Abstract
Practical cell-based assays can accelerate anti-Zika (ZIKV) and anti-dengue (DENV) virus drug discovery. We developed an immunodetection assay (IA), using a pan-flaviviral monoclonal antibody recognizing a conserved envelope domain. The final protocol includes a direct virus yield reduction assay (YRA) carried out in the human Huh7 cell line, followed by transfer of the supernatant to a secondary Huh7 culture to characterize late antiviral effects. Sofosbuvir and ribavirin were used to validate the assay, while celgosivir was used to evaluate the ability to discriminate between early and late antiviral activity. In the direct YRA, at 100, 50, and 25 TCID50, sofosbuvir IC50 values were 5.0 ± 1.5, 2.7 ± 0.5, 2.5 ± 1.1 µM against ZIKV and 16.6 ± 2.8, 4.6 ± 1.4, 2.6 ± 2.2 µM against DENV; ribavirin IC50 values were 6.8 ± 4.0, 3.8 ± 0.6, 4.5 ± 1.4 µM against ZIKV and 17.3 ± 4.6, 7.6 ± 1.2, 4.1 ± 2.3 µM against DENV. Sofosbuvir and ribavirin IC50 values determined in the secondary YRA were reproducible and comparable with those obtained by direct YRA and plaque reduction assay (PRA). In agreement with the proposed mechanism of late action, celgosivir was active against DENV only in the secondary YRA (IC50 11.0 ± 1.0 µM) and in PRA (IC50 10.1 ± 1.1 µM). The assay format overcomes relevant limitations of the gold standard PRA, allowing concurrent analysis of candidate antiviral compounds against different viruses and providing preliminary information about early versus late antiviral activity.
Introduction
Dengue (DENV) and Zika (ZIKV) viruses are related members of the Flaviviridae family, transmitted by mosquitoes of the Aedes genus.1–3 Multiple factors, such as globalization, 4 environmental changes favoring reproduction of the vector, 5 and viral adaptation to the urban setting, 6 have recently spread these viruses to novel areas. DENV is the most prevalent arboviral infection in humans, as indicated by the World Health Organization (WHO) (https://www.who.int/denguecontrol/disease/en/), causing severe flu-like illness and occasionally lethal dengue hemorrhagic fever or dengue shock syndrome. Over the last 50 years, the incidence of DENV has increased dramatically with an estimated 400 million new infections per year occurring mainly in tropical and subtropical areas. 1 Since the first recognized large outbreak of ZIKV in Micronesia in 2007, ZIKV has also spread rapidly to many countries in the Americas affecting millions of individuals. The association of ZIKV infection with Guillain-Barré syndrome in adults and congenital brain abnormalities in newborn infants, 7 established during the last Brazilian outbreak, has renewed the interest in ZIKV. Consequently, the WHO has ranked DENV as the most critical mosquito-borne viral disease and ZIKV as an international public health emergency.
Despite the urgent need for effective treatment, no specific antiviral therapy is available to control ZIKV or DENV infection and transmission.8,9 In addition, increasing rates of co-infections with different flaviviruses co-circulating within the same vector complicate the clinical outcome and treatment options. 10 Potential targets for antiflavivirus compounds include viral proteins, such as protease or polymerase, and host cell functions essential for virus replication, such as α-glucosidase and proteins involved in nucleoside biosynthesis.11,12
High-throughput screening (HTS) of libraries of small molecules is a powerful tool to identify novel flavivirus inhibitors;13–15 however, measurement of virus replication can be cumbersome, expensive, and prone to inaccuracy. To date, a variety of methods have been developed, including the classical plaque reduction assay (PRA),16–18 microscopy monitoring of cytopathic effect (CPE), 19 and immunofluorescence-based assays such as the fluorescence focus assay and the most advanced fluorescence-activated cell sorting assay.20,21 Cell-based assays using live viruses, such as PRA or CPE, are indicated as the reference standard for antiviral screening, despite poor reproducibility, the requirement of experienced technicians, and high-turnaround times. 8 Consequently, the development of accurate, easy-to-perform, and fast cell-based assays is highly valuable to test candidate inhibitors of ZIKV and DENV replication.
In this study, we describe a fast and accurate cell-based flavivirus immunodetection assay (IA) allowing quantification of ZIKV and/or DENV antigen by a specific monoclonal antibody to the fusion loop of the E protein domain II, which is shared among different flaviviruses. The assay is applied as a readout of a direct yield reduction assay (YRA) measuring inhibition of virus replication in the initially infected cell culture. In addition, viral stocks generated in the direct YRA can be transferred to a second cell culture in the absence of drug, to better characterize antiviral activity exerted at steps occurring later than envelope expression. To validate the assay, sofosbuvir and ribavirin half-maximal inhibitory concentrations (IC50) were determined and compared with values obtained by a standardized PRA 22 and with values previously reported in the literature.23–26 To evaluate the ability of the system to discriminate between early and late antiviral effects, the IC50 of celgosivir, an α-glucosidase inhibitor acting at late steps of DENV infection and recently evaluated in a phase Ib/IIa randomized clinical trial (NCT01619969),27,28 was determined by both a direct and a secondary YRA, as well as by the reference PRA against both viruses. In the literature, celgosivir anti-DENV effects were also determined in vitro29,30 and in animal models. 31 Even though a possible activity of celgosivir against ZIKV has been hypothesized based on the high similarity between ZIKV and DENV, 28 in a recently published work 32 celgosivir was not active in vitro against ZIKV when a monkey cell line (VERO) was used.
