Abstract
The development of the first open-source automated peptide synthesizer, PepSy, using Arduino UNO and readily available components is reported. PepSy was primarily designed to synthesize small peptides in a relatively small scale (<100 µmol). Scripts to operate PepSy in a fully automatic or manual mode were written in Python. Fully automatic script includes functions to carry out resin swelling, resin washing, single coupling, double coupling, Fmoc deprotection, ivDde deprotection, on-resin oxidation, end capping, and amino acid/reagent line cleaning. Several small peptides and peptide conjugates were successfully synthesized on PepSy with reasonably good yields and purity depending on the complexity of the peptide.
Introduction
Significant advancement has been happening since the past decade in the development of very low-cost open-source microcontrollers such as Arduino and Raspberry Pi.1,2 The use of these microcontrollers in combination with the powerful general-purpose and easy to learn programming languages such as Python allows scientists to develop certain automated instruments relatively easily at a small fraction of the cost of their commercially (if) available counterparts. 3 In addition, these instruments can be fully customized for their laboratory needs.4–7 Here, the development of the first open-source automated peptide synthesizer, PepSy, developed using Arduino UNO and readily available components, is reported. The scripts to operate PepSy were written in Python.
PepSy uses the solid-phase peptide synthesis (SPPS) method employing the traditional Fmoc chemistry. 8 The SPPS method and its application in the automated synthesis of peptides was first reported by Merrifield in the mid-1960s.9,10 Later after the introduction of the Fmoc-based SPPS and the availability of improved hardware as well as software simplified the design of automated peptide synthesizers.11,12 Since then, several automated peptide synthesizers with a variety of options were developed and became commercially available.13,14 However, even today they remain relatively expensive (>$30,000) and require recurring service contracts, which limits their use in the laboratories with limited funding. In this regard, PepSy will serve as a low-cost (<$4000) do-it-yourself alternative to the commercially available automated peptide synthesizers.
Materials and Methods
All chemicals obtained commercially were used without further purification. Bestatin was obtained from Acros Organics (Geel, Belgium). Fmoc-bestatin was synthesized as previously reported. 15 Rink amide MBHA resin (100–200 mesh, 0.3–0.8 mmol/g) and Fmoc-Thr(tBu)-Wang resin (100–200 mesh) were obtained from EMD Chemicals (Gibbstown, NJ). Fmoc-8-amino-3,6-dioxaoctanoic acid (PEG2) was obtained from Peptides International (Louisville, KY). Fmoc-DAP(ivDde)-OH was obtained from Bachem Americas (Torrance, CA). Other Fmoc-protected amino acids and coupling agents were obtained from Advanced ChemTech (Louisville, KY) or Chem-Impex International (Wood Dale, IL).
General SPPS Procedure
Swelling of the resin
The resin (50 µmol) was initially swelled for 15 min in 2 mL of 1:1 methylene chloride (DCM)/N,N-dimethylformamide (DMF) and drained. The resin was then washed once with 2 mL DMF.
Coupling of the amino acid (primary amine)
A Fmoc-protected amino acid was coupled to the deprotected N-terminal amine on the resin (1 eq.) using Fmoc-protected amino acid (3.3 eq.) in 0.5 mL DMF, 2-(1H-benzotriazol-1-yl)-1,1,3,3-tetramethyluronium hexafluorophosphate (HBTU) (3.3 eq.) in 0.5 mL DMF, 1-hydroxybenzotriazole (HOBt) (3.3 eq.) in 0.25 mL DMF and N,N-diisopropylethylamine (DIPEA) (6.6 eq.) in 0.25 mL DMF for 1 h at room temperature. The resin was then washed five times with 2 mL DMF.
Coupling of the amino acid (secondary amine)
An Fmoc-protected amino acid was coupled to the deprotected N-terminal amine of the resin (1 eq.) using Fmoc-protected amino acid (3.3 eq.) in 0.5 mL DMF, HBTU (3.3 eq.) in 0.5 mL DMF and DIPEA (6.6 eq.) in 0.25 mL DMF for 1 h at room temperature. The solvent was then drained and the same procedure was repeated once more. The resin was then washed five times with 2 mL DMF.
Deprotection of the Fmoc group
Fmoc deprotection after each amino acid coupling was accomplished using 1 mL of 20% piperidine in DMF for 1 × 10 min and 1 × 20 min at room temperature. The resin was then washed five times with 2 mL DMF.
Deprotection of the ivDde group
ivDde deprotection was accomplished using 1 mL of 2% hydrazine hydrate in DMF for 1 × 10 min and 1 × 20 min at room temperature. The resin was then washed five times with 2 mL DMF. 16
On-resin disulfide bond formation
On-resin oxidation of cysteine thiols was carried out by treating the acm-protected cysteines containing peptide-resin with thallium (III) trifluoroacetate (2 eq.) in 2 mL DMF for 1 × 60 min and 1 × 60 min at room temperature. The resin was then washed five times with 2 mL DMF. 17
End capping
End capping of the N-terminal amine was accomplished using 50 eq. of acetic anhydride and pyridine in 1 mL DMF for 30 min at room temperature. The resin was then washed five times with 2 mL DMF.
