Abstract
In modern laboratories, software can drive the full experimental process from data acquisition to storage, processing, and analysis. The automation of laboratory data acquisition is an important consideration for every laboratory. When implementing a laboratory automation scheme, important parameters include its reliability, time to implement, adaptability, and compatibility with software used at other stages of experimentation. In this article, we present an open-source, flexible, and extensible Python package for Laboratory Automation, Control, and Experimentation (PLACE). The package uses modular organization and clear design principles; therefore, it can be easily customized or expanded to meet the needs of diverse laboratories. We discuss the organization of PLACE, data-handling considerations, and then present an example using PLACE for laser-ultrasound experiments. Finally, we demonstrate the seamless transition to post-processing and analysis with Python through the development of an analysis module for data produced by PLACE automation.
Introduction
The open-source software (OSS) movement traces back to the 1950s, 1 when voluntary organizations such as Share—an intermediary between IBM and its users—pursued collaboration and reduced programming costs by leveraging diverse expertise and practices of academic exchange. 2 However, notable developments, such as the launch of the GNU Project in 1984 3 and Netscape’s decision to release the source code for its Internet browser in 1998, 1 significantly increased interest in the movement. More recently, examples such as the Linux operating system, 4 WordPress for blogging, 5 and Google Chrome for Internet browsing 6 have further demonstrated the potential for successful open-source projects. A majority of code developed under the umbrella of “open source” is distributed either free or for a nominal charge, but the primary distinguishing feature is the freedom to use, modify, and distribute source code under a public, unrestricted license.1,7
Laboratories built on the principles of the open-source movement have been in existence for decades. 8 Beyond the obvious benefit of reducing laboratory running costs, there are many advantages of an open-source laboratory. Collaborative development fuels software capabilities to be expanded beyond what is plausible through the efforts of a single development team. 9 When open-source code is effectively disseminated, there is a community of programmers available for testing. 10 Therefore, some of the burden of validation is taken over by the community, errors are reduced, and reliability and traceability are improved. 9 In the spirit of Eric Raymond’s proclaimed Linus’s Law, “given enough eyeballs, all bugs are shallow.” 11
Perspectives toward OSS are changing, as the tools we use become increasingly open source. Still, laboratory automation remains an uncommon field of open software development. 9 When proprietary automation packages are used, specialized software is often required to both acquire and read the data. This causes difficulties in exchanging data across computers or operating systems and incurs extra cost and licensing issues. 12 For collaborative research, it is beneficial for all contributors to use the same software. Open-source systems facilitate this with a minimal amount of cost and inconvenience. Altogether, it is favorable to use a software platform that is completely open source, from the acquisition software and data storage through the data analysis and postprocessing.
Here we present PLACE, an open-source Python package for Laboratory Automation, Control, and Experimenta-tion. 13 The package is available at https://www.github.com/johjam/PLACE and contains a core set of instrument driver modules for hardware used in the Physical Acoustics Laboratory ( Table 1 ). The software is currently used to automate laser-generated ultrasound experiments, including laser ultrasonics of heterogeneous materials, 14 biomedical photoacoustic imaging, 15 and fracture characterization. 16 However, PLACE accommodates a wide range of applications and laboratory schemes, as unique modules are written for each instrument.
Hardware Controlled by PLACE Automation.