Materials and Methods
Cells
Vero E6 (African green monkey kidney cell line; ATCC, Manassas, VA, USA, CRL-1586), A549 (human lung carcinoma cell line; ATCC CCL-185), Huh7 (human hepatoma cell line; kindly provided by Istituto Toscano Tumori, Core Research Laboratory, Siena, Italy), and LN-18 (glioblastoma cell line; ATCC CRL-2610) cells were used to titrate ZIKV and DENV viral stocks by IA. The C6/36 (Aedes albopictus mosquito; ATCC CRL-1660) cell line was used to expand DENV, and the VERO E6 cell line was used to expand ZIKV. The cell propagation medium was Dulbecco’s modified Eagle’s medium (DMEM), high glucose with sodium pyruvate, and
Viruses
The H/PF/2013 ZIKV strain, belonging to the Asian lineage, and the New Guinea C DENV serotype 2 strain were kindly provided by the Istituto Superiore di Sanità, Rome, Italy. Once expanded in VERO E6 (ZIKV) and C6/36 (DENV) cells, viral stocks were titrated by plaque assay 22 in A549 and VERO E6 cells, yielding viral titers of 400,000 and 20,000 plaque-forming units (PFU) per milliliter, respectively. Briefly, confluent cells in six-well plate format were infected with three 10-fold dilutions of viral stock, and after 1 h viral adsorption at 37 °C with 5% CO2, cells were washed with PBS and infection medium with 0.75% Sea Plaque Agarose (Lonza, Rockland, ME, USA) was added to each well. After 5 days’ incubation at 37 °C, the monolayers were fixed with 10% formaldehyde (Carlo Erba Chemicals, Milan, Italy) and stained with 0.1% crystal violet (Carlo Erba Chemicals). After at least 3 h of incubation, the agar overlay was removed by water washing and PFU were counted.
Antivirals
The FDA-approved anti-hepatitis C virus compounds so-fosbuvir (β-
Cytotoxicity Assay
Serial twofold dilutions of antivirals in infection medium (propagation medium supplemented with 1% FBS) were added to Huh7 cells seeded at 7000 cells/well in a 96-well plate. After 72 h of incubation, drug cytotoxicity was measured by using the CellTiter-Glo 2.0 Luminescent Cell Viability Assay (Promega, Madison, WI, USA) according to the manufacturer’s protocol. The luminescent signal generated by cells treated with the test compound was compared with that generated by cells treated with DMSO/water to determine the half-maximal cytotoxic concentration (CC50).
Setup of the Immunodetection Assay
Optimal experimental conditions for the detection of viral antigen by IA were defined by growing viral stocks in human cell lines (A549, Huh7, and LN-18) and in the reference monkey line (VERO E6) that were titrated at 48, 72, and 96 h. The day before infection, each cell line was seeded in a 96-well plate format at the appropriate concentration to obtain 90% confluence at the time of antigen detection. Serial twofold dilutions of viral stocks were adsorbed to target cells in quadruplicate for 1 h at 37 °C in a humidified incubator with 5% CO2. After removal of the virus inoculum, DMEM infection medium with 1% or 3% FBS was added to cultures to be maintained for 48/72 h or 96 h, respectively.
