies conventionally employ 2D monolayer-cultured cancer cells that may not accurately reflect in vivo pathobiology. For instance, Lovitt and colleagues demonstrated that monolayer breast cancer cells displayed increased drug sensitivity compared with corresponding 3D tumor spheroids. 57 Specifically, the authors tested several standard-of-care neoplastic agents, including epirubicin, paclitaxel, and vinorelbine, and observed decreased activity of these drugs in 3D compared with the 2D monolayer-cultured cells, underscoring the importance of the third dimension in drug testing. However, several challenges have to be addressed in the transition from simple monolayer cultures to complex 3D tissues for HCS.
Going beyond the Simple, Scaffold-Less Cancer Spheroid
Although gaining popularity, 3D cultures have not been widely embraced for HCS due to several technical challenges, including the added difficulty (compared with 2D cultures) of generating consistent 3D cultures and the need to handle large and complex datasets. 58 Traditional methods of generating spheroids, including the rotary cell culture system, 59 hanging drop, or use of ultra-low attachment plates, 60 are often tedious and produce spheroids of inconsistent sizes, making spheroid cultures difficult to scale up for HCS studies. Advances in microfluidics and microfabrication technology over the past decades have enabled improved spheroid formation efficiency and consistency in spheroid size.29,61 More importantly, the compatibility of 3D tumor spheroids with existing liquid-handling robotics for HCS can greatly enhance the drug screening process. For instance, Tung et al. described the use of a multiplexed 384-well hanging drop array platform to produce uniformly sized spheroids compatible with HCS instrumentation and showed that 3D tumor spheroids responded differently than conventional 2D monolayer cultures. 26 Specifically, they showed that 5-fluorouracil (5-FU) specifically targeted proliferating cells while sparing quiescent cells in the spheroids, and that tirapazamine (TPZ), a hypoxia-activated cytotoxin, is more effective in 3D tumor spheroids. Furthermore, combinatorial drug treatment with both TPZ and 5-FU exerted synergistic effects on viability. More recently, Cavnar et al. developed a complementary spheroid transfer and imaging (TRIM) plate for their 384-well hanging drop spheroid cultures. The authors demonstrated that the low-aspect ratio of wells in the TRIM plate can facilitate high-fidelity spheroid capture and several downstream analyses, such as immersion-based imaging, bulk spheroid collection, and timely enzyme substrate kinetic studies. 62 Furthermore, incorporation of liquid-handling automation and micropattern designs for the generation of consistent 3D tumor spheroids was described by Monjaret et al.; the authors demonstrated the ability to multiplex and carry out analyses in a single plate, thereby enabling the reliable reproduction of samples and imaging in an easy one-step method, an aspect critical for the successful implementation of automated HCS for drug testing. 63
While these advances demonstrate the merging of simple 3D spheroids with HCS amenable for personalized drug screening, continued efforts are needed to merge more advanced 3D culture technologies with HCS for ultimate use in personalized treatment planning. As described above, while advances have been made to enable high-throughput studies with spheroid cultures (without ECM), these systems not only do not support the study of ECM influence on the cancer cell phenotype, but also do not enable the spatial control of heterogeneous cell types (cells self-assemble). Moreover, imaging of untethered cells remains challenging. To overcome this problem, a few groups have demonstrated the feasibility of incorporating matrices to generate 3D scaffold-based systems suitable for automated 3D high-throughput drug screening. In one example, Engel et al. used a high-throughput benchtop dispensing system to generate hyaluronic acid–based 3D cultures of prostate and endometrial cancers for drug screening, where it was shown that these models were particularly able to identify drugs that failed in clinical trials. 64 In another example, Li et al. developed an off-the-shelf microscaffold (based on macroporous gelatin sponges) array chip that was designed according to the dimensions of a commercialized 384-multi-well plate, thereby making it compatible with existing benchtop equipment for high-throughput cell-based assays. 65 Compared with cells cultured on 2D, cancer cells in this platform were observed to exhibit greater drug resistance. On a similar note, Fan et al. demonstrated the feasibility of creating tumor-on-a-chip using PEG diacrylate hydrogels to form brain cancer tissues, amenable to high-throughput drug screening applications. 66 These described studies represent early (and few) efforts in the field recognizing the need to automate and adapt 3D culture technologies for high-throughput mechanistic studies and drug screening. With rapidly increasing interest in developing more physiologically relevant human microtissues to facilitate drug discovery, 67 more efforts are needed for the construction of complex 3D tumor tissues incorporating primary tumor tissue (with the relevant ECM) that are amenable for high-throughput personalized oncology studies.
Defining the Tumor Unit to Be Recapitulated In Vitro
Should the bottom-up or top-down approach be adopted for personalized drug screening? If the bottom-up approach were to be taken where components of the tumor microenvironment are pieced together, to what extent does the tumor microenvironment need to be recapitulated in vitro? That is, what constitutes the minimum 3D tumor unit required for functional drug testing? For example, tumor microenvironment-targeting agents, such as antiangiogenic or immunological checkpoint blockade therapies, would require the presence of vasculature and immune cells. As described above, it is indeed possible to incorporate desired cellular interactions with spatiotemporal control to elicit in vivo–like tumor–stromal cross talk. 42 However, it may be challenging to obtain, identify, and expand patient-specific stromal cells for this bottom-up (reconstruction) approach. The tumor slice culture system and histocultures represent top-down approaches that take into full consideration the importance of the tumor microenvironment on drug response68,69; however, limited patient tissue may hinder the feasibility and implementation of these approaches.
Development of New Approaches to Assess Cellular Response in Heterogeneous Tumors
Given the heterogeneity of primary tumor tissues, what methods should be employed to assess the degree of heterogeneity in drug response? This heterogeneity in response refers to that arising from the presence of stroma and different tumor clones. Notably, drug efficacy in vitro has traditionally been assessed as an averaged global response, evaluated using standard assays such as those based on tetrazolium or resazurin reduction or by measuring the amount of cellular ATP. Such an approach formed the basis for early chemosensitivity assays using patient cells.69,70 The use of more precise analytical methods, such as high-content imaging, to identify specifically the subpopulations of cells that are sensitive or resistant to drug treatment may require the creative generation of novel drug response evaluation criteria to correlate in vitro drug response with clinical efficacy.71–74
In summary, we have made significant advances to an unprecedented level to model cancer with much greater accuracy for precision medicine, in moving from the use of simplistic monolayer culture methods to the use of primary tumor sources and sophisticated 3D systems. These advances are fundamentally built on the recognition that the tumor microenvironment and tumor heterogeneity play a critical role in tumor progression and therapeutic resistance. With further improvements in 3D tumor modeling, we are likely to ultimately have the means to develop reliable 3D ‘patient avatars’ that can guide therapeutic selection in the clinic for cancer patients.
Footnotes
Acknowledgements
We would like to thank Wong Chun Xi (Mechanobiology Institute, Singapore) for his help in illustrating the figure in this manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: H.Y. was supported by funding from the Institute of Bioengineering and Nanotechnology (A*STAR, Singapore). E.L.S.F. was supported by funding from the National Medical Research Council (Ministry of Health, Singapore; grant number R-185-000-294-511). T.T. and E.C. were supported by funding from the Ministry of Education, Singapore; grant MOE2015-T2-2-126 and T1-BSRG 2014-5.