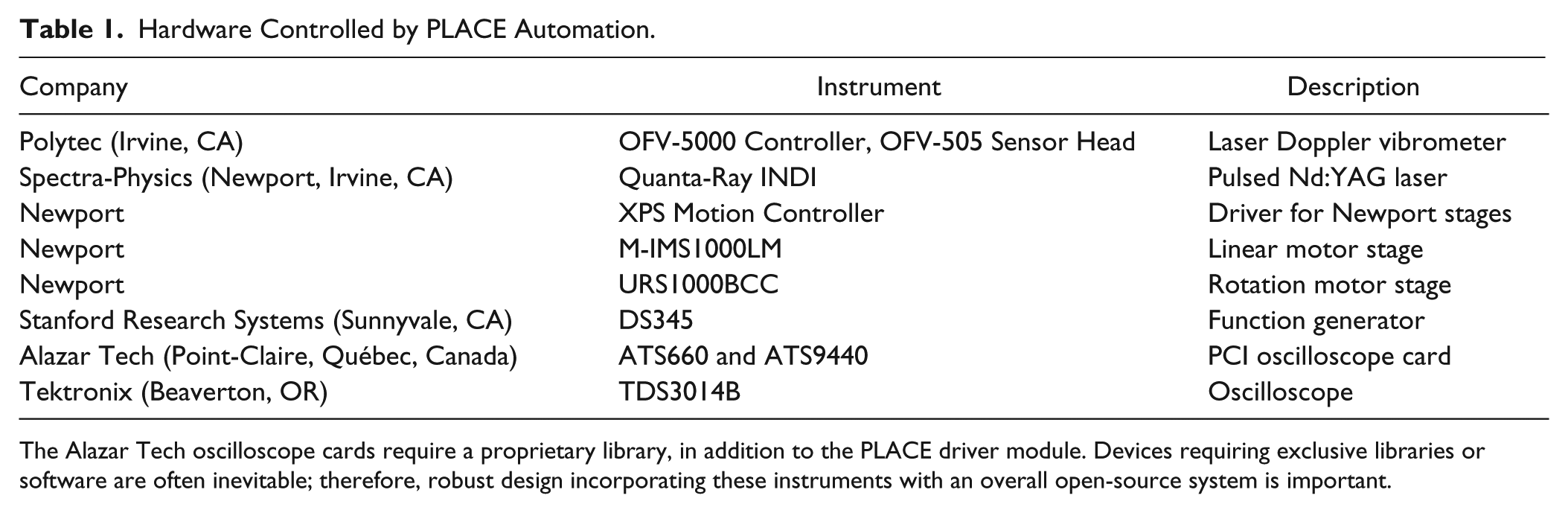
The Alazar Tech oscilloscope cards require a proprietary library, in addition to the PLACE driver module. Devices requiring exclusive libraries or software are often inevitable; therefore, robust design incorporating these instruments with an overall open-source system is important.
Python-based automation packages are in development, such as PyDAQmx, 17 pyVISA, 18 and Lantz, 19 which are designed to facilitate communication between a PC and laboratory instruments with Python. PLACE’s approach differs in that we extend our open-source approach to encourage compatibility across all aspects of experimentation. In addition to instrument drivers, a collection of visualization functions and processing software has been developed for data produced by PLACE automation. These functions can be called with a single line of code and provide interactive plotting and data-processing capabilities. PLACE drivers use communication protocols standard to most personal computers, further supporting portable and flexible use. The PLACE package is extensible, such that new instruments, packages, or protocols can be easily incorporated. For example, parallel processes can be incorporated by implementing a package such as Parallel Python. 20 Devices requiring proprietary software or libraries often cannot be avoided. However, whenever possible, we aim to develop open-source drivers that stand alone, rather than as wrappers for proprietary code. Naturally, this also requires careful hardware consideration. It is our goal to expand this repository through continued internal development and contributions from the broader community.
PLACE for Laboratory Automation
Python is a powerful, mature tool that possesses many advantageous qualities for laboratory automation. It is open source, freely available, and continually growing through the contribution of an international community of developers. Guido van Rossum, the founder of Python, was motivated by the theory that “readability counts” 21 ; thus, Python syntax was designed with clarity at the forefront. 22 The standard library is well debugged and equipped with extensive built-in modules. Python is flexible and extensible, competes among the fastest languages for computation speed, and requires minimal lines of code for high productivity. 22 The cost for numerical computing of large data sets has been shown to be less than 25% higher than pure Fortran code. 23 Furthermore, automation packages written in Python are virtually platform independent, as most operating systems—including Linux platforms, Windows, and Mac OS X—are supported.
The organization of complex code can be streamlined using Python’s object-oriented approach. As illustrated in Figure 1 , PLACE automation is organized with a unique driver module for each instrument. These modules contain the fundamental code for communication and control with a given instrument. Further hierarchical organization is accomplished with the use of classes, subclasses, and functions. A master script imports the driver modules and coordinates the activity of all instruments to perform the tasks required for a given protocol. This object-oriented approach seamlessly incorporates new instruments or methodologies as a laboratory grows and advances.