For the immunodetection of virus antigen, the supernatant was removed and cells were fixed for 30 min with 10% formaldehyde (Carlo Erba Chemicals), rinsed with 1% PBS, and permeabilized for 10 min with 1% Triton X-100 (Carlo Erba). Following washing with PBS containing 0.05% Tween 20 (Carlo Erba Chemicals), cells were incubated for 1 h with monoclonal antiflavivirus mouse antibody (clone D1-4G2-4-15; Novus Biologicals, Centennial, CO, USA, NBP2-52709) diluted 1:400 in blocking buffer (PBS containing 1% BSA and 0.1% Tween 20). After washing four times, cells were incubated for 1 h with a polyclonal horseradish peroxidase (HRP)-coupled anti-mouse IgG secondary antibody (Novus Biologicals NB7570) diluted 1:10,000 in blocking buffer. Next, cells were washed five times and the 3,3′,5,5′-tetramethylbenzidine substrate (Sigma Aldrich) was added to each well. After 15 min of incubation in the dark, the reaction was stopped with one volume of 0.5 M sulfuric acid. All incubation steps were performed at room temperature. Absorbance was measured at 450 nm optical density (OD450) using the Absorbance Module of the GloMax Discover Multimode Microplate Reader (Promega) and adjusted by subtracting the background value, established as twofold the mean OD450 value of quadruplicate uninfected cells. The 50% tissue culture infectious dose (TCID50) of each virus was calculated according to Reed and Muench. 33
Direct Yield Reduction Assay
The direct YRA is based on the infection of cells in the presence of serial drug dilutions followed by absorbance measurement by IA. Since the readout is based on the detection of the E protein, the system allows us to measure interference with the virus life cycle up to protein production but not at later steps. To define the optimal virus inoculum, 7000 Huh7 cells/well were