Cleavage of the peptide from the resin
Before cleavage, resin was washed five times with 2 mL DCM and dried for 30 min by nitrogen purging. Cleavage was performed by treating the peptide-resin with 2 mL of cocktail containing trifluoroacetic acid (TFA), ethanedithiol or triisopropylsilane, water, and thioanisol in a ratio of 90:5:2.5:2.5 for 4 h at room temperature. The peptide-containing cocktail was separated and collected in a test tube. The resin was then washed with 0.2 mL TFA, and the washings were added to the cocktail. The crude peptide was precipitated out by adding 0.5 mL of the cocktail to 50 mL of ice-cold methyl tert-butyl ether (MTBE) in a 50-mL falcon centrifuge tube. The tube was centrifuged at 2000 rpm for 10 min, and MTBE was carefully decanted each time. Finally, the crude peptide was washed twice with 50 mL MTBE and separated by centrifugation. The crude peptide was then dried by purging nitrogen for 2 h at room temperature and purified by high-performance liquid chromatography (HPLC).
HPLC conditions
HPLC solvents consisted of water containing 0.1% v/v TFA (solvent A) and acetonitrile containing 0.1% v/v TFA (solvent B). A Sonoma C18 column (ES Industries, West Berlin, NJ; 5 µm, 100 Å, 4.6 mm × 250 mm) was used with a flow rate of 1.5 mL/min. The HPLC gradient system began with an initial solvent composition of 95% A and 5% B for 2 min followed by a linear gradient to 50% A and 50% B in 15 min, after which the column was reequilibrated.