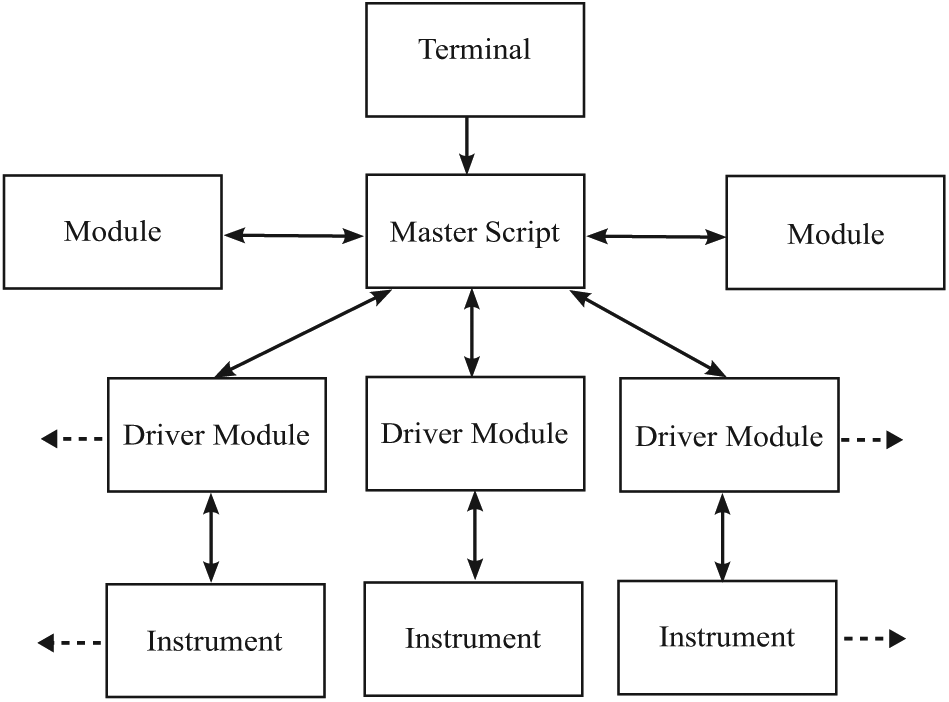
Basic organization of PLACE scripts and modules. Experimental protocols are defined in the terminal. The master script communicates with driver modules developed inhouse to automate laboratory instruments. Python modules available from repositories or built in to Python are used to complete tasks such as plotting, saving data, and serial communication.
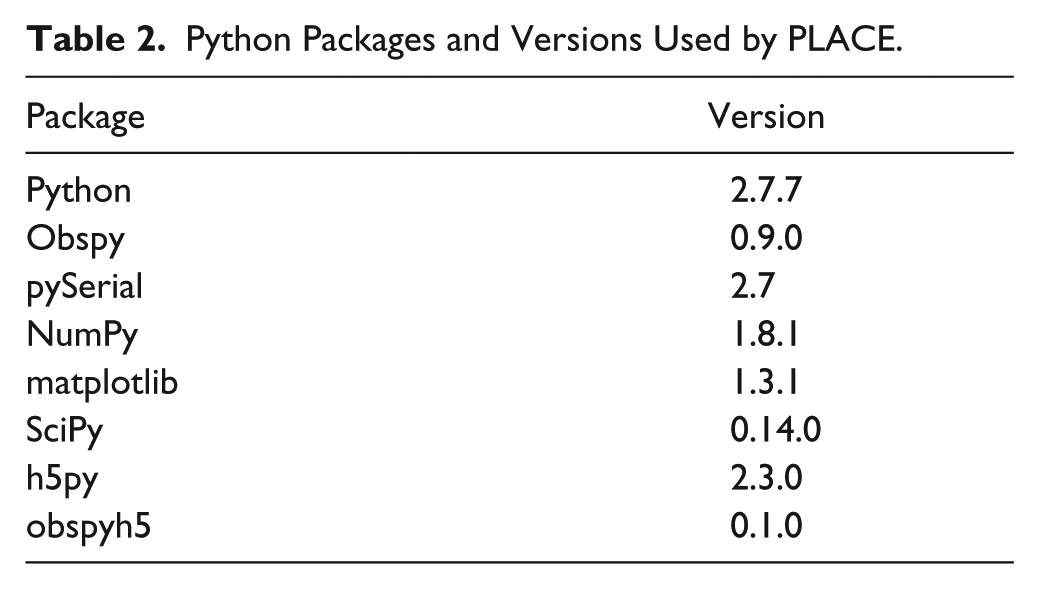
The PLACE modules developed inhouse rely on well-established modules in Python repositories ( Table 2 ). These include NumPy for scientific computing 24 and matplotlib for plotting. 25 Data are transmitted between the acquisition PC and instruments via RS-232, ethernet, and USB using pySerial. 26 ObsPy is used to define and save header information (metadata), store data in a list-like object known as a stream, 27 and append data to a file dynamically. Saving data in an ObsPy-compatible format during acquisition allows us to make full use of ObsPy’s capabilities for postprocessing and analysis. ObsPy is designed for processing seismological data but has powerful functionality for processing waveforms or time series of all kinds. The package continues to develop at a rapid pace, yet ObsPy’s test-driven approach 28 has yielded stable releases on a regular basis.