infected with ZIKV or DENV at 100, 50, and 25 TCID50, as determined by the IA described above. Viral adsorption was performed in 96-well plates for 1 h at 37 °C with 5% CO2. After virus removal, serial dilutions of sofosbuvir or ribavirin were added to the cell media at final concentrations ranging from 0.03 to 100 µM and the plates were incubated at 37 °C with 5% CO2. All drug concentrations were tested in triplicate and three independent experiments at each TCID50 used were performed to determine the assay reproducibility. Infected and uninfected cells without antivirals were used to calculate 100% and 0% of viral replication, respectively. After 72 h, supernatants were harvested and stored at −80 °C for subsequent analysis, and IA was performed on cell monolayers as described above. Based on initial experiments, each IA run was validated when the OD450 value in the virus control culture was above 1. This value was taken as 100% replication and IC50 values were calculated based on this reference by a nonlinear regression analysis of the dose–response curves generated with the GraphPad PRISM software version 6.01 (La Jolla, CA, USA). The activity of celgosivir against ZIKV and DENV was determined by YRA with 50 TCID50 as described above.
Secondary Yield Reduction Assay
The secondary YRA is designed to measure viral protein production driven by the virus generated in the first round in the presence of drug. Thus, antiviral effects exerted at late steps of the virus life cycle, for example, virus glycosylation and assembly, not detected by the direct YRA, can be measured. The secondary YRA was carried out by infecting 7000 Huh7 cells/well in a 96-well plate with ZIKV and DENV viral supernatants generated by direct YRA with reference compounds. Triplicate viral stocks derived from the direct