PepSy Design and Assembly
The fluidic flow diagram of PepSy is shown in Figure 1 . Figure 2 shows the electrical wiring schematic used for connecting all the electrical/electronic components of PepSy. All the components used for assembling PepSy are shown in Table 1 . Scripts to operate PepSy in a fully automatic mode or manual mode ( Fig. 3 ) were written in Python.
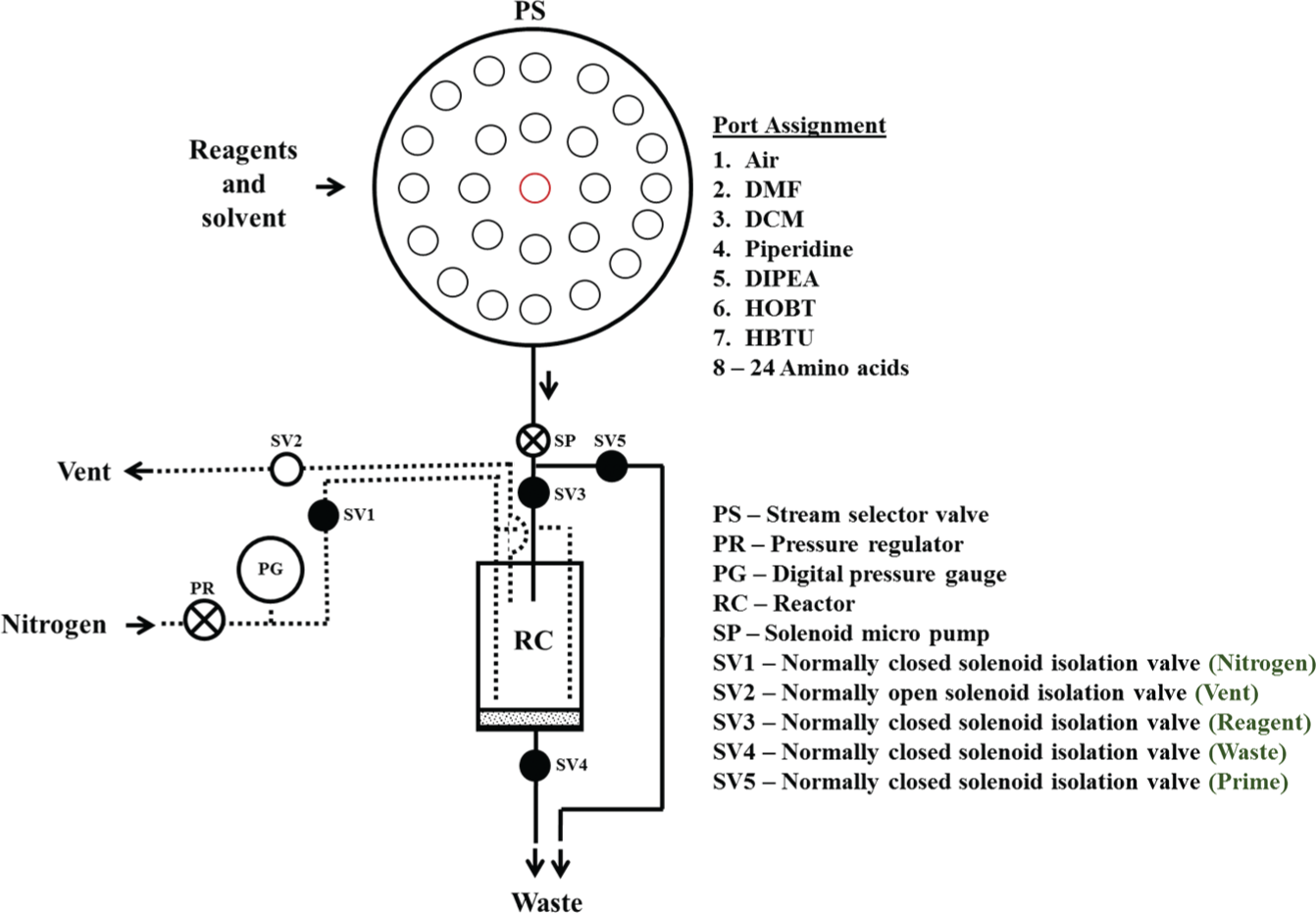
A fluidic flow diagram of PepSy. The reagents and solvents are delivered in precise volumes to the reactor (RC) by the solenoid micro pump (SP) via the VICI stream selector valve (PS). Mixing of the reaction mixture is achieved by the nitrogen bubbling. The nitrogen is also used to push the reagents and solvents from the reactor.
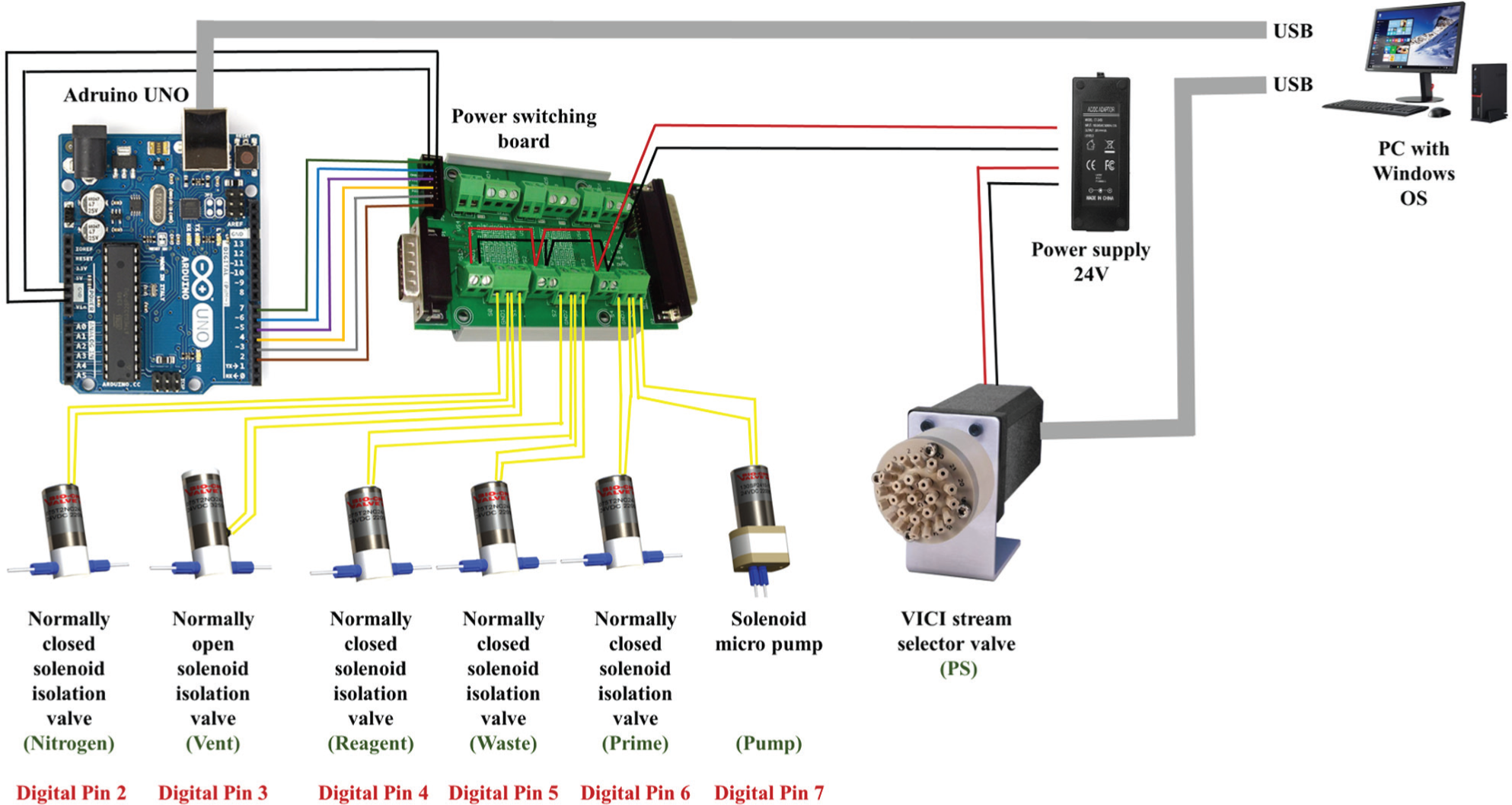
An electrical wiring diagram of PepSy. The solenoid isolation valves and the micro pump are connected to the digital I/O pins of the Arduino board via the power switching board. Both the Arduino board and the VICI stream selector valve are connected to the USB ports of a PC running Windows OS. The Arduino board is powered by the USB port itself, whereas the power switching board/solenoid valves/pump and the VICI stream selector valve are powered by an external 24-V DC power supply.