Python Packages and Versions Used by PLACE.
Data Handling
Robust data handling is essential for automated laboratory processes to ensure integrity, accountability, and preservation of data. 28 We ensure effective data management by incorporating detailed header information, using an efficient and well-established data format, and dynamically saving data to file as it is acquired.
A custom header format was created, which saves the unique parameters for each trace (A-scan) as it is acquired. We chose to save metadata in a header during acquisition, as opposed to using a database, with goals of convenience and robust coupling of the metadata to the data it describes. Headers standardize laboratory records, simplify postprocessing, and essentially serve as an electronic laboratory notebook. As an added benefit, both PLACE and ObsPy functions automatically read and use header parameters, minimizing the need to enter arguments manually. For example, a voltage is acquired from our vibrometer receiver. The calibration factor required to convert the voltage to physical units, such as displacement or particle velocity, is dependent on the choice of decoder and settings of the vibrometer receiver. These values are held by the vibrometer hardware, which is queried during acquisition and recorded in the header. The ObsPy read function automatically applies this calibration factor upon import. All ObsPy processing that is applied to a stream is also recorded in the header, which further supports accurate record keeping and reproducibility. Example header information for a single trace is shown below.
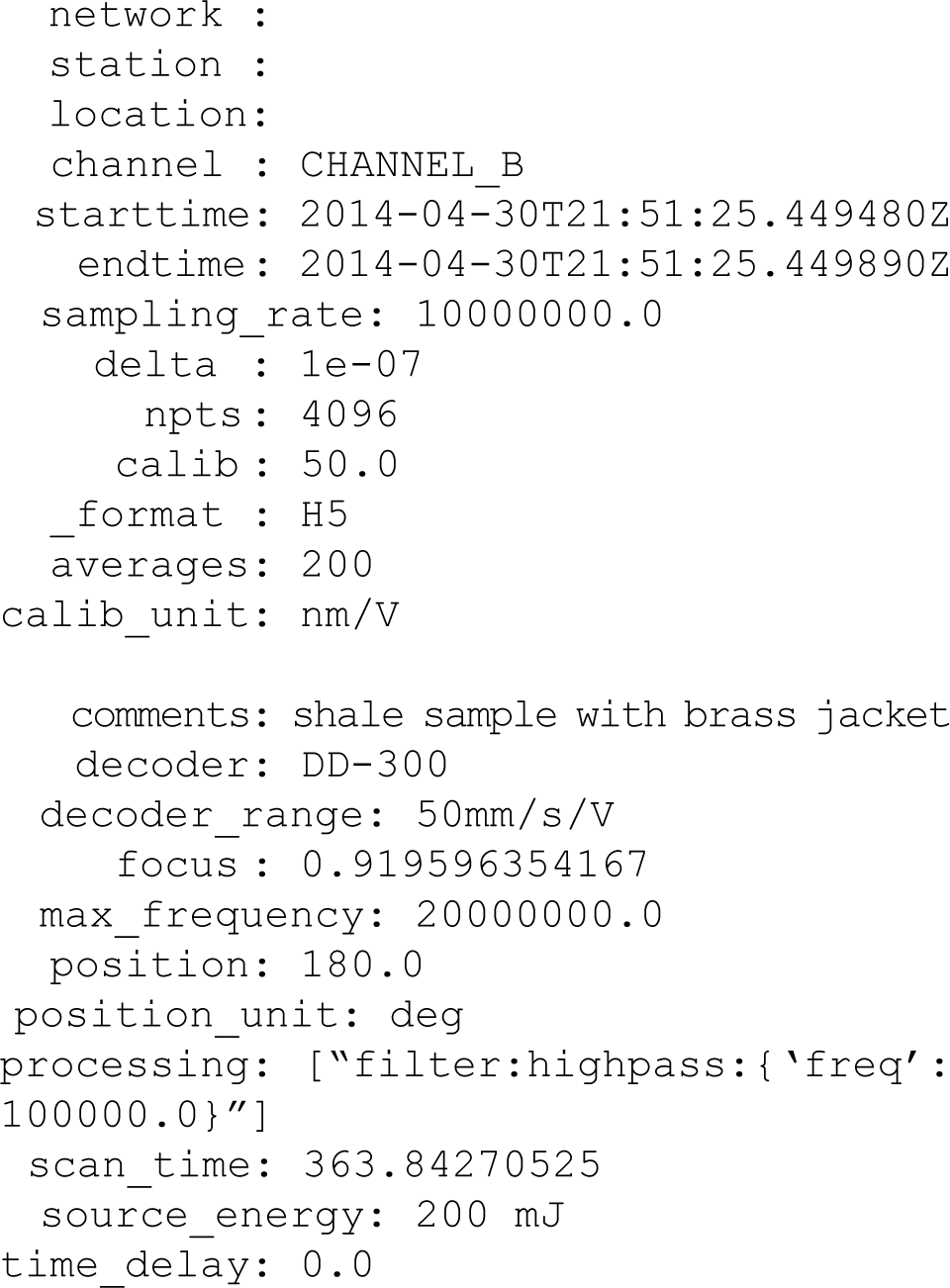
Data are saved in HDF5, the free and open standard hierarchical data format. HDF5 is designed for large quantities of complex data and efficient I/O. In addition, HDF5 is portable across operating systems 29 and supported by most programming languages, including C++, java, MATLAB, and Scilab.