YRA were used and two independent runs of the secondary YRA were performed to assess the reproducibility of results. After 72 h of incubation at 37 °C with 5% CO2, cells were fixed, and IA was performed to determine the IC50 value for each drug as described in the “Direct Yield Reduction Assay” section (
Plaque Reduction Assay of ZIKV and DENV on Reference Compounds
The PRA on reference compounds was performed as previously described.
22
Briefly, Huh7 cells were infected with ZIKV or DENV at 0.1 multiplicity of infection (MOI), as determined by plaque assay quantification, in the presence of serial fivefold drug dilutions, with a final drug concentration ranging from 0.03 to 100 µM for sofosbuvir and ribavirin and from 0.02 to 50 µM for celgosivir. After 72 h of incubation, three 10-fold dilutions of cell supernatant were used to infect in duplicate A549 (ZIKV) and VERO E6 (DENV) cells. Each experiment included a positive control (original viral stock) and a mock-infected well with infection medium only (
Results
Choice of Cell System and Incubation Time for IA
Titration of ZIKV and DENV viral stocks by IA was possible at 48, 72, and 96 h in VERO E6 and Huh7 cell lines (
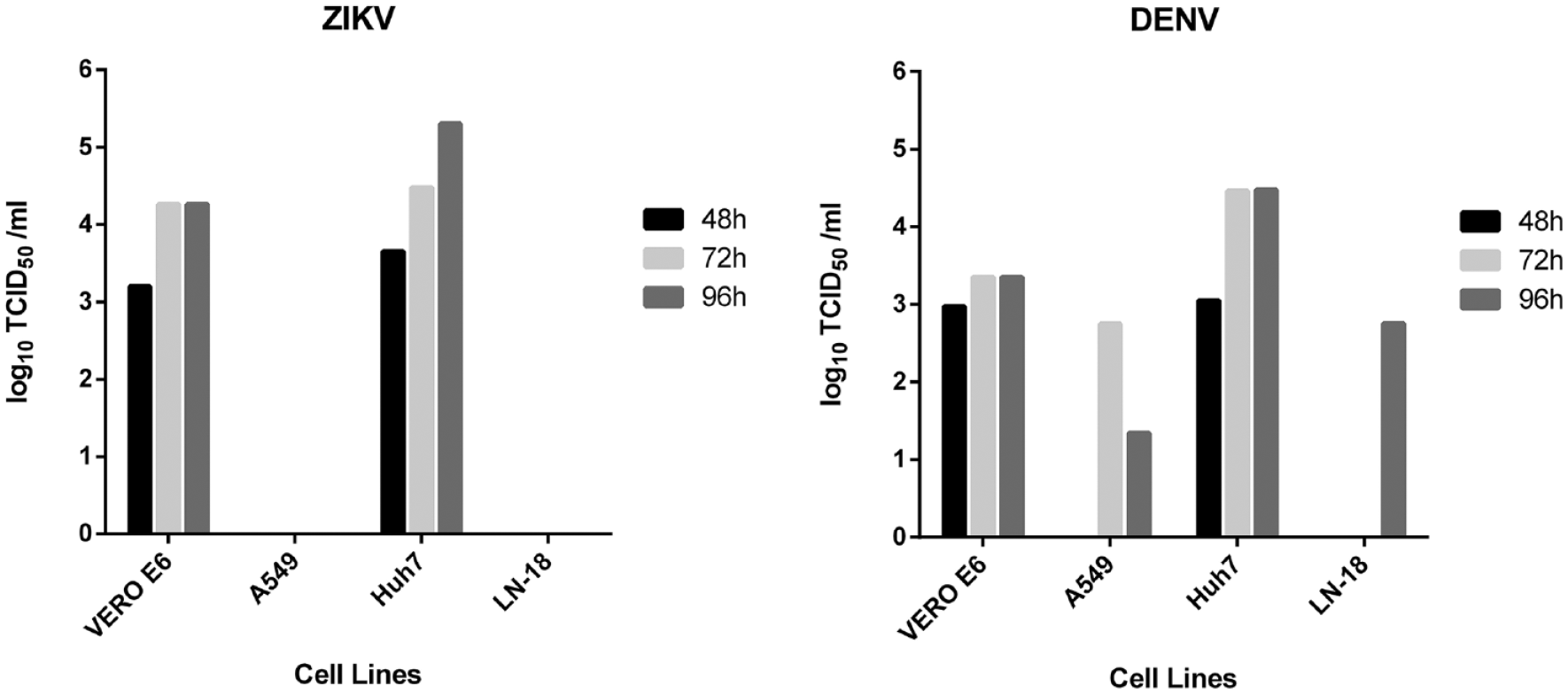
Titration of ZIKV and DENV viral stocks in Huh7, A549, LN-18, and VERO E6 cells at 48, 72, and 96 h by IA.
Performance of the Direct and Secondary YRA in Determining the Antiviral Activity of Reference Compounds
Reference compounds showed no cytotoxicity in the tested concentration range (0.78–200 µM) (
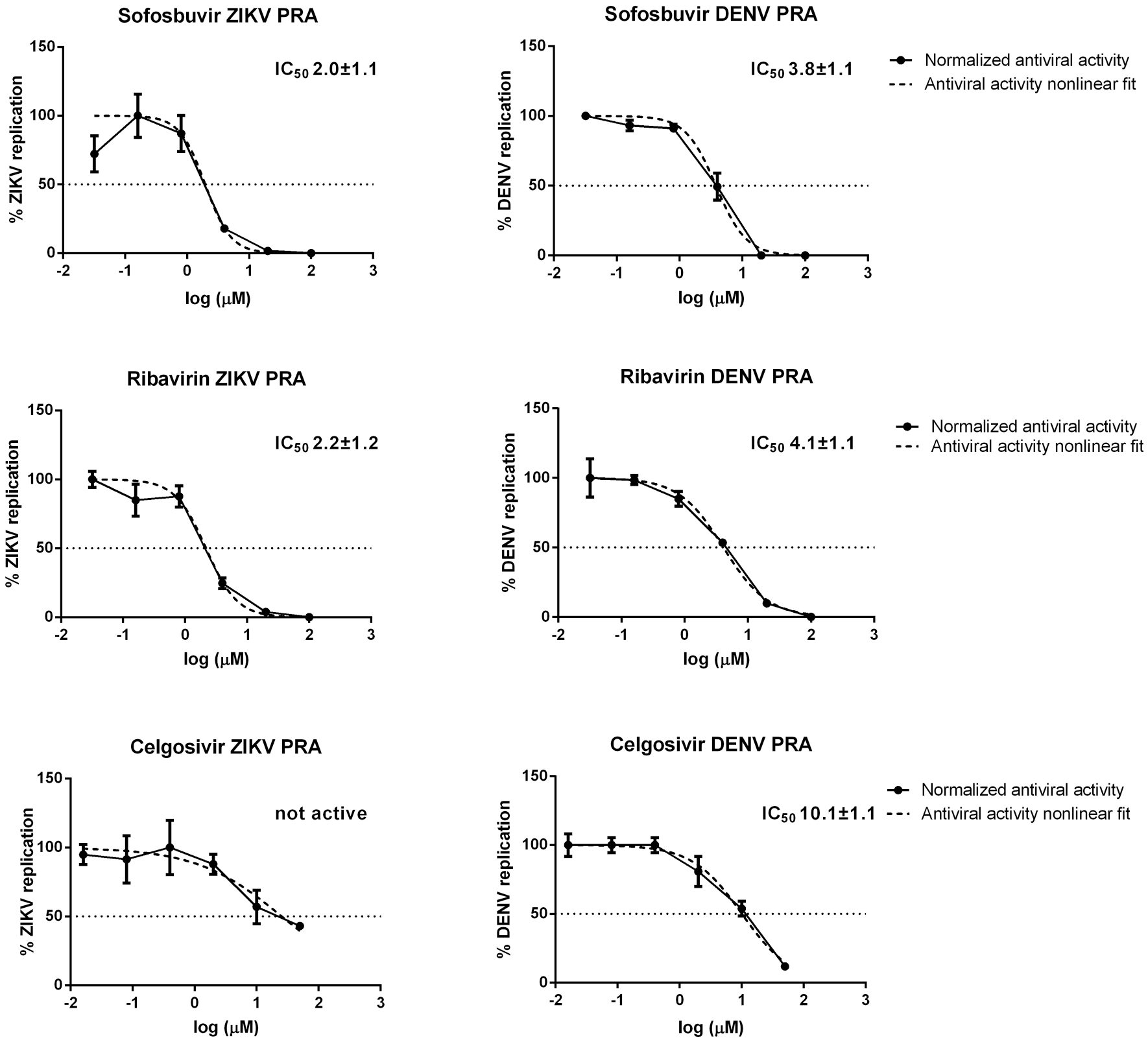
Activity of sofosbuvir and ribavirin against ZIKV and DENV as determined by PRA at 0.1 MOI.
IC50 of Sofosbuvir and Ribavirin against ZIKV and DENV.
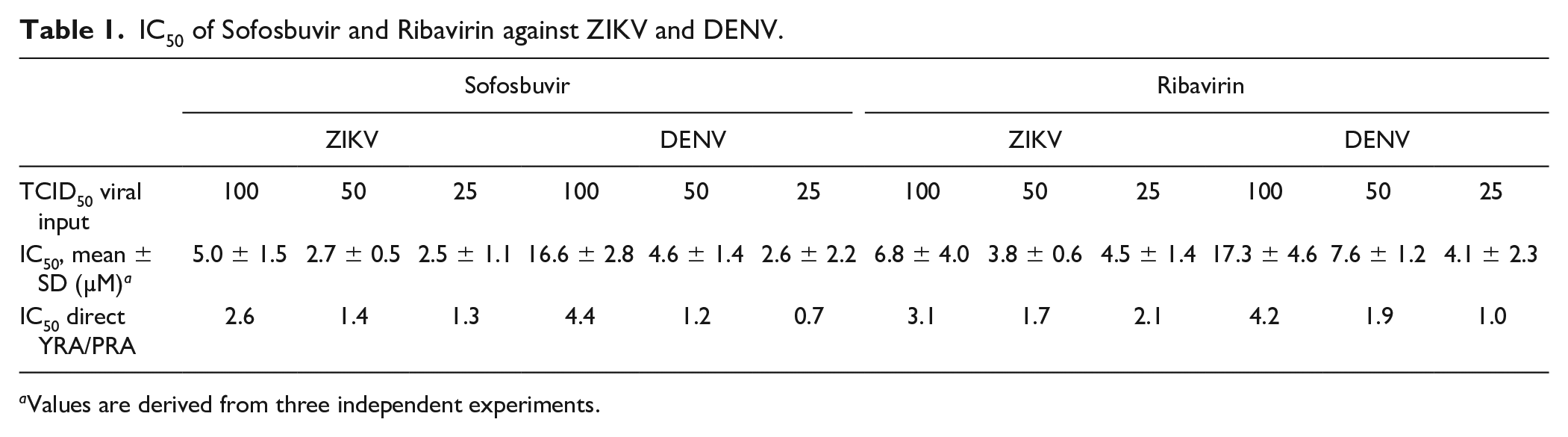
Values are derived from three independent experiments.