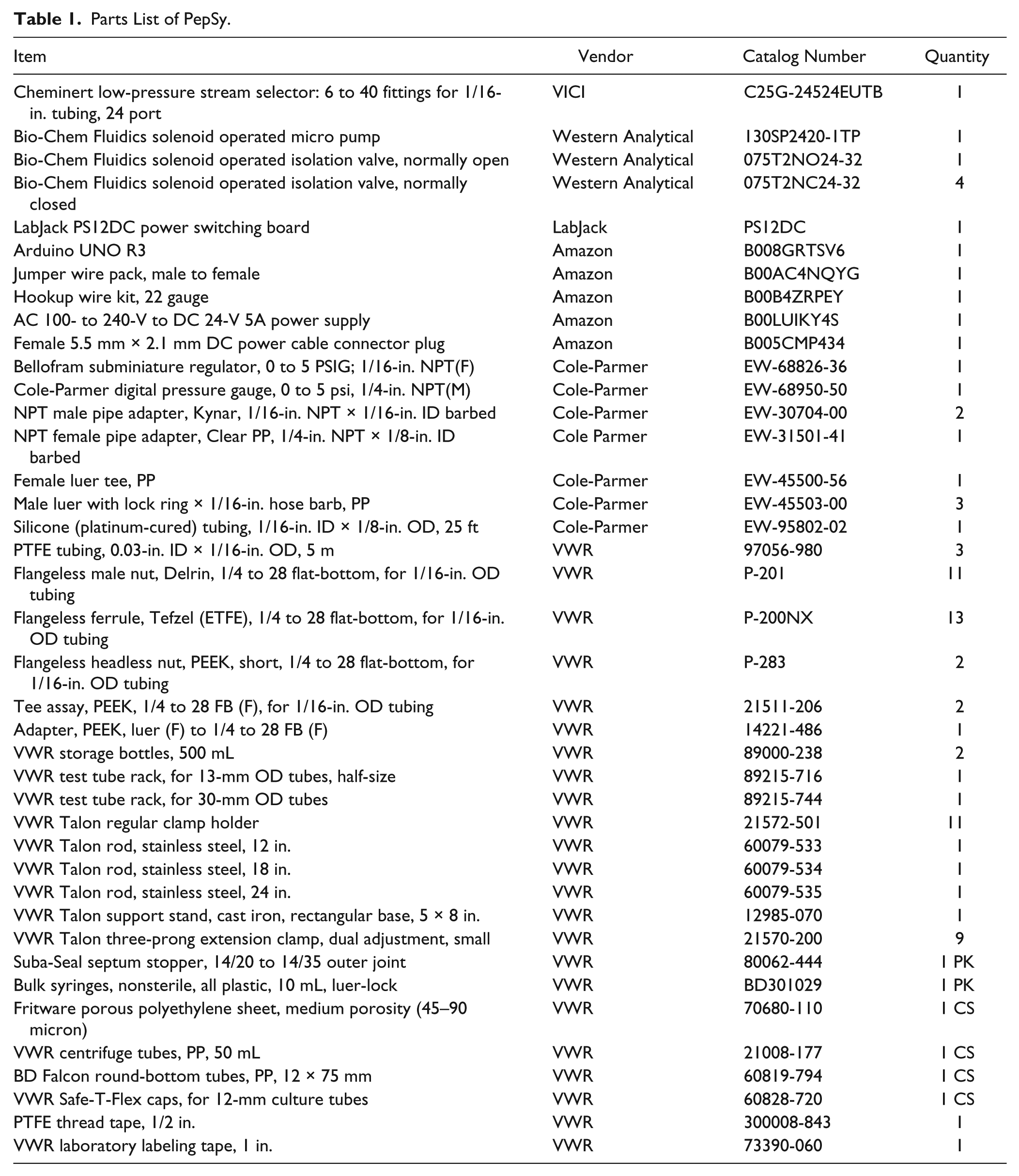
Parts List of PepSy.