Traces with headers are saved dynamically to an HDF5 file through the ObsPy, H5py, 30 and obspyh5 31 packages. Within the file, traces are appended to a stream. Saving continuously reduces concerns with limited buffer memory and ensures data are preserved if acquisition is disturbed. If an experiment must be interrupted, automation can be paused and restarted from the last position without creating a new data file. In addition, multiple experiments with different protocols can be saved to a single file. The unique header for each trace ensures the discernibility of the experiments. Furthermore, when only the metadata are required, Obspy can efficiently load the trace headers, without unpacking the waveform data.
Implementation of PLACE Automation
An example using PLACE for a laser-ultrasonics experiment is presented in Figure 2 . First, all options are defined in the terminal and imported to the master script. Subsequently, the instruments are initialized and header information is defined. To perform specific tasks, the master script interfaces with individual modules. For example, prior to recording a trace, the master script reads the signal strength of the vibrometer sensor head through the oscilloscope card module. If the signal is suboptimal, the vibrometer module is employed to focus the sensor head until a satisfactory signal level is reached. Subsequently, the trace is recorded with the oscilloscope card module. The frequency content of laser-generated ultrasound experiments ranges across the ultrasound spectrum, with up to tens of megahertz generated in applications such as biomedical photoacoustic imaging. We are limited strictly by the receiver hardware, as acquisition with PLACE and the oscilloscope card allow frequencies up to 125 MHz to be recorded. Once acquired, the trace and header are saved to an HDF5 file and displayed.
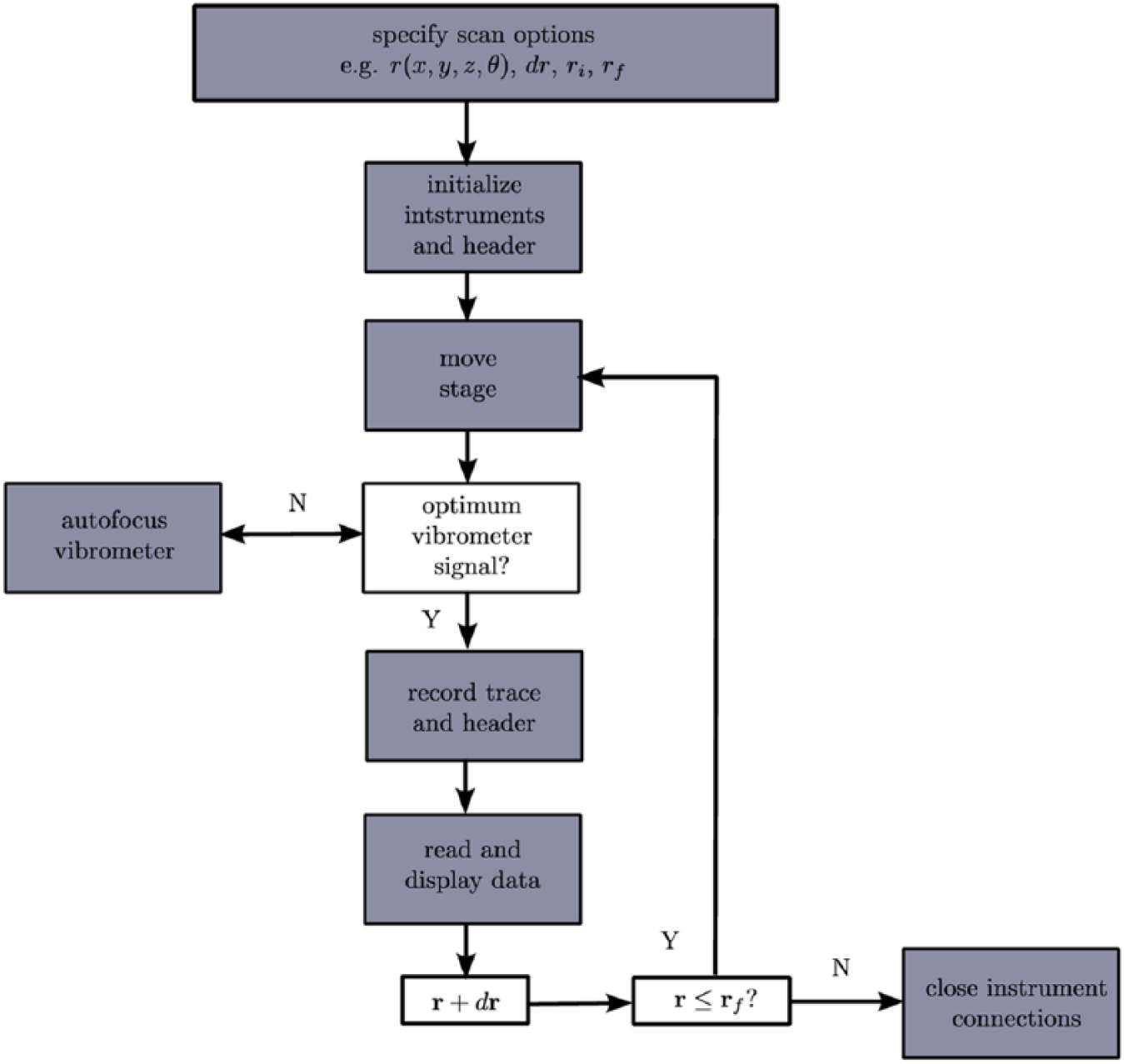
Example implementation of PLACE automation for a laser-ultrasound scan, where
Throughout an experiment, progress can be monitored both via the terminal output and graphically ( Figure 3 ). After each trace is acquired, the most recent trace and the cumulative wavefield are displayed with matplotlib. The master script prints updates to the terminal with each trace, including the approximate time remaining, current trace number and stage position, and notifications when the vibrometer is refocused. The experiment continues until conditions for completion are met.
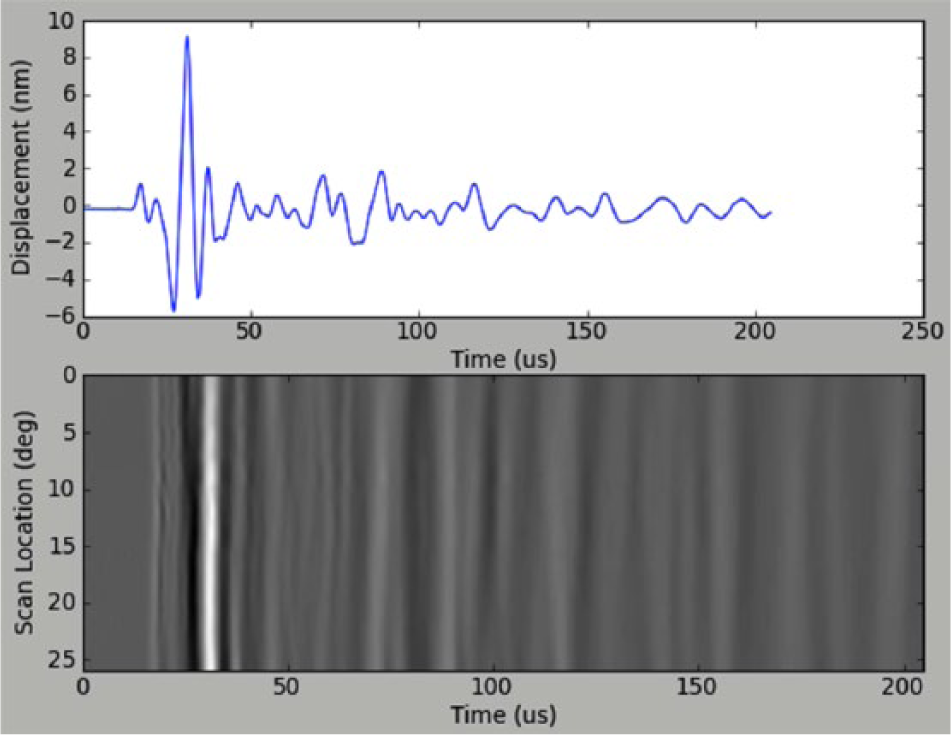
Screenshot of the most recent trace and cumulative wavefield during laser-ultrasound scan of a sandstone sample.