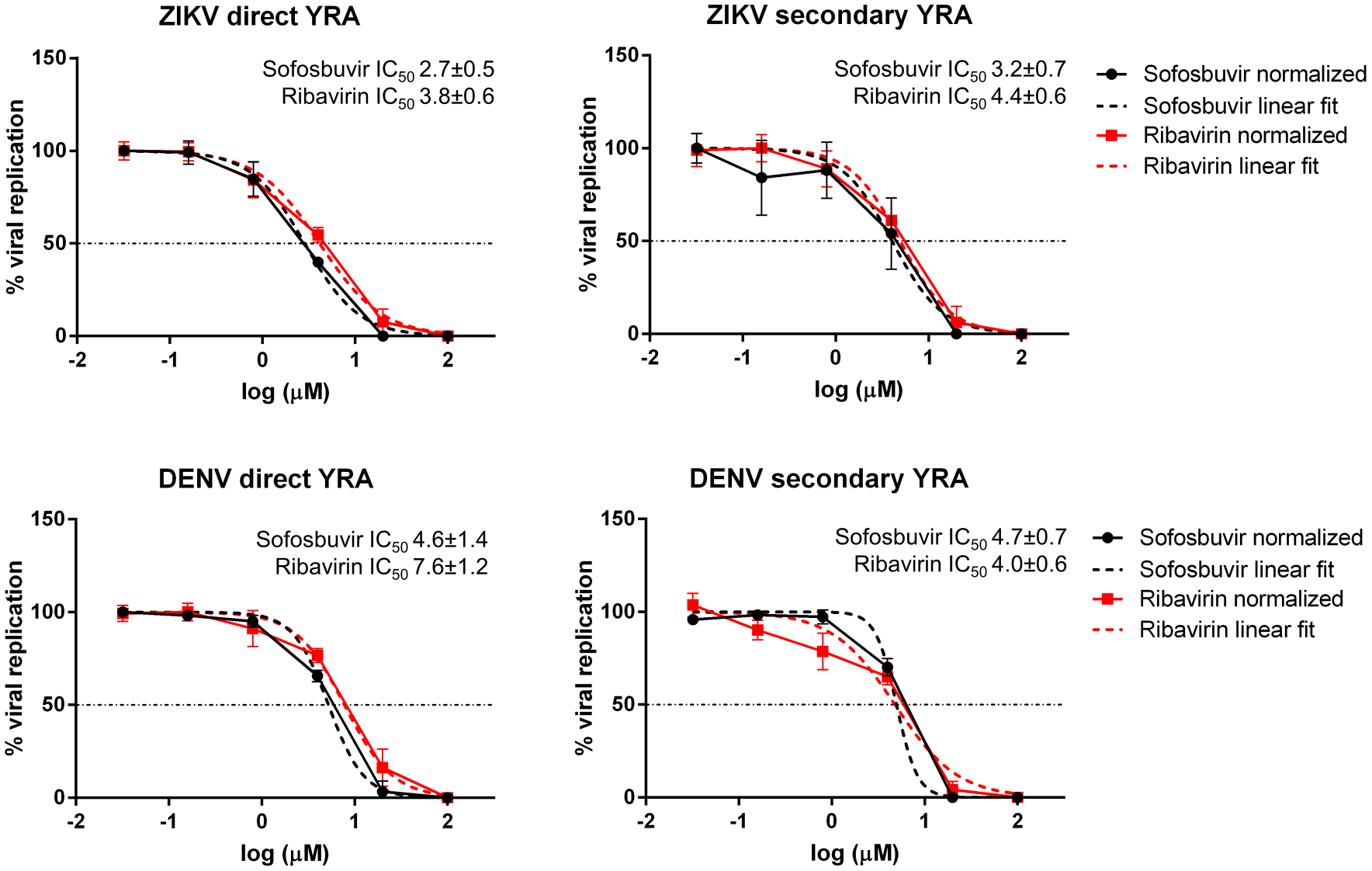
Activity of sofosbuvir and ribavirin against ZIKV and DENV in the direct and secondary YRA.
IC50 Values of Sofosbuvir, Ribavirin, and Celgosivir against ZIKV and DENV.
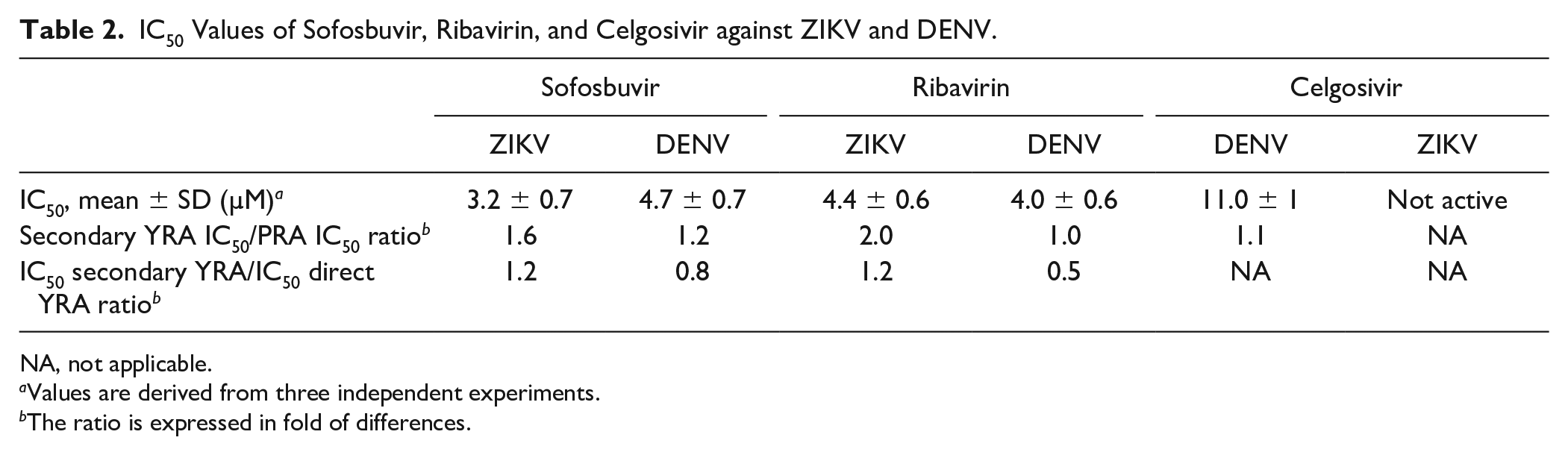
NA, not applicable.