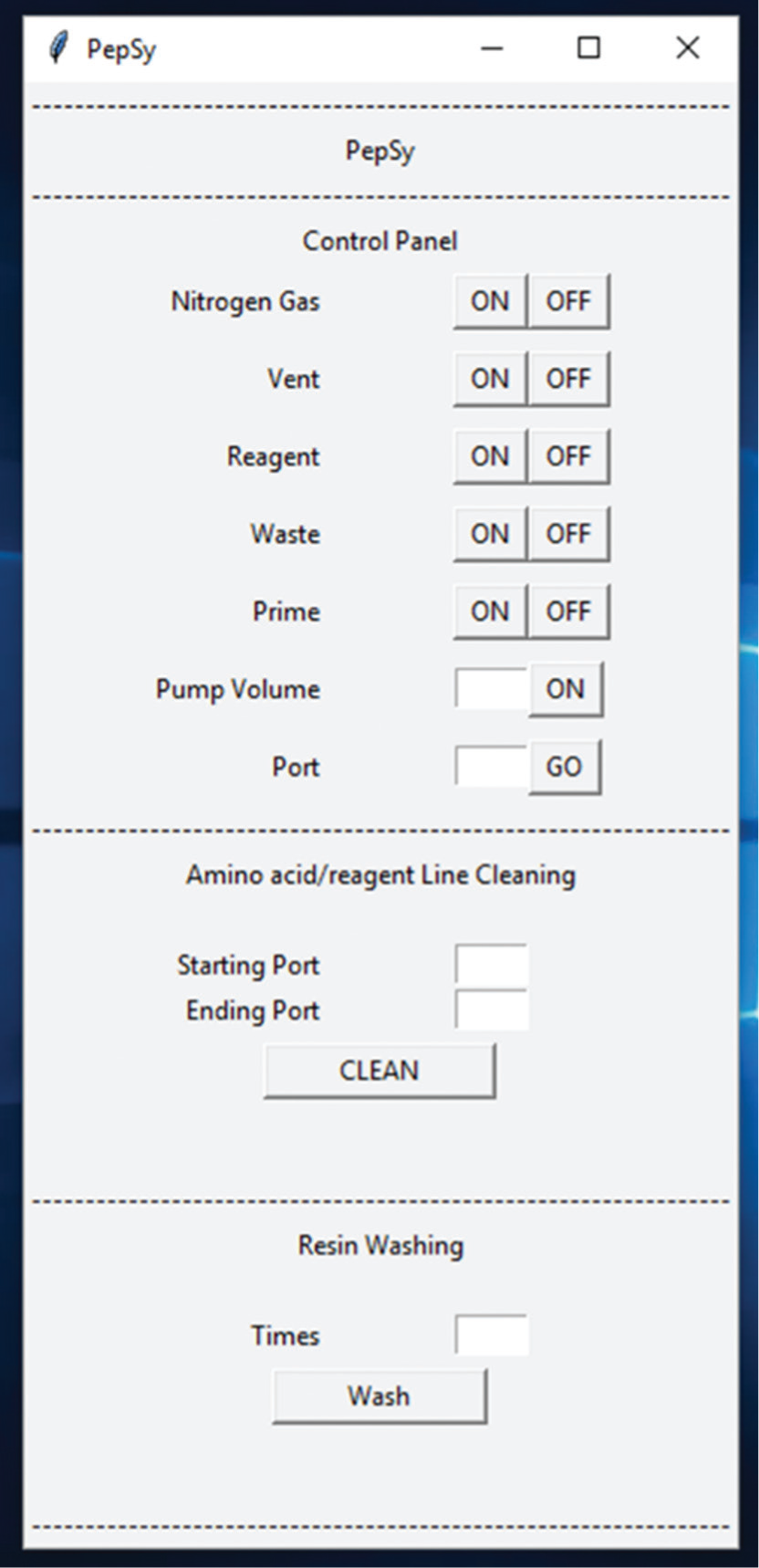
A Python/tkinter-based graphical user interface to manually control PepSy. The manual mode script was developed mainly to turn on or off a specific solenoid isolation valve, to change the VICI stream selector valve position, and to pump a specific volume of the reagent. It can be used to perform any one of the four unit operations (priming, reagent/solvent addition, mixing, or removing the reactants/solvents from the reactor and drying) by turning on the specific valve(s) and/or the pump.