The setup for a rotational scan of a shale cylinder with P-wave anisotropy is shown in Figure 4 . The source and receiver were aligned and stationary. The shale was mounted on a 360° continuous rotational stage, and waveforms were acquired over 180°. All experimental parameters were defined in the terminal, and the master script was executed using the following line of code:
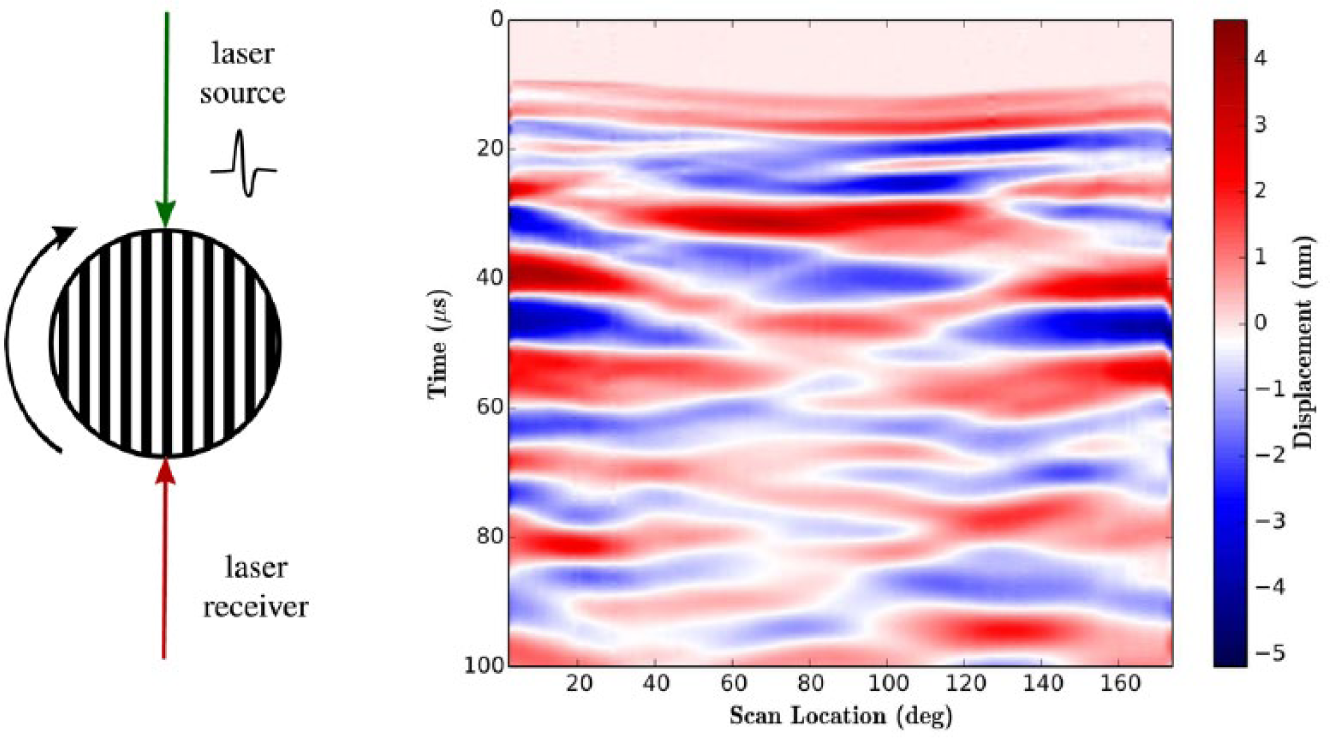
(

where each parameter is defined in the example Scan.py file provided with the PLACE package. The resulting waveform data are shown in Figure 4 .
PLACE Data Analysis
The first release of PLACE includes an analysis module, PALplots. Several visualization functions are incorporated: a contour plot for viewing the raw waveforms ( Fig. 4 ), a “wiggle” plot that highlights the location of dominant events, and a function for viewing the frequency-wavenumber (f-k) spectrum. In addition, an f-k filter is included for suppressing unwanted or interfering waves within a range of apparent velocities. A wiggle plot and f-k spectrum for the shale data are shown in Figure 5 .