Values are derived from three independent experiments.
The ratio is expressed in fold of differences.
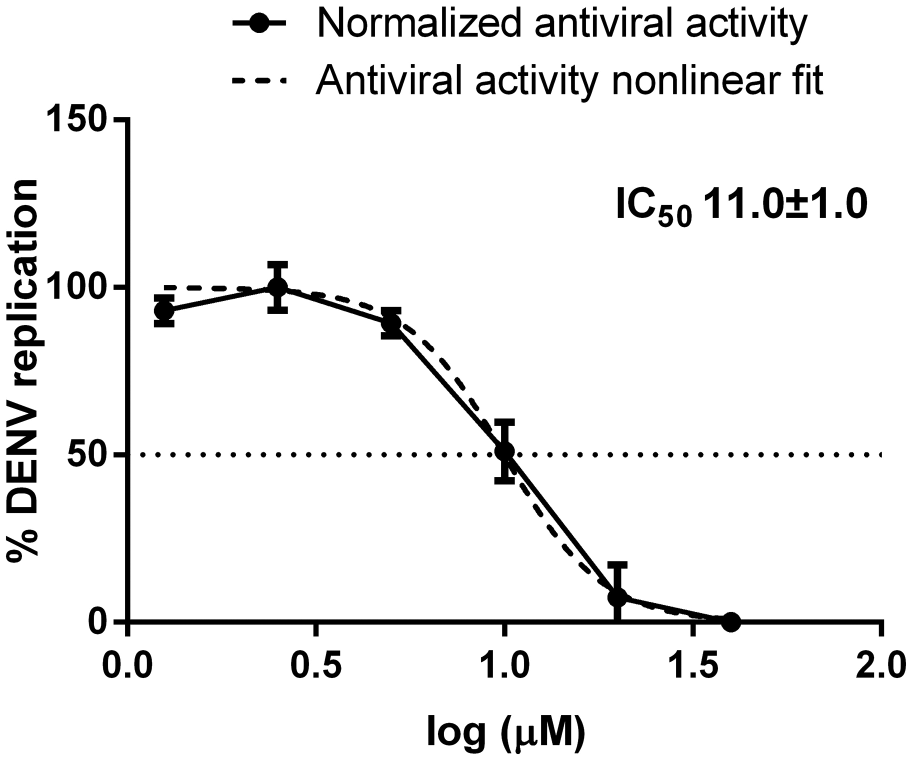
Activity of celgosivir against DENV as determined by the secondary YRA.
Discussion
In the absence of effective vaccines and therapeutic options, supportive care is the only available option for the treatment of flavivirus infections. 34 Assessment of antiviral effects in cultured cells is a key approach for screening candidate compounds. Several cell-based phenotypic assays have been developed, including assays using live virus, subgenomic viral replicons, or virus-like particles. 35 The main disadvantage of the live-virus assays is the obvious necessity for high-level biosafety containment. Subgenomic viral replicons and virus-like particles can overcome safety concerns and are prevalently based on convenient readouts, such as luminescence and fluorescence; however, they do not recapitulate the complete virus life cycle and thus are not amenable for the screening of compounds with unknown targets. Moreover, these assays must be validated carefully to avoid false-positive hits resulting from cytotoxicity or interaction with the luciferase readout. 8 Among live-virus assays, PRA has long been considered the gold standard for antiviral screening and is commonly used for anti-DENV and anti-ZIKV antibody titration in plaque reduction neutralization tests. 36 However, PRA has several drawbacks, including high labor, long-turnaround time, and low throughput, making it not suitable for the analysis of large numbers of compounds or sera.
This study describes the development and validation of an IA-based yield reduction test to simultaneously determine the antiviral activity of candidate compounds against ZIKV and DENV in vitro. To define the best experimental conditions, both viruses were propagated in four different cell lines (Huh7, A549, LN-18, and VERO E6) and the viral titer was determined by IA at different time points. The most effective combination of shorter propagation time and better maintenance of cell health was obtained with Huh7 cells, a widely used human hepatoma cell line, and with VERO E6, the monkey cell line mostly used for the propagation and titration of flaviviruses. However, differences in drug metabolism in monkey cells with respect to human cells 37 impact the activity of sofosbuvir and ribavirin against ZIKV and DENV,25,38 as well as West Nile virus (WNV). 39 Thus, Huh7 was chosen as the model cell line for assay validation. In addition, human cell lines are clearly preferred when assaying candidate host targeting agents for a possible antiviral effect.