Results and Discussion
PepSy was primarily designed to synthesize small peptides on a relatively small scale (<100 µmol). The current reaction conditions were optimized for a 50-µmol scale in our laboratory and tested for synthesizing peptides with up to 30 amino acid residues. 18 As shown in Figure 1 , all the reagents and solvents are delivered in precise volumes to the reactor (RC, a 10-mL disposable polypropylene syringe barrel fitted with a porous polyethylene frit) by a solenoid micro pump (Bio-Chem Fluidics, Boonton, NJ), labeled SP, via a VICI stream selector valve (Valco Instruments, Houston, TX), labeled PS. Mixing of the reaction mixture is achieved by bubbling nitrogen through the reaction mixture. Two nitrogen lines were used to achieve a homogeneous mixing during the reaction and washing steps. The nitrogen is also used for removing reactants/solvents from the reactor after each reaction and washing step by pushing them to the waste.
The SPPS method used by PepSy is mainly composed of four unit operations: (1) priming, (2) reagent/solvent addition, (3) mixing, and (4) removing the reactants/solvents from the RC and drying. For priming (filling the lines between the reagent/solvent bottle/tube and the SP), the actuator on the PS is moved to the corresponding position, and the solenoid isolation valve (Bio-Chem Fluidics), labeled Prime (SV5), is opened to enable flow of the reagent/solvent to the waste; other valves remain in their normal state. For the reagent/solvent addition, the actuator on the PS is moved to the corresponding position, and the solenoid isolation valve, labeled Reagent (SV3), is opened to enable the flow of the reagent/solvent to the RC; other valves remain in their normal state. For mixing, the solenoid isolation valve, labeled Nitrogen (SV1), is opened to enable flow of the nitrogen to the RC and escape through the vent while sparing the reaction mixture, and other valves remain in their normal state. For removing the reactants/solvents from the RC and drying, the SV1 is opened; the solenoid isolation valve, labeled Vent (SV2), is closed; the solenoid isolation valve, labeled Waste (SV4), is opened; and other valves remains in their normal state. This arrangement of solenoid valve states allows nitrogen to pass through the RC into the waste. A nitrogen pressure of ~2 psi was found to be optimal for mixing, removing reactants/solvents, and resin drying for 50-µmol scale synthesis. The PTFE tubing internal diameter (1/16 in.) and their lengths were kept as minimum as possible to minimize the reagent wastage during the priming step.
All solenoid isolation valves and the micro pump were connected to the digital I/O pins (only six used) of the Arduino UNO (http://arduino.cc/) via a power switching board (LabJack Corporation, Lakewood, CO) ( Fig. 2 ). The power switching board contains 12 digitally controlled switches (only six used) with LED indicators, resettable fuse protection, and flyback protection. Arduino board’s digital pins 2 to 7 were connected to the power switching board’s P3-14 pin header (pins DI0 to DI5, respectively), and both boards’ grounding pins were connected together. The SV1 to SV5 and SP were connected to the power switching board pins S0 to S5, respectively. Both Arduino board and the PS were connected to the USB ports of a PC running Windows OS. The Arduino board was powered by the USB port itself, whereas the power switching board/solenoid valves/pump and the PS were powered by an external 24-V DC power supply.
The open-source software used to operate PepSy in a fully automatic or manual mode was developed in Python. Python was chosen for writing scripts mainly because it is a dynamic high-level programming language that is easy to learn; scientists with no prior software programming knowledge can learn it fast, and it is freely available. 3 In addition, the Python scripts can be run on the most operating systems. Firmata was used to interact the Python script with the Arduino board, whereas the PS was directly controlled by the Python script via serial communication. The Python scripts to operate PepSy in a fully automatic (PepSy.py) and manual mode (PepSy-manual.py) are available to download at GitHub. 19 Instructions, scripts, device configuration file, and an example of the sequence configuration file are included in the supplemental material. Fully automatic mode script generates an output file for each run with a timestamp for each step. In addition, real-time status update of the step being carried out is displayed on the computer screen. Examples of an output file and the information displayed on the screen during a run are included in the supplemental material. The manual mode script displays a graphical user interface, as shown in Figure 3 . This script was developed mainly to turn on or off a specific valve, to change the VICI stream selector valve position, and to pump a specific volume of the reagent during the troubleshooting of any mechanical or electrical issues. However, it can be used to perform any one of the four unit operations (priming, reagent/solvent addition, mixing, or removing the reactants/solvents from the reactor and drying) by turning on the specific valve(s) and/or the pump. The peptide synthesis is carried out mainly in the automated mode. However, if the PepSy experiences an unexpected error at any stage while running in an automated mode, then the automated mode script can be closed and the sequence configuration file can be modified accordingly to begin the synthesis from where it stopped in the previous run. Fully automatic mode script includes functions to carry out resin swelling, resin washing, single coupling, double coupling, Fmoc deprotection, ivDde deprotection, on-resin oxidation, end capping, and amino acid/reagent line cleaning. Additional functions to carry out specific reactions can be added to the script by the user as needed.