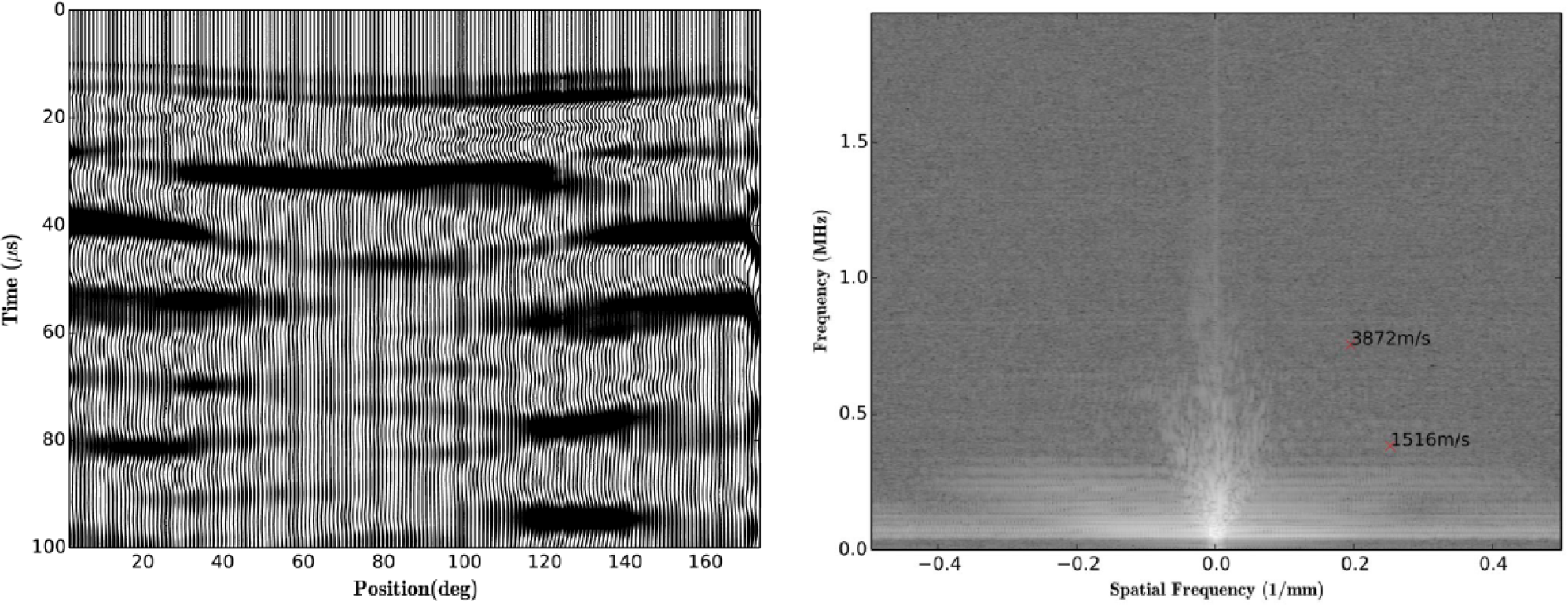
Example plots for the shale sample with brass jacket created with the PALplots module. (
Each PALplots function provides the user with the ability to interactively select points in the figure. For the
Conclusions and Future Work
PLACE is an open source, freely available Python package for laboratory automation and analysis. The clean, object-oriented design provides the freedom to expand and customize PLACE to meet the needs of diverse laboratories and experiments. Future work includes the development of driver modules for scanning mirrors, amplifiers, and additional stages for three-dimensional scanning. We plan to expand the PLACE library of processing software for laser-generated ultrasound experiments, as well as integrate code for resonant ultrasound spectroscopy. Our open-source approach spans across the spectrum of experimentation and has also fueled the development of open-source hardware. This includes projects such as seismometers for primary and secondary education 32 and an open-source laser Doppler vibrometer. PLACE has been thoroughly tested under the Linux operating system and has shown functionality across platforms. However, future work includes additional testing under Windows and Mac OS X operating systems. We aim to actively develop additional modules and software and we encourage contribution from the community.
Footnotes
Acknowledgements
J.L.J. thanks the University of Auckland Doctoral Scholarship for doctoral study support. The authors would like to give a special thanks to Dr. John Scales, Colorado School of Mines, for input that has improved this article and PLACE, as well as the naming of the package. In addition, thanks to Dr. Thomas Lecocq, the Royal Observatory of Belgium, for his helpful suggestions.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: J.L.J. thanks the University of Auckland Doctoral Scholarship for doctoral study support.