The antiviral activity of sofosbuvir and ribavirin was determined by a direct YRA in which the immunodetection of the E protein is directly performed on cells infected with viral stocks and subjected to drug pressure. In the secondary YRA, the antiviral activity is determined by measuring the infectivity of viral stocks generated in the direct YRA. Both drugs were shown to be active against ZIKV and DENV in the low-micromolar range with IC50 values that were comparable in both the direct and secondary YRA performed in this work and in previously reported studies.11,23,24,26 The secondary YRA can additionally screen compounds exerting antiviral activity at the late stage of the viral cycle (i.e., assembly and maturation of viral particles) that would go undetected or only partially detected by direct YRA. For example, a similar two-step system is adopted to measure the anti-HIV activity of drugs acting at different steps of virus replication.40,41 Thus, the combined use of the direct and secondary YRA can not only measure antiviral activity but also help characterize the mechanism of action. As proof of concept, we tested celgosivir, an inhibitor of endoplasmic reticulum (ER) α-glycosidases, found to be active against DENV both in vitro, with IC50 values ranging from the sub- (0.2 µM) to low- (5.7 µM) micromolar range,30,42 and in vivo in a mouse model, demonstrating the reduction of viremia and inducement of protection against virus-induced mortality.30,31 Celgosivir impairs viral protein glycosylation affecting virus assembly and egress, inducing ER stress and the unfolded protein response.
43
We observed that celgosivir did not interfere with the expression of viral E protein at each drug concentration tested in the direct YRA, while a dose-dependent effect of celgosivir on the expression of the E protein was detected in the secondary YRA (
Importantly, the IA format overcomes relevant limitations of the gold standard PRA. The direct YRA and the secondary YRA are completed in 72 and 144 h, respectively, compared with 192 h for ZIKV and 312 h for DENV required by PRA. In addition, the readout is automated through microplate reading as opposed to manual and error-prone counting in PRA. The use of a pan-flaviviral monoclonal antibody allows use of the same system for different viruses, and indeed similar systems have been described for screening antiviral candidates against DENV.26,44 However, several of these procedures rely on high-content fluorescence imaging, which may be not easily available, and none are designed to simultaneously screen multiple viruses or to distinguish between early and late antiviral effects.44–46 Some published protocols were adapted to HTS of large libraries of compounds.32,47 However, these systems are based on CPE readout, an indirect measurement of viral infectivity possibly confounded by cell death caused by candidate compounds, as opposed to direct estimates of virus activity like PRA and IA. In terms of turnaround time (about 4 h for 12 compounds analyzed simultaneously for ZIKV and DENV), our system can be defined as a medium-throughput screening assay suitable for testing small to medium libraries of candidate compounds. In summary, the system described here combines several advantages with respect to previously published work, including (1) the use of the same protocol for two different viruses, (2) the ability to distinguish between early and late antiviral effects, (3) a readout directly proportional to virus production and consequently to virus inhibition, and (4) the completion of the assay within 6 days. Thus, the system provides an opportunity to expand the potential for fast cell-based screening of multiple compounds for antiflavivirus therapy.
Supplemental Material
Supplemental_Matherial_Cell_based_Immunodetection_Assay_DENV_ZIKV_Antivirals_by_Vicenti_et_al_rev_2 – Supplemental material for Development of a Cell-Based Immunodetection Assay for Simultaneous Screening of Antiviral Compounds Inhibiting Zika and Dengue Virus Replication
Supplemental material, Supplemental_Matherial_Cell_based_Immunodetection_Assay_DENV_ZIKV_Antivirals_by_Vicenti_et_al_rev_2 for Development of a Cell-Based Immunodetection Assay for Simultaneous Screening of Antiviral Compounds Inhibiting Zika and Dengue Virus Replication by Ilaria Vicenti, Filippo Dragoni, Alessia Giannini, Federica Giammarino, Michele Spinicci, Francesco Saladini, Adele Boccuto and Maurizio Zazzi in SLAS Discovery
Footnotes
Acknowledgements
We would like to thank Giulietta Venturi, from Istituto Superiore della Sanità (Rome, Italy), for providing the DENV and ZIKV strains used in this study.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: M.Z. reports consultancy for ViiV Healthcare, Gilead Sciences, and Janssen-Cilag, and grants for his institution from ViiV Healthcare and Gilead outside the submitted work.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was partially supported by the PANVIR Project (“Preclinical Development of Novel Panviral Compounds within a Specialized Regional Network,” FESR 2014–2020 Program, Tuscany Region, Italy).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