The total cost of the components ( Table 1 ) to assemble PepSy ( Fig. 4a ) is <$4,000. PepSy uses disposables commonly available in the most chemistry laboratories such as culture tubes for holding amino acid solutions, and the reactor ( Fig. 4b ) is made of a disposable 10-mL BD syringe barrel, a 70-µm polyethylene frit, and a rubber septa. Operating cost of PepSy is very low due to use of the amino acids and other reagents purchased in bulk and low-cost disposables. In addition, all the components are readily available online and can be replaced very easily.
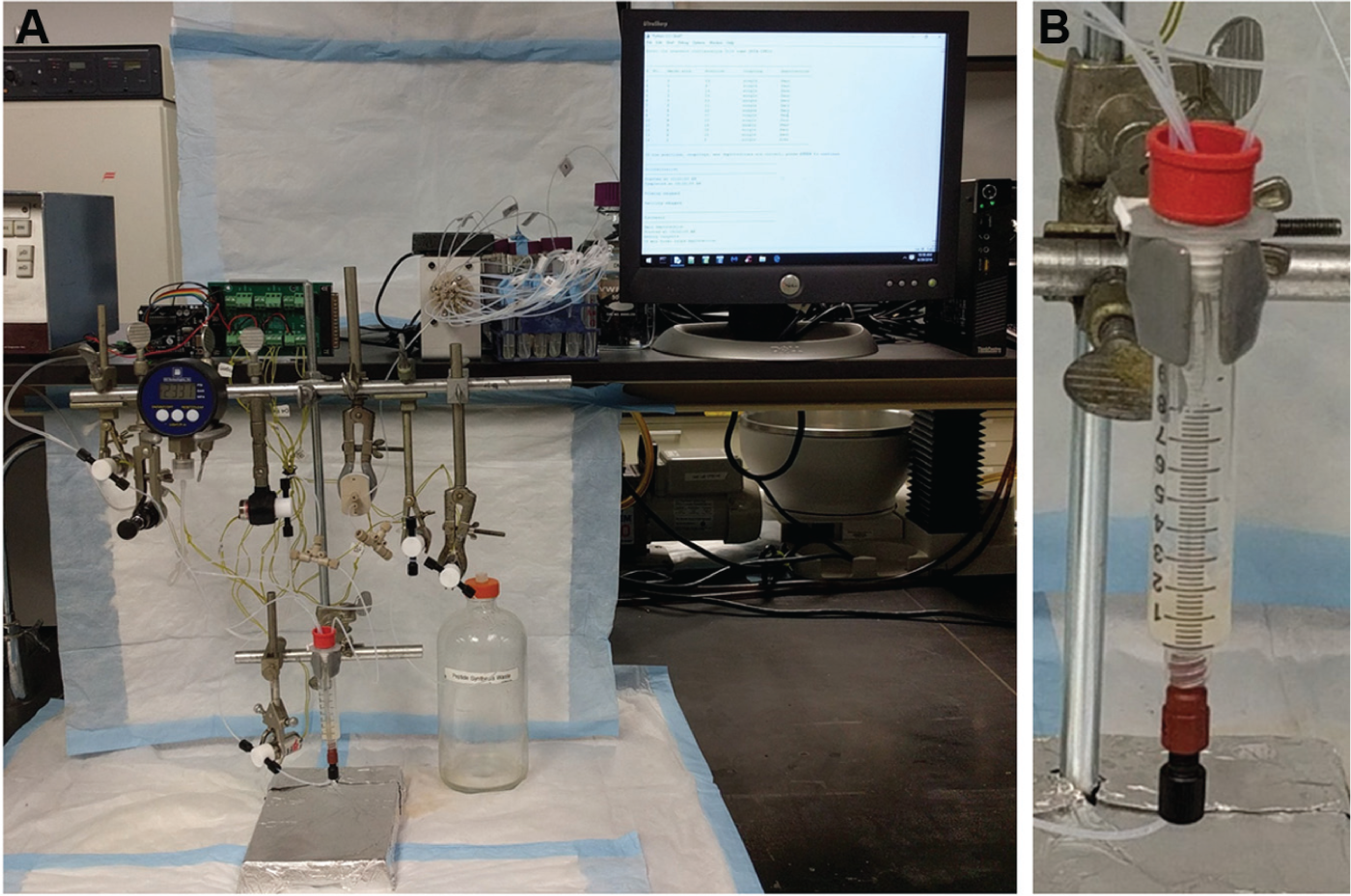

A photograph of (
Several small peptides and peptide conjugates were successfully synthesized on PepSy with reasonably good yields and purity depending upon the complexity of the peptide. Three examples are shown in Figure 5 . DOTA-PEG2-Bombesin(7-14) is a 10-residue linear peptide, whereas DOTATATE is a 9-residue cyclic peptide and used on-resin oxidation. Similarly, DOTA-PEG2-Probestin used orthogonal ivDde deprotection. The crude yields and purity were much better than the method done manually.
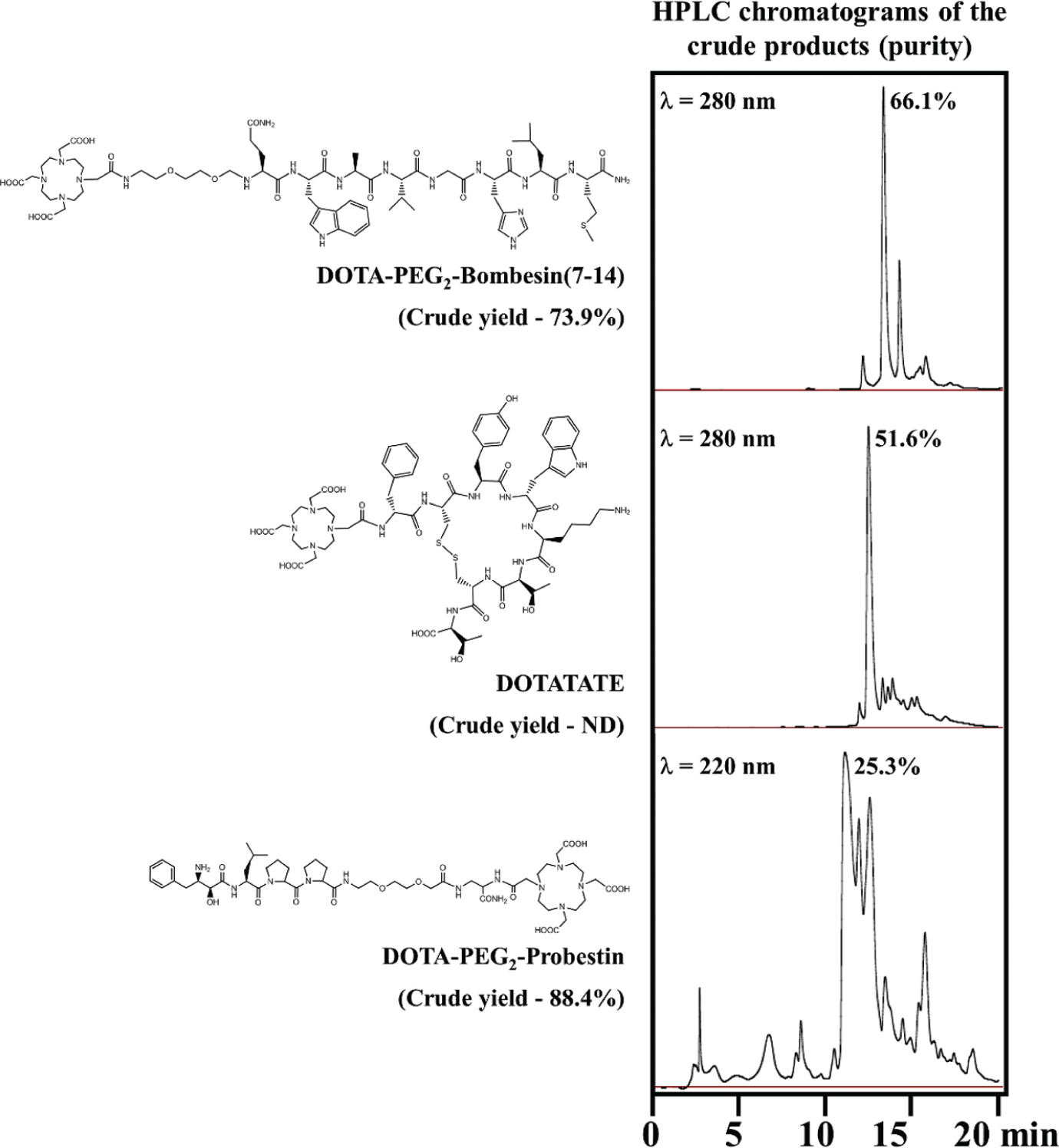
Examples of the peptides synthesized on PepSy. The crude product purity and yield of the peptides synthesized by the PepSy were similar or better than the peptides synthesized manually.
This open-source automated peptide synthesizer provides an encouragement to chemists to develop various automated synthesizers (e.g., to carry out radiochemistry).
Footnotes
Acknowledgements
The author gratefully acknowledges the assistance of Dr. Gopal Pathuri, Mr. Benjamin C. Anderson, and Ms. Vy Ho for synthesizing the peptides on the PepSy.
Supplementary material is available online with this article.
Declaration of Conflicting Interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the OU College of Pharmacy startup grant and an Institutional Development Award (IDeA) from the National Institute of General Medical Sciences of the National Institutes of Health under grant number 8P20GM103447.